Abstract
Background
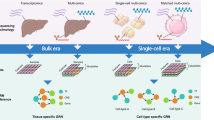
More and more high-throughput datasets are available from multiple levels of measuring gene regulations. The reverse engineering of gene regulatory networks from these data offers a valuable research paradigm to decipher regulatory mechanisms. So far, numerous methods have been developed for reconstructing gene regulatory networks.
Results
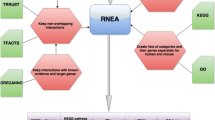
In this paper, we provide a review of bioinformatics methods for inferring gene regulatory network from omics data. To achieve the precision reconstruction of gene regulatory networks, an intuitive alternative is to integrate these available resources in a rational framework. We also provide computational perspectives in the endeavors of inferring gene regulatory networks from heterogeneous data. We highlight the importance of multi-omics data integration with prior knowledge in gene regulatory network inferences.
Conclusions
We provide computational perspectives of inferring gene regulatory networks from multiple omics data and present theoretical analyses of existing challenges and possible solutions. We emphasize on prior knowledge and data integration in network inferences owing to their abilities of identifying regulatory causality.

Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Marx, V. (2013) Biology: the big challenges of big data. Nature, 498, 255–260
Babu, M. M., Luscombe, N. M., Aravind, L., Gerstein, M. and Teichmann, S. A. (2004) Structure and evolution of transcriptional regulatory networks. Curr. Opin. Struct. Biol., 14, 283–291
Liu, Z. P. (2015) Reverse engineering of genome-wide gene regulatory networks from gene expression data. Curr. Genomics, 16, 3–22
Lee, T. I. and Young, R. A. (2013) Transcriptional regulation and its misregulation in disease. Cell, 152, 1237–1251
Bandyopadhyay, S., Mehta, M., Kuo, D., Sung, M. K., Chuang, R., Jaehnig, E. J., Bodenmiller, B., Licon, K., Copeland, W., Shales, M., et al. (2010) Rewiring of genetic networks in response to DNA damage. Science, 330, 1385–1389
Johnson, D. S., Mortazavi, A., Myers, R. M. and Wold, B. (2007) Genome-wide mapping of in vivo protein-DNA interactions. Science, 316, 1497–1502
Park, P. J. (2009) ChIP-seq: advantages and challenges of a maturing technology. Nat. Rev. Genet., 10, 669–680
Edgar, R., Domrachev, M. and Lash, A. E. (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res., 30, 207–210
Brazma, A., Parkinson, H., Sarkans, U., Shojatalab, M., Vilo, J., Abeygunawardena, N., Holloway, E., Kapushesky, M., Kemmeren, P., Lara, G. G., et al. (2003) ArrayExpress—a public repository for microarray gene expression data at the EBI. Nucleic Acids Res., 31, 68–71
Jaenisch, R. and Bird, A. (2003) Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat. Genet., 33, 245–254
Song, C. X., Yi, C. and He, C. (2012) Mapping recently identified nucleotide variants in the genome and transcriptome. Nat. Biotechnol., 30, 1107–1116
Schena, M., Shalon, D., Davis, R. W. and Brown, P. O. (1995) Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science, 270, 467–470
Wang, Z., Gerstein, M. and Snyder, M. (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet., 10, 57–63
Blackham, S., Baillie, A., Al-Hababi, F., Remlinger, K., You, S., Hamatake, R. and McGarvey, M. J. (2010) Gene expression profiling indicates the roles of host oxidative stress, apoptosis, lipid metabolism, and intracellular transport genes in the replication of hepatitis C virus. J. Virol., 84, 5404–5414
