Abstract
Knowledge of the vastness of microbial diversity associated with plants is still limited. Plant microbiome structure and functions are shaped by several factors, including host genotype and developmental stage, the presence or absence of diseases, and environmental conditions. These factors may lead to distinct microbial communities in the rhizosphere, endosphere, and phyllosphere. Studies directed to microbial interactions in plant compartments are fundamental for understanding the microbial ecology of phytobiomes, enabling the development of microbiome-based technologies in the search for sustainable agriculture. In this chapter, we describe plant compartments, i.e., the rhizosphere, phyllosphere and endosphere, and the more common bacterial composition of each compartment. We also discuss manipulation of the plant microbiome toward improved plant health. Advances in this field will lead to strategies where the manipulation of the plant microbiome will allow the reduction of pesticide and fertilizer use in field crops, paving the way to a more sustainable agriculture.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Microbial communities
- Rhizosphere
- Endosphere
- Phyllosphere
- Phytobiome manipulation
- Plant-microbiome interaction
- Endophytes
Introduction
The concept of the microbiome was described for the first time as the "ecological community of commensal microorganisms, symbionts or pathogens, which literally occupy a space in our body" [1]. Recently, this term has been used for different environments inhabited by microorganisms [2,3,4]. This term has also been used in the plant context as "an environment, which consists of the plant and all microbes associated with it" [3].
The relationship between plants and their surroundings, especially those plant-microbe interactions with a beneficial output, has been the center of attention of various studies [5]. Traditionally, many researchers have tried to understand these interactions, looking to individual plant-microbe relationships, i.e., a one vs. one approach, but these interactions are much more complex, as they involve a vast diversity of microbes and environmental factors [6].
Plant, soil, soil-borne microbes, and environmental factors together influence the various changes that cooperate to create plant health and productivity. Recent advances in “-omics” research have shed light on microbiome compositions and interactions with the environment [7]. These advances have contributed to the development of novel approaches that seek to improve plant fitness through the artificial selection of microbes with specific effects on host performance. The selection of microbial communities occurs indirectly through host traits that have coevolved together with the microorganisms and influence the microbiomes [8].
In this chapter, firstly, we define each plant compartment, i.e., the rhizosphere, phyllosphere, and endosphere, and within each compartment we describe “who” is there (microbiome structure), “what” they are doing (microbiome functions), and what are the major drivers shaping the assembly of the microbiome. Finally, we discuss the advances in microbiome manipulation and the possibilities of using such manipulation to improve and optimize crop productivity.
The Rhizosphere Ecosystem
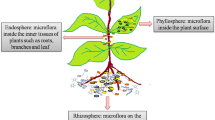
The term rhizosphere was coined by the soil bacteriologist Lorenz Hiltner in 1904 [9]. This term is derived from the Greek word rhiza (root) and the Latin word sphaera (sphere), referring to an environment or compartment that encloses the inhabited “microbial world” on the plant roots. The rhizosphere is the narrow zone of soil surrounding the root system where plants and microorganisms interact [10,11,12,13,14] (Fig. 1) and it is characterized by a chemical, biological, and physical gradient that changes radially and longitudinally along the roots [15].
Schematic representation of plant microbiome compartments and frequency of studies describing bacterial phyla in each compartment, i.e., phyllosphere, endosphere, and rhizosphere. Each pie graph shows the frequency of studies reporting bacterial phyla per plant compartment. For example, 18% of 15 studies on the phyllosphere detected Actinobacteria in the bacterial community. Seventy-one studies were surveyed, 15 for the phyllosphere, 29 for the endosphere, and 27 for the rhizosphere. Searches were performed in the Scopus database between February 03, 2016 and March 15, 2016. The search used a combination of words describing plant compartments (“rhizosphere”, “phyllosphere”, “endophytic”, “endosphere”) and investigative techniques (“sequencing”, “metagenomic”, “next-generation sequencing”). Studies using cultivation-dependent approaches were not included in the survey. Phyla cited in only one manuscript were included in the “Others” category
The idea of microbial colonization of the rhizosphere seems to be supported by the niche theory of species diversity, which is driven by various abiotic and biotic factors, such as plant genotype and soil [5, 13, 16,17,18,19]. Changes in the rhizosphere microbial community begin when the soil microbiota is exposed to rhizodeposits, which are influenced by the plant genotype, including glucose, amino acids, organic acids, polysaccharides, and proteins [10, 13]. Rhizodeposition increases the microbial populations in the rhizosphere, known as the “rhizosphere effect” [11,12,13, 16]. Later, the plant genotype selects and assembles a closely associated microbial community in the rhizoplane and within the plant roots [13, 16, 20]. It has been hypothesized that each plant species selects specific microbial populations as a result of the high degree of host specificity in the coevolution of plants and microbes [5, 13, 21].
Plants release exudates into their direct surroundings to attract, stimulate, or repel microorganisms on the roots. The amount and composition of the rhizodeposits, which structure and modulate the rhizosphere microbial community throughout the plant life cycle, may vary among different plant species [22] and throughout their growth [23], as well as in different stages of root development [5]. Microbial succession starts with the release of carbon from seeds during the germination stage, and microorganisms in the rhizosphere are distributed according to root type and zones, as well as according to their movement through the soil during root growth [13]. In the early stages of plant development (seedlings), alcohol and sugars are released, while in the later stages, amino acids and phenolic compounds predominate [23]. This phenomenon suggests the attraction of a large diversity of microorganisms in the early stages of plant development, while later the release of specific substrates selects certain microorganisms in the rhizosphere [5, 21, 23].
