Abstract
Protein misfolding leading to aggregation and amyloid fibril formation has been implicated in a variety of neurodegenerative disorders. Under suitably designed in vitro conditions, intermolecular contacts between polypeptide chains mediated by various non-covalent interactions result in the formation of oligomeric species that are eventually sequestered into β-sheet-rich amyloid fibrils. Owing to the inherent heterogeneity and complexity of protein aggregation processes, detection and structural characterization of the early, transiently-populated cytotoxic oligomeric intermediates during the amyloid fibrillation cascade still poses a formidable challenge. Fluorescence spectroscopy is an extremely sensitive multiparametric technique that provides simultaneous information about the conformational- and size changes for both early oligomeric species as well as for the large-sized aggregates. In this review, we emphasize recent and selected examples on the application of various fluorescence spectroscopic techniques in the study of protein aggregation. Additionally, we also summarize the recent results on protein aggregation studies using fluorescence spectroscopy from our laboratory.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Fluorescence spectroscopy
- Protein misfolding
- Protein aggregation
- Amyloid fibrils
- Oligomers
- Single-molecule fluorescence
1.1 A Brief Introduction to Protein Misfolding and Aggregation
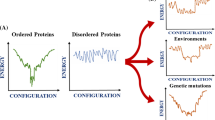
According to the energy landscape theory, an unfolded polypeptide chain folds into its native state by sampling through an ensemble of conformations as described by the protein folding funnel [1]. During the folding process, interplay of a variety of non-covalent interactions such as hydrogen bonding, hydrophobic interactions, electrostatic interactions play key roles in the formation of a correctly folded native structure. However, it has been conjectured that the protein energy landscape is significantly complex accompanied by a “dramatic increase” in its ruggedness due to the localization of the protein in a crowded cellular milieu [2]. Consequently, at higher protein concentrations in the cells, this confinement results in frequent collisions between the polypeptide chains initiating both intra- and intermolecular contacts. Under suitable conditions, when intermolecular interactions predominate over the intramolecular interactions responsible for correct folding, aberrant protein folding occurs that lead to the misfolding and aggregation. Protein aggregation that results in the formation of amyloid fibrils has been implicated in a variety of neurodegenerative disorders (Alzheimer’s, Parkinson’s, Huntington’s and prion diseases), localized and systemic amyloidoses (type II diabetes and dialysis-related, respectively) [3]. The primary agent in protein aggregation and amyloid assembly is identified to be a partially destabilized conformer of a natively folded or a partially stabilized conformer of a natively unfolded protein that accumulates, undergoes conformational changes and ultimately results in the formation of ordered (amyloid) aggregates or self-associate non-specifically to form disordered (amorphous) aggregates depending on the solution conditions namely, pH, ionic strength, temperature etc. [4–6]. The fact that every protein can form aggregates without having a predisposition towards amyloid assembly indicates that the protein aggregation is a generic phenomenon and is likely to be independent of the native structure [7, 8]. However, it has been documented that the amino acid sequence and environmental conditions have a significant impact on the fibrillation kinetics, fibril morphology and architecture along with the fibril stability [9].
It is now recognized that tailoring the solution conditions conducive to protein aggregation in vitro results in the accumulation of a multitude of non-native conformers that serve as precursors to amorphous aggregates and amyloid fibrils . Detection and structural characterization of these non-native oligomeric species in a heterogeneous and complex mixture along with the elucidation of a molecular mechanism, especially during the early stages of protein aggregation still proves to be a formidable task. A number of experimental approaches have been used to detect and characterize the early intermediates since the oligomers have been suggested to be more cytotoxic than the fibrils [10, 11] but an in-depth comprehension of the initial structural details and protein aggregation mechanisms at the molecular level still remain elusive. In this review, we discuss various fluorescence spectroscopic techniques that are utilized to study protein aggregation. Additionally, our efforts directed towards investigation and elucidation of aggregation mechanisms in a few model proteins such as, lysozyme, serum albumins and casein using fluorescence spectroscopy will also be described.
