Abstract
Retrotransposons are an abundant class of mobile genetic elements in mammalian genomes that contribute to genetic instability and variation in the population by integrating at new sites in the genome. Retrotransposons need to be active in the germline so that new retrotransposon integrations can accumulate in the genome during evolution and therefore retrotransposons contain sequences to drive their expression in cells in the germline. While mammals appear to have evolved mechanisms in the germline to limit retrotransposon activity and the resulting mutational load, retrotransposons also appear to be contributing to the regulation of gene expression and driving evolution of transcriptional networks in germline cells. This review will discuss the interplay between retrotransposons and their host cells in the mammalian germline, the genome defence systems that germ cells and pluripotent cells use to limit the mutagenic activity of retrotransposons, and the impact that retrotransposons are having on the biology of mammalian germ cells and pluripotent cells.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Retrotransposons in Mammalian Genomes
Retrotransposons are a class of mobile genetic elements that make up around 40 % of the sequenced mammalian genome (Chinwalla et al. 2002; Lander et al. 2001). Retrotransposons contribute to genomic instability in mammalian genomes by providing interspersed repeats of homologous sequences that can act as substrates for recombination causing deletions, duplications and structural rearrangements in the genome (Romanish et al. 2010). Retrotransposons are thought to be the only active class of mobile genetic element in most mammalian genomes, and can also cause genome instability through jumping to new locations in the genome. These de novo retrotransposon insertions have been reported as the causal mutation in various human genetic diseases (Crichton et al. 2014; Hancks and Kazazian 2012). The copy-and-paste mechanism that retrotransposons use to jump to new locations in the genome involves reverse-transcription of retrotransposon RNA, and integration of the resulting cDNA into new locations in the genome. There are typically a few hundred different types of retrotransposon annotated in each mammalian genome, with each type of retrotransposon being present in up to 10,000 copies. However, the types of retrotransposon, their copy numbers and their genomic locations vary significantly between species.
Mammalian retrotransposons are classified into LIN E (long interspersed nuclear elements ), SINE (short interspersed nuclear elements ) and LTR (long terminal repeat ) retrotransposon classes (Chinwalla et al. 2002; Lander et al. 2001). Each class of retrotransposons can be further subdivided into families, and each family into individual types. The LINE class of retrotransposons in mammals is primarily represented by the LINE-1 family. Only the human-specific L1HS-Ta subfamily of LINE-1 is still active in the human genome, whereas the L1MdA, L1MdTf and L1MdGf types of LINE-1 are all active in the mice (Beck et al. 2011; Hancks and Kazazian 2012; Sookdeo et al. 2013). Full-length LINE-1s are typically 6–7 kb long and are transcribed from an internal promoter located in the 5′ untranslated region (UTR) of these elements (Beck et al. 2011; Hancks and Kazazian 2012; Swergold 1990). The LINE-1 promoter also generates an antisense transcript that can extend into the adjacent flanking cellular DNA (Cruickshanks et al. 2013; Li et al. 2014; Macia et al. 2011; Speek 2001). The LINE-1 5′ UTR varies significantly between species, and even between individual types of LINE-1 within species (Khan et al. 2006; Lee et al. 2010; Sookdeo et al. 2013). LINE-1 encodes two open reading frames in mice and rodents, and three open reading frames in humans and primates (Beck et al. 2011; Denli et al. 2015; Hancks and Kazazian 2012). The recently discovered primate-specific LINE-1 ORF0 protei n is localised to nuclear bodies and enhances LINE-1 retrotransposition activity, although its mechanism of action is not currently understood (Denli et al. 2015). LINE-1 ORF1 protein encodes an RNA-binding protein that forms a particle with LINE-1 RNA and is required for LINE-1 retrotransposition activity (Khazina and Weichenrieder 2009; Martin and Branciforte 1993; Moran et al. 1996). The LINE-1-encoded ORF2 protein is also required for LINE-1 retrotransposition and encodes an endonuclease and reverse transcriptase that nicks the host genomic DNA and catalyses target-primed reverse transcription of LINE-1 RNA into DNA at the site of genomic integration (Cost et al. 2002; Feng et al. 1996; Mathias et al. 1991; Moran et al. 1996). LINE-1 elements display a strong cis-preference where LINE-1 ORF1p and ORF2p tend to associate with the same mRNA molecule from which they are translated (Esnault et al. 2000; Kulpa and Moran 2006; Wei et al. 2001).
The SINE class of retrotransposons includes a group of elements that are typically 100–300 bp long and are derived from small non-coding cellular RNAs including 7SL RNA, 5S rRNA and tRNAs (Kramerov and Vassetzky 2011). SINE retrotransposons are transcribed from internal RNA polymerase III promoters, and include the Alu family in humans and primates. The human genome contains hundreds of active Alu elements, particularly those belonging to AluY and AluS subfamilies (Bennett et al. 2008). Alu elements have been proposed to utilise a ‘stealth’ mode of amplification where elements mobilise at low frequencies for a long time, occasionally generating hyperactive copies that expand aggressively but rapidly become extinct (Han et al. 2005). Alu elements, and SINEs in general, are non-autonomous retrotransposons that rely on LINE-1-encoded proteins to catalyse their retrotransposition (Dewannieux et al. 2003; Hancks et al. 2011; Raiz et al. 2012). The crystal structure of the Alu ribonucleoprotein particle suggests that Alu elements hijack LINE-1 reverse transcriptase by binding to ribosomes that are stalled when LINE-1 ORF2p reverse transcribes its encoding mRNA (Ahl et al. 2015). Other active SINEs in the human genome include SVA elements which can be up to 2 kb in length and contain regions derived from SINE-R and Alu retrotransposons (Hancks and Kazazian 2010; Wang et al. 2005).
The LTR retrotransposon class, whose members are also known as endogenous retroviruses (ERVs ), typically encodes the Gag, Pol, Pro and sometimes also Env proteins that are found in retrovirus genomes (Bannert and Kurth 2006). These retroviral proteins function in the production of retroviral capsid proteins, retroviral DNA synthesis and integration into the host genome, processing of retroviral proteins, and forming the surface envelope on the retroviral capsid, respectively. The 5′ LTR acts as a promoter for these elements. LTR retrotransposons typically reverse-transcribe their RNA in the cytoplasm of the host, then translocate the cDNA into the nucleus and use a LTR retrotransposon-encoded integrase to insert the DNA into the genome. Some LTR retrotransposons are autonomous and encode the proteins required to catalyse their own retrotransposition, whereas others use proteins encoded by other types of LTR retrotransposon to retrotranspose in trans (Dewannieux et al. 2004; Ribet et al. 2004). LTR retrotransposons actively mobilise in rodents, and de novo retrotransposition of LTR retrotransposons accounts for around 10 % of spontaneously occurring mutations in mice (Maksakova et al. 2006). However, the vast majority of LTR retrotransposons in human genomes are probably extinct, and mobilisation of LTR retrotransposons in humans is extremely limited (Wildschutte et al. 2016).
Despite the diversity in retrotransposon structure and life cycle, the reason why each and every one of these elements has been able to accumulate multiple copies in the genome during evolution is because they are able to retrotranspose in the cells that can transmit those new genomic copies to the next generation. These crucial cells that play a key role in the life cycle and biology of retrotransposons in mammals belong to the germline.
2 The Mammalian Germline
In mammals, genetic information is transmitted from generation to generation by germ cells. Germ cells are conceptually distinct from the somatic cells that populate organs like the liver, brain, kidney, lungs and heart in that any genetic change that arises in germ cells can potentially be transmitted to subsequent generations whereas those that arise in somatic cells cannot (Fig. 1). This defining distinction between germ cells and somatic cells, as first proposed by the evolutionary biologist August Weismann (1834–1914) (Weismann 1889), means that it is events and activities that occur within germ cells that shape the landscape of mammalian genomes during evolution. Thus, while any individual retrotransposon might retrotranspose in any specific somatic tissue, all successful retrotransposons must retrotranspose in the germline.
The germline cycle . A schematic diagram showing the mammalian germline cycle. Genetic information, variation and mutations are inherited in the zygote from the parental gametes. The zygote gives rise to an individual containing soma (white) and germline (grey) tissues. Genetic information, variation and mutation in germline tissues can be incorporated into the gametes and transmitted to the next generation, whereas that in the soma cannot
Weismann’s distinction between germline and soma means that genetic information and mutations that arise in somatic cells cannot be transmitted to the germ cells and subsequent generations, but this barrier between germ cells and soma is unidirectional (Fig. 1). Weismann realised that the soma must originate from the germline during early development, and therefore genetic information and mutations that arise in the germline can be propagated into the soma. These early embryonic cells that can give rise to both the germline and all somatic tissues would correspond to totipotent and pluripotent cells in modern terminology (Nichols and Smith 2009), while the term ‘germ cell’ is now typically reserved for lineage-restricted cells that have the capacity to differentiate into the mature gametes, but that do not normally contribute to somatic tissues in that individual (McLaren 2003). Weismann’s germline encompasses totipotent cells, pluripotent cells and germ cells as they are all able to transmit genetic information and mutations to the next generation. Similarly, I will include the totipotent and pluripotent cells that are present in early mammalian development as part of the germline for the purposes of this review.
The major stages in mammalian germline development are outlined in Fig. 2. As many of these events are best characterised in mice, the timings and stages of germline development will be described for this species although there are likely to be broad similarities with other mammalian species. Mouse development initiates when mature germ cells, that is, female eggs and male sperm, fuse during fertilisation to generate a single-celled zygote. This zygote is totipotent in that it has the potential to differentiate into all extra-embryonic and embryonic cell types in the conceptus, which includes the germ cells. As pre-implantation development proceeds to blastocyst stage, some cells in the embryo differentiate into trophectoderm and primitive endoderm lineages that contribute only to extra-embryonic tissues. The remaining cells differentiate into pluripotent epiblast cells which retain the capacity to differentiate into all cell types in the embryo proper, including the germ cells (Magnúsdóttir and Surani 2014; Nichols and Smith 2009). After the blastocyst implants into the uterus, some pluripotent epiblast cells located close to the extra-embryonic ectoderm are induced by extracellular signals to differentiate into primordial germ cells. This germ cell specification event occurs around 6.25–7.25 days post coitum (dpc) in mouse embryos, and generates a founding population of around 40 primordial germ cells (Lawson and Hage 1994; Ohinata et al. 2005). The nascent primordial germ cells then embark on a phase of proliferation as they migrate through the embryo to reach the emerging gonads around 10.5 dpc. The germ cells are typically referred to as primordial germ cells during the early stages of their development until they reach the gonad, and primordial germ cell development at this point occurs similarly in male and female embryos. Once the primordial germ cells colonise the gonad, they continue to proliferate for a few more days, then initiate sex-specific differentiation into either male prospermatogonia or female oocytes (Kocer et al. 2009).
Germline development in mice . A schematic diagram summarising the main stages of germline development in mice. Germ cells and their developmental precursors (totipotent and pluripotent cells) are coloured grey. For pre-implantation stages, a totipotent zygote and cleavage stage embryo, and a blastocyst containing pluripotent epiblast cells are shown. Post-implantation embryos before and after gastrulation are also shown, with pluripotent epiblast cells and primordial germ cells coloured grey. For foetal stages, primordial germ cells are shown within the gonads, and meiotic cells are indicated by a pair of homologous chromosomes in their nuclei. Pachytene stage of meiosis is depicted by the ladder-like synaptonemal complex between homologous chromosomes, which is absent in dictyate oocytes. Products of the first and second meiotic divisions are depicted by nuclei with a single replicated chromosome and an individual chromatid, respectively. Note that the second meiotic division in oocytes is typically only completed at fertilisation. Chromosomes are not shown in diploid mitotic cells for clarity, and the distinctive morphologies of fully grown oocytes and mature sperm are also indicated
The germ cells’ decision to differentiate along a male or a female pathway depends on sex-determining cues present in the gonadal environment rather than the sex chromosome constitution of the germ cells themselves (Kocer et al. 2009). In mice, germ cells in a foetal ovary become committed to develop down a female pathway between 12.5 dpc and 13.5 dpc (Adams and McLaren 2002). These oocytes initiate meiosis around 13.5 dpc and progress through most of the first meiotic prophase during late foetal development, then arrest as dictyate oocytes a few days after birth. The dictyate oocytes will eventually be stimulated to grow and resume meiosis in response to hormonal cues in adult animals. Fully grown oocytes then arrest meiosis during the second meiotic division before being ovulated and potentially fertilised (MacLennan et al. 2015).