The number of microorganisms in the rhizosphere is higher than that in bulk soil, due to the carbon availability in the rhizosphere. Generally, gram-negative bacteria are stimulated by rhizodeposition, whereas gram-positive bacteria are inhibited [10]. Proteobacteria (α, β, γ), Firmicutes, Actinobacteria, Bacteroidetes, Crenarchaeota, Acidobacteria, Ascomycota, and Glomeromycota, and also unclassified bacteria, represent relatively large groups detected in the rhizosphere [5, 12, 13] (Fig. 1).
The microorganisms found in the rhizosphere can have beneficial or deleterious effects on the growth and health of the plant [13]. The beneficial microbes, among others, include mycorrhizal fungi and rhizobia, which provide phosphorus and nitrogen; siderophore-producing bacteria, which facilitate iron acquisition; and plant-growth-promoting rhizobacteria (PGPR), which promote plant growth [12, 14, 24]. PGPR can suppress disease by mechanisms such as competition for nutrients and microsites, parasitism and antibiosis, or by inducing systemic resistance to pathogens in the plant [13]. There are some examples of microorganisms that promote plant adaptation to abiotic stresses such as drought, flooding, saline stress, temperature or pH extremes, and high concentrations of toxic compounds, and these cases reveal complex associations of microorganisms with plants as a result of coevolution in their native habitats [13, 25]. Biotic stress includes the presence of phytopathogenic microorganisms such as nematodes, fungi, and oomycetes, which have agronomic importance because they reduce the yields of food, feed, fiber, and fuel crops [12].
Given that root exudates are strongly linked to the recruitment of the microorganisms that comprise the rhizosphere microbial community, it can be seen that the rhizosphere is closely involved with plant health and growth; therefore, the understanding of rhizosphere functioning and ecology is key to increasing crop yield.
The Phyllosphere
The second compartment of the plant microbiome is the phyllosphere, or aerial plant surface, which is characterized as being nutrient poor when compared with the rhizosphere [26]. The phyllosphere is composed of microbial cells that are able to colonize the aerial plant surfaces [27, 28] that are dominated by the leaves, although the term phyllosphere can be used for any aerial part of the plant [29] (Fig. 1).
The microbial habitat on the surfaces of leaves may be one of the largest microbial habitats on earth, with the terrestrial leaf surface area estimated to exceed 108 km2 globally [30]. The phyllosphere microbiome is composed of viruses, bacteria, filamentous fungi, yeasts, algae, and, occasionally, protozoa and nematodes [26]. Bacteria are the most abundant of the cellular organisms in the phyllosphere community, present in numbers between 106 and 107 cells cm−2 of leaf tissue [26, 29]. Fungi and archaea are apparently less abundant; however, their population has not been estimated yet [26, 30, 31].
Overall, species richness in phyllosphere communities is high [32]; however, the bacterial community diversity is lower than the diversity of the communities in the rhizosphere or bulk soil [31, 33]. Advances in sequencing technologies have vastly expanded our understanding of plant microbiome structure, including that in the phyllosphere [34]. At the phylum level, the phyllosphere bacterial communities are composed mainly of Actinobacteria, Bacteroidetes, Firmicutes, and Proteobacteria [35], with a predominance of the classes Alphaproteobacteria and Gammaproteobacteria [36, 37] (Fig. 1). Further analysis of community composition at the genus level suggests that Pseudomonas, Sphingomonas, Methylobacterium, Bacillus, Massilia, Arthrobacter, and Pantoea are consistently found as part of the phyllosphere microbiome across a wide range of plant species [35].
The colonization of plant leaf surfaces, in large part, occurs through the immigration of bacteria, fungi, and other microorganisms from air, soil, water, seeds, or through animal sources [29]. Furthermore, studies have shown that some of these microorganisms of the foliar microbiome can be transferred not only through environmental exchange, but also vertically, through generations of plants [38]. Neighboring environmental ecosystems can also randomly contribute to the assembly of the foliar microbiome [39]. Even after the stabilization of phyllosphere microbial communities, variations may occur, caused by nutritional heterogeneity in different regions on the leaf surface, where the carbon sources (e.g., glucose, fructose, and sucrose) are spatially heterogeneous, leading to distinct microbial assemblages on the leaf veins, which are regions near the stomata and surface appendages [26, 29]. Large fluxes in temperature, moisture, and radiation throughout the day and night also cause changes in the phyllosphere microbiome structure [26, 29, 40]. In some cases, this spatial heterogeneity is promoted by the organization of microbial cells into biofilms, which are a common feature of organisms in the phyllosphere, acting as aggregators and protectors of the microbial cells under the frequently inhospitable conditions [26, 41].
The microbial communities found in the phyllosphere may perform key processes related to plant development; for example, nitrogen fixation [42, 43], protection from invading pathogens [44], modification of metabolites, and the biosynthesis of phytohormones [45]. Metagenomic and metaproteomic studies showed that microbes in the phyllosphere could produce proteins that promote substrate uptake, via porins and ABC transporters; resistance to stresses, including reactive oxygen species (ROS); and nutrient cycling [31]. Methylobacteria are involved in methanotrophy and are often detected in phyllosphere communities [46, 47].
The interactions between the plant and the phyllosphere microbial communities, and the variations in their distinct environmental factors, modulate the assemblage of these microbial communities in the phyllosphere and contribute to the heterogeneity in their abundance and structure in distinct plant species. New molecular technologies have shown the importance of microbial functions in the phyllosphere and have provided new insights into the major drivers of microbial community composition. The combination of multiple “omics” technologies will lead us to a system-level understanding of the phyllosphere microbial communities and their physiological potential.