1.2 Experimental Techniques to Study Protein Aggregation
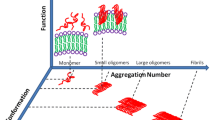
Various biophysical techniques that are used to investigate protein aggregation have indicated that the fibril formation is a stochastic process preceded by the generation of transient species possessing heterogeneous conformations. These transient species, that are eventually sequestered into β-sheet-rich amyloids , have been characterized as oligomeric intermediates that share structural similarities with the amyloid fibrils but are shown to possess higher cytotoxicity than the fibril themselves [10, 11]. Therefore, there is a pressing need to detect and characterize these oligomers that undergo conformational changes and size growth enroute to amyloid fibrillation (Fig. 1.1) . Here, we describe several in vitro spectroscopic techniques very briefly that are either used in isolation or more routinely, in combination with other techniques. Solid-state nuclear magnetic resonance (NMR) [12, 13] and H/D-exchange NMR experiments [14] are extensively used to investigate the structural details of the amyloid fibrils both at the residue-specific as well as at the core levels. Dynamic light scattering (DLS) allows the estimation of the size (hydrodynamic radius) of a protein (and/or protein complexes) [15]. Circular dichroism (CD) is another essential spectroscopic technique that is routinely used to analyze the secondary structural elements in protein monomers and aggregates. As amyloid fibrils are composed of a cross-β sheet-rich structure, CD is commonly utilized to monitor the conformational change from any native conformation (α, α + β etc.) to a predominantly β-sheet amyloid at 215–218 nm. Recently, an extension of CD namely, vibrational circular dichroism (VCD) has been shown to be extremely sensitive towards the detection of amyloid fibrils whereby the inherent chirality of the supramolecular amyloid fibrils is characterized [16, 17]. Another prevalent technique in structural amyloid biology is the X-ray diffraction method [18, 19] that reveals the cross-β structure of amyloids whereby the β-sheets are oriented parallel to the fiber axis but the constituent β-strands are oriented perpendicular to the axis. The X-ray diffraction pattern of amyloid fibrils exhibit two distinct intense reflections, namely, 4.7–4.8 Å meridional and 10 Å equatorial which denote the spacing between the adjacent β-strands and the “face-to-face separation” between the β-sheets, respectively. The X-ray diffraction technique serves as a direct and definitive proof of amyloid fibrils. Additionally, electrospray ionization mass spectrometry (ESI-MS) has been shown to successfully detect various stages of fibril formation whereby the temporarily-formed oligomeric species can be directly observed [20, 21]. Transmission electron microscopy (TEM) and Atomic force microscopy (AFM) are imaging techniques that are used to shed light into the morphologies of the protein aggregates and fibrils on the nanometer length-scale [22, 23]. TEM gives information about the sample surface in two-dimensions and involves staining of the sample to enhance the signal whereas AFM does not require any extraneous markers and provides a three-dimensional nanoscale morphology of aggregates and fibrils.
1.3 Fluorescence Spectroscopy and Protein Aggregation
Fluorescence spectroscopy has proved to be a powerful methodology to extract information about the structural and mechanistic characterization of protein aggregates and amyloid fibrils. The uniqueness of fluorescence spectroscopy lies in the fact that in addition to extreme sensitivity, the protein conformational changes occurring concurrently with size changes can be observed which is very important for monitoring the structural variations occurring during early misfolding and assembly events as well as during the later stages of aggregation (Table 1.1 and Fig. 1.2). Probing the conformational changes in the early oligomeric precursors is particularly useful and important since the fluorescence lifetime of the probes is in the nanosecond timescale which is much faster than the timescales of most conformational changes and interconversions. Information gathered through these observations aid in gaining insights into the complex mechanistic pathway of amyloid formation. Moreover, site-specific fluorophore labeling strategies allow one to selectively label a protein of interest (at ε–amino group of lysine or thiol group of cysteine ; details are discussed later) covalently with a desired fluorescent probe to perform the aggregation studies either in the wild-type protein or in its mutants. The following sections describe the utilization of several oligomer- and amyloid-sensitive fluorescent probes followed by various fluorescence spectroscopic techniques that have exhibited their potential in the systematic investigation of protein aggregation.