In contrast, germ cells in a foetal mouse testis become committed to male development between 11.5 dpc and 12.5 dpc (Adams and McLaren 2002). The resulting prospermatogonia, also termed gonocytes, enter a period of quiescence and differentiation during late foetal development, then resume mitotic proliferation a few days after birth (Rossitto et al. 2015). The prospermatogonia give rise to mitotic spermatogonia and to a pool of spermatogonial stem cells which will self-renew and differentiate into more mitotic spermatogonia, thereby maintaining spermatogenesis throughout adulthood (Yang and Oatley 2014). The spermatogonia undergo a number of mitotic divisions before initiating meiosis which, in contrast to oocytes, typically proceeds without interruption. During spermiogenesis, the post-meiotic spermatids undergo a series of morphological changes that include condensation of the chromatin, elongation of the nucleus, specialisation of the Golgi membranes into an acrosome, generation of a flagella and elimination of residual cytoplasm (O’Donnell 2015). The sperm chromatin is delivered, along with a limited amount of cytoplasmic material, into the oocyte at fertilisation. The highly condensed and specialised sperm chromatin is predominantly associated with protamines rather than histones, and is reprogrammed with oocyte-derived histones shortly after fertilisation (Hogg and Western 2015).
3 Retrotransposon Expression in the Mammalian Germline
For a retrotransposon to accumulate new genomic integrations during evolution it needs to be active in the mammalian germline. While each retrotransposon does not need to be expressed and active at all stages in the germline cycle, each retrotransposon needs to be expressed and active at least at one point in the germline cycle. As outlined in the previous section, the germline cycle involves multiple distinct phases of development, and germline cells appear to use distinct transcription factor networks during these phases. Thus, it is rare to find germline-specific genes or transcription factors that are expressed throughout pre-implantation development, primordial germ cell development, foetal gametogenesis , oogenesis and spermatogenesis . One might expect individual retrotransposon expression profiles to behave similarly.
LINE-1 element transcripts , for example, are reported to be present in pre-implantation embryos, and in primordial germ cells in foetal gonads from 11.5 onwards (Fadloun et al. 2013; Hayashi et al. 2008; Molaro et al. 2014; Seisenberger et al. 2012). Although LINE-1 transcripts are present in foetal germ cells from 11.5 dpc, LINE-1 ORF1 protein is not detected in these cells until 15.5 dpc (Trelogan and Martin 1995). In female germ cells, LINE-1 ORF1p protein is expressed during early meiotic prophase but does not appear to be as abundant during post-natal oocyte development (Malki et al. 2014; Trelogan and Martin 1995). In male germ cells, LINE-1 ORF1 protein levels decrease after birth and, somewhat analogously to females, increase transiently during early meiotic prophase, then decrease as spermatogenesis proceeds (Branciforte and Martin 1994). Thus, early meiotic prophase appears to be a point in the germline cycle that LINE-1 retrotransposons target for expression. Interestingly, different types of LINE-1 retrotransposon have distinct RNA expression profiles during mouse spermatogenesis (Zamudio et al. 2015), which presumably reflects differences in the transcription factors binding to the distinct 5′ UTRs that each type of LINE-1 possesses.
LTR retrotransposons have a rich diversity of expression patterns during the germline cycle. Multiple types of LTR retrotransposon are highly expressed in pre-implantation and early post-implantation embryos, and retroviral-like particles are abundant in the totipotent and pluripotent cells present in these stages, although each LTR retrotransposon has quite specific expression profiles within these developmental stages (Brûlet et al. 1985; Dupressoir and Heidmann 1996; Fadloun et al. 2013; Macfarlan et al. 2012; Peaston et al. 2004; Piko et al. 1984; Reichmann et al. 2012; Ribet et al. 2008; Yotsuyanagi and Szöllösi 1981). The complete germline expression patterns of many types of LTR retrotransposon have not been reported, and additional intricacies are likely to emerge from next generation sequencing of RNA isolated from germline cells. As the retrotransposon LTRs will contain binding sites for transcription factors that are expressed in the germline, understanding how retrotransposons are expressed at specific stages in the germline cycle may help decipher some aspects of the transcriptional regulatory networks operating in these cells and can potentially help identify developmentally distinct subpopulations in the germline cycle (Macfarlan et al. 2012).
4 Retrotransposon Activity in the Mammalian Germline
The rate of de novo retrotransposition in the germline is presumably subject to evolutionary constraint. Although retrotransposons and de novo retrotransposition provide a rich source of genetic material and genetic variation, the insertional mutations that arise from these events contribute to genome instability and high de novo retrotransposition rates could prove to be deleterious for both the host and the retrotransposon. Rates of de novo retrotransposition in the germline have been estimated from sequencing genomic DNA, and from identification of disease and phenotype-causing mutations in mice and humans (Hancks and Kazazian 2012). In humans, de novo LINE-1 insertions are estimated to arise in 1 in every 100 births, with de novo Alu insertions estimated to occur five times more frequently. SVA elements have a somewhat lower de novo retrotransposition rate of approximately 1 in every 1000 births. LTR retrotransposons are not thought to be retrotranspositionally active in humans, but in mice de novo LTR retrotransposition events account for around 10–15 % of sequenced spontaneous mutant alleles (Maksakova et al. 2006). However, in general the stages of germline development during which the retrotransposition can occur are not known.
Retrotransposition at different stages of germline development can have distinct consequences for the host. Retrotransposition in the one cell zygote immediately after fertilisation and before the first round of DNA replication would theoretically result in the de novo retrotransposition event being present in a heterozygous state in all germ cells and somatic cells in that individual. However, retrotransposition at later times during pre-implantation development or during early post-implantation development would likely result in a mosaic conceptus containing some cells that have new heterozygous retrotransposition events, and some that do not. The characterisation of a mutagenic LINE-1 insertion in humans suggests that LINE-1 retrotransposition can generate extensive somatic mosaicism consistent with retrotransposition occurring early in development (van den Hurk et al. 2007). Importantly, if a de novo retrotransposition event has phenotypic consequences , the genetically distinct cells in the conceptus could potentially compete with each other, and select for or against cells carrying the de novo retrotransposition event. Lastly, de novo retrotransposition within the germ cells themselves once they are specified will generate mosaicism and potential competition and selection in the germ cell population, but these events will not be present in the soma.
The timing of de novo retrotransposition is probably best studied for LINE-1 elements. Experiments using transgenic mice and rats carrying human or mouse LINE-1 retrotransposition reporter cassettes suggest that de novo LINE-1 retrotransposition occurs infrequently in the germ cells themselves, and is more readily detectable in pre-implantation embryos (Kano et al. 2009). It is not clear if the inner cell mass, trophectoderm and primitive endoderm layers present in mouse blastocysts have differential susceptibilities to LINE-1 retrotransposition. Even though expression of these LINE-1 reporter transgenes was significantly higher in spermatogenic cells than somatic cells, the relative abundance of cells carrying de novo retrotransposition events in sperm was an order of magnitude lower than that in somatic tissues (Kano et al. 2009). Thus germ cells may possess host defence mechanisms that inhibit LINE-1 retrotransposition at a post-transcriptional level. Intriguingly, this study also indicates that pre-implantation embryos can inherit LINE-1 RNA from both the mature parental sperm and egg, and that this parentally transcribed RNA can retrotranspose in pre-implantation embryos (Kano et al. 2009). The finding that transgenic LINE-1 reporters can retrotranspose in early pre-implantation embryos when pluripotent cells are present in mice is consistent with the observation that transgenic LINE-1 reporter and endogenous LINE-1 and SINE retrotransposition occurs in human induced pluripotent stem cells and human embryonic stem (ES) cells in culture (Garcia-Perez et al. 2010; Klawitter et al. 2016; Wissing et al. 2011). However, data on retrotransposition rates of endogenous LINE-1 elements at different stages of the germline cycle is still lacking. Similarly, the rates of de novo retrotransposition of LTR retrotransposons at different stages of the mouse germline cycle also remain poorly understood. The application and adaptation of new methodologies to identify de novo retrotransposition events in genomic DNA (Baillie et al. 2011; Evrony et al. 2012; Ewing and Kazazian 2010; Upton et al. 2015) should help characterise the natural retrotransposition rates of individual retrotransposons during different stages of the germline cycle.
Although viable retrotransposons must be able to retrotranspose in the germline cycle, some retrotransposons may also be active in somatic tissues . In recent years, evidence has accumulated for de novo LINE-1 retrotransposition in human and mouse brain tissue, and de novo retrotransposition has been shown to be a mutational mechanism that can inactivate tumour suppressor genes in cancer (Baillie et al. 2011; Coufal et al. 2009; Evrony et al. 2012; Muotri et al. 2005; Shukla et al. 2013; Solyom et al. 2012; Upton et al. 2015). This aspect is detailed in chapters ‘Retrotransposon Contribution to Genomic Plasticity’, ‘The Mobilisation of Processed Transcripts in Germline and Somatic Tissues’, ‘Neuronal Genome Plasticity: Retrotransposons, Environment and Disease’ and ‘Activity of Retrotransposons in Stem Cells and Differentiated Cells’ of this book. Similarly, various types of LTR retrotransposon are also expressed in specific somatic tissues (Chuong et al. 2013; Faulkner et al. 2009; Gimenez et al. 2010; Seifarth et al. 2005). Some retrotransposon expression in somatic tissues may represent additional somatic roles for the transcription factors that individual retrotransposons are using to drive their expression in the germline cycle. For example, SOX2, which associates with the LINE-1 5′UTR in human cells (Coufal et al. 2009), is an integral component of the transcription factor network in pluripotent cells, but it also maintains the identity of neural progenitor cells (Avilion et al. 2003; Boyer et al. 2005; Graham et al. 2003). For other retrotransposons, expression in somatic tissues may represent the transcription of a small number of specific integration events that have occurred at genomic loci that promote their expression in somatic tissues. The liver-specific expression of a subset of IAP elements in mice appears to fall into this category (Puech et al. 1997), as does expression of the active L1HS-Ta LINE-1 subfamily in human cell lines (Philippe et al. 2016).
It is often not clear whether retrotransposon expression in somatic tissues has a functional role or if it is evolutionarily neutral. Retrotransposition itself can generate mosaicism in an individual, and it is possible that this provides some evolutionary advantage in some somatic tissues (Muotri et al. 2007). Some retrotransposons can influence expression of nearby host genes, for example, the LTRs of some types of LTR retrotransposon appears to act as enhancers in the placenta, and the activity of these elements in the placenta could be being selected for (Chuong et al. 2013). In some cases, specific copies of retrotransposon-encoded proteins appear to have been co-opted into the host genome to provide key functions in somatic cells, with the repeated independent co-option of LTR retrotransposon proteins to promote cell–cell fusions in the placenta during mammalian evolution being a good example of this (Dupressoir et al. 2011; Mi et al. 2000) (see chapter ‘Roles of Endogenous Retrovirus-Encoded Syncytins in Human Placentation’). The domestication of an ancient LINE-like retrotransposon to generate telomerase, the enzyme that maintains telomeres at the ends of chromosomes in eukaryotes, suggests that retrotransposon-derived sequences can evolve to have functions in somatic tissues (Belfort et al. 2011; Curcio and Belfort 2007; Eickbush 1997). Thus, although viable retrotransposons must be expressed and active in the germline cycle, this requirement is not incompatible with potential roles for these elements in somatic tissues.
5 The Impact of Retrotransposons on the Mammalian Germline
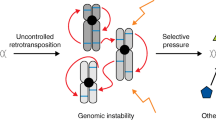
Retrotransposons are able to impact on the germline as a source of trans-generational genomic instability that can cause insertional mutations due to jumping to new locations in the genome, and due to recombination between homologous retrotransposon loci causing genetic deletions, segmental duplications and other chromosomal rearrangements (Fig. 3). However, retrotransposons also influence the biology of germline cells in other ways (Fig. 3). Retrotransposons use germline transcription factors to drive their expression, therefore each retrotransposon locus provides clusters of binding sites for transcription factors that are active in the germline that can provide a useful source of DNA sequence modules for evolution (Bourque et al. 2008; Rebollo et al. 2012). For example, retrotransposon sequences, particularly those from the ERV1 family of LTR retrotransposons, account for around 15–20 % of the genomic locations occupied by either OCT4 or NANOG pluripotency-associated transcription factors in human ES cells, and can drive expression of nearby genes in human ES cells (Kunarso et al. 2010). The ERVL family of LTR retrotransposons similarly appears to strongly influence the oocyte transcriptome and a subset of oocyte transcripts are chimaeras between ERVL retrotransposons and host genes (Peaston et al. 2004). One gene that has an oocyte-specific isoform driven by an ERVL retrotransposon promoter in oocytes is DICER1, which encodes an RNA endonuclease that is involved in the production of endogenous short interfering RNAs (Flemr et al. 2013). This oocyte-specific isoform of DICER1 influences the abundance of retrotransposon transcripts in mouse oocytes, presumably through endogenous siRNAs targeting retrotransposon transcripts (Flemr et al. 2013). Thus, retrotransposon sequences in the genome are even promoting expression of retrotransposon defence mechanisms in the germline. Chimaeric oocyte transcripts originating from ERVL-derived promoters can encode retrotransposon-host gene fusion proteins that are translated in the oocytes (Peaston et al. 2004). Similarly, the recently discovered primate LINE-1 ORF0 transcript can run through to adjacent host genes and generate fusions between the ORF0 polypeptide and host-encoded proteins in human ES cells (Denli et al. 2015). Non-coding RNAs derived from LTR retrotransposons are also reported to be essential to maintain human ES cells in a pluripotent state, possibly through facilitating the binding of some transcriptional co-activator proteins to chromatin (Lu et al. 2014; Wang et al. 2014). Thus retrotransposons appear to be a rich source of genetic material that is contributing to the evolution of the transcriptome and the proteome of germline cells.
Impact of retrotransposons on germline biology. A schematic diagram summarising some of the ways that retrotransposons impact on mammalian germ cells. A region of a chromosome containing a gene (grey) flanked by retrotransposons (RPNs, black) is indicated. Transcription is indicated by corner arrows, and active enhancers by asterisks. RNA transcripts are indicated by wavy lines. For details see main text
Mutations in retrotransposon defence mechanisms can result in high levels of retrotransposon expression during the germline cycle (Crichton et al. 2014; Ollinger et al. 2010; Zamudio and Bourc’his 2010). Mutations in the PIWI-piRNA pathway (Fu and Wang 2014), or in the accessory de novo methyltransferase DNMT3L, can result in high levels of LINE-1 expression in male germ cells, particularly during meiotic prophase (De Fazio et al. 2011; Di Giacomo et al. 2013; Soper et al. 2008; Zamudio et al. 2015). In general, these mutants arrest spermatogenesis during meiosis, typically around the pachytene stage (Crichton et al. 2014; Ollinger et al. 2010; Zamudio and Bourc’his 2010). Mice that have mutations in MAEL, a component of the PIWI-piRNA pathway, are reported to arrest during meiosis with high levels of meiosis-independent DNA damage that could potentially be caused by high levels of de novo retrotransposition of the de-repressed retrotransposons (Soper et al. 2008). In contrast, DNMT3L -/- mice are reported to have no detectable increase in meiosis-independent DNA damage, and have been proposed to arrest during meiosis due to histone modifications at transcriptionally active LINE-1 retrotransposon loci recruiting the meiotic recombination machinery, and disrupting the pairing of homologous chromosomes that characterise meiotic prophase (Zamudio et al. 2015). However, mice that de-repress LINE-1 post-transcriptionally also exhibit chromosome asynapsis and pachytene arrest (Di Giacomo et al. 2013), suggesting that there may be additional aspects to this phenotype that are not currently understood. Importantly, although de-repression of retrotransposons has been reported in various germline genome defence mutants, it remains to be determined whether de novo retrotransposition events are accumulating in the mutant germ cells.
De-repression of retrotransposons in oocytes is also associated with defects in progression through meiotic prophase. Mutations in LSH, a gene implicated in the establishment or maintenance of DNA methylation at retrotransposons and some single copy genes, result in loss of methylation at IAP retrotransposon sequences in oocytes, a failure of oocytes to progress through pachytene, and foetal oocyte death (De La Fuente et al. 2006). Mutating LSH in somatic cells also results in loss of DNA methylation and de-repression of retrotransposons (Dunican et al. 2013). This loss of DNA methylation appears to have indirect effects on other chromatin modifications in the genome as polycomb repressive complexes re-localise to sites normally occupied by DNA methylation, and sequestering polycomb repressive complexes away from their normal targets (Dunican et al. 2013). A similar phenomenon is seen in somatic cells with mutations in the maintenance DNA methyltransferase DNMT1 (Reddington et al. 2013), and in hypomethylated embryonic stem cells (Lynch et al. 2012). It is not clear if relocalisation of polycomb repressive complexes also happens in LSH mutant oocytes or in PIWI-piRNA mutant spermatocytes, but DNA hypomethylation at abundant retrotransposon sequences does have the potential to cause significant effects on the genome-wide distribution of other histone modifications that could be contributing more to the mutant phenotypes than retrotransposon de-repression itself.
Mutations in MAEL result in de-repression of retrotransposons, meiotic abnormalities and foetal oocyte death (Malki et al. 2014). The MAEL -/- oocyte phenotype appears to be related to de-repression of LINE-1 retrotransposons, and differences in the level of LINE-1 expression between individual oocytes in wild-type mice has been proposed to influence foetal oocyte attrition and the number of oocytes present in the ovarian pool at birth (Malki et al. 2014). The rate of foetal oocyte attrition is accelerated in transgenic mice carrying an active LINE-1 transgene, and delayed by treating pregnant mice with an anti-retroviral drug, although these manipulations do not change the final number of oocytes in the ovarian pool (Malki et al. 2014). There are fundamental differences in the way that the oocyte pool influences fertility and menopause between humans and mice, and it will be of interest to determine if manipulating LINE-1 activity can influence the size of the oocyte pool in humans. Human oocytes do, however, contain the host factors to support LINE-1 retrotransposition, at least using engineered LINE-1 reporter constructs (Georgiou et al. 2009).
6 Genome Defence Mechanisms Operating in the Mammalian Germline
The mammalian germline possesses a number of defence mechanisms that suppress the potentially mutagenic activity of retrotransposons in these cells (Crichton et al. 2014; Friedli and Trono 2015; Zamudio and Bourc’his 2010). Histone modification appears to play an important role in repressing retrotransposons in mouse ES cells, with H3K9me3 and H3K27me3 chromatin marks frequently associating with silenced retrotransposons in these cells (Day et al. 2010). Canonical and alternative polycomb repressive complexes , which catalyse trimethylation of H3K27, are involved in repressing multiple families of LTR retrotransposons in mouse ES cells (Hisada et al. 2012; Leeb and Wutz 2007; Reichmann et al. 2012). SETDB1 and SUV39H1/SUV39H2, which trimethylate H3K9, are similarly implicated in repressing LINE-1 and multiple families of LTR retrotransposons on mouse ES cells (Bulut-Karslioglu et al. 2014; Karimi et al. 2011; Matsui et al. 2010; Reichmann et al. 2012). The lysine demethylase KDM1A also plays a role in repressing LINE-1 elements and ERVL LTR retrotransposons (Macfarlan et al. 2012), and histone deacetylases are implicated in repression of ERVK LTR retrotransposons in mouse ES cells (Reichmann et al. 2012), and in suppressing de novo but not pre-existing LINE-1 integrations in human embryonal carcinoma cells (Garcia-Perez et al. 2010). The variety of different mechanisms operating in ES cells may reflect the diversity of retrotransposons in the genome and the multiple strategies that these elements use to drive their transcription in the germline.
As retrotransposons hijack the transcriptional networks present in the host to drive their transcription, it is possible that some of the silencing mechanisms operating in ES cells reflect the mechanisms normally used to regulate the developmental timing of host genes whose expression is driven by the same transcription factors. For example, endogenous gene transcripts that have a role in the zygote might be downregulated as pre-implantation development proceeds such that they are not expressed in pluripotent epiblast cells or ES cells. Similarly, retrotransposons that are using this transcription factor network to drive their expression in zygotes would be expected to be downregulated in ES cells. Some of the mechanisms involved in repressing retrotransposons in ES cells will presumably reflect the normal mechanisms that mediate developmental changes in host gene transcription in these cells, with retrotransposon sequences being repressed ‘by association’ due to their co-regulation with host genes that are downregulated at this stage of development.
In contrast, the specific targeting of repressive chromatin marks to retrotransposons that is mediated by Krüppel-associated box zinc finger proteins (KRAB-ZFPs ) appears to represent a more active and directed defence mechanism against these elements. Specific KRAB-ZFPs bind to specific retrotransposon sequences, recruiting the co-repressor KAP1 (also known as TRIM28) and H3K9me3 chromatin modifications to these sites (Friedman et al. 1996; Rowe et al. 2010; Wolf and Goff 2009, 2007). KAP1 interacts with the histone H3K9 methyltransferase SETDB1, which appears to be the major H3K9 histone methyltransferase involved in silencing LTR retrotransposons in mouse ES cells (Karimi et al. 2011; Matsui et al. 2010; Sripathy et al. 2006). Some LINE-1 elements are silenced by SUV39H1/SUV39H2 H3K9 histone methyltransferases rather than SETDB1 (Bulut-Karslioglu et al. 2014; Matsui et al. 2010), but it is not clear why different histone methyltransferases are being used to silence different retrotransposons. For KAP1 to target a retrotransposon for silencing, a KRAB-ZFP must evolve to bind to that retrotransposon sequence, and KRAB-ZFPs appear to be evolving rapidly for this purpose (Jacobs et al. 2014). Furthermore, retrotransposon loci that mutate or delete their KRAB-ZFP binding sites can escape from KAP1-dependent repression and evolve into new retrotransposon sub-types (Jacobs et al. 2014). Thus while some LINE-1 elements recruit and are repressed by KAP1 in mouse and human ES cells, the youngest types of LINE-1 are repressed by alternative mechanisms (Castro-Diaz et al. 2014).
DNA methylation plays an important role in repressing retrotransposons in germ cells and somatic cells (Bourc’his and Bestor 2004; Davis et al. 1989; De La Fuente et al. 2006; Dunican et al. 2013; Jackson-Grusby et al. 2001; Walsh et al. 1998), although its role in repressing retrotransposons in mouse ES cells may be more restricted than either KAP1 or SETDB1 (Karimi et al. 2011; Matsui et al. 2010; Rowe et al. 2010). The ability of mouse ES cells to induce compensatory histone modifications at some retrotransposons in response to DNA hypomethylation (Walter et al. 2016) may contribute to this. Pluripotent epiblast cells in mouse embryos undergo a wave of de novo DNA methylation during post-implantation development (Borgel et al. 2010) and it is possible that DNA methylation is recruited to retrotransposons that are already silenced by histone modifications at this stage to reinforce and stabilise the repressed state (Rowe et al. 2013). This mechanism bears some resemblance to the observation that bulk of the de novo DNA methylation that occurs during tumourigenesis is located at genes that are already silenced by other mechanisms (Sproul et al. 2011). However, at least for some retrotransposons, persistent KAP1/KRAB-ZFP activity is required to maintain repression of some retrotransposon loci in differentiating and adult somatic cells (Ecco et al. 2016), and the interplay between histone modifications and DNA methylation in establishing and maintaining silencing at different retrotransposon sequences in different cell types is likely to be complex and requires further study.
In the developing germ cells, DNA methylation is globally lost from the genome from 8.5 dpc through to 11.5 dpc as the primordial germ cells migrate to and colonise the genital ridge (Hajkova et al. 2008, 2002). This global loss of DNA methylation includes retrotransposon sequences, although IAP elements are amongst the most resistant to this phenomenon (Popp et al. 2010; Seisenberger et al. 2012). This loss of DNA methylation is part of a more extensive epigenetic reprogramming event that also involves genome-wide loss of various repressive histone modifications including H3K9me1 and H3K9me2 (Hajkova et al. 2008). It is not clear how retrotransposons are transcriptionally repressed during this stage of germ cell development, but the histone arginine methyltransferase PRMT5 localises to the nucleus and symmetric dimethylation of histones H2AR3 and/or H4R3 is upregulated during the early part of this reprogramming event and are present at LINE-1 and IAP retrotransposon sequences (Ancelin et al. 2006; Kim et al. 2014). PRMT5 -/- primordial germ cells have undetectable levels of H2A/H4R3me2s in their nuclei, and de-repress LINE-1, IAP and other retrotransposons suggesting that PRMT5-dependent symmetric dimethylation of histones H2AR3 and H4R3 may contribute directly to the repression of retrotransposons in hypomethylated primordial germ cells (Kim et al. 2014). There may of course be additional, presently uncharacterised, histone modifications associated with specific types of retrotransposon that are helping to repress transcription of these elements in hypomethylated germ cells.
At around 11.5 dpc, PRMT5 is re-localised from the primordial germ cell nucleus into the cytoplasm and levels of H2A/H4R3me2s concomitantly decrease (Ancelin et al. 2006). Additional mechanisms likely become important to limit retrotransposon activity in hypomethylated germ cells at this stage. Analysis of retrotransposon transcript abundance in the transcriptome of 13.5 dpc primordial germ cell from wild-type mice supports widespread low level transcriptional de-repression of retrotransposons in hypomethylated germ cells at this stage (Molaro et al. 2014). The DNA hypomethylation that occurs in the developing primordial germ cells is not restricted to retrotransposons and extends to most genomic features including endogenous gene promoters (Popp et al. 2010; Seisenberger et al. 2012). Thus this global DNA hypomethylation event can influence expression of host genes in the developing germline (Hackett et al. 2012). Interestingly, many genes that are primarily and causally regulated by DNA methylation in mice are germline-specific genes that are involved in suppressing retrotransposon activity (Hackett et al. 2012). DNA hypomethylation in the developing germline therefore appears to induce expression of a group of retrotransposon-suppressing genome defence genes to compensate for the increase in potential retrotransposon activity caused by loss of this repressive epigenetic mark.
One of the genome defence genes that is most sensitive to DNA hypomethylation is TEX19.1. TEX19.1 is required to repress MMERVK10C LTR retrotransposons in spermatocytes, and also to repress LINE-1 elements and LTR retrotransposons in the hypomethylated somatic cells present in the placenta (Öllinger et al. 2008; Reichmann et al. 2013). Most of the other germline genome defence genes induced in response to global DNA hypomethylation are components of the PIWI-piRNA pathway for repressing retrotransposons. The PIWI-piRNA pathway uses small RNAs encoded in the genome to target retrotransposons for suppression by epigenetic and post-transcriptional mechanisms in the germline (Fu and Wang 2014; Iwasaki et al. 2015). There are three PIWI proteins in mouse and rat genomes, but four PIWI proteins in many other mammals including humans. The three mouse PIWI proteins have distinct expression profile during spermatogenesis. These PIWI proteins physically interact with small single-stranded PIWI-interacting RNAs (piRNAs ) whose sequence is thought to target the PIWI proteins to retrotransposon sequences (Aravin et al. 2006, 2008, 2007; Carmell et al. 2007; Kuramochi-Miyagawa et al. 2001). Genomic piRNAs are derived from long RNA precursors that undergo a number of processing events to generate mature piRNAs. These primary piRNAs can facilitate processing of complementary precursor sequences, such as retrotransposon transcripts, into secondary piRNAs which in turn can promote processing of genomic precursors into primary piRNAs. This ping-pong cycle can amplify groups of piRNAs and is important for generating an effective piRNA response against LINE-1 elements in male mouse germ cells (De Fazio et al. 2011). The slicer RNA endonuclease activity of PIWI proteins plays an important role in processing piRNA precursors, and PIWI-piRNA-directed slicing of retrotransposon RNAs by PIWIL1 and PIWIL2 contribute to the PIWI-piRNA defence against retrotransposons (De Fazio et al. 2011; Di Giacomo et al. 2013; Reuter et al. 2011). PIWIL2 and PIWIL4 are required for male germ cells to establish de novo DNA methylation at retrotransposon sequences (Aravin et al. 2008; Kuramochi-Miyagawa et al. 2008). De novo methylation of retrotransposons occurs from 16.5 dpc onwards in foetal male germ cells, and it is possible that sequence information present in the PIWI-piRNA pathway is being used to direct the de novo DNA methylation machinery to these sequences. It is not clear if PIWI-piRNA complexes regulate DNA methylation directly, or indirectly through other chromatin modifications. H3K9 methylation has been implicated in PIWI-dependent retrotransposon silencing in Drosophila, and in silencing retrotransposons in post-natal male germ cells in mice (Di Giacomo et al. 2014, 2013; Huang et al. 2013; Pezic et al. 2014). Perhaps one of the reasons that genome-wide loss of DNA methylation occurs in the developing germ cells is to expose retrotransposon loci to the PIWI-piRNA pathway so that retrotransposons can be identified and epigenetic repression of these elements established de novo in preparation for transmission of the genome to the next generation. Removing and resetting epigenetic marks on these sequences may be preferable to propagating existing marks in order to prevent epimutations from being transmitted across multiple generations. There may also be some analogies to the mechanisms operating in Arabidopsis, where programmed loss of DNA demethylation in the pollen’s vegetative nucleus results in de-repression of retrotransposons whose transcripts are processed into small RNAs, transported to the pollen’s germline nucleus, and used to direct epigenetic silencing of retrotransposons in the germline DNA (Calarco et al. 2012; Slotkin et al. 2009). The DNA methylation-sensitive coupling of expression of post-transcriptional genome defence mechanisms and components of the PIWI-piRNA pathway to transcriptional de-repression of retrotransposons in mouse germ cells may similarly allow mouse germ cells to generate the retrotransposon RNA transcripts needed to direct de novo identification and silencing of retrotransposon loci in the mammalian germline (Fig. 4).
Potential role of genome defence genes in the male germline. A schematic diagram outlining the potential role of genome defence genes during epigenetic reprogramming in the male germline. Germ cells losing DNA methylation (filled grey circles → filled white circles) transcribe RNA (wavy lines) encoding genome defence genes and retrotransposons, which can be translated into protein (filled triangles and squares, respectively). Genome defence proteins, including components of the PIWI-piRNA pathway, that inhibit any post-transcriptional stages of the retrotransposon life cycle (grey) can limit mutations caused by retrotransposition, while allowing retrotransposon RNA transcripts to prime the PIWI-piRNA pathway (broken wavy lines). The PIWI-piRNA pathway slices retrotransposon RNA transcripts to generate piRNAs and can potentially use sequence information in the piRNAs to direct de novo DNA methylation onto retrotransposon sequences (indicated by question mark)
Components of the PIWI-piRNA system also appear to play roles in post-transcriptional suppression of retrotransposons in oocytes, and PIWIL2 suppresses LINE-1 mobility in human induced pluripotent cells (Lim et al. 2013; Malki et al. 2014; Marchetto et al. 2013; Watanabe et al. 2008). In mice, PIWI function in oocytes is primarily provided by PIWIL2 although additional members of the PIWI family may also contribute to PIWI function in oocytes in other mammalian species including human (Roovers et al. 2015). Mutating PIWIL2 in mice does not have the severe consequences for fertility in females that it does in males (Kuramochi-Miyagawa et al. 2004; Lim et al. 2013), which may in part reflect differences in the way that de novo DNA methylation is regulated between spermatogenesis and oogenesis (Smallwood and Kelsey 2012). De novo DNA methylation occurs post-natally during oocyte growth in the female germline, and oocytes are therefore in a DNA hypomethylated state throughout their prolonged dictyate arrest and for much of their adult life. In the absence of PIWIL2, the abundance of some retrotransposon transcripts is elevated in oocytes, potentially reflecting PIWIL2-dependent post-transcriptional suppression of these elements during oogenesis (Lim et al. 2013; Watanabe et al. 2008). However, the PIWI-piRNA system is not the only mechanism that operates in oocytes to post-transcriptionally suppress retrotransposons, and DICER1-dependent endogenous siRNAs also make a significant contribution (Flemr et al. 2013; Stein et al. 2015; Tam et al. 2008; Watanabe et al. 2008). MARF1 may represent another mechanism regulating retrotransposons at a post-transcriptional level in these cells (Su et al. 2012a, b).
The distinct post-transcriptional suppression mechanisms operating in mouse oocytes appear to complement each other to target different types of retrotransposon (Watanabe et al. 2008). Pools of endogenous siRNA and piRNA present in fully grown oocytes can potentially be transmitted to the next generation to provide some protection against retrotransposons in pre-implantation embryos. However, retrotransposon-encoded transcripts, proteins, and ribonucleoprotein particles that are expressed during the oocyte’s prolonged dictyate arrest or post-natal growth can similarly be transmitted in the oocyte cytoplasm and can cause retrotransposition in the next generation (Kano et al. 2009). The presence of multiple overlapping and complementary genome defence mechanisms in oocytes may therefore provide some protection against retrotransposon mobilisation during oogenesis, and also help to limit maternal transmission of retrotransposon-derived ribonucleoprotein particles than can retrotranspose in the next generation.
7 Concluding Remarks
As described in this chapter, the mammalian germline and retrotransposons are intrinsically linked in multiple ways. All retrotransposons need to be expressed and active in the mammalian germline in order to accumulate in the genome during evolution, and while the germline appears to have evolved multiple defence mechanisms to limit the mutagenic activity of these elements, these mechanisms are helping to drive evolution of retrotransposons to escape suppression. This situation is analogous to the Red Queen hypothesis (van Valen 1973) as both retrotransposons and germline defence mechanisms need to continue to evolve simply to keep up with each other. However, in addition to this antagonistic relationship, retrotransposons appear to be participating in the transcriptional and proteomic networks of germline cells and providing regulatory modules for gene expression that are being repurposed by the germline to help it evolve. Additional intricacies will likely emerge in the coming years as the interplay between retrotransposons and the germline becomes better understood, but it would appear that retrotransposons can be viewed as having both beneficial and deleterious effects on their germline hosts.
References
Adams IR, McLaren A (2002) Sexually dimorphic development of mouse primordial germ cells: switching from oogenesis to spermatogenesis. Development 129:1155–1164
Ahl V, Keller H, Schmidt S, Weichenrieder O (2015) Retrotransposition and crystal structure of an Alu RNP in the ribosome-stalling conformation. Mol Cell 60:715–727. doi:10.1016/j.molcel.2015.10.003
Ancelin K, Lange UC, Hajkova P, Schneider R, Bannister AJ, Kouzarides T, Surani MA (2006) Blimp1 associates with Prmt5 and directs histone arginine methylation in mouse germ cells. Nat Cell Biol 8:623–630. doi:10.1038/ncb1413
Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31:785–799. doi:10.1016/j.molcel.2008.09.003
Aravin AA, Sachidanandam R, Girard A, Fejes-Toth K, Hannon GJ (2007) Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316:744–747. doi:10.1126/science.1142612
Aravin A, Gaidatzis D, Pfeffer S, Lagos-Quintana M, Landgraf P, Iovino N, Morris P, Brownstein MJ, Kuramochi-Miyagawa S, Nakano T, Chien M, Russo JJ, Ju J, Sheridan R, Sander C, Zavolan M, Tuschl T (2006) A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442:203–207. doi:10.1038/nature04916
Avilion AA, Nicolis SK, Pevny LH, Perez L, Vivian N, Lovell-Badge R (2003) Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev 17:126–140. doi:10.1101/gad.224503
Baillie JK, Barnett MW, Upton KR, Gerhardt DJ, Richmond TA, De Sapio F, Brennan P, Rizzu P, Smith S, Fell M, Talbot RT, Gustincich S, Freeman TC, Mattick JS, Hume DA, Heutink P, Carninci P, Jeddeloh JA, Faulkner GJ (2011) Somatic retrotransposition alters the genetic landscape of the human brain. Nature 479:534–537. doi:10.1038/nature10531
Bannert N, Kurth R (2006) The evolutionary dynamics of human endogenous retroviral families. Annu Rev Genomics Hum Genet 7:149–173. doi:10.1146/annurev.genom.7.080505.115700
Beck CR, Garcia-Perez JL, Badge RM, Moran JV (2011) LINE-1 elements in structural variation and disease. Annu Rev Genomics Hum Genet 12:187–215. doi:10.1146/annurev-genom-082509-141802
Belfort M, Curcio MJ, Lue NF (2011) Telomerase and retrotransposons: reverse transcriptases that shaped genomes. Proc Natl Acad Sci U S A 108:20304–20310. doi:10.1073/pnas.1100269109
Bennett EA, Keller H, Mills RE, Schmidt S, Moran JV, Weichenrieder O, Devine SE (2008) Active Alu retrotransposons in the human genome. Genome Res 18:1875–1883. doi:10.1101/gr.081737.108
Borgel J, Guibert S, Li Y, Chiba H, Schübeler D, Sasaki H, Forné T, Weber M (2010) Targets and dynamics of promoter DNA methylation during early mouse development. Nat Genet 42:1093–1100. doi:10.1038/ng.708
Bourc’his D, Bestor TH (2004) Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431:96–99. doi:10.1038/nature02886
Bourque G, Leong B, Vega VB, Chen X, Lee YL, Srinivasan KG, Chew J-L, Ruan Y, Wei C-L, Ng HH, Liu ET (2008) Evolution of the mammalian transcription factor binding repertoire via transposable elements. Genome Res 18:1752–1762. doi:10.1101/gr.080663.108
Boyer LA, Lee TI, Cole MF, Johnstone SE, Levine SS, Zucker JP, Guenther MG, Kumar RM, Murray HL, Jenner RG, Gifford DK, Melton DA, Jaenisch R, Young RA (2005) Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122:947–956. doi:10.1016/j.cell.2005.08.020
Branciforte D, Martin SL (1994) Developmental and cell type specificity of LINE-1 expression in mouse testis: implications for transposition. Mol Cell Biol 14:2584–2592
Brûlet P, Condamine H, Jacob F (1985) Spatial distribution of transcripts of the long repeated ETn sequence during early mouse embryogenesis. Proc Natl Acad Sci U S A 82:2054–2058
Bulut-Karslioglu A, De La Rosa-Velázquez IA, Ramirez F, Barenboim M, Onishi-Seebacher M, Arand J, Galán C, Winter GE, Engist B, Gerle B, O’Sullivan RJ, Martens JHA, Walter J, Manke T, Lachner M, Jenuwein T (2014) Suv39h-dependent H3K9me3 marks intact retrotransposons and silences LINE elements in mouse embryonic stem cells. Mol Cell 55:277–290. doi:10.1016/j.molcel.2014.05.029
Calarco JP, Borges F, Donoghue MTA, Van Ex F, Jullien PE, Lopes T, Gardner R, Berger F, Feijó JA, Becker JD, Martienssen RA (2012) Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151:194–205. doi:10.1016/j.cell.2012.09.001
Carmell MA, Girard A, van de Kant HJG, Bourc’his D, Bestor TH, de Rooij DG, Hannon GJ (2007) MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell 12:503–514. doi:10.1016/j.devcel.2007.03.001
Castro-Diaz N, Ecco G, Coluccio A, Kapopoulou A, Yazdanpanah B, Friedli M, Duc J, Jang SM, Turelli P, Trono D (2014) Evolutionally dynamic L1 regulation in embryonic stem cells. Genes Dev 28:1397–1409. doi:10.1101/gad.241661.114
Chinwalla AT, Cook LL, Delehaunty KD, Fewell GA, Fulton LA, Fulton RS, Graves TA, Hillier LW, Mardis ER, McPherson JD, Miner TL, Nash WE, Nelson JO, Nhan MN, Pepin KH, Pohl CS, Ponce TC, Schultz B, Thompson J, Trevaskis E, Waterston RH, Wendl MC, Wilson RK, Yang S-P, An P, Berry E, Birren B, Bloom T, Brown DG, Butler J, Daly M, David R, Deri J, Dodge S, Foley K, Gage D, Gnerre S, Holzer T, Jaffe DB, Kamal M, Karlsson EK, Kells C, Kirby A, Kulbokas EJ, Lander ES, Landers T, Leger JP, Levine R, Lindblad-Toh K, Mauceli E, Mayer JH, McCarthy M, Meldrim J, Mesirov JP, Nicol R, Nusbaum C, Seaman S, Sharpe T, Sheridan A, Singer JB, Santos R, Spencer B, Stange-Thomann N, Vinson JP, Wade CM, Wierzbowski J, Wyman D, Zody MC, Birney E, Goldman N, Kasprzyk A, Mongin E, Rust AG, Slater G, Stabenau A, Ureta-Vidal A, Whelan S, Ainscough R, Attwood J, Bailey J, Barlow K, Beck S, Burton J, Clamp M, Clee C, Coulson A, Cuff J, Curwen V, Cutts T, Davies J, Eyras E, Grafham D, Gregory S, Hubbard T, Hunt A, Jones M, Joy A, Leonard S, Lloyd C, Matthews L, McLaren S, McLay K, Meredith B, Mullikin JC, Ning Z, Oliver K, Overton-Larty E, Plumb R, Potter S, Quail M, Rogers J, Scott C, Searle S, Shownkeen R, Sims S, Wall M, West AP, Willey D, Williams S, Abril JF, Guigó R, Parra G, Agarwal P, Agarwala R, Church DM, Hlavina W, Maglott DR, Sapojnikov V, Alexandersson M, Pachter L, Antonarakis SE, Dermitzakis ET, Reymond A, Ucla C, Baertsch R, Diekhans M, Furey TS, Hinrichs A, Hsu F, Karolchik D, Kent WJ, Roskin KM, Schwartz MS, Sugnet C, Weber RJ, Bork P, Letunic I, Suyama M, Torrents D, Zdobnov EM, Botcherby M, Brown SD, Campbell RD, Jackson I, Bray N, Couronne O, Dubchak I, Poliakov A, Rubin EM, Brent MR, Flicek P, Keibler E, Korf I, Batalov S, Bult C, Frankel WN, Carninci P, Hayashizaki Y, Kawai J, Okazaki Y, Cawley S, Kulp D, Wheeler R, Chiaromonte F, Collins FS, Felsenfeld A, Guyer M, Peterson J, Wetterstrand K, Copley RR, Mott R, Dewey C, Dickens NJ, Emes RD, Goodstadt L, Ponting CP, Winter E, Dunn DM, von Niederhausern AC, Weiss RB, Eddy SR, Johnson LS, Jones TA, Elnitski L, Kolbe DL, Eswara P, Miller W, O’Connor MJ, Schwartz S, Gibbs RA, Muzny DM, Glusman G, Smit A, Green ED, Hardison RC, Yang S, Haussler D, Hua A, Roe BA, Kucherlapati RS, Montgomery KT, Li J, Li M, Lucas S, Ma B, McCombie WR, Morgan M, Pevzner P, Tesler G, Schultz J, Smith DR, Tromp J, Worley KC, Green ED (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562. doi:10.1038/nature01262
Chuong EB, Rumi MAK, Soares MJ, Baker JC (2013) Endogenous retroviruses function as species-specific enhancer elements in the placenta. Nat Genet 45:325–329. doi:10.1038/ng.2553
Cost GJ, Feng Q, Jacquier A, Boeke JD (2002) Human L1 element target-primed reverse transcription in vitro. EMBO J 21:5899–5910
Coufal NG, Garcia-Perez JL, Peng GE, Yeo GW, Mu Y, Lovci MT, Morell M, O’Shea KS, Moran JV, Gage FH (2009) L1 retrotransposition in human neural progenitor cells. Nature 460:1127–1131. doi:10.1038/nature08248
Crichton JH, Dunican DS, Maclennan M, Meehan RR, Adams IR (2014) Defending the genome from the enemy within: mechanisms of retrotransposon suppression in the mouse germline. Cell Mol Life Sci 71:1581–1605. doi:10.1007/s00018-013-1468-0
Cruickshanks HA, Vafadar-Isfahani N, Dunican DS, Lee A, Sproul D, Lund JN, Meehan RR, Tufarelli C (2013) Expression of a large LINE-1-driven antisense RNA is linked to epigenetic silencing of the metastasis suppressor gene TFPI-2 in cancer. Nucleic Acids Res 41:6857–6869. doi:10.1093/nar/gkt438
Curcio MJ, Belfort M (2007) The beginning of the end: links between ancient retroelements and modern telomerases. Proc Natl Acad Sci U S A 104:9107–9108. doi:10.1073/pnas.0703224104
Davis CM, Constantinides PG, van der Riet F, van Schalkwyk L, Gevers W, Parker MI (1989) Activation and demethylation of the intracisternal A particle genes by 5-azacytidine. Cell Differ Dev 27:83–93
Day DS, Luquette LJ, Park PJ, Kharchenko PV (2010) Estimating enrichment of repetitive elements from high-throughput sequence data. Genome Biol 11:R69–R69. doi:10.1186/gb-2010-11-6-r69
De Fazio S, Bartonicek N, Di Giacomo M, Abreu-Goodger C, Sankar A, Funaya C, Antony C, Moreira PN, Enright AJ, O’Carroll D (2011) The endonuclease activity of Mili fuels piRNA amplification that silences LINE1 elements. Nature 480:259–263. doi:10.1038/nature10547
De La Fuente R, Baumann C, Fan T, Schmidtmann A, Dobrinski I, Muegge K (2006) Lsh is required for meiotic chromosome synapsis and retrotransposon silencing in female germ cells. Nat Cell Biol 8:1448–1454. doi:10.1038/ncb1513
Denli AM, Narvaiza I, Kerman BE, Pena M, Benner C, Marchetto MCN, Diedrich JK, Aslanian A, Ma J, Moresco JJ, Moore L, Hunter T, Saghatelian A, Gage FH (2015) Primate-specific ORF0 contributes to retrotransposon-mediated diversity. Cell 163:583–593. doi:10.1016/j.cell.2015.09.025
Dewannieux M, Dupressoir A, Harper F, Pierron G, Heidmann T (2004) Identification of autonomous IAP LTR retrotransposons mobile in mammalian cells. Nat Genet 36:534–539. doi:10.1038/ng1353
Dewannieux M, Esnault C, Heidmann T (2003) LINE-mediated retrotransposition of marked Alu sequences. Nat Genet 35:41–48. doi:10.1038/ng1223
Di Giacomo M, Comazzetto S, Saini H, De Fazio S, Carrieri C, Morgan M, Vasiliauskaite L, Benes V, Enright AJ, O’Carroll D (2013) Multiple epigenetic mechanisms and the piRNA pathway enforce LINE1 silencing during adult spermatogenesis. Mol Cell 50:601–608. doi:10.1016/j.molcel.2013.04.026
Di Giacomo M, Comazzetto S, Sampath SC, Sampath SC, O’Carroll D (2014) G9a co-suppresses LINE1 elements in spermatogonia. Epigenetics Chromatin 7:24. doi:10.1186/1756-8935-7-24
Dunican DS, Cruickshanks HA, Suzuki M, Semple CA, Davey T, Arceci RJ, Greally J, Adams IR, Meehan RR (2013) Lsh regulates LTR retrotransposon repression independently of Dnmt3b function. Genome Biol 14:R146. doi:10.1186/gb-2013-14-12-r146
Dupressoir A, Heidmann T (1996) Germ line-specific expression of intracisternal A-particle retrotransposons in transgenic mice. Mol Cell Biol 16:4495–4503
Dupressoir A, Vernochet C, Harper F, Guégan J, Dessen P, Pierron G, Heidmann T (2011) A pair of co-opted retroviral envelope syncytin genes is required for formation of the two-layered murine placental syncytiotrophoblast. Proc Natl Acad Sci U S A 108:E1164–E1173. doi:10.1073/pnas.1112304108
Ecco G, Cassano M, Kauzlaric A, Duc J, Coluccio A, Offner S, Imbeault M, Rowe HM, Turelli P, Trono D (2016) Transposable elements and their KRAB-ZFP controllers regulate gene expression in adult tissues. Dev Cell 36:611–623. doi:10.1016/j.devcel.2016.02.024
Eickbush TH (1997) Telomerase and retrotransposons: which came first? Science 277:911–912
Esnault C, Maestre J, Heidmann T (2000) Human LINE retrotransposons generate processed pseudogenes. Nat Genet 24:363–367. doi:10.1038/74184
Evrony GD, Cai X, Lee E, Hills LB, Elhosary PC, Lehmann HS, Parker JJ, Atabay KD, Gilmore EC, Poduri A, Park PJ, Walsh CA (2012) Single-neuron sequencing analysis of L1 retrotransposition and somatic mutation in the human brain. Cell 151:483–496. doi:10.1016/j.cell.2012.09.035
Ewing AD, Kazazian HH (2010) High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res 20:1262–1270. doi:10.1101/gr.106419.110
Fadloun A, Le Gras S, Jost B, Ziegler-Birling C, Takahashi H, Gorab E, Carninci P, Torres-Padilla M-E (2013) Chromatin signatures and retrotransposon profiling in mouse embryos reveal regulation of LINE-1 by RNA. Nat Struct Mol Biol 20:332–338. doi:10.1038/nsmb.2495
Faulkner GJ, Kimura Y, Daub CO, Wani S, Plessy C, Irvine KM, Schroder K, Cloonan N, Steptoe AL, Lassmann T, Waki K, Hornig N, Arakawa T, Takahashi H, Kawai J, Forrest ARR, Suzuki H, Hayashizaki Y, Hume DA, Orlando V, Grimmond SM, Carninci P (2009) The regulated retrotransposon transcriptome of mammalian cells. Nat Genet 41:563–571. doi:10.1038/ng.368
Feng Q, Moran JV, Kazazian HH Jr, Boeke JD (1996) Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell 87:905–916
Flemr M, Malik R, Franke V, Nejepinska J, Sedlacek R, Vlahovicek K, Svoboda P (2013) A retrotransposon-driven dicer isoform directs endogenous small interfering RNA production in mouse oocytes. Cell 155:807–816. doi:10.1016/j.cell.2013.10.001
Friedli M, Trono D (2015) The developmental control of transposable elements and the evolution of higher species. Annu Rev Cell Dev Biol 31:429–451. doi:10.1146/annurev-cellbio-100814-125514
Friedman JR, Fredericks WJ, Jensen DE, Speicher DW, Huang XP, Neilson EG, Rauscher FJ (1996) KAP-1, a novel corepressor for the highly conserved KRAB repression domain. Genes Dev 10:2067–2078
Fu Q, Wang PJ (2014) Mammalian piRNAs: biogenesis, function, and mysteries. Spermatogenesis 4, e27889. doi:10.4161/spmg.27889
Garcia-Perez JL, Morell M, Scheys JO, Kulpa DA, Morell S, Carter CC, Hammer GD, Collins KL, O’Shea KS, Menendez P, Moran JV (2010) Epigenetic silencing of engineered L1 retrotransposition events in human embryonic carcinoma cells. Nature 466:769–773. doi:10.1038/nature09209
Georgiou I, Noutsopoulos D, Dimitriadou E, Markopoulos G, Apergi A, Lazaros L, Vaxevanoglou T, Pantos K, Syrrou M, Tzavaras T (2009) Retrotransposon RNA expression and evidence for retrotransposition events in human oocytes. Hum Mol Genet 18:1221–1228. doi:10.1093/hmg/ddp022
Gimenez J, Montgiraud C, Pichon J-P, Bonnaud B, Arsac M, Ruel K, Bouton O, Mallet F (2010) Custom human endogenous retroviruses dedicated microarray identifies self-induced HERV-W family elements reactivated in testicular cancer upon methylation control. Nucleic Acids Res 38:2229–2246. doi:10.1093/nar/gkp1214
Graham V, Khudyakov J, Ellis P, Pevny L (2003) SOX2 functions to maintain neural progenitor identity. Neuron 39:749–765
Hackett JA, Reddington JP, Nestor CE, Dunican DS, Branco MR, Reichmann J, Reik W, Surani MA, Adams IR, Meehan RR (2012) Promoter DNA methylation couples genome-defence mechanisms to epigenetic reprogramming in the mouse germline. Development 139:3623–3632. doi:10.1242/dev.081661
Hajkova P, Ancelin K, Waldmann T, Lacoste N, Lange UC, Cesari F, Lee C, Almouzni G, Schneider R, Surani MA (2008) Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 452:877–881. doi:10.1038/nature06714
Hajkova P, Erhardt S, Lane N, Haaf T, El-Maarri O, Reik W, Walter J, Surani MA (2002) Epigenetic reprogramming in mouse primordial germ cells. Mech Dev 117:15–23. doi:10.1016/S0925-4773(02)00181-8
Hancks DC, Goodier JL, Mandal PK, Cheung LE, Kazazian HH (2011) Retrotransposition of marked SVA elements by human L1s in cultured cells. Hum Mol Genet 20:3386–3400. doi:10.1093/hmg/ddr245
Hancks DC, Kazazian H (2010) SVA retrotransposons: evolution and genetic instability. Semin Cancer Biol 20:234–245. doi:10.1016/j.semcancer.2010.04.001
Hancks DC, Kazazian HH Jr (2012) Active human retrotransposons: variation and disease. Curr Opin Genet Dev 22:191–203. doi:10.1016/j.gde.2012.02.006
Han K, Xing J, Wang H, Hedges DJ, Garber RK, Cordaux R, Batzer MA (2005) Under the genomic radar: the stealth model of Alu amplification. Genome Res 15:655–664. doi:10.1101/gr.3492605
Hayashi K, Chuva de Sousa Lopes SM, Kaneda M, Tang F, Hajkova P, Lao K, O’Carroll D, Das PP, Tarakhovsky A, Miska EA, Surani MA (2008) MicroRNA biogenesis is required for mouse primordial germ cell development and spermatogenesis. PLoS One 3:e1738. doi:10.1371/journal.pone.0001738
Hisada K, Sánchez C, Endo TA, Endoh M, Román-Trufero M, Sharif J, Koseki H, Vidal M (2012) RYBP represses endogenous retroviruses and preimplantation- and germ line-specific genes in mouse embryonic stem cells. Mol Cell Biol 32:1139–1149. doi:10.1128/MCB.06441-11
Hogg K, Western PS (2015) Refurbishing the germline epigenome: out with the old, in with the new. Semin Cell Dev Biol 45:104–113. doi:10.1016/j.semcdb.2015.09.012
Huang XA, Yin H, Sweeney S, Raha D, Snyder M, Lin H (2013) A major epigenetic programming mechanism guided by piRNAs. Dev Cell 24:502–516. doi:10.1016/j.devcel.2013.01.023
Iwasaki YW, Siomi MC, Siomi H (2015) PIWI-interacting RNA: its biogenesis and functions. Annu Rev Biochem 84:405–433. doi:10.1146/annurev-biochem-060614-034258
Jackson-Grusby L, Beard C, Possemato R, Tudor M, Fambrough D, Csankovszki G, Dausman J, Lee P, Wilson C, Lander E, Jaenisch R (2001) Loss of genomic methylation causes p53-dependent apoptosis and epigenetic deregulation. Nat Genet 27:31–39. doi:10.1038/83730
Jacobs FMJ, Greenberg D, Nguyen N, Haeussler M, Ewing AD, Katzman S, Paten B, Salama SR, Haussler D (2014) An evolutionary arms race between KRAB zinc-finger genes ZNF91/93 and SVA/L1 retrotransposons. Nature 516:242–245. doi:10.1038/nature13760
Kano H, Godoy I, Courtney C, Vetter MR, Gerton GL, Ostertag EM, Kazazian HH Jr (2009) L1 retrotransposition occurs mainly in embryogenesis and creates somatic mosaicism. Genes Dev 23:1303–1312. doi:10.1101/gad.1803909
Karimi MM, Goyal P, Maksakova IA, Bilenky M, Leung D, Tang JX, Shinkai Y, Mager DL, Jones S, Hirst M, Lorincz MC (2011) DNA methylation and SETDB1/H3K9me3 regulate predominantly distinct sets of genes, retroelements, and chimeric transcripts in mESCs. Cell Stem Cell 8:676–687. doi:10.1016/j.stem.2011.04.004
Khan H, Smit A, Boissinot S (2006) Molecular evolution and tempo of amplification of human LINE-1 retrotransposons since the origin of primates. Genome Res 16:78–87. doi:10.1101/gr.4001406
Khazina E, Weichenrieder O (2009) Non-LTR retrotransposons encode noncanonical RRM domains in their first open reading frame. Proc Natl Acad Sci U S A 106:731–736. doi:10.1073/pnas.0809964106
Kim S, Günesdogan U, Zylicz JJ, Hackett JA, Cougot D, Bao S, Lee C, Dietmann S, Allen GE, Sengupta R, Surani MA (2014) PRMT5 protects genomic integrity during global DNA demethylation in primordial germ cells and preimplantation embryos. Mol Cell 56:564–579. doi:10.1016/j.molcel.2014.10.003
Klawitter S, Fuchs NV, Upton KR, Muñoz-Lopez M, Shukla R, Wang J, Garcia-Cañadas M, Lopez-Ruiz C, Gerhardt DJ, Sebe A, Grabundzija I, Merkert S, Gerdes P, Pulgarin JA, Bock A, Held U, Witthuhn A, Haase A, Sarkadi B, Löwer J, Wolvetang EJ, Martin U, Ivics Z, Izsvák Z, Garcia-Perez JL, Faulkner GJ, Schumann GG (2016) Reprogramming triggers endogenous L1 and Alu retrotransposition in human induced pluripotent stem cells. Nat Commun 7:10286. doi:10.1038/ncomms10286
Kocer A, Reichmann J, Best D, Adams IR (2009) Germ cell sex determination in mammals. Mol Hum Reprod 15:205–213. doi:10.1093/molehr/gap008
Kramerov DA, Vassetzky NS (2011) SINEs. Wiley Interdiscip Rev RNA 2:772–786. doi:10.1002/wrna.91
Kulpa DA, Moran JV (2006) Cis-preferential LINE-1 reverse transcriptase activity in ribonucleoprotein particles. Nat Struct Mol Biol 13:655–660. doi:10.1038/nsmb1107
Kunarso G, Chia N-Y, Jeyakani J, Hwang C, Lu X, Chan Y-S, Ng H-H, Bourque G (2010) Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat Genet 42:631–634. doi:10.1038/ng.600
Kuramochi-Miyagawa S, Kimura T, Ijiri TW, Isobe T, Asada N, Fujita Y, Ikawa M, Iwai N, Okabe M, Deng W, Lin H, Matsuda Y, Nakano T (2004) Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development 131:839–849. doi:10.1242/dev.00973
Kuramochi-Miyagawa S, Kimura T, Yomogida K, Kuroiwa A, Tadokoro Y, Fujita Y, Sato M, Matsuda Y, Nakano T (2001) Two mouse piwi-related genes: miwi and mili. Mech Dev 108:121–133
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Totoki Y, Toyoda A, Ikawa M, Asada N, Kojima K, Yamaguchi Y, Ijiri TW, Hata K, Li E, Matsuda Y, Kimura T, Okabe M, Sakaki Y, Sasaki H, Nakano T (2008) DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev 22:908–917. doi:10.1101/gad.1640708
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann N, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs RA, Muzny DM, Scherer SE, Bouck JB, Sodergren EJ, Worley KC, Rives CM, Gorrell JH, Metzker ML, Naylor SL, Kucherlapati RS, Nelson DL, Weinstock GM, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Smith DR, Doucette-Stamm L, Rubenfield M, Weinstock K, Lee HM, Dubois J, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang H, Yu J, Wang J, Huang G, Gu J, Hood L, Rowen L, Madan A, Qin S, Davis RW, Federspiel NA, Abola AP, Proctor MJ, Myers RM, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson MV, Kaul R, Raymond C, Shimizu N, Kawasaki K, Minoshima S, Evans GA, Athanasiou M, Schultz R, Roe BA, Chen F, Pan H, Ramser J, Lehrach H, Reinhardt R, McCombie WR, de la Bastide M, Dedhia N, Blöcker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey JA, Bateman A, Batzoglou S, Birney E, Bork P, Brown DG, Burge CB, Cerutti L, Chen HC, Church D, Clamp M, Copley RR, Doerks T, Eddy SR, Eichler EE, Furey TS, Galagan J, Gilbert JG, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang W, Johnson LS, Jones TA, Kasif S, Kaspryzk A, Kennedy S, Kent WJ, Kitts P, Koonin EV, Korf I, Kulp D, Lancet D, Lowe TM, McLysaght A, Mikkelsen T, Moran JV, Mulder N, Pollara VJ, Ponting CP, Schuler G, Schultz J, Slater G, Smit AF, Stupka E, Szustakowski J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf YI, Wolfe KH, Yang SP, Yeh RF, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand KA, Patrinos A, Morgan MJ, de Jong P, Catanese JJ, Osoegawa K, Shizuya H, Choi S, Chen YJ, Szustakowki J, International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921. doi:10.1038/35057062
Lawson KA, Hage WJ (1994) Clonal analysis of the origin of primordial germ cells in the mouse. Ciba Found Symp 182:68–84, discussion 84–91
Leeb M, Wutz A (2007) Ring1B is crucial for the regulation of developmental control genes and PRC1 proteins but not X inactivation in embryonic cells. J Cell Biol 178:219–229. doi:10.1083/jcb.200612127
Lee S-H, Cho S-Y, Shannon MF, Fan J, Rangasamy D (2010) The impact of CpG island on defining transcriptional activation of the mouse L1 retrotransposable elements. PLoS One 5, e11353. doi:10.1371/journal.pone.0011353
Li J, Kannan M, Trivett AL, Liao H, Wu X, Akagi K, Symer DE (2014) An antisense promoter in mouse L1 retrotransposon open reading frame-1 initiates expression of diverse fusion transcripts and limits retrotransposition. Nucleic Acids Res 42:4546–4562. doi:10.1093/nar/gku091
Lim AK, Lorthongpanich C, Chew TG, Tan CWG, Shue YT, Balu S, Gounko N, Kuramochi-Miyagawa S, Matzuk MM, Chuma S, Messerschmidt DM, Solter D, Knowles BB (2013) The nuage mediates retrotransposon silencing in mouse primordial ovarian follicles. Development 140:3819–3825. doi:10.1242/dev.099184
Lu X, Sachs F, Ramsay L, Jacques P-É, Göke J, Bourque G, Ng H-H (2014) The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat Struct Mol Biol 21:423–425. doi:10.1038/nsmb.2799
Lynch MD, Smith AJH, De Gobbi M, Flenley M, Hughes JR, Vernimmen D, Ayyub H, Sharpe JA, Sloane-Stanley JA, Sutherland L, Meek S, Burdon T, Gibbons RJ, Garrick D, Higgs DR (2012) An interspecies analysis reveals a key role for unmethylated CpG dinucleotides in vertebrate Polycomb complex recruitment. EMBO J 31:317–329. doi:10.1038/emboj.2011.399
Macfarlan TS, Gifford WD, Driscoll S, Lettieri K, Rowe HM, Bonanomi D, Firth A, Singer O, Trono D, Pfaff SL (2012) Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487:57–63. doi:10.1038/nature11244
Macia A, Muñoz-Lopez M, Cortes JL, Hastings RK, Morell S, Lucena-Aguilar G, Marchal JA, Badge RM, Garcia-Perez JL (2011) Epigenetic control of retrotransposon expression in human embryonic stem cells. Mol Cell Biol 31:300–316. doi:10.1128/MCB.00561-10
MacLennan M, Crichton JH, Playfoot CJ, Adams IR (2015) Oocyte development, meiosis and aneuploidy. Semin Cell Dev Biol 45:68–76. doi:10.1016/j.semcdb.2015.10.005
Magnúsdóttir E, Surani MA (2014) How to make a primordial germ cell. Development 141:245–252. doi:10.1242/dev.098269
Maksakova IA, Romanish MT, Gagnier L, Dunn CA, van de Lagemaat LN, Mager DL (2006) Retroviral elements and their hosts: insertional mutagenesis in the mouse germ line. PLoS Genet 2:e2. doi:10.1371/journal.pgen.0020002
Malki S, van der Heijden GW, O’Donnell KA, Martin SL, Bortvin A (2014) A role for retrotransposon LINE-1 in fetal oocyte attrition in mice. Dev Cell 29:521–533. doi:10.1016/j.devcel.2014.04.027
Marchetto MCN, Narvaiza I, Denli AM, Benner C, Lazzarini TA, Nathanson JL, Paquola ACM, Desai KN, Herai RH, Weitzman MD, Yeo GW, Muotri AR, Gage FH (2013) Differential L1 regulation in pluripotent stem cells of humans and apes. Nature 503:525–529. doi:10.1038/nature12686
Martin SL, Branciforte D (1993) Synchronous expression of LINE-1 RNA and protein in mouse embryonal carcinoma cells. Mol Cell Biol 13:5383–5392
Mathias SL, Scott AF, Kazazian HH Jr, Boeke JD, Gabriel A (1991) Reverse transcriptase encoded by a human transposable element. Science 254:1808–1810
Matsui T, Leung D, Miyashita H, Maksakova IA, Miyachi H, Kimura H, Tachibana M, Lorincz MC, Shinkai Y (2010) Proviral silencing in embryonic stem cells requires the histone methyltransferase ESET. Nature 464:927–931. doi:10.1038/nature08858
McLaren A (2003) Primordial germ cells in the mouse. Dev Biol 262:1–15
Mi S, Lee X, Li X, Veldman GM, Finnerty H, Racie L, LaVallie E, Tang X-Y, Edouard P, Howes S, Keith JC, McCoy JM (2000) Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature 403:785–789. doi:10.1038/35001608
Molaro A, Falciatori I, Hodges E, Aravin AA, Marran K, Rafii S, McCombie WR, Smith AD, Hannon GJ (2014) Two waves of de novo methylation during mouse germ cell development. Genes Dev 28:1544–1549. doi:10.1101/gad.244350.114
Moran JV, Holmes SE, Naas TP, DeBerardinis RJ, Boeke JD, Kazazian HH Jr (1996) High frequency retrotransposition in cultured mammalian cells. Cell 87:917–927
Muotri AR, Chu VT, Marchetto MCN, Deng W, Moran JV, Gage FH (2005) Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature 435:903–910. doi:10.1038/nature03663
Muotri AR, Marchetto MCN, Coufal NG, Gage FH (2007) The necessary junk: new functions for transposable elements. Hum Mol Genet 16:R159–R167. doi:10.1093/hmg/ddm196
Nichols J, Smith A (2009) Naive and primed pluripotent states. Cell Stem Cell 4:487–492. doi:10.1016/j.stem.2009.05.015
O’Donnell L (2015) Mechanisms of spermiogenesis and spermiation and how they are disturbed. Spermatogenesis 4:e979623. doi:10.4161/21565562.2014.979623
Ohinata Y, Payer B, O’Carroll D, Ancelin K, Ono Y, Sano M, Barton SC, Obukhanych T, Nussenzweig M, Tarakhovsky A, Saitou M, Surani MA (2005) Blimp1 is a critical determinant of the germ cell lineage in mice. Nature 436:207–213. doi:10.1038/nature03813
Öllinger R, Childs AJ, Burgess HM, Speed RM, Lundegaard PR, Reynolds N, Gray NK, Cooke HJ, Adams IR (2008) Deletion of the pluripotency-associated Tex19.1 gene causes activation of endogenous retroviruses and defective spermatogenesis in mice. PLoS Genet 4:e1000199. doi:10.1371/journal.pgen.1000199
Ollinger R, Reichmann J, Adams IR (2010) Meiosis and retrotransposon silencing during germ cell development in mice. Differentiation 79:147–158. doi:10.1016/j.diff.2009.10.004
Peaston AE, Evsikov AV, Graber JH, de Vries WN, Holbrook AE, Solter D, Knowles BB (2004) Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev Cell 7:597–606. doi:10.1016/j.devcel.2004.09.004
Pezic D, Manakov SA, Sachidanandam R, Aravin AA (2014) piRNA pathway targets active LINE1 elements to establish the repressive H3K9me3 mark in germ cells. Genes Dev 28:1410–1428. doi:10.1101/gad.240895.114
Philippe C, Vargas-Landin DB, Doucet AJ, van Essen D, Vera-Otarola J, Kuciak M, Corbin A, Nigumann P, Cristofari G (2016) Activation of individual L1 retrotransposon instances is restricted to cell-type dependent permissive loci. Elife 5:e13926. doi:10.7554/eLife.13926
Piko L, Hammons MD, Taylor KD (1984) Amounts, synthesis, and some properties of intracisternal A particle-related RNA in early mouse embryos. Proc Natl Acad Sci U S A 81:488–492
Popp C, Dean W, Feng S, Cokus SJ, Andrews S, Pellegrini M, Jacobsen SE, Reik W (2010) Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463:1101–1105. doi:10.1038/nature08829
Puech A, Dupressoir A, Loireau MP, Mattei MG, Heidmann T (1997) Characterization of two age-induced intracisternal A-particle-related transcripts in the mouse liver. Transcriptional read-through into an open reading frame with similarities to the yeast ccr4 transcription factor. J Biol Chem 272:5995–6003
Raiz J, Damert A, Chira S, Held U, Klawitter S, Hamdorf M, Löwer J, Strätling WH, Löwer R, Schumann GG (2012) The non-autonomous retrotransposon SVA is trans-mobilized by the human LINE-1 protein machinery. Nucleic Acids Res 40:1666–1683. doi:10.1093/nar/gkr863
Rebollo R, Romanish MT, Mager DL (2012) Transposable elements: an abundant and natural source of regulatory sequences for host genes. Annu Rev Genet 46:21–42. doi:10.1146/annurev-genet-110711-155621
Reddington JP, Perricone SM, Nestor CE, Reichmann J, Youngson NA, Suzuki M, Reinhardt D, Dunican DS, Prendergast JG, Mjoseng H, Ramsahoye BH, Whitelaw E, Greally JM, Adams IR, Bickmore WA, Meehan RR (2013) Redistribution of H3K27me3 upon DNA hypomethylation results in de-repression of Polycomb target genes. Genome Biol 14:R25. doi:10.1186/gb-2013-14-3-r25
Reichmann J, Crichton JH, Madej MJ, Taggart M, Gautier P, Garcia-Perez JL, Meehan RR, Adams IR (2012) Microarray analysis of LTR retrotransposon silencing identifies Hdac1 as a regulator of retrotransposon expression in mouse embryonic stem cells. PLoS Comput Biol 8:e1002486. doi:10.1371/journal.pcbi.1002486
Reichmann J, Reddington JP, Best D, Read D, Öllinger R, Meehan RR, Adams IR (2013) The genome-defence gene Tex19.1 suppresses LINE-1 retrotransposons in the placenta and prevents intra-uterine growth retardation in mice. Hum Mol Genet 22:1791–1806. doi:10.1093/hmg/ddt029
Reuter M, Berninger P, Chuma S, Shah H, Hosokawa M, Funaya C, Antony C, Sachidanandam R, Pillai RS (2011) Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480:264–267. doi:10.1038/nature10672
Ribet D, Dewannieux M, Heidmann T (2004) An active murine transposon family pair: retrotransposition of “master” MusD copies and ETn trans-mobilization. Genome Res 14:2261–2267. doi:10.1101/gr.2924904
Ribet D, Louvet-Vallée S, Harper F, de Parseval N, Dewannieux M, Heidmann O, Pierron G, Maro B, Heidmann T (2008) Murine endogenous retrovirus MuERV-L is the progenitor of the “orphan” epsilon viruslike particles of the early mouse embryo. J Virol 82:1622–1625. doi:10.1128/JVI.02097-07
Romanish MT, Cohen CJ, Mager DL (2010) Potential mechanisms of endogenous retroviral-mediated genomic instability in human cancer. Semin Cancer Biol 20:246–253. doi:10.1016/j.semcancer.2010.05.005
Roovers EF, Rosenkranz D, Mahdipour M, Han C-T, He N, Chuva de Sousa Lopes SM, van der Westerlaken LAJ, Zischler H, Butter F, Roelen BAJ, Ketting RF (2015) Piwi proteins and piRNAs in mammalian oocytes and early embryos. Cell Rep 10:2069–2082. doi:10.1016/j.celrep.2015.02.062
Rossitto M, Philibert P, Poulat F, Boizet-Bonhoure B (2015) Molecular events and signalling pathways of male germ cell differentiation in mouse. Semin Cell Dev Biol 45:84–93. doi:10.1016/j.semcdb.2015.09.014
Rowe HM, Friedli M, Offner S, Verp S, Mesnard D, Marquis J, Aktas T, Trono D (2013) De novo DNA methylation of endogenous retroviruses is shaped by KRAB-ZFPs/KAP1 and ESET. Development 140:519–529. doi:10.1242/dev.087585
Rowe HM, Jakobsson J, Mesnard D, Rougemont J, Reynard S, Aktas T, Maillard PV, Layard-Liesching H, Verp S, Marquis J, Spitz F, Constam DB, Trono D (2010) KAP1 controls endogenous retroviruses in embryonic stem cells. Nature 463:237–240. doi:10.1038/nature08674
Seifarth W, Frank O, Zeilfelder U, Spiess B, Greenwood AD, Hehlmann R, Leib-Mösch C (2005) Comprehensive analysis of human endogenous retrovirus transcriptional activity in human tissues with a retrovirus-specific microarray. J Virol 79:341–352. doi:10.1128/JVI.79.1.341-352.2005
Seisenberger S, Andrews S, Krueger F, Arand J, Walter J, Santos F, Popp C, Thienpont B, Dean W, Reik W (2012) The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol Cell 48:849–862. doi:10.1016/j.molcel.2012.11.001
Shukla R, Upton KR, Muñoz-Lopez M, Gerhardt DJ, Fisher ME, Nguyen T, Brennan PM, Baillie JK, Collino A, Ghisletti S, Sinha S, Iannelli F, Radaelli E, Dos Santos A, Rapoud D, Guettier C, Samuel D, Natoli G, Carninci P, Ciccarelli FD, Garcia-Perez JL, Faivre J, Faulkner GJ (2013) Endogenous retrotransposition activates oncogenic pathways in hepatocellular carcinoma. Cell 153:101–111. doi:10.1016/j.cell.2013.02.032
Slotkin RK, Vaughn M, Borges F, Tanurdzić M, Becker JD, Feijó JA, Martienssen RA (2009) Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136:461–472. doi:10.1016/j.cell.2008.12.038
Smallwood SA, Kelsey G (2012) De novo DNA methylation: a germ cell perspective. Trends Genet 28:33–42. doi:10.1016/j.tig.2011.09.004
Solyom S, Ewing AD, Rahrmann EP, Doucet T, Nelson HH, Burns MB, Harris RS, Sigmon DF, Casella A, Erlanger B, Wheelan S, Upton KR, Shukla R, Faulkner GJ, Largaespada DA, Kazazian HH Jr (2012) Extensive somatic L1 retrotransposition in colorectal tumors. Genome Res 22:2328–2338. doi:10.1101/gr.145235.112
Sookdeo A, Hepp CM, McClure MA, Boissinot S (2013) Revisiting the evolution of mouse LINE-1 in the genomic era. Mob DNA 4:3. doi:10.1186/1759-8753-4-3
Soper SFC, van der Heijden GW, Hardiman TC, Goodheart M, Martin SL, de Boer P, Bortvin A (2008) Mouse maelstrom, a component of nuage, is essential for spermatogenesis and transposon repression in meiosis. Dev Cell 15:285–297. doi:10.1016/j.devcel.2008.05.015
Speek M (2001) Antisense promoter of human L1 retrotransposon drives transcription of adjacent cellular genes. Mol Cell Biol 21:1973–1985. doi:10.1128/MCB.21.6.1973-1985.2001
Sproul D, Nestor C, Culley J, Dickson JH, Dixon JM, Harrison DJ, Meehan RR, Sims AH, Ramsahoye BH (2011) Transcriptionally repressed genes become aberrantly methylated and distinguish tumors of different lineages in breast cancer. Proc Natl Acad Sci U S A 108:4364–4369. doi:10.1073/pnas.1013224108
Sripathy SP, Stevens J, Schultz DC (2006) The KAP1 corepressor functions to coordinate the assembly of de novo HP1-demarcated microenvironments of heterochromatin required for KRAB zinc finger protein-mediated transcriptional repression. Mol Cell Biol 26:8623–8638. doi:10.1128/MCB.00487-06
Stein P, Rozhkov NV, Li F, Cárdenas FL, Davydenko O, Davydenk O, Vandivier LE, Gregory BD, Hannon GJ, Schultz RM (2015) Essential role for endogenous siRNAs during meiosis in mouse oocytes. PLoS Genet 11:e1005013. doi:10.1371/journal.pgen.1005013
Su Y-Q, Sugiura K, Sun F, Pendola JK, Cox GA, Handel MA, Schimenti JC, Eppig JJ (2012a) MARF1 regulates essential oogenic processes in mice. Science 335:1496–1499. doi:10.1126/science.1214680
Su Y-Q, Sun F, Handel MA, Schimenti JC, Eppig JJ (2012b) Meiosis arrest female 1 (MARF1) has nuage-like function in mammalian oocytes. Proc Natl Acad Sci U S A 109:18653–18660. doi:10.1073/pnas.1216904109
Swergold GD (1990) Identification, characterization, and cell specificity of a human LINE-1 promoter. Mol Cell Biol 10:6718–6729
Tam OH, Aravin AA, Stein P, Girard A, Murchison EP, Cheloufi S, Hodges E, Anger M, Sachidanandam R, Schultz RM, Hannon GJ (2008) Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature 453:534–538. doi:10.1038/nature06904
Trelogan SA, Martin SL (1995) Tightly regulated, developmentally specific expression of the first open reading frame from LINE-1 during mouse embryogenesis. Proc Natl Acad Sci U S A 92:1520–1524
Upton KR, Gerhardt DJ, Jesuadian JS, Richardson SR, Sánchez-Luque FJ, Bodea GO, Ewing AD, Salvador-Palomeque C, van der Knaap MS, Brennan PM, Vanderver A, Faulkner GJ (2015) Ubiquitous L1 mosaicism in hippocampal neurons. Cell 161:228–239. doi:10.1016/j.cell.2015.03.026
van den Hurk JAJM, Meij IC, Seleme M d C, Kano H, Nikopoulos K, Hoefsloot LH, Sistermans EA, de Wijs IJ, Mukhopadhyay A, Plomp AS, de Jong PTVM, Kazazian HH, Cremers FPM (2007) L1 retrotransposition can occur early in human embryonic development. Hum Mol Genet 16:1587–1592. doi:10.1093/hmg/ddm108
van Valen L (1973) A new evolutionary law. Evol Theory 1:1–30
Walsh CP, Chaillet JR, Bestor TH (1998) Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat Genet 20:116–117. doi:10.1038/2413
Walter M, Teissandier A, Pérez-Palacios R, Bourc’his D (2016) An epigenetic switch ensures transposon repression upon dynamic loss of DNA methylation in embryonic stem cells. Elife 5:e11418. doi:10.7554/eLife.11418
Wang H, Xing J, Grover D, Hedges DJ, Han K, Walker JA, Batzer MA (2005) SVA elements: a hominid-specific retroposon family. J Mol Biol 354:994–1007. doi:10.1016/j.jmb.2005.09.085
Wang J, Xie G, Singh M, Ghanbarian AT, Raskó T, Szvetnik A, Cai H, Besser D, Prigione A, Fuchs NV, Schumann GG, Chen W, Lorincz MC, Ivics Z, Hurst LD, Izsvák Z (2014) Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516:405–409. doi:10.1038/nature13804
Watanabe T, Totoki Y, Toyoda A, Kaneda M, Kuramochi-Miyagawa S, Obata Y, Chiba H, Kohara Y, Kono T, Nakano T, Surani MA, Sakaki Y, Sasaki H (2008) Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature 453:539–543. doi:10.1038/nature06908
Weismann A (1889) Essays upon heredity. Clarendon Press, Oxford
Wei W, Gilbert N, Ooi SL, Lawler JF, Ostertag EM, Kazazian HH, Boeke JD, Moran JV (2001) Human L1 retrotransposition: cis preference versus trans complementation. Mol Cell Biol 21:1429–1439. doi:10.1128/MCB.21.4.1429-1439.2001
Wildschutte JH, Williams ZH, Montesion M, Subramanian RP, Kidd JM, Coffin JM (2016) Discovery of unfixed endogenous retrovirus insertions in diverse human populations. Proc Natl Acad Sci U S A 113:E2326–E2334. doi:10.1073/pnas.1602336113
Wissing S, Montano M, Garcia-Perez JL, Moran JV, Greene WC (2011) Endogenous APOBEC3B restricts LINE-1 retrotransposition in transformed cells and human embryonic stem cells. J Biol Chem 286:36427–36437. doi:10.1074/jbc.M111.251058
Wolf D, Goff SP (2009) Embryonic stem cells use ZFP809 to silence retroviral DNAs. Nature 458:1201–1204. doi:10.1038/nature07844
Wolf D, Goff SP (2007) TRIM28 mediates primer binding site-targeted silencing of murine leukemia virus in embryonic cells. Cell 131:46–57. doi:10.1016/j.cell.2007.07.026
Yang Q-E, Oatley JM (2014) Spermatogonial stem cell functions in physiological and pathological conditions. Curr Top Dev Biol 107:235–267. doi:10.1016/B978-0-12-416022-4.00009-3
Yotsuyanagi Y, Szöllösi D (1981) Early mouse embryo intracisternal particle: Fourth type of retrovirus-like particle associated with the mouse. J Natl Cancer Inst 67:677–685
Zamudio N, Barau J, Teissandier A, Walter M, Borsos M, Servant N, Bourc’his D (2015) DNA methylation restrains transposons from adopting a chromatin signature permissive for meiotic recombination. Genes Dev 29:1256–1270. doi:10.1101/gad.257840.114
Zamudio N, Bourc’his D (2010) Transposable elements in the mammalian germline: a comfortable niche or a deadly trap? Heredity 105:92–104. doi:10.1038/hdy.2010.53
Acknowledgements
I thank the MRC for funding research in this area in my laboratory through an intramural programme grant from MRC Human Genetics Unit. I am extremely grateful to Chris Playfoot and Jose Luis Garcia-Perez (both MRC Human Genetics Unit, Edinburgh) for their critical comments and excellent suggestions on this chapter.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Adams, I.R. (2017). Retrotransposons and the Mammalian Germline. In: Cristofari, G. (eds) Human Retrotransposons in Health and Disease. Springer, Cham. https://doi.org/10.1007/978-3-319-48344-3_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-48344-3_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-48343-6
Online ISBN: 978-3-319-48344-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)