The Endosphere
The endosphere consists of the inner plant tissues, inhabited by microorganisms intimately interacting with the host plant [28, 48, 49]. This compartment is composed of the internal root tissue (endorhizosphere), internal shoot and leaf tissue (endophyllosphere), internal plant reproductive tissue, and the internal seed tissue [50,51,52,53,54,55]. Endophytic microorganisms are organisms that reside internally in plant tissues for at least part of their life cycle [48] without causing visible disease symptoms [56] and they can be accessed from the plant after surface disinfection by cultivation-dependent and/or molecular approaches [57,58,59] (Fig. 1). Although this concept is one of the most commonly accepted ones and is currently applied, it is important to note that there are niches on the surfaces of aerial parts and roots where microorganisms may remain protected from the action of the chemical products usually used for surface disinfection. Recent studies have used sonication to remove surface layers of the plant tissue and to access the endophytic microorganisms on the remaining tissue [17, 20].
Endophytes are beneficial or commensal, and they can shift between parasitic and mutualistic life strategies [60, 61]. Their beneficial role in plant development and health can be mediated and is characterized by metabolic interactions, including the production of plant growth hormones [62,63,64], antibiotics, and toxicants [65, 66]; the improvement of nutrient uptake; and/or increasing the plant tolerance to biotic and abiotic stresses [62, 67, 68]. In addition to these characteristics, the lifestyle of endophytes can also involve altering/inducing the gene expression of plants’ defense and metabolic pathways [66, 69, 70], and, depending on the type of interaction, members of the endosphere microbiome can induce both local and systemic alterations in the host [71]. As an example of these alterations, genome analysis of Bacillus pumilus INR7, an endophytic bacterium that promotes plant growth and induces systemic resistance against several plant patogens, revealed the presence of non-ribosomal peptide synthetase gene clusters for the production of antibacterial compounds such as surfactin, bacillibactin, and bacilysin, as well as genes for the biosynthesis of growth promoters such as indole-3-acetaldehyde and 2,3-butanediol [72].
The endosphere microbiome structure is driven by soil type, host phylogeny, and/or microbes. The soil traits that affect microbial recruitment from bulk soil are soil type [20, 53, 73], soil pH [53, 74], local edaphic conditions [75], and anthropogenic management factors, such as fertilizer and pesticide application and soil preparation [76, 77] The endosphere microbiome structure is also variously affected by plant species [78], plant life stage [77, 79], and plant health, as a result of the differences in root architecture and types of exudates [16]. Finally, the capacity of microbes to reach inner plant tissues and establish themselves there also affects the microbial composition of the endosphere. Endophytes need to have the capacity to reach the root surface, and to express genes for the invasion of plant tissue and the colonization of a niche within the plant tissue [80]. Studies have shown that the endosphere is mainly composed of bacterial phyla, such as Proteobacteria, Actinobacteria, Bacteriodetes, and Firmicutes [17, 20, 77, 81], and fungi, including Ascomycota and Basidiomycota [35, 82,83,84] (Fig. 1).
Endophytes are classified as systemic/true and transient/nonsystemic [56] or as obligate and facultative [48]. Systemic or obligate endophytes are dependent on the plant metabolism, and are disseminated among plants by vertical transmission or by vector activity [48]. In addition, systemic endophytes do not produce any visible symptoms of disease in the host at any life stage [56]. Because they live in a low-competition and low-predation environment, obligate endophytes have evolved to produce specific metabolites that support their interaction with the host [85]. In contrast, facultative or transient endophytes live inside plant tissues for at least part of their life cycle, without producing any apparent disease symptoms in the plants, but they become pathogenic when the host plant faces resource-limited conditions [86]. Transient endophytes vary both in diversity and abundance, depending on changes in the environment [83] and they face high levels of competition in the rhizosphere before entering the plant [80], therefore producing many metabolites that are involved in both their defense and in interactions with the plant [85].
The microbiomes associated with above-ground (phyllosphere), below-ground (rhizosphere), and internal (endosphere) tissues are distinct, especially considering that the endosphere is where specific metabolic capacities are required to survive. Endophytes have a significant effect on the host plant by modulating its health, growth, and development. Naveed et al. [87] observed that Enterobacter sp. strain FD17 promoted the growth and health of maize grown under natural conditions, increasing grain yield by 42% and reducing the time until flowering. Mendes et al. [62] reported that the endophytic Burkholderia spp. showed ability to control the growth of the sugarcane pathogen Fusarium moniliforme. Khan et al. [88] have shown that tomato plants inoculated with endophytic Sphingomonas sp. LK11 showed increases in shoot length, chlorophyll content, and shoot and root dry weights, indicating that the phyto-hormones produced by this strain may help in increasing crop growth. Although there are still gaps in our knowledge of endophytes, the investigation of these microbes as a bioresource for plant growth-promoting regulators and as biocontrol agents for disease and pest management represents opportunities for improving crop yield and health in a sustainable way.
Manipulation of the Plant Microbiome Toward Improved Plant Health
According to the latest United Nations projections, the world population will exceed ten billion by 2100 [89]. In order to meet the demand for food, both the land area used by agriculture and productivity must increase in the near future. In this scenario, intentional manipulation of the plant microbiome may be an alternative way to improve agriculture sustainability. This would be done by exploiting rhizosphere microorganisms with beneficial traits to, for example, make nutrients more available for plants or increase plant tolerance to biotic and abiotic stresses, consequently decreasing the dependence on chemical input in agriculture.
Manipulating the plant microbiome can be achieved simply by promoting good management of soil. Crop rotations increase the diversity of microorganisms in soil, promoting high resilience to plant pathogens [90]. Bakker et al. [91] showed that where resource changes altered the bulk soil microbial community, the effects were observed in the rhizosphere of two different cultivars of corn, suggesting that rhizosphere microbial communities are altered depending on the site history and selective events.
The stimulation of certain microorganisms or the introduction of inoculants is another strategy for plant microbiome manipulation. This approach aims to establish a beneficial community that competitively excludes plant pathogens. Reducing the time of niche exploration is crucial for enhancing microbial root-colonizing capacity [80, 92]; this can be achieved by the co-inoculation of several beneficial strains, including endophytes. The inoculation of a bacterial consortium might also promote the release of antimicrobial compounds [93] that improve the suppression of soil-borne pathogens [94].
The inoculation of microorganisms also has the potential to improve plant nutritional status. Rhizobium spp. are some of the most common microorganisms used as inoculants in legumes and their use can supply almost all of the nitrogen required by legume crops [95]. Phosphorus-solubilizing microorganisms can also be applied as inoculants, either alone or in association with rock phosphate [96]. A limitation in the use of inoculants is that the densities of the inoculated microorganisms are subject to decline over time, and the inoculants have to be able to survive under different field conditions. It is also important to consider that inoculants must be free of metabolites that are hazardous for humans, animals, and plants [97].
The plant genotype, in interaction with environmental conditions, is responsible for regulating the release of exudates in the rhizosphere soil, and this exudate release is one of the main drivers of the microbiome structure. In this context, the microbiome may be manipulated by changing the amount and quality of root exudates through plant breeding or genetic modification [98,99,100]. However, it is important to note that this strategy is limited in many ways: (a) the methods are very time-consuming and are restricted to the target species/cultivar; (b) traditionally, breeding programs do not consider the interaction among plants and microorganisms when new cultivars are being developed [101]; and (c) the quantity and quality of the exudates vary tremendously among soil types and physiological conditions of plants, making the exudates difficult to manipulate [102].
Although less commonly studied, manipulation of the microbiome of aerial plant parts can also be a strategy for improving plant growth and health. Falk et al. [103] suggested that it is possible to reduce the severity of powdery mildew infections caused by Uncinula necator on grapevines by releasing the conidia of the mycoparasite fungus Ampelomyces quisqualis. Several pesticides applied in agriculture have the potential to affect the natural occurrence of a microbial community [104, 105], while it has already been shown that the natural leaf microbiome is beneficial to the plant. Perazzolli et al. [106] showed that the naturally occurring microbiomes of grapevine leaves could reduce signs of powdery mildew on the surfaces of the leaves under controlled conditions.
Optimizing plant-microbiome interactions through microbiome manipulation has the potential to improve crop sustainability, reducing the impacts of traditional agricultural practices. Although many efforts have been made to understand the factors controlling microbiome assemblage, manipulating the microbiome is still a challenge to be addressed.
References
Lederberg J, McCray AT (2001) ‘Ome Sweet Omics’ - a genealogical treasury of words. The Scientist 15:8
Boon E, Meehan CJ, Whidden C, Wong DH, Langille MG, Beiko RG (2014) Interactions in the microbiome: communities of organisms and communities of genes. FEMS Microbiol Rev 38(1):90–118. doi:10.1111/1574-6976.12035
Melcher U, Verma R, Schneider WL (2014) Metagenomic search strategies for interactions among plants and multiple microbes. Front Plant Sci 5:268. doi:10.3389/fpls.2014.00268
Ofek M, Voronov-Goldman M, Hadar Y, Minz D (2014) Host signature effect on plant root-associated microbiomes revealed through analyses of resident vs active communities. Environ Microbiol 16(7):2157–2167. doi:10.1111/1462-2920.12228
Berg G, Smalla K (2009) Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol Ecol 68(1):1–3. doi:10.1111/j.1574-6941.2009.00654.x
Mendes R, Kruijt M, de Bruijn I, Dekkers E, van der Voort M, Schneider JH, Piceno YM, DeSantis TZ, Andersen GL, Bakker PA, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332(6033):1097–1100. doi:10.1126/science.1203980
Morales SE, Holben WE (2011) Linking bacterial identities and ecosystem processes: can ‘omic’ analyses be more than the sum of their parts? FEMS Microbiol Ecol 75(1):2–16. doi:10.1111/j.1574-6941.2010.00938.x
Mueller UG, Sachs JL (2015) Engineering microbiomes to improve plant and animal health. Trends Microbiol 23(10):606–617. doi:10.1016/j.tim.2015.07.009
Curl EA, Truelove B (1986) The rhizosphere. Springer-Verlag, New York, pp 1–8
Nannipieri P, Ascher J, Ceccherini MT, Landi L, Pietramellara G, Renella G, Valori F (2007) Microbial diversity and microbial activity in the rhizosphere. Ci Suelo 25:89–97
Aira M, Gómez-Brandón M, Lazcano C, Baath E, Domínguez J (2010) Plant genotype strongly modifies the structure and growth of maize rhizosphere microbial communities. Soil Biol Biochem 42:2276–2281. doi:10.1016/j.soilbio.2010.08.029
Mendes R, Garbeva P, Raaijmakers JM (2013) The rhizosphere microbiome: significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol Rev 37:634–663. doi:10.1111/1574-6976.12028
Philippot L, Raaijmakers JM, Lemanceau P, Puttem WH van der (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799. doi:10.1038/nrmicro3109
Sugiyama A, Ueda Y, Zushi T, Takase H, Yazaki K (2014) Changes in the bacterial community of soybean rhizospheres during growth in the field. PLoS One. doi:10.1371/journal.pone.0100709
McNear Jr DH (2013) The rhizosphere–roots, soil and everything in between. Nat Educ Knowl 4(3):1. http://www.nature.com/scitable/knowledge/library/the-rhizosphere-roots-soil-and-67500617. Accessed 11 Oct 2015
Berendsen RL, Pieterse CM, Bakker PA (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17(8):478–486. doi:10.1016/j.tplants.2012.04.001
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, Del Rio TG, Edgar RC (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488(7409):86–90. doi:10.1038/nature11237
Mendes LW, Kuramae EE, Navarrete AA, Veen JA, van Tsai SM (2014) Taxonomical and functional microbial community selection in soybean rhizosphere. ISME J. doi:10.1038/ismej.2014.17
Lebeis SL (2015) Greater than the sum of their parts: characterizing plant microbiomes at the community level. Curr Opin Plant Biol 24:82–86. doi:10.1016/j.pbi.2015.02.004
Bulgarelli D, Rott M, Schlaeppi K, van Themaat EV, Ahmadinejad N, Assenza F, Rauf P, Huettel B, Reinhardt R, Schmelzer E, Peplies J (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488(7409):91–95. doi:10.1038/nature11336
Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266. doi:10.1146/annurev.arplant.57.032905.105159
Haichar FZ, Marol C, Berge O, Rangel-Castro JI, Prosser JI, Balesdent J, Heulin T, Achouak W (2008) Plant host habitat and root exudates shape soil bacterial community structure. ISME J 2(12):1221–1230. doi:10.1038/ismej.2008.80
Chaparro JM, Bradi DV, Vivanco JM (2014) Rhizosphere microbiome assemblage is affected by plant development. ISME J 8:790–803. doi:10.1038/ismej.2013.196
Schenk PM, Carnalhais LC, Kazan K (2012) Unraveling plant-microbe interactions: can multi-species transcriptomics help? Trends Biotechnol 30:177–184. doi:10.1016/j.tibtech.2011.11.002
Lau JA, Lennon J (2011) Evolutionary ecology of plant-microbe interactions: soil microbial structure alters selection on plant traits. New Phytol 192:215–224. doi:10.1111/j.1469-8137.2011.03790.x
Lindow SE, Brandl MT (2003) Microbiology of the phyllosphere. Appl Environ Microbiol 69(4):1875–1883. doi:10.1128/AEM.69.4.1875-1883.2003
Lambais MR, Crowley DE, Cury JC, Büll RC, Rodrigues RR (2006) Bacterial diversity in tree canopies of the Atlantic forest. Science 312(5782):1917. doi: 10.1126/science.1124696
Andreote FD, Gumiere T, Durrer A (2014) Exploring interactions of plant microbiomes. Sci Agric 71(6):528–539. doi:10.1590/0103-9016-2014-0195
Vorholt JA (2012) Microbial life in the phyllosphere. Nat Rev Microbiol 10(12):828–840. doi:10.1038/nrmicro2910
Baldotto LE, Olivares FL (2008) Phylloepiphytic interaction between bacteria and different plant species in a tropical agricultural system. Can J Microbiol 54(11):918–931. doi:10.1139/W08-087
Knief C, Delmotte N, Chaffron S, Stark M, Innerebner G, Wassmann R, von Mering C, Vorholt JA (2012) Metaproteogenomic analysis of microbial communities in the phyllosphere and rhizosphere of rice. ISME J 6(7):1378–1390. doi:10.1038/ismej.2011.192
Jumpponen A, Jones KL (2009) Massively parallel 454 sequencing indicates hyperdiverse fungal communities in temperate Quercus macrocarpa phyllosphere. New Phytol 184(2):438–448. doi:10.1111/j.1469-8137.2009.02990.x
Delmotte N, Knief C, Chaffron S, Innerebner G, Roschitzki B, Schlapbach R, von Mering C, Vorholt JA (2009) Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc Natl Acad Sci 106(38):16428–16433. doi:10.1073/pnas.0905240106
Guttman DS, McHardy AC, Schulze-Lefert P (2014) Microbial genome-enabled insights into plant-microorganism interactions. Nat Rev Genet 15(12):797–813. doi:10.1038/nrg3748
Bulgarelli D, Schlaeppi K, Spaepen S, van Themaat EV, Schulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64:807–838. doi:10.1146/annurev-arplant-050312-120106
Rastogi G, Sbodio A, Tech JJ, Suslow TV, Coaker GL, Leveau JH (2012) Leaf microbiota in an agroecosystem: spatiotemporal variation in bacterial community composition on field-grown lettuce. ISME J 6(10):1812–1822. doi:10.1038/ismej.2012.32
Whipps JM, Hand P, Pink D, Bending GD (2008) Phyllosphere microbiology with special reference to diversity and plant genotype. J Appl Microbiol 105:1744–1755. doi:10.1111/j.1365-2672.2008.03906.x
Zilber-Rosenberg I, Rosenberg E (2008) Role of microorganisms in the evolution of animals and plants: the hologenome theory of evolution. FEMS Microbiol Rev 32(5):723–735. doi:10.1111/j.1574-6976.2008.00123.x
Berlec A (2012) Novel techniques and findings in the study of plant microbiota: search for plant probiotics. Plant Sci 193:96–102. doi:10.1016/j.plantsci.2012.05.010
Lindow SE (1996) Role of immigration and other processes in determining epiphytic bacterial populations. Aerial Plant Surface Microbiol 155–168. doi:10.1007/978-0-585-34164-4_10
Davey ME, O'Toole GA (2000) Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64(4):847–867. doi:10.1128/MMBR.64.4.847-867.2000
Jones K (1970) Nitrogen fixation in the phyllosphere of the Douglas fir, Pseudotsuga douglasii. Ann Bot 34(1):239–244
Freiberg E (1998) Microclimatic parameters influencing nitrogen fixation in the phyllosphere in a Costa Rican premontane rain forest. Oecologia 117(1–2):9–18. doi:10.1007/s004420050625
Kishore GK, Pande S, Podile AR (2005) Biological control of late leaf spot of peanut (Arachis hypogaea) with chitinolytic bacteria. Phytopathology 95(10):1157–1165. doi:10.1094/PHYTO-95-1157
Brandl MT, Quinones B, Lindow SE (2001) Heterogeneous transcription of an indoleacetic acid biosynthetic gene in Erwinia herbicola on plant surfaces. Proc Natl Acad Sci 98(6):3454–3459. doi:10.1073/pnas.061014498
Maignien L, DeForce EA, Chafee ME, Eren AM, Simmons SL (2014) Ecological succession and stochastic variation in the assembly of Arabidopsis thaliana phyllosphere communities. mBio 5(1):e00682–e00613. doi:10.1128/mBio.00682-13
Knief C, Ramette A, Frances L, Alonso-Blanco C, Vorholt JA (2010) Site and plant species are important determinants of the Methylobacterium community composition in the plant phyllosphere. ISME J 4(6):719–728. doi:10.1038/ismej.2010.9
Hardoim PR, van Overbeek LS, van Elsas JD (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol 16(10):463–471. doi:10.1016/j.tim.2008.07.008
Berg G, Grube M, Schloter M, Smalla K (2014) Unraveling the plant microbiome: looking back and future perspectives. Front Microbiol 5(148). doi:10.3389/fmicb.2014.00148
Coombs JT, Franco CM (2003) Visualization of an endophytic Streptomyces species in wheat seed. Appl Environ Microbiol 69(7):4260–4262. doi:10.1128/AEM.69.7.4260-4262.2003
Compant S, Reiter B, Sessitsch A, Nowak J, Clément C, Barka EA (2005) Endophytic colonization of Vitis vinifera L. by plant growth-promoting bacterium Burkholderia sp. strain PsJN. Appl Environ Microbiol 71(4):1685–1693. doi:10.1128/AEM.71.4.1685-1693.2005
Compant S, Mitter B, Colli-Mull JG, Gangl H, Sessitsch A (2011) Endophytes of grapevine flowers, berries, and seeds: identification of cultivable bacteria, comparison with other plant parts, and visualization of niches of colonization. Microb Ecol 62(1):188–197. doi:10.1007/s00248-011-9883-y
Hardoim PR, Hardoim CC, Van Overbeek LS, Van Elsas JD (2012) Dynamics of seed-borne rice endophytes on early plant growth stages. PLoS One 7(2):e30438. doi:10.1371/journal.pone.0030438
Hameed A, Yeh MW, Hsieh YT, Chung WC, Lo CT, Young LS (2015) Diversity and functional characterization of bacterial endophytes dwelling in various rice (Oryza sativa L.) tissues, and their seed-borne dissemination into rhizosphere under gnotobiotic P-stress. Plant Soil 394(1–2):177–197. doi:10.1007/s11104-015-2506-5
Truyens S, Weyens N, Cuypers A, Vangronsveld J (2015) Bacterial seed endophytes: genera, vertical transmission and interaction with plants. Environ Microbiol Rep 7(1):40–50
Mostert L, Crous PW, Petrini O (2000) Endophytic fungi associated with shoots and leaves of Vitis vinifera, with specific reference to the Phomopsis viticola complex. Sydowia 52(1):46–58
Sessitsch A, Reiter B, Berg G (2004) Endophytic bacterial communities of field-grown potato plants and their plant-growth-promoting and antagonistic abilities. Can J Microbiol 50(4):239–249. doi:10.1139/w03-118
de Melo Pereira GV, Magalhães KT, Lorenzetii ER, Souza TP, Schwan RF (2012) A multiphasic approach for the identification of endophytic bacterial in strawberry fruit and their potential for plant growth promotion. Microb Ecol 63(2):405–417. doi:10.1007/s00248-011-9919-3
Eevers N, Beckers B, de Beeck MO, White JC, Vangronsveld J, Weyens N (2015) Comparison between cultivated and total bacterial communities associated with Cucurbita pepo using cultivation-dependent techniques and 454 pyrosequencing. Syst Appl Microbiol 23
Schulz B, Boyle C (2005) The endophytic continuum. Mycol Res 109(06):661–686. doi: 10.1017/S095375620500273X
Malcolm GM, Kuldau GA, Gugino BK, Jiménez-Gasco MD (2013) Hidden host plant associations of soilborne fungal pathogens: an ecological perspective. Phytopathology 103(6):538–544. doi:10.1094/PHYTO-08-12-0192-LE
Mendes R, Pizzirani-Kleiner AA, Araujo WL, Raaijmakers JM (2007) Diversity of cultivated endophytic bacteria from sugarcane: genetic and biochemical characterization of Burkholderia cepacia complex isolates. Appl Environ Microbiol 73(22):7259–7267. doi:10.1128/AEM.01222-07