Shi, L., Reid, L. H., Jones, W. D., Shippy, R., Warrington, J. A., Baker, S. C., Collins, P. J., de Longueville, F., Kawasaki, E. S., Lee, K. Y., et al.. (2006) The MicroArray Quality Control (MAQC) project shows inter-and intraplatform reproducibility of gene expression measurements. Nat. Biotechnol., 24, 1151–1161
Wu, S., Liu, Z. P., Qiu, X. and Wu, H. (2014) Modeling genomewide dynamic regulatory network in mouse lungs with influenza infection using high-dimensional ordinary differential equations. PLoS One, 9, e95276
Raue, A., Kreutz, C., Maiwald, T., Bachmann, J., Schilling, M., Klingmüller, U. and Timmer, J. (2009) Structural and practical identifiability analysis of partially observed dynamical models by exploiting the profile likelihood. Bioinformatics, 25, 1923–1929
Leinonen, R., Sugawara, H. and Shumway, M., and the International Nucleotide Sequence Database Collaboration. (2011) The sequence read archive. Nucleic Acids Res., 39, D19–D21
The Cancer Genome Atlas Research Network. (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature, 455, 1061–1068
Hudson, T. J., Anderson, W., Artez, A., Barker, A. D., Bell, C., Bernabé, R. R., Bhan, M. K., Calvo, F., Eerola, I., Gerhard, D. S., et al. (2010) International network of cancer genome projects. Nature, 464, 993–998
Benson, D. A., Cavanaugh, M., Clark, K., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. and Sayers, E. W. (2013) GenBank. Nucleic Acids Res., 41, D36–D42
Maher, B. (2012) ENCODE: the human encyclopaedia. Nature, 489, 46–48
Muers, M. (2011) Functional genomics: the modENCODE guide to the genome. Nat. Rev. Genet., 12, 80
Bernstein, B. E., Stamatoyannopoulos, J. A., Costello, J. F., Ren, B., Milosavljevic, A., Meissner, A., Kellis, M., Marra, M. A., Beaudet, A. L., Ecker, J. R., et al. (2010) The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol., 28, 1045–1048
Fingerman, I. M., McDaniel, L., Zhang, X., Ratzat, W., Hassan, T., Jiang, Z., Cohen, R. F. and Schuler, G. D. (2011) NCBI Epigenomics: a new public resource for exploring epigenomic data sets. Nucleic Acids Res., 39, D908–D912
Cantara, W. A., Crain, P. F., Rozenski, J., McCloskey, J. A., Harris, K. A., Zhang, X., Vendeix, F. A., Fabris, D. and Agris, P. F. (2011) The RNA Modification Database, RNAMDB: 2011 update. Nucleic Acids Res., 39, D195–D201
Machnicka, M. A., Milanowska, K., Osman Oglou, O., Purta, E., Kurkowska, M., Olchowik, A., Januszewski, W., Kalinowski, S., Dunin-Horkawicz, S., Rother, K. M., et al. (2013) MODOMICS: a database of RNA modification pathways—2013 update. Nucleic Acids Res., 41, D262–D267
Bujold, D., de Lima Morais, D.A., Gauthier, C., Côté, C., Caron, M., Kwan, T., Chen, K.T., Laperle, J., Markovits, A. N., Pastinen, T., et al. (2016) The International Human Epigenome Consortium Data Portal. Cell Syst., 3, 496–499
Ardlie, K. G., Deluca, D. S., Segre, A. V., Sullivan, T. J., Young, T. R., Gelfand, E. T., Trowbridge, C. A., Maller, J. B., Tukiainen, T., Lek, M., et al. (2015) The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science, 348, 648–660
Matys, V., Fricke, E., Geffers, R., Gössling, E., Haubrock, M., Hehl, R., Hornischer, K., Karas, D., Kel, A. E., Kel-Margoulis, O. V., et al. (2003) TRANSFAC: transcriptional regulation, from patterns to profiles. Nucleic Acids Res., 31, 374–378
Bryne, J. C., Valen, E., Tang, M. H., Marstrand, T., Winther, O., da Piedade, I., Krogh, A., Lenhard, B. and Sandelin, A. (2008) JASPAR, the open access database of transcription factor-binding profiles: new content and tools in the 2008 update. Nucleic Acids Res., 36, D102–D106