1.4 Fluorescence Techniques to Study Protein Oligomers and Amyloids During Protein Aggregation
The advantage of fluorescence spectroscopy also lies in the fact that it is a multiparametric technique . This means that alterations in various parameters e.g. fluorescence intensity, fluorescence anisotropy, spectral shifts, lifetimes etc. of fluorophore(s) can be successfully utilized as distinct structural determinants of the monomeric, oligomeric and large-sized aggregated species of a protein under investigation. These possibilities enable one to monitor a number of different observables from the same reaction mixture populated by an ensemble of structurally diverse molecular species. Such multiparametric data collection offers an unambiguous way to monitor the molecular events in protein aggregation that is commonly endowed with high variability. It is important to mention here that the measurements of fluorescence properties can be carried out either in the steady state or in the time-resolved format. The fluorescence intensity, emission maxima, spectral shifts and fluorescence anisotropy of intrinsic/extrinsic fluorophores can be observed and analyzed conveniently using steady-state fluorescence spectroscopy . In the time-resolved format, it is possible to monitor distributions and subpopulations of molecular species that are on the pathway to amyloid. Additionally, the time-resolved measurements are independent of concentrations of the fluorophores . The time-resolved fluorescence spectroscopy measurements are particularly useful in anisotropy, fluorescence resonance energy transfer (FRET) and quenching experiments. In the following paragraphs, we will discuss briefly about the various fluorescence observables and the usefulness of such parameters in extracting the molecular details during amyloid aggregation . As per the scope of the review, we have restricted to recent results available in the literature with an emphasis on detection of early oligomeric intermediates that are known to be more cytotoxic than the matured fibrils.
1.4.1 Fluorescence Intensity, Spectrum and Lifetime
The fluorescence intensity and lifetime are related to the fluorescence quantum yield and are sensitive to microenvironments [24]. This implies that any change in the protein conformation that affects the surroundings of the intrinsic and/or extrinsic fluorophores, that are sensitive to environmental changes, will be reflected as a change in the fluorescence intensity. Moreover, a shift in the fluorophore emission spectrum is observed depending on whether the fluorophore(s) are exposed or buried due to the protein conformational changes.
1.4.1.1 Intrinsic Fluorophores
Changes in the steady state fluorescence intensity of intrinsic fluorophores such as tryptophan and tyrosine have been commonly used as structural markers in the protein aggregation studies to observe the conformational changes [25–31]. It has been observed that usually, the tryptophan fluorescence intensity increases as the aggregation proceeds forward with a concomitant blue-shift in the emission maximum. Such observations suggest that the average environment around the tryptophan residue(s) progressively becomes non-polar as a function of aggregation. In few cases, a decrease in tryptophan fluorescence intensity has been reported during aggregation suggesting quenching of tryptophan(s) by proximal histidines, phenylalanines, disulfide bonds etc upon protein association [31]. Tyrosine is generally used as an aggregation marker in those cases where the protein is devoid of any tryptophan residue. Though the quantum yield of tyrosine is much lower than that of tryptophan, the absence of tryptophan residues limits any possibility of energy transfer from tyrosine to tryptophan and hence, the fluorescence emission of tyrosine is not quenched. Like tryptophan, the fluorescence intensity of tyrosine is dependent on the polarity of its surroundings but its emission maximum at ~305 nm remains unchanged unless there is a change in pH. An increase in pH results in the formation of tyrosinate ion and a red-shifted emission at ~340 nm. Quenching of the tyrosine fluorescence intensity occurs upon exposure to solvent or due to the presence of aspartic and/or glutamic acid carboxylate side chains in its vicinity [28]. In addition to fluorescence emission from intrinsic tryptophan and tyrosine, an intrinsic blue fluorescence emanating from protein aggregates and fibrils has been observed [32, 33]. As protein aggregates and β-sheet-rich amyloid fibrils comprise of extensive backbone hydrogen bonding networks (>C=O---H-N), it has been suggested that electron delocalization along the peptide backbone due to the presence of these intra- and intermolecular hydrogen bonds give rise to such fluorescence emission and the aromatic side chains do not play any role in the observed blue fluorescence. It was also shown that the fluorescence emission depends largely on the “retention” of water molecules under ambient conditions [33].