Khan AL, Hamayun M, Kang SM, Kim YH, Jung HY, Lee JH, Lee IJ (2012) Endophytic fungal association via gibberellins and indole acetic acid can improve plant growth under abiotic stress: an example of Paecilomyces formosus LHL10. BMC Microbiol 12(1):1. doi:10.1186/1471-2180-12-3
Waqas M, Khan AL, Lee IJ (2014) Bioactive chemical constituents produced by endophytes and effects on rice plant growth. J Plant Interact 9(1):478–487. doi:10.1080/17429145.2013.860562
Schardl CL, Florea S, Pan J, Nagabhyru P, Bec S, Calie PJ (2013) The epichloae: alkaloid diversity and roles in symbiosis with grasses. Curr Opin Plant Biol 16(4):480–488. doi:10.1016/j.pbi.2013.06.012
Gond SK, Bergen MS, Torres MS, White JF Jr (2015) Endophytic Bacillus spp. produce antifungal lipopeptides and induce host defence gene expression in maize. Microbiol Res 172:79–87. doi:10.1016/j.micres.2014.11.004
Carvalho TL, Balsemão-Pires E, Saraiva RM, Ferreira PC, Hemerly AS (2014) Nitrogen signalling in plant interactions with associative and endophytic diazotrophic bacteria. J Exp Bot. eru319. doi:10.1093/jxb/eru319
Yaish MW, Antony I, Glick BR (2015) Isolation and characterization of endophytic plant growth-promoting bacteria from date palm tree (Phoenix dactylifera L.) and their potential role in salinity tolerance. Antonie Van Leeuwenhoek 107(6):1519–1532. doi:10.1007/s10482-015-0445-z
Rosenblueth M, Martínez-Romero E (2006) Bacterial endophytes and their interactions with hosts. Mol Plant Microbe Interact 19(8):827–837. doi:10.1094/MPMI-19-0827
Mathys J, De Cremer K, Timmermans P, Van Kerckhove S, Lievens B, Vanhaecke M, Cammue B, De Coninck B (2012) Genome-wide characterization of ISR induced in Arabidopsis thaliana by Trichoderma hamatum T382 against Botrytis cinerea infection. Front Plant Sci 3:1–25. doi:10.3389/fpls.2012.00108
Ownley BH, Gwinn KD, Vega FE (2009) Endophytic fungal entomopathogens with activity against plant pathogens: ecology and evolution. In: The Ecology of Fungal Entomopathogens pp. 113–128. Springer Netherlands. doi: 10.1007/978-90-481-3966-8_9
Jeong H, Choi SK, Kloepper JW, Ryu CM (2014) Genome sequence of the plant endophyte Bacillus pumilus INR7, triggering induced systemic resistance in field crops. Genome Announc 2(5):e01093–e01014
Long HH, Sonntag DG, Schmidt DD, Baldwin IT (2010) The structure of the culturable root bacterial endophyte community of Nicotiana attenuata is organized by soil composition and host plant ethylene production and perception. New Phytol 185(2):554–567. doi:10.1111/j.1469-8137.2009.03079.x
Baker KL, Langenheder S, Nicol GW, Ricketts D, Killham K, Campbell CD, Prosser JI (2009) Environmental and spatial characterisation of bacterial community composition in soil to inform sampling strategies. Soil Biol Biochem 41(11):2292–2298. doi:10.1016/j.soilbio.2009.08.010
Yandigeri MS, Meena KK, Singh D, Malviya N, Singh DP, Solanki MK, Yadav AK, Arora DK (2012) Drought-tolerant endophytic actinobacteria promote growth of wheat (Triticum aestivum) under water stress conditions. Plant Growth Regul 68(3):411–420. doi:10.1007/s10725-012-9730-2
Seghers D, Wittebolle L, Top EM, Verstraete W, Siciliano SD (2004) Impact of agricultural practices on the Zea mays L. endophytic community. Appl Environ Microbiol 70(3):1475–1482. doi:10.1128/AEM.70.3.1475-1482.2004
Robinson RJ, Fraaije BA, Clark IM, Jackson RW, Hirsch PR, Mauchline TH (2015) Endophytic bacterial community composition in wheat (Triticum aestivum) is determined by plant tissue type, developmental stage and soil nutrient availability. Plant Soil 4:1–6. doi:10.1007/s11104-015-2495-4
Li CH, Shi L, Han Q, Hu HL, Zhao MW, Tang CM, Li SP (2012) Biocontrol of Verticillium wilt and colonization of cotton plants by an endophytic bacterial isolate. J Appl Microbiol 113(3):641–651. doi:10.1111/j.1365-2672.2012.05371.x
Marques JM, da Silva TF, Vollú RE, de Lacerda JR, Blank AF, Smalla K, Seldin L (2015) Bacterial endophytes of sweet potato tuberous roots affected by the plant genotype and growth stage. Appl Soil Ecol 96:273–281. doi:10.1016/j.apsoil.2015.08.020
Compant S, Clément C, Sessitsch A (2010) Plant growth-promoting bacteria in the rhizo- and endosphere of plants: their role, colonization, mechanisms involved and prospects for utilization. Soil Biol Biochem 42(5):669–678. doi:10.1016/j.soilbio.2009.11.024
Manter DK, Delgado JA, Holm DG, Stong RA (2010) Pyrosequencing reveals a highly diverse and cultivar-specific bacterial endophyte community in potato roots. Microb Ecol 60(1):157–166. doi:10.1007/s00248-010-9658-x
Toju H, Yamamoto S, Sato H, Tanabe AS, Gilbert GS, Kadowaki K (2013) Community composition of root-associated fungi in a Quercus-dominated temperate forest:“codominance” of mycorrhizal and root-endophytic fungi. Ecol Evol 3(5):1281–1293. doi:10.1002/ece3.546
Higgins KL, Arnold AE, Coley PD, Kursar TA (2014) Communities of fungal endophytes in tropical forest grasses: highly diverse host- and habitat generalists characterized by strong spatial structure. Fungal Ecol 8:1–1. doi:10.1016/j.funeco.2013.12.005