Liu, Z. P., Wu, C., Miao, H. and Wu, H. (2015) RegNetwork: an integrated database of transcriptional and post-transcriptional regulatory networks in human and mouse. Database (Oxford), 2015, bav095
Xie, C., Yuan, J., Li, H., Li, M., Zhao, G., Bu, D., Zhu, W., Wu, W., Chen, R. and Zhao, Y. (2014) NONCODEv4: exploring the world of long non-coding RNA genes. Nucleic Acids Res., 42, D98–D103
The RNAcentral Consortium. (2015) RNAcentral: an international database of ncRNA sequences. Nucleic Acids Res., 43, D123–D129
Sethupathy, P., Corda, B. and Hatzigeorgiou, A. G. (2006) TarBase: a comprehensive database of experimentally supported animal microRNA targets. RNA, 12, 192–197
Volders, P. J., Helsens, K., Wang, X., Menten, B., Martens, L., Gevaert, K., Vandesompele, J. and Mestdagh, P. (2013) LNCipedia: a database for annotated human lncRNA transcript sequences and structures. Nucleic Acids Res., 41, D246–D251
Amaral, P. P., Clark, M. B., Gascoigne, D. K., Dinger, M. E. and Mattick, J. S. (2011) lncRNAdb: a reference database for long noncoding RNAs. Nucleic Acids Res., 39, D146–D151
Griffiths-Jones, S., Saini, H. K., van Dongen, S. and Enright, A. J. (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res., 36, D154–D158
Glažar, P., Papavasileiou, P. and Rajewsky, N. (2014) circBase: a database for circular RNAs. RNA, 20, 1666–1670
Yang, J. H., Li, J. H., Jiang, S., Zhou, H. and Qu, L. H. (2013) ChIPBase: a database for decoding the transcriptional regulation of long non-coding RNA and microRNA genes from ChIP-Seq data. Nucleic Acids Res., 41, D177–D187
Wang, Q., Huang, J., Sun, H., Liu, J.,Wang, J.,Wang, Q., Qin, Q., Mei, S., Zhao, C., Yang, X., et al. (2014) CR Cistrome: a ChIPSeq database for chromatin regulators and histone modification linkages in human and mouse. Nucleic Acids Res., 42, D450–D458
Berman, H. M., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T. N., Weissig, H., Shindyalov, I. N. and Bourne, P. E. (2000) The Protein Data Bank. Nucleic Acids Res., 28, 235–242
The UniProt Consortium. (2008) The universal protein resource (UniProt). Nucleic Acids Res., 36, D190–D195
von Mering, C., Jensen, L. J., Kuhn, M., Chaffron, S., Doerks, T., Krüger, B., Snel, B. and Bork, P. (2007) STRING 7—recent developments in the integration and prediction of protein interactions. Nucleic Acids Res., 35, D358–D362
Zhang, X., Zhao, X. M., He, K., Lu, L., Cao, Y., Liu, J., Hao, J. K., Liu, Z. P. and Chen, L. (2012) Inferring gene regulatory networks from gene expression data by path consistency algorithm based on conditional mutual information. Bioinformatics, 28, 98–104
Zhang, B. and Horvath, S. (2005) A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol., 4, Article17
Meyer, P. E., Lafitte, F. and Bontempi, G. (2008) minet: a R/Bioconductor package for inferring large transcriptional networks using mutual information. BMC Bioinformatics, 9, 461
Margolin, A. A., Nemenman, I., Basso, K., Wiggins, C., Stolovitzky, G., Dalla Favera, R. and Califano, A. (2006) ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinformatics, 7, S7
Wilczyński, B. and Dojer, N. (2009) BNFinder: exact and efficient method for learning Bayesian networks. Bioinformatics, 25, 286–287
Scutari, M. (2010) Learning Bayesian Networks with the bnlearn R Package. J. Stat. Softw., 35, 1–22