1.4.1.2 Extrinsic Fluorophores
A large number of extrinsic fluorophores (either non-covalently bound or covalently attached to proteins) have been used to investigate protein aggregation and amyloid formation (Figs. 1.2 and 1.3) [34]. Generally, thioflavin-T (ThT; amyloid-specific dye) fluorescence assay is routinely used to determine the formation of cross-β-rich amyloid fibrils whereby an increase in the fluorescence intensity of ThT at ~480 nm is observed [35, 36]. It has been reported in a few studies that ThT binding assay is unresponsive to prefibrillar species and the fluorescence enhancement is solely due to the binding of ThT to mature fibrils [37]. Also, ThT binding assay is carried out at neutral or slightly higher pH wherein a few protein amyloids formed at low pH have been reported to dissociate thus, limiting the efficiency of fibril detection [38]. Nile red has been reported as a better amyloid reporter compared to ThT, especially, if the aggregation studies are carried out under acidic conditions [39]. Unbound Nile red emission band exhibits a blue shift with a concomitant increase in the fluorescence intensity upon binding to amyloid fibrils. However, like ThT, Nile red is also more sensitive towards amyloid fibrils compared to the oligomeric species. Therefore, attempts have been made to synthesize new fluorophores to detect the oligomeric species formed at initial stages. DCVJ (4-(dicyanovinyl)-julolidine) [37] and a pentameric oligothiophene derivative (4´,3´´´-bis(carboxymethyl) [2,2´;5´,2´´;5´´,2´´´;5´´´,2´´´´] quinquethiophene-5,5´´´´-dicarboxylic acid; p-FTAA) [40] have been shown to successfully detect and bind efficiently to oligomeric, prefibrillar intermediates even under low pH conditions (Fig. 1.3) [41]. DCVJ is a molecular rotor and an “intrinsically quenched” fluorophore wherein the fluorescence from the julolidine group is quenched by the freely rotating dicyano functionality in solution [37]. When it binds to prefibrillar oligomeric species, the torsional flexibility of the dicyano moiety is lost that results in an increase in DCVJ’s quantum yield. Consequently, a blue-shift in the emission maximum (from ~510 to ~500 nm) of DCVJ is observed with a substantial increase in the fluorescence intensity. The free oligothiophene derivative, p-FTAA, forms π-stacked aggregates in solution which shows a blue-shift of ~100 nm (from 630 to 530 nm) upon binding to prefibrillar species implying a disruption of the p-FTAA π-aggregates [40]. Moreover, binding of p-FTAA to the aggregates causes hindrance in the conformational flexibility of the oligothiophene backbone, thus, affecting the fluorescence emission properties of the dye which can serve as an indirect readout for conformational changes of protein aggregates. Additionally, both the probes are found to be efficient for investigating aggregation kinetics without influencing the rate of amyloid aggregation even at higher concentration of the dye.
In addition to the conjugated fluorescent dyes discussed above, non-covalently bound naphthalene-based dyes such as ANS (1,8-anilinonaphthalene-sulfonate) and bis-ANS (Fig. 1.3) have been utilized in probing prefibrillar and fibrillar species during protein aggregation [30, 31, 37, 42–44]. ANS is weakly fluorescent in aqueous environment but fluoresces strongly (with a concurrent blue-shift in its emission maximum from ~510 to ~475 nm) when located in a hydrophobic environment [45]. Therefore, as expected, an increase in ANS fluorescence intensity accompanied by a blue-shift in the emission is observed upon oligomerization and fibril formation . ANS has also been found to be a good reporter of the aggregation kinetics without perturbing the aggregation rate.
Another strategy of monitoring the emergence of oligomeric species and fibrils during protein aggregation studies involves covalent modification of specific sites of a protein (or its mutants) by a desired fluorophore. The site-specific fluorophore labeling approach offers a major advantage of precisely determining different segments or domains of the polypeptide chain that either participate in forming the hydrophobic interior of aggregates at different stages of fibril formation or remain isolated throughout the whole process. However, caution must be exercised during the external fluorophore labeling procedure to ensure that the secondary structure remains unaltered, especially in the mutant protein, as well as the attached fluorophore should not affect the kinetics of overall aggregation process. The former can be easily checked by collecting CD spectra of both unmodified and covalently-modified protein whereas the latter can be examined by observing any change in fluorescence properties in a concentration-dependent manner by altering the ratio of the fluorescently-labeled to the unlabeled protein. The labeling of ε-amino (−NH2) side chain of lysine by the isothiocyanate (−N=C=S) and thiol (−SH) group of cysteine by the maleimide or iodoacetamide derivatives of fluorescein, rhodamine B, pyrene, acrylodan, Alexa-488, Cy3 etc. are quite common practices. Pyrene is a rigid, polycyclic aromatic hydrocarbon fluorophore whose fluorescence emