Abstract
Recent advances in science have clarified the biosynthesis pathway and functional role of secondary metabolites. They play a major role not only for completion of the plant life cycle but also for communication with other organisms. In Brassicaceae , including radish, the most well-characterized secondary metabolite is glucosinolate . Glucosinolates are sulfur-containing metabolite and their associated degradation products have distinctive benefits for human diet and defense against pests. Plants produce approximately 200 types of different glucosinolates and those from different species show great diversity, with their contents being affected by the environment, cultivation conditions, and genetic background. The profile of glucosinolates in radish is attractive, but its biosynthesis pathway remains unclear. Here, we highlight recent progress in glucosinolate research of model plant Arabidopsis thaliana . To compare researches on glucosinolate between radish and A. thaliana, we further discuss with specificity the nature of glucosinolate in radish.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
10.1 Introduction
Glucosinolates are well-characterized plant secondary metabolites found in Brassicaceae (e.g., radish, cabbage, rapeseed, and Arabidopsis) and related plant families. An overview of the biosynthesis pathway of Arabidopsis thaliana was presented based on the observed natural variation in glucosinolate composition and genetic analysis. Furthermore, the advent of next-generation sequencing techniques allows for transcriptome analysis of Brassicaceae plants, which reveal the similarities and differences in the glucosinolates biosynthesis pathway between species.
The degradation products of glucosinolates , primarily including isothiocyanates , are pungent compounds specific to Brassicaceae plants, which not only influence the taste of vegetables but also act as an attractant or repellent to certain insects. Isothiocyanates including sulforaphane are reported to possess bioactivity beneficial to humans, for example, anticarcinogenic activity or induction of detoxification enzymes. Therefore, the consumption of glucosinolate-containing Brassicaceae vegetables is important in human diet.
Radish (Raphanus sativus L., 2n = 18) is an important Brassicaceae root vegetable, which has been cultivated since ancient times. The composition of glucosinolate shows a simple profile despite the root shape and weight showing large variations among germplasms. The most abundant glucosinolate in roots is glucoraphasatin (4-methylthio-3-butenyl glucosinolate), derived from methionine, and is essential for flavor and nutritional quality of the taproot.
This chapter presents a summary of topics related to the genes involved in glucosinolate biosynthesis in Brassicaceae, especially in radish, along with a summary of the chemical structures, biosynthesis pathways, and importance in plant breeding.
10.2 Structural Variations in Glucosinolates
Glucosinolates are sulfur- and nitrogen-comprising secondary metabolites having a common basic structure containing a β-D-thioglucose group, a sulfonated aldoxime moiety, and a variable side chain (=R) derived from amino acids (Fig. 10.1). Based on the precursor amino acid, glucosinolates can be classified into three groups: (1) aliphatic glucosinolates derived from methionine, isoleucine, leucine, or valine; (2) indolic glucosinolates derived from tryptophan; and (3) aromatic glucosinolates derived from phenylalanine or tryptophan. Thus, the side-chain structure primarily results in a diversity of structure; more than 200 structures have been identified (Clarke 2010; Fahey et al. 2001). Glucosinolates from different species show great diversity, with their contents being affected by the environment, cultivation conditions, and genetic background (Fig. 10.2). Genotype is the most important factor affecting glucosinolate profiles, for example, 39 accessions of A. thaliana possessed 34 types of glucosinolates containing aliphatic, indolic, and aromatic glucosinolates in leaves and seed, which are useful for the identification of genes encoding the enzymes involved in biosynthesis pathways (Kliebenstein et al. 2001a). Brassica vegetables contain various aliphatic glucosinolates that vary in length and modifications of the side chains (Ishida et al. 2014). These variations are explained based on the genomic structure of Brassica species.
Typical glucosinolate profile of radish (a) and broccoli (b). Preparation of desulfoglucosinolates with sulfatase digestion was conducted according to the method described by Bjerg and Sørensen (1987) for HPLC analysis. Asterisk shows internal control, sinigrin
The number of chromosomes varies in different species of Brassica. The genome relationship between three monogenomic species (comprising A, B, and C genomes) and three digenomic species is known as Triangle of U . In general, Brassica nigra (BB, 2n = 16) contains glucosinolates with three carbon side chains derived from a single elongation reaction; B. oleracea (CC, 2n = 18) contains glucosinolates with three or four carbon side chains; and B. rapa (AA, 2n = 20) contains glucosinolates with either four or five carbon side chains. The glucosinolate composition of three amphidiploid Brassica species reflects two elementary species, for example, B. napus (AACC, 2n = 38), an amphidiploid having the B. rapa and B. oleracea genomes, contains three, four, or five carbon side chains.