Glynou K, Ali T, Buch AK, Haghi Kia S, Ploch S, Xia X, Çelik A, Thines M, Maciá-Vicente JG (2016) The local environment determines the assembly of root endophytic fungi at a continental scale. Environ Microbiol 18(8):2418–2434. doi:10.1111/1462-2920.13112
Brader G, Compant S, Mitter B, Trognitz F, Sessitsch A (2014) Metabolic potential of endophytic bacteria. Curr Opin Biotechnol 27:30–37. doi:10.1016/j.copbio.2013.09.012
Wani ZA, Ashraf N, Mohiuddin T, Riyaz-Ul-Hassan S (2015) Plant-endophyte symbiosis, an ecological perspective. Appl Microbiol Biotechnol 99(7):2955–2965. doi:10.1007/s00253-015-6487-3
Naveed M, Mitter B, Yousaf S, Pastar M, Afzal M, Sessitsch A (2014) The endophyte Enterobacter sp. FD17: a maize growth enhancer selected based on rigorous testing of plant beneficial traits and colonization characteristics. Biol Fertil Soils 50(2):249–262. doi:10.1007/s00374-013-0854-y
Khan AL, Waqas M, Kang SM, Al-Harrasi A, Hussain J, Al-Rawahi A, Al-Khiziri S, Ullah I, Ali L, Jung HY, Lee IJ (2014) Bacterial endophyte Sphingomonas sp. LK11 produces gibberellins and IAA and promotes tomato plant growth. J Microbiol 52(8):689–695. doi:10.1007/s12275-014-4002-7
ONU (2013) Demographic components of future population growth. Tech Pap. (3)
Hwang SF, Ahmed HU, Gossen BD, Kutcher HR, Brandt SA, Strelkov SE, Chang KF, Turnbull GD (2009) Effect of crop rotation on soil pathogen population and dynamics and canola seedlings establishment. Plant Pathol J 8(3):106–112. doi:10.3923/ppj.2009.106.112
Bakker MG, Chaparro JM, Manter DK, Vivanco JM (2015) Impacts of bulk soil microbial community structure on rhizosphere microbiomes of Zea mays. Plant Soil 392(1–2):115–126. doi:10.1007/s11104-015-2446-0
Nelson LM (2004) Plant growth promoting rhizobacteria (PGPR): prospects for new inoculants. Crop Mana 3:1
de Boer W, Wagenaar A-M, Klein Gunnewiek PJ, van Veen J (2007) In vitro suppression of fungi caused by combinations of apparently non-antagonistic soil bacteria. FEMS Microbiol Ecol 59(1):177–185. doi:10.1111/j.1574-6941.2006.00197.x
Garbeva P, de Boer W (2009) Inter-specific interactions between carbon-limited soil bacteria affect behavior and gene expression. Microb Ecol 58(1):36–46. doi:10.1007/s00248-009-9502-3
Catroux G, Hartmann A, Revellin C (2001) Trends in rhizobial inoculant production and use. Plant Soil 230:21–30. doi:10.1023/A:1004777115628
Sharma SB, Sayyed RZ, Trivedi MH, Gobi T (2013) Phosphate solubilizing microbes: sustainable approach for managing phosphorus deficiency in agricultural soils. Springer Plus 2(1):587
Bashan Y, De-Bashan LE, Prabhu SR, Hernandez J-P (2014) Advances in plant growth-promoting bacterial inoculant technology: formulations and practical perspectives [1998–2013]. Plant Soil 378(1–2):1–33. doi:10.1007/s11104-013-1956-x
Ryan PR, Dessaux Y, Thomashow LS, Weller DM (2009) Rhizosphere engineering and management for sustainable agriculture. Plant Soil 321(1–2):363–383. doi:10.1007/s11104-009-0001-6
Bakker MG, Manter DK, Sheflin AM, Weir TL, Vivanco JM (2012) Harnessing the rhizosphere microbiome through plant breeding and agricultural management. Plant Soil 360(1–2):1–13. doi:10.1007/s11104-012-1361-x
Quiza L, St-Arnaud M, Yergeau E (2015) Harnessing phytomicrobiome signaling for rhizosphere microbiome engineering. Front Plant Sci 6:507. doi:10.3389/fpls.2015.00507
Wissuwa M, Mazzola M, Picard C (2008) Novel approaches in plant breeding for rhizosphere-related traits. Plant Soil 321(1–2):409–430. doi:10.1007/s11104-008-9693-2
Phillips RP, Erlitz Y, Bier R, Bernhardt ES (2008) New approach for capturing soluble root exudates in forest soils. Funct Ecol 22(6):990–999. doi:10.1111/j.1365-2435.2008.01495.x
Falk SP, Gadoury DM, Pearson RC, Seem RC (1995) Partial control of grape powdery mildew by the mycoparasite Ampelomyces quisqualis. Plant Dis 79(5):483–490
Gu L, Bai Z, Jin B, Hu Q, Wang H, Zhuang G et al (2010) Assessing the impact of fungicide enostroburin application on bacterial community in wheat phyllosphere. J Environ Sci 22(1):134–141. doi:10.1016/S1001-0742(09]60084-X
Zhang B, Bai Z, Hoefel D, Tang L, Wang X, Li B et al (2009) The impacts of cypermethrin pesticide application on the non-target microbial community of the pepper plant phyllosphere. Sci Total Environ 407(6):1915–1922. doi:10.5897/AJB2013.1
Perazzolli M, Antonielli L, Storari M, Puopolo G, Pancher M, Giovannini O et al (2014) Resilience of the natural phyllosphere microbiota of the grapevine to chemical and biological pesticides. Appl Environ Microbiol 80(12):3585–3596. doi:10.1128/AEM.00415-14
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Rossmann, M., Sarango-Flores, S.W., Chiaramonte, J.B., Kmit, M.C.P., Mendes, R. (2017). Plant Microbiome: Composition and Functions in Plant Compartments. In: Pylro, V., Roesch, L. (eds) The Brazilian Microbiome. Springer, Cham. https://doi.org/10.1007/978-3-319-59997-7_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-59997-7_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59995-3
Online ISBN: 978-3-319-59997-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)