Shmulevich, I., Dougherty, E. R., Kim, S. and Zhang, W. (2002) Probabilistic Boolean Networks: a rule-based uncertainty model for gene regulatory networks. Bioinformatics, 18, 261–274
Müssel, C., Hopfensitz, M. and Kestler, H. A. (2010) BoolNe—an R package for generation, reconstruction and analysis of Boolean networks. Bioinformatics, 26, 1378–1380
Schaffter, T., Marbach, D. and Floreano, D. (2011) GeneNet-Weaver: in silico benchmark generation and performance profiling of network inference methods. Bioinformatics, 27, 2263–2270
Bonneau, R., Reiss, D. J., Shannon, P., Facciotti, M., Hood, L., Baliga, N. S. and Thorsson, V. (2006) The Inferelator: an algorithm for learning parsimonious regulatory networks from systems-biology data sets de novo. Genome Biol., 7, R36
Liu, Z. P., Zhang,W., Horimoto, K. and Chen, L. (2013) Gaussian graphical model for identifying significantly responsive regulatory networks from time course high-throughput data. IET Syst. Biol., 7, 143–152
Liu, Z. P., Wu, H., Zhu, J. and Miao, H. (2014) Systematic identification of transcriptional and post-transcriptional regulations in human respiratory epithelial cells during influenza A virus infection. BMC Bioinformatics, 15, 336
Haury, A. C., Mordelet, F., Vera-Licona, P. and Vert, J. P. (2012) TIGRESS: trustful inference of gene regulation using stability selection. BMC Syst. Biol., 6, 145
Huynh-Thu, V. A., Irrthum, A., Wehenkel, L. and Geurts, P. (2010) Inferring regulatory networks from expression data using tree-based methods. PLoS One, 5, e12776
Langfelder, P. and Horvath, S. (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics, 9, 559
Liu, Z. P. (2017) Quantifying gene regulatory relationships with association measures: a comparative study. Front. Genet., 8, 96
Basso, K., Margolin, A. A., Stolovitzky, G., Klein, U., Dalla-Favera, R. and Califano, A. (2005) Reverse engineering of regulatory networks in human B cells. Nat. Genet., 37, 382–390
Friedman, N. (2004) Inferring cellular networks using probabilistic graphical models. Science, 303, 799–805
Zou, M. and Conzen, S. D. (2005) A new dynamic Bayesian network (DBN) approach for identifying gene regulatory networks from time course microarray data. Bioinformatics, 21, 71–79
Amit, I., Garber, M., Chevrier, N., Leite, A. P., Donner, Y., Eisenhaure, T., Guttman, M., Grenier, J. K., Li,W., Zuk, O., et al. (2009) Unbiased reconstruction of a mammalian transcriptional network mediating pathogen responses. Science, 326, 257–263
Thomas, R. (1973) Boolean formalization of genetic control circuits. J. Theor. Biol., 42, 563–585
Akutsu, T., Miyano, S. and Kuhara, S. (1999) Identification of genetic networks from a small number of gene expression patterns under the Boolean network model. Pac. Symp. Biocomput., 99, 17–28
Saito, S., Aburatani, S. and Horimoto, K. (2008) Network evaluation from the consistency of the graph structure with the measured data. BMC Syst. Biol., 2, 84
Jordan, M. I. and Mitchell, T. M. (2015) Machine learning: trends, perspectives, and prospects. Science, 349, 255–260
Marbach, D., Roy, S., Ay, F., Meyer, P. E., Candeias, R., Kahveci, T., Bristow, C. A. and Kellis, M. (2012) Predictive regulatory models in Drosophila melanogaster by integrative inference of transcriptional networks. Genome Res., 22, 1334–1349
Bartel, D. P. (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell, 116, 281–297
Pefanis, E., Wang, J., Rothschild, G., Lim, J., Kazadi, D., Sun, J., Federation, A., Chao, J., Elliott, O., Liu, Z. P., et al. (2015) RNA exosome-regulated long non-coding RNA transcription controls super-enhancer activity. Cell, 161, 774–789
Memczak, S., Jens, M., Elefsinioti, A., Torti, F., Krueger, J., Rybak, A., Maier, L., Mackowiak, S. D., Gregersen, L. H., Munschauer, M., et al. (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature, 495, 333–338
Garber, M., Grabherr, M. G., Guttman, M. and Trapnell, C. (2011) Computational methods for transcriptome annotation and quantification using RNA-seq. Nat. Methods, 8, 469–477
Irizarry, R. A., Hobbs, B., Collin, F., Beazer-Barclay, Y. D., Antonellis, K. J., Scherf, U. and Speed, T. P. (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics, 4, 249–264
Elowitz, M. B., Levine, A. J., Siggia, E. D. and Swain, P. S. (2002) Stochastic gene expression in a single cell. Science, 297, 1183–1186
Gibcus, J. H. and Dekker, J. (2012) The context of gene expression regulation. F1000 Biol. Rep., 4, 8
Ideker, T., Dutkowski, J. and Hood, L. (2011) Boosting signal-tonoise in complex biology: prior knowledge is power. Cell, 144, 860–863
de la Fuente, A., Bing, N., Hoeschele, I. and Mendes, P. (2004) Discovery of meaningful associations in genomic data using partial correlation coefficients. Bioinformatics, 20, 3565–3574
Zheng, G., Xu, Y., Zhang, X., Liu, Z. P., Wang, Z., Chen, L. and Zhu, X. G. (2016) CMIP: a software package capable of reconstructing genome-wide regulatory networks using gene expression data. BMC Bioinformatics, 17, 535
Burchard, J., Zhang, C., Liu, A. M., Poon, R. T., Lee, N. P., Wong, K. F., Sham, P. C., Lam, B. Y., Ferguson, M. D., Tokiwa, G., et al. (2010) microRNA-122 as a regulator of mitochondrial metabolic gene network in hepatocellular carcinoma. Mol. Syst. Biol., 6, 402
Liu, Z. P. (2014) Systematic identification of local structure binding motifs in protein-RNA recognition. In Proceedings of 8th International Conference on Systems Biology, pp. 74–80
Cheng, C., Yan, K. K., Hwang, W., Qian, J., Bhardwaj, N., Rozowsky, J., Lu, Z. J., Niu, W., Alves, P., Kato, M., et al. (2011) Construction and analysis of an integrated regulatory network derived from high-throughput sequencing data. PLoS Comput. Biol., 7, e1002190
The ENCODE Project Consortium. (2012) An integrated encyclopedia of DNA elements in the human genome. Nature, 489, 57–74
Amaral, P. P., Dinger, M. E., Mercer, T. R. and Mattick, J. S. (2008) The eukaryotic genome as an RNA machine. Science, 319, 1787–1789
Spitz, F. and Furlong, E. E. (2012) Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet., 13, 613–626
Hecker, M., Lambeck, S., Toepfer, S., van Someren, E. and Guthke, R. (2009) Gene regulatory network inference: data integration in dynamic models-a review. Biosystems, 96, 86–103
Jensen, S. T., Chen, G. and Stoeckert, C. J. Jr (2007) Bayesian variable selection and data integration for biological regulatory networks. Ann. Appl. Stat., 1, 612–633
Yeung, M. K., Tegnér, J. and Collins, J. J. (2002) Reverse engineering gene networks using singular value decomposition and robust regression. Proc. Natl. Acad. Sci. USA, 99, 6163–6168
Tegner, J., Yeung, M. K., Hasty, J. and Collins, J. J. (2003) Reverse engineering gene networks: integrating genetic perturbations with dynamical modeling. Proc. Natl. Acad. Sci. USA, 100, 5944–5949
Lam, K. Y., Westrick, Z. M., Müller, C. L., Christiaen, L. and Bonneau, R. (2016) Fused regression for multi-source gene regulatory network inference. PLoS Comput. Biol., 12, e1005157
Werhli, A. V. and Husmeier, D. (2007) Reconstructing gene regulatory networks with bayesian networks by combining expression data with multiple sources of prior knowledge. Stat. Appl. Genet. Mol. Biol., 6, Article15
Zhu, J., Zhang, B., Smith, E. N., Drees, B., Brem, R. B., Kruglyak, L., Bumgarner, R. E. and Schadt, E. E. (2008) Integrating large-scale functional genomic data to dissect the complexity of yeast regulatory networks. Nat. Genet., 40, 854–861
Santra, T. (2014) A Bayesian framework that integrates heterogeneous data for inferring gene regulatory networks. Front. Bioeng. Biotechnol., 2, 13
De Smet, R. and Marchal, K. (2010) Advantages and limitations of current network inference methods. Nat. Rev. Microbiol., 8, 717–729
Mordelet, F. and Vert, J. P. (2008) SIRENE: supervised inference of regulatory networks. Bioinformatics, 24, i76–i82
Patel, A. P., Tirosh, I., Trombetta, J. J., Shalek, A. K., Gillespie, S. M., Wakimoto, H., Cahill, D. P., Nahed, B. V., Curry, W. T., Martuza, R. L., et al. (2014) Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science, 344, 1396–1401
Djebali, S., Davis, C. A., Merkel, A., Dobin, A., Lassmann, T., Mortazavi, A., Tanzer, A., Lagarde, J., Lin, W., Schlesinger, F., et al. (2012) Landscape of transcription in human cells. Nature, 489, 101–108
Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. and Elowitz, M. B. (2005) Gene regulation at the single-cell level. Science, 307, 1962–1965
Marbach, D., Costello, J. C., Küffner, R., Vega, N. M., Prill, R. J., Camacho, D. M., Allison, K. R., Kellis, M., Collins, J. J. and Stolovitzky, G., et al. (2012) Wisdom of crowds for robust gene network inference. Nat. Methods, 9, 796–804
Moignard, V., Woodhouse, S., Haghverdi, L., Lilly, A. J., Tanaka, Y., Wilkinson, A. C., Buettner, F., Macaulay, I. C., Jawaid, W., Diamanti, E., et al. (2015) Decoding the regulatory network of early blood development from single-cell gene expression measurements. Nat. Biotechnol., 33, 269–276
Graham, J. E., Marians, K. J. and Kowalczykowski, S. C. (2017) Independent and stochastic action of DNA polymerases in the replisome. Cell, 169, 1201–1213
Acknowledgements
Thanks are due to the three anonymous reviewers for their constructive comments. This work was partially supported by the National Natural Science Foundation of China (Nos. 61572287 and 61533011), the Shandong Provincial Key Research and Development Program (2018GSF118043), the Natural Science Foundation of Shandong Province, China (ZR2015FQ001), the Fundamental Research Funds of Shandong University (Nos. 2015QY001 and 2016JC007), the Scientific Research Foundation for the Returned Overseas Chinese Scholars, Ministry of Education of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Author summary: In this paper, we summarize and comment recent progresses in the important bioinformatics field of gene regulatory network inference from quantitative gene expression profiles, especially focus on the endeavors for improving the inference precision by integrating multiple resources. The paper will potentially facilitate scientists who are interested in reversely engineering gene regulatory network to quickly obtain an integrative overview and follow the start-of-the-art computational techniques.
Rights and permissions
About this article
Cite this article
Liu, ZP. Towards precise reconstruction of gene regulatory networks by data integration. Quant Biol 6, 113–128 (2018). https://doi.org/10.1007/s40484-018-0139-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-018-0139-4