Glucoraphasatin (4-methylthio-3-butenyl glucosinolate) , also known as dehydroerucin, glucodehydroerucin, or 4MTB-GSL, is an aliphatic glucosinolate predominantly found in radish roots, and comprises >90% of the total glucosinolates in Japanese white radish (Ishida et al. 2012). Although the presence of glucoraphasatin was reported in five genera of the Brassicaceae family, such as Brassica (Newkirk and Classen 2002; Thacker and Newkirk 2005), Bunias (Bennett et al. 2006), Matthiola (Bennett et al. 2004), Raphanus (Carlson et al. 1985; Ishii et al. 1989), and Rapistrum (Curto et al. 2005), a species with >90% of the total glucosinolate content is restricted to R. sativus. In the 632 radish cultivars, glucoraphasatin content ranges from 43.8 to 475.5 µmol g−1 dry weight at the cotyledon stage (Ishida et al. 2015). Most cultivars contain two other related glucosinolates , glucoraphenin (4-methylsulfinyl-3-butenyl glucosinolate) and glucoerucin (4-methylthiobutyl glucosinolate) , in smaller quantities. Glucoraphasatin is detected in various radish organs such as the seeds, seedlings, mature leaves, stems, and roots (Beevi et al. 2009; Ciska et al. 2008; Griffiths et al. 2001; Yamada et al. 2003).
10.3 Degradation of Glucosinolates
When plant tissue is damaged, glucosinolates are rapidly hydrolyzed by endogenous thioglucosidases called myrosinases (Fig. 10.3) (Rask et al. 2000). Myrosinases are accumulated in myrosin cells, which specifically form along the leaf veins (Andreasson et al. 2001; Rask et al. 2000; Ueda et al. 2006). In contrast, glucosinolates are stored in S-cells that are physically separated from myrosin cells (Koroleva et al. 2000, 2010). Once the tissue is mechanically damaged, glucosinolates are hydrolyzed intensively by myrosinases . As a result of myrosinase activity, glucose and sulfate are released and some toxic/bioactive products, such as isothiocyanates , thiocyanates, nitriles, and epithionitriles, are formed depending on the chemical structure of the side chain, reaction conditions such as pH, presence of ferrous ions, and the modifier proteins (Kissen et al. 2009; Wittstock and Halkier 2002). The major degradation products, i.e., isothiocyanates, impart the characteristic taste and flavor to Brassicaceae vegetables. Several in vitro and in vivo studies have shown that sulforaphane (4-methylsulfinylbutane isothiocyanate), a degradation product of glucoraphanin abundant in broccoli sprouts, is associated with potential health-promoting activity (Juge et al. 2007). In radish, isothiocyanates derived from glucoraphasatin are pungent compounds that react with water and produce a yellow pigment and methanethiol , which are involved in the color and smell of pickles made from Asian big radish (Ozawa et al. 1990a, b).
Scheme of glucosinolate hydrolysis and structure of possible degradation products. Glucosinolates are hydrolyzed by myrosinase when tissues are damaged. At neutral pH (pH 5–7), the unstable aglucones change to isothiocyanates. The interaction of epithiospecifier protein (ESP) with myrosinase alters the reaction toward the production of nitriles depending on the glucosinolate structure and pH. Some glucosinolates can be hydrolyzed to thiocyanates. R, variable side chain
10.4 Biosynthesis Pathway
Genetic and biochemical analyses show that the biosynthesis pathway involves the natural variation in glucosinolate profiles among A. thaliana accessions. Here, we focus on the biosynthesis pathway of aliphatic glucosinolate , which is abundant in A. thaliana, and outline the function of each enzyme involved in the pathway (Fig. 10.4).
Biosynthesis pathway of aliphatic glucosinolate. a Side-chain elongation steps. b Biosynthesis of core glucosinolate structure. A. thaliana gene identifiers are as follows: BCAT4 (At3g19710); MAM1 (At5g23010), MAM3 (At5g23020); IPMI LSU1 (At4g13430), IPMI LSU2 (At2g43100), IPMI LSU3 (At3g58990); IPMDH1 (At5g14200), IPMDH3 (At1g31180); BCAT3 (At3g49680); CYP79F1 (At1g16410), CYP79F2 (At1g16400); CYP83A1 (At4g13770); GSTF11 (At3g03190), GSTU20 (At1g78370), GGP1 (At4g30530); SUR1 (At2g20610); UGT74C1 (At2g31790); SOT17 (At1g18590), SOT18 (At1g74090)
The biosynthesis of aliphatic glucosinolates can be divided into three independent steps. First, the carbon chain of methionine is elongated by a chain elongation cycle in the chloroplast. Second, the core structure, comprising a β-D-thioglucose group and a sulfonated aldoxime moiety, is constructed. Third, the side chain is modified. Polymorphism or the absence/presence of respective genes determines the glucosinolate profile.
10.4.1 Chain Elongation
The side-chain elongation step in A. thaliana and Eruca sativa was confirmed by a feeding experiment using radio-labeled acetate (Graser et al. 2001, 2000). The side-chain elongation is initiated by transamination of methionine to form the corresponding 2-oxo acid. An A. thaliana bcat4 mutant shows a decrease in aliphatic glucosinolate levels and an increase in free methionine levels, suggesting that BCAT4 catalyzes this step (Schuster et al. 2006). The newly formed 2-oxo acid is transported to the chloroplast via plastid-localized bile acid transporter 5 (BAT5) and BAT5/bile acid: sodium symporter family protein 5 (BASS5) (Gigolashvili et al. 2009; Sawada et al. 2009b). The transported 2-oxo acid is then involved in a cycle comprising three transformations: condensation with acetyl-CoA catalyzed by a methylthioalkylmalate (MAM) synthase, isomerization by an isopropylmalate isomerase (IPMI), and oxidative decarboxylation by an isopropylmalate dehydrogenase (IPMDH) (Field et al. 2004; Knill et al. 2009; Kroymann et al. 2001; Sawada et al. 2009a; Textor et al. 2007). The final step of the elongation cycle is the transamination by BCAT3 (Knill et al. 2008). The product of these reactions is a 2-oxo acid that is elongated by a single methylene (–CH2–) moiety at each cycle.
10.4.2 Core Structure Formation
The aliphatic glucosinolate core structure is formed in five biochemical steps catalyzed by 11 enzymes (Grubb and Abel 2006). The first step is oxidation; precursor amino acids, which have elongated side chains, are converted to aldoximes by cytochrome P450 of the CYP79 family. CYP79F1 oxidizes all elongated methionines (1–6 carbons), while CYP79F2 only oxidizes long-chained methionines (5 and 6 carbons) (Chen et al. 2003; Hansen et al. 2001). The aldoximes are oxidized by CYP83A1 of the CYP79 family to an aci-nitro compound (Bak and Feyereisen 2001; Hemm et al. 2003). The activated forms are transformed to thiohydroximates via glutathione conjugation and a C–S lyase (SUR1) reaction (Mikkelsen et al. 2004). Thiohydroximates are, in turn, S-glucosylated by glucosyltransferases of the UGT74 family to form desulfoglucosinolates. UGT74C1 was shown to metabolize the methionine-derived thiohydroximates (Grubb et al. 2014). Finally, the thiohydroximates are converted to the glucosinolate structure by S-glucosyltransferases of the sulfotransferases (SOTs) (Piotrowski et al. 2004). After the glucosinolate structure is formed, the side chains are modified by oxygenation, hydroxylation, alkenylation, benzoylation, and methoxylation.
10.4.3 Side-Chain Modification
S-oxygenation is the first modification of aliphatic glucosinolates (Fig. 10.5). S-oxygenation of aliphatic glucosinolates is a common modification undergone by flavin-containing monooxygenase (FMOGS-OXs). Phylogenetic analysis of plant FMOs revealed the presence of the Brassicaceae-specific clade, including FMOGS-OXs, which are involved in the S-oxygenation of glucosinolates (Hansen et al. 2007; Li et al. 2008). Five FMOGS-OXs are encoded by the A. thaliana genome and are closely linked at chromosome 1. Using a recombinant enzyme assay, it was revealed that FMOGS-OX1–FMOGS-OX4 are able to catalyze the S-oxygenation independent of the side-chain length (Hansen et al. 2007; Li et al. 2008). In contrast, FMOGS-OX5 is specific to long-chain (8C) glucosinolate (Li et al. 2008).
The next step of the side-chain modification is performed by three 2-oxoglutarate-dependent dioxygenases, AOP2 and AOP3, and GS-OH. Kliebenstein et al. (2001b) demonstrated the absence of methylsulfinylalkyl side-chain modification in null aop2 accessions among 39 A. thaliana accessions. The over-expression line, which is a transgenic Columbia (Col-0) line having a functional AOP2 transgene with the non-functional AOP2 and AOP3, accumulated a methylsulfinylalkyl form (Neal et al. 2010). Interestingly, the introduction of a functional AOP2 into Col-0, which has a null AOP2, showed that the total content of aliphatic glucosinolate was increased (Wentzell et al. 2007). AOP3 catalyzes the formation of hydroxyalkyl glucosinolates from methylsulfinylalkyl glucosinolates. None of the 21 A. thaliana accessions express both AOPs (Kliebenstein et al. 2001b). The GS-OH locus is responsible for the production of 2-hydroxybut-3-butenyl glucosinolate (Mithen et al. 1995; Parkin et al. 1994). The 2-oxoglutarate-dependent dioxygenase, required for 2-hydroxybut-3-butenyl glucosinolate formation, was identified using positional cloning of GS-OH (Hansen et al. 2008).
Another modification is the benzoylation. The high accumulation of benzoylated glucosinolates in A. thaliana seeds is known (Kliebenstein et al. 2007). In silico analysis, it was revealed that the three serine carboxypeptidase-like (SCPL) genes are co-expressed with AOP3 and BZO1, which synthesize the benzoate precursor cinnamoyl CoA in the seeds. A benzoate feeding experiment suggested that SCPL17 is involved in the final step of benzoylated glucosinolate biosynthesis (Lee et al. 2012).
10.5 Transcription Factors
The family of avian myeloblastosis virus (MYB) domain transcription factors includes the conserved MYB DNA-binding domain, which can control the transcriptional state of target genes in various plant secondary metabolism pathways (Jin and Martin 1999). In A. thaliana, MYB28, 29, and 76 orchestrate the transcription of aliphatic glucosinolates biosynthesis genes, while MYB34, 51, and 122 regulate the indolic glucosinolate pathway (Frerigmann and Gigolashvili 2014; Gigolashvili et al. 2008; Hirai et al. 2007; Malitsky et al. 2008; Sonderby et al. 2010). The functional role of MYB28 and MYB29 was characterized using the omics approach. Co-expression analysis demonstrated the relationship between metabolic pathway genes and transcription factor genes (Hirai et al. 2007). Double mutants in MYB28 and MYB29 showed complete absence of short- and long-chain aliphatic glucosinolates , while some indolic glucosinolates increased to a small extent. Interestingly, the biosynthesis of long-chain aliphatic glucosinolates was blocked by the absence of MYB28; however, short-chain aliphatic glucosinolates were reduced by approximately 50% in both the myb28 and myb29 single mutants (Beekwilder et al. 2008; Sonderby et al. 2007). In the case of indolic glucosinolate regulation, MYB34, MYB51, and MYB122 reciprocally regulate their expression and perform a central role in the gene expression of indolic glucosinolates pathway under the influence of various signals (Frerigmann and Gigolashvili 2014).
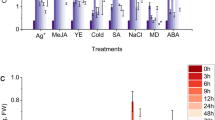
Because glucosinolate metabolism has evolved as a result of plant–environment interaction, its biosynthesis is regulated through various signals, including touch, injury, plant hormones, and insect infestation (Textor and Gershenzon 2008). Although glucosinolates perform the role of a constitutive defense compound during plant life cycle, insect herbivore feeding increases their levels (Wittstock and Halkier 2002); for example, mechanical wounding induces the expression of MYB29 and MYB76 within 1 min. Aliphatic glucosinolates are accumulated by the activation of MYB28, MYB29, and MYB76, with each of these genes upregulating the expression of the others. By contrast, these MYBs downregulate the expression of genes involved in the control of indolic glucosinolate biosynthesis (Gigolashvili et al. 2008). Therefore, external stimuli are believed to be amplified by the expression control of transcription factors.
10.6 Transcriptome Analysis in Radish
Next-generation sequencing-based RNA sequencing of the transcriptome enables comprehensive gene expression analysis for non-model plants such as radish. Two independent studies revealed the expression profile of genes involved in the glucosinolate biosynthesis of radish (Mitsui et al. 2015; Wang et al. 2013). Mitsui et al. (2015) performed a comparison between the transcriptional profiles of glucosinolate biosynthesis in four tissues (root, root tip, cortex, and xylem) at six developmental stages. Interestingly, although the radish transcripts pool contained the majority of genes required for glucosinolate biosynthesis, nine genes (i.e., RsMAM3, RsIPMI-SSU3, RsIPMDH3, RsCYP79F2, RsCYP81F1, RsFMO GS-OX3 , RsFMO GS-OX4 , RsAOP2, and RsAOP3) were not identified. They suggested that the absence of these genes in the genome might explain the specific glucosinolate profile of radish. Wang et al. (2013) constructed a de novo assembly data set at three developmental stages (seedling, taproot thickening, and mature stages). Assembled unigenes included almost ortholog genes for A. thaliana glucosinolate biosynthesis genes and their transcription factors, without APOs and GS-OH (Wang et al. 2013). Accumulation of glucoraphasatin is observed in the root tips and outer zones, including the peels (Ishii 1991). Gene expression profiles reflect the distribution of glucoraphasatin. The results indicate that the numerous biosynthesis-related genes were strongly expressed in the root tip and cortex (Mitsui et al. 2015). Moreover, the intensity of gene expression appears to be more likely at a younger stage. These observations indicate that the spacial and developmental controls of biosynthesis genes are involved in glucoraphasatin accumulation. Genetic analysis revealed that three loci containing RsMAM3, RsIPMDH1, and RsBCAT4 responsible for controlling the glucosinolate content in taproots were identified by the quantitative trait loci analysis using biparental F2 populations. Furthermore, a clear difference in gene expression levels of these candidate genes was observed between parental lines at the taproot stage (Zou et al. 2013). From the transcriptome data, it is evident that there are no ortholog genes for 2-oxoglutarate-dependent dioxygenases, AOP2 and AOP3, which catalyze the hydroxylation of side chains, in the radish genome (Mitsui et al. 2015). Thus, the end product of glucosinolate biosynthesis in radish might be glucoraphenin alone. These observations, such as the absence of AOP2 and AOP3, and the presence of desaturase (see below section), could explain the dominant accumulation of 4-carbon side-chain glucosinolates in radish.
10.7 A Radish Mutant with an Altered Glucosinolate Profile
Glucoraphasatin is the predominant glucosinolate found in radish taproots (Carlson et al. 1985) and accounts for >90% of the total GSLs in Japanese cultivars (Ishida et al. 2012). Using a comprehensive analysis of glucosinolate profiles of global radish accessions and cultivars, a spontaneous mutant having significantly low glucoraphasatin was identified, and a completely glucoraphasatin-free line was generated by the self-pollination of the genetic resource (Fig. 10.6). Genetic analysis revealed that the glucoraphasatin-free trait was controlled by a single recessive locus, which is located at the edge of the linkage group 1 (Ishida et al. 2015).
a Typical HPLC profile obtained for the wild-type radish (black line) and glucoraphasatin-free mutant (red line). Peak 1, glucoerucin; peak 2, glucoraphasatin. Asterisk shows internal control, sinigrin. b A putative pathway of glucosinolate biosynthesis in radish. GRS1 encoding an enzyme catalyzing the desaturation of glucoerucin side chain is deficient in glucoraphasatin-free mutant
The glucoraphasatin-free line contains glucoerucin, which accounts for >85% of the total glucosinolates , and 2% of glucoraphanin instead of glucoraphasatin and glucoraphenin (Ishida et al. 2015). Glucoraphasatin differs from glucoerucin only by the presence of a double bond in the side chain (Fig. 10.1b). These genetic data and glucosinolate profile show that the single gene, which encodes the enzyme involved in the desaturation of the side chain of glucoerucin, has lesions in the mutant line. In other words, radish might have obtained a gene that encodes desaturase, which is not present in other Brassicaceae plants. Similarly, a mutant which predominantly generates 4-methythiobutyl isothiocyanate derived from glucoerucin was identified in Japanese landrace ‘Shibori-daikon’ (Hori et al. 1999). Recently, we have identified a gene encoding a desaturase of side chain, GLUCORAPHASATIN SYNTHASE 1 (GRS1) , by genetic mapping using a mutant that genetically lacks glucoraphasatin (Fig. 10.6b) (Kakizaki et al. 2017). GRS1 is a member of the 2-oxoglutarate and Fe(II)-dependent dioxygenase superfamily and transgenic A. thaliana which overexpressed GRS1 cDNA, accumulated glucoraphasatin in the leaves. These data present interesting observation that partly explains the specific GSL profile for radish.
10.8 Conclusion
Radish is an important vegetable that is consumed in a variety of ways: cooked, raw, pickled, brined, or dried. Glucosinolates in the taproots significantly affect the flavor and quality of radish. The glucosinolate profile of radish is found to be amazingly simple when compared with the other Brassicaceae , because glucoraphasatin accounts for more than 90% of the total glucosinolates in Japanese cultivars (Ishida et al. 2012). The breakdown product of glucoraphasatin by myrosinase generates yellow pigment and a methanethiol component that is responsible for the color and flavor of pickles (Takahashi et al. 2015). In our preliminary data, pickles made from the mutant having lesion in GRS1 was it did not generate appreciable methanethiol . Utilization of non-functional GRS1 gene should be useful in metabolite engineering for breeding of high-value vegetables.
References
Andreasson E, Jorgensen LB, Hoglund AS, Rask L, Meijer J (2001) Different myrosinase and idioblast distribution in Arabidopsis and Brassica napus. Plant Physiol 127:1750–1763
Bak S, Feyereisen R (2001) The involvement of two P450 enzymes, CYP83B1 and CYP83A1, in auxin homeostasis and glucosinolate biosynthesis. Plant Physiol 127:108–118
Beekwilder J, van Leeuwen W, van Dam NM, Bertossi M, Grandi V, Mizzi L, Soloviev M, Szabados L, Molthoff JW, Schipper B, Verbocht H, de Vos RCH, Morandini P, Aarts MGM, Bovy A (2008) The impact of the absence of aliphatic glucosinolates on insect herbivory in Arabidopsis. PLoS ONE 3:e2068
Beevi SS, Mangamoori LN, Dhand V, Ramakrishna DS (2009) Isothiocyanate profile and selective antibacterial activity of root, stem, and leaf extracts derived from Raphanus sativus L. Foodborne Pathog Dis 6:129–136
Bennett RN, Mellon FA, Kroon PA (2004) Screening crucifer seeds as sources of specific intact glucosinolates using ion-pair high-performance liquid chromatography negative ion electrospray mass spectrometry. J Agric Food Chem 52:428–438
Bennett RN, Rosa EA, Mellon FA, Kroon PA (2006) Ontogenic profiling of glucosinolates, flavonoids, and other secondary metabolites in Eruca sativa (salad rocket), Diplotaxis erucoides (wall rocket), Diplotaxis tenuifolia (wild rocket), and Bunias orientalis (Turkish rocket). J Agric Food Chem 54:4005–4015
Bjerg B, Sørensen H (1987) Quantitative analysis of glucosinolates and HPLC of intact glucosinolates. In: Wathelet J-P (ed.) Glucosinolates in Rapeseeds: analytical Aspects, Martinus Nijhoff Publishers, Dordrecht, Netherlands, pp. 125–150
Carlson DG, Daxenbichler ME, Vanetten CH, Hill CB, Williams PH (1985) Glucosinolates in radish cultivars. J Am Soc Hortic Sci 110:634–638
Chen SX, Glawischnig E, Jorgensen K, Naur P, Jorgensen B, Olsen CE, Hansen CH, Rasmussen H, Pickett JA, Halkier BA (2003) CYP79F1 and CYP79F2 have distinct functions in the biosynthesis of aliphatic glucosinolates in Arabidopsis. Plant J 33:923–937
Ciska E, Honke J, Kozlowska H (2008) Effect of light conditions on the contents of glucosinolates in germinating seeds of white mustard, red radish, white radish, and rapeseed. J Agric Food Chem 56:9087–9093
Clarke DB (2010) Glucosinolates, structures and analysis in food. Anal Methods 2:310–325
Curto G, Dallavalle E, Lazzeri L (2005) Life cycle duration of Meloidogyne incognita and host status of Brassicaceae and Capparaceae selected for glucosinate content. Nematology 7:203–212
Fahey JW, Zalcmann AT, Talalay P (2001) The chemical diversity and distribution of glucosinolates and isothiocyanates among plants. Phytochemistry 56:5–51
Field B, Cardon G, Traka M, Botterman J, Vancanneyt G, Mithen R (2004) Glucosinolate and amino acid biosynthesis in Arabidopsis. Plant Physiol 135:828–839
Frerigmann H, Gigolashvili T (2014) MYB34, MYB51, and MYB122 distinctly regulate indolic glucosinolate biosynthesis in Arabidopsis thaliana. Mol Plant 7:814–828
Gigolashvili T, Engqvist M, Yatusevich R, Muller C, Flugge UI (2008) HAG2/MYB76 and HAG3/MYB29 exert a specific and coordinated control on the regulation of aliphatic glucosinolate biosynthesis in Arabidopsis thaliana. New Phytol 177:627–642
Gigolashvili T, Yatusevich R, Rollwitz I, Humphry M, Gershenzon J, Flugge UI (2009) The plastidic bile acid transporter 5 is required for the biosynthesis of methionine-derived glucosinolates in Arabidopsis thaliana. Plant Cell 21:1813–1829
Graser G, Schneider B, Oldham NJ, Gershenzon J (2000) The methionine chain elongation pathway in the biosynthesis of glucosinolates in Eruca sativa (Brassicaceae). Arch Biochem Biophys 378:411–419
Graser G, Oldham NJ, Brown PD, Temp U, Gershenzon J (2001) The biosynthesis of benzoic acid glucosinolate esters in Arabidopsis thaliana. Phytochemistry 57:23–32
Griffiths DW, Deighton N, Birch AN, Patrian B, Baur R, Stadler E (2001) Identification of glucosinolates on the leaf surface of plants from the Cruciferae and other closely related species. Phytochemistry 57:693–700
Grubb CD, Abel S (2006) Glucosinolate metabolism and its control. Trends Plant Sci 11:89–100
Grubb CD, Zipp BJ, Kopycki J, Schubert M, Quint M, Lim EK, Bowles DJ, Pedras MS, Abel S (2014) Comparative analysis of Arabidopsis UGT74 glucosyltransferases reveals a special role of UGT74C1 in glucosinolate biosynthesis. Plant J 79:92–105
Hansen CH, Wittstock U, Olsen CE, Hick AJ, Pickett JA, Halkier BA (2001) Cytochrome P450 CYP79F1 from Arabidopsis catalyzes the conversion of dihomomethionine and trihomomethionine to the corresponding aldoximes in the biosynthesis of aliphatic glucosinolates. J Biol Chem 276:11078–11085
Hansen BG, Kliebenstein DJ, Halkier BA (2007) Identification of a flavin-monooxygenase as the S-oxygenating enzyme in aliphatic glucosinolate biosynthesis in Arabidopsis. Plant J 50:902–910
Hansen BG, Kerwin RE, Ober JA, Lambrix VM, Mitchell-Olds T, Gershenzon J, Halkier BA, Kliebenstein DJ (2008) A novel 2-oxoacid-dependent dioxygenase involved in the formation of the goiterogenic 2-hydroxybut-3-enyl glucosinolate and generalist insect resistance in Arabidopsis. Plant Physiol 148:2096–2108
Hemm MR, Ruegger MO, Chapple C (2003) The Arabidopsis ref2 mutant is defective in the gene encoding CYP83A1 and shows both phenylpropanoid and glucosinolate phenotypes. Plant Cell 15:179–194
Hirai MY, Sugiyama K, Sawada Y, Tohge T, Obayashi T, Suzuki A, Araki R, Sakurai N, Suzuki H, Aoki K, Goda H, Nishizawa OI, Shibata D, Saito K (2007) Omics-based identification of Arabidopsis Myb transcription factors regulating aliphatic glucosinolate biosynthesis. Proc Natl Acad Sci USA 104:6478–6483
Hori K, Ohisa N, Suzuki M, Sato T, Kikawa A (1999) Chemical structure and quantitative determination of mustard oil in Shibori Daikon (Studies on special food resources in Akita part I). J Jpn Soc Food Sci 46:528–534 (In Japanese)
Ishida M, Nagata M, Ohara T, Kakizaki T, Hatakeyama K, Nishio T (2012) Small variation of glucosinolate composition in Japanese cultivars of radish (Raphanus sativus L.) requires simple quantitative analysis for breeding of glucosinolate component. Breed Sci 62:63–70
Ishida M, Hara M, Fukino N, Kakizaki T, Morimitsu Y (2014) Glucosinolate metabolism, functionality and breeding for the improvement of Brassicaceae vegetables. Breed Sci 64:48–59
Ishida M, Kakizaki T, Morimitsu Y, Ohara T, Hatakeyama K, Yoshiaki H, Kohori J, Nishio T (2015) Novel glucosinolate composition lacking 4-methylthio-3-butenyl glucosinolate in Japanese white radish (Raphanus sativus L.). Theor Appl Genet 128:2037–2046
Ishii G (1991) Glucosinolate in Japanese Radish, Raphanus-Sativus L. JARQ 24:273–279
Ishii G, Saijo R, Nagata M (1989) The difference of glucosinolate content in different cultivar of daikon roots (Raphanus sativus L.). J Jpn Soc Food Sci 36:739–742 (In Japanese)
Jin H, Martin C (1999) Multifunctionality and diversity within the plant MYB-gene family. Plant Mol Biol 41:577–585
Juge N, Mithen RF, Traka M (2007) Molecular basis for chemoprevention by sulforaphane: a comprehensive review. Cell Mol Life Sci 64:1105–1127
Kakizaki T, Kitashiba H, Zou Z, Li F, Fukino N, Ohara T, Nishio T, Ishida M (2017) A 2-oxoglutarate-dependent dioxygenase mediates the biosynthesis of glucoraphasatin in radish. Plant Physiol 173:1583–1593
Kissen R, Pope TW, Grant M, Pickett JA, Rossiter JT, Powell G (2009) Modifying the alkylglucosinolate profile in Arabidopsis thaliana alters the tritrophic interaction with the herbivore Brevicoryne brassicae and parasitoid Diaeretiella rapae. J Chem Ecol 35:958–969
Kliebenstein DJ, Kroymann J, Brown P, Figuth A, Pedersen D, Gershenzon J, Mitchell-Olds T (2001a) Genetic control of natural variation in Arabidopsis glucosinolate accumulation. Plant Physiol 126:811–825
Kliebenstein DJ, Lambrix VM, Reichelt M, Gershenzon J, Mitchell-Olds T (2001b) Gene duplication in the diversification of secondary metabolism: tandem 2-oxoglutarate-dependent dioxygenases control glucosinolate biosynthesis in Arabidopsis. Plant Cell 13:681–693
Kliebenstein DJ, D’Auria JC, Behere AS, Kim JH, Gunderson KL, Breen JN, Lee G, Gershenzon J, Last RL, Jander G (2007) Characterization of seed-specific benzoyloxyglucosinolate mutations in Arabidopsis thaliana. Plant J 51:1062–1076
Knill T, Schuster J, Reichelt M, Gershenzon J, Binder S (2008) Arabidopsis branched-chain aminotransferase 3 functions in both amino acid and glucosinolate biosynthesis. Plant Physiol 146:1028–1039
Knill T, Reichelt M, Paetz C, Gershenzon J, Binder S (2009) Arabidopsis thaliana encodes a bacterial-type heterodimeric isopropylmalate isomerase involved in both Leu biosynthesis and the Met chain elongation pathway of glucosinolate formation. Plant Mol Biol 71:227–239
Koroleva OA, Davies A, Deeken R, Thorpe MR, Tomos AD, Hedrich R (2000) Identification of a new glucosinolate-rich cell type in Arabidopsis flower stalk. Plant Physiol 124:599–608
Koroleva OA, Gibson TM, Cramer R, Stain C (2010) Glucosinolate-accumulating S-cells in Arabidopsis leaves and flower stalks undergo programmed cell death at early stages of differentiation. Plant J 64:456–469
Kroymann J, Textor S, Tokuhisa JG, Falk KL, Bartram S, Gershenzon J, Mitchell-Olds T (2001) A gene controlling variation in Arabidopsis glucosinolate composition is part of the methionine chain elongation pathway. Plant Physiol 127:1077–1088
Lee S, Kaminaga Y, Cooper B, Pichersky E, Dudareva N, Chapple C (2012) Benzoylation and sinapoylation of glucosinolate R-groups in Arabidopsis. Plant J 72:411–422
Li J, Hansen BG, Ober JA, Kliebenstein DJ, Halkier BA (2008) Subclade of flavin-monooxygenases involved in aliphatic glucosinolate biosynthesis. Plant Physiol 148:1721–1733
Malitsky S, Blum E, Less H, Venger I, Elbaz M, Morin S, Eshed Y, Aharoni A (2008) The transcript and metabolite networks affected by the two clades of Arabidopsis glucosinolate biosynthesis regulators. Plant Physiol 148:2021–2049
Mikkelsen MD, Naur P, Halkier BA (2004) Arabidopsis mutants in the C–S lyase of glucosinolate biosynthesis establish a critical role for indole-3-acetaldoxime in auxin homeostasis. Plant J 37:770–777
Mithen R, Clarke J, Lister C, Dean C (1995) Genetics of aliphatic glucosinolates. III. Side-chain structure of aliphatic glucosinolates in Arabidopsis thaliana. Heredity 74:210–215
Mitsui Y, Shimomura M, Komatsu K, Namiki N, Shibata-Hatta M, Imai M, Katayose Y, Mukai Y, Kanamori H, Kurita K, Kagami T, Wakatsuki A, Ohyanagi H, Ikawa H, Minaka N, Nakagawa K, Shiwa Y, Sasaki T (2015) The radish genome and comprehensive gene expression profile of tuberous root formation and development. Sci Rep 5:10835
Neal CS, Fredericks DP, Griffiths CA, Neale AD (2010) The characterisation of AOP2: a gene associated with the biosynthesis of aliphatic alkenyl glucosinolates in Arabidopsis thaliana. BMC Plant Biol 10:170
Newkirk RW, Classen HL (2002) The effects of toasting canola meal on body weight, feed conversion efficiency, and mortality in broiler chickens. Poult Sci 81:815–825
Ozawa Y, Kawakishi S, Uda Y, Maeda Y (1990a) Isolation and identification of a novel beta-carboline derivative in salted radish roots, Raphanus sativus L. Agric Biol Chem 54:1241–1245
Ozawa Y, Uda Y, Kawakishi S (1990b) Generation of a beta-carboline derivative, the yellowish precursor of processed radish roots, from 4-methylthio-3-butenyl isothiocyanate and L-tryptophan. Agric Biol Chem 54:1849–1851
Parkin I, Magrath R, Keith D, Sharpe A, Mithen R, Lydiate D (1994) Genetics of aliphatic glucosinolates. II. Hydroxylation of alkenyl glucosinolates in Brassica napus. Heredity 72:594–598
Piotrowski M, Schemenewitz A, Lopukhina A, Muller A, Janowitz T, Weiler EW, Oecking C (2004) Desulfoglucosinolate sulfotransferases from Arabidopsis thaliana catalyze the final step in the biosynthesis of the glucosinolate core structure. J Biol Chem 279:50717–50725
Rask L, Andreasson E, Ekbom B, Eriksson S, Pontoppidan B, Meijer J (2000) Myrosinase: gene family evolution and herbivore defense in Brassicaceae. Plant Mol Biol 42:93–113
Sawada Y, Kuwahara A, Nagano M, Narisawa T, Sakata A, Saito K, Hirai MY (2009a) Omics-based approaches to methionine side chain elongation in Arabidopsis: characterization of the genes encoding methylthioalkylmalate isomerase and methylthioalkylmalate dehydrogenase. Plant Cell Physiol 50:1181–1190
Sawada Y, Toyooka K, Kuwahara A, Sakata A, Nagano M, Saito K, Hirai MY (2009b) Arabidopsis bile acid:sodium symporter family protein 5 is involved in methionine-derived glucosinolate biosynthesis. Plant Cell Physiol 50:1579–1586
Schuster J, Knill T, Reichelt M, Gershenzon J, Binder S (2006) branched-chain aminotransferase4 is part of the chain elongation pathway in the biosynthesis of methionine-derived glucosinolates in Arabidopsis. Plant Cell 18:2664–2679
Sonderby IE, Hansen BG, Bjarnholt N, Ticconi C, Halkier BA, Kliebenstein DJ (2007) A systems biology approach identifies a R2R3 MYB gene subfamily with distinct and overlapping functions in regulation of aliphatic glucosinolates. PLoS ONE 2:e1322
Sonderby IE, Burow M, Rowe HC, Kliebenstein DJ, Halkier BA (2010) A complex interplay of three R2R3 MYB transcription factors determines the profile of aliphatic glucosinolates in Arabidopsis. Plant Physiol 153:348–363
Takahashi A, Yamada T, Uchiyama Y, Hayashi S, Kumakura K, Takahashi H, Kimura N, Matsuoka H (2015) Generation of the antioxidant yellow pigment derived from 4-methylthio-3-butenyl isothiocyanate in salted radish roots (takuan-zuke). Biosci Biotechnol Biochem 79:1512–1517
Textor S, Gershenzon J (2008) Herbivore induction of the glucosinolate-myrosinase defense system: major trends, biochemical bases and ecological significance. Phytochem Rev 8:149–170
Textor S, de Kraker JW, Hause B, Gershenzon J, Tokuhisa JG (2007) MAM3 catalyzes the formation of all aliphatic glucosinolate chain lengths in Arabidopsis. Plant Physiol 144:60–71
Thacker PA, Newkirk RW (2005) Performance of growing-finishing pigs fed barley-based diets containing toasted or non-toasted canola meal. Can J Anim Sci 85:53–59
Ueda H, Nishiyama C, Shimada T, Koumoto Y, Hayashi Y, Kondo M, Takahashi T, Ohtomo I, Nishimura M, Hara-Nishimura I (2006) AtVAM3 is required for normal specification of idioblasts, myrosin cells. Plant Cell Physiol 47:164–175
Wang Y, Pan Y, Liu Z, Zhu X, Zhai L, Xu L, Yu R, Gong Y, Liu L (2013) De novo transcriptome sequencing of radish (Raphanus sativus L.) and analysis of major genes involved in glucosinolate metabolism. BMC Genom 14:836
Wentzell AM, Rowe HC, Hansen BG, Ticconi C, Halkier BA, Kliebenstein DJ (2007) Linking metabolic QTLs with network and cis-eQTLs controlling biosynthetic pathways. PLoS Genet 3:1687–1701
Wittstock U, Halkier BA (2002) Glucosinolate research in the Arabidopsis era. Trends Plant Sci 7:263–270
Yamada K, Hasegawa T, Minami E, Shibuya N, Kosemura S, Yamamura S, Hasegawa KI (2003) Induction of myrosinase gene expression and myrosinase activity in radish hypocotyls by phototropic stimulation. J Plant Physiol 160:255–259
Zou Z, Ishida M, Li F, Kakizaki T, Suzuki S, Kitashiba H, Nishio T (2013) QTL analysis using SNP markers developed by next-generation sequencing for identification of candidate genes controlling 4-methylthio-3-butenyl glucosinolate contents in roots of radish Raphanus sativus L. PLoS One 8:e53541
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Kakizaki, T., Ishida, M. (2017). Genetic Profile of Glucosinolate Biosynthesis. In: Nishio, T., Kitashiba, H. (eds) The Radish Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-319-59253-4_10
Download citation
DOI: https://doi.org/10.1007/978-3-319-59253-4_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-59252-7
Online ISBN: 978-3-319-59253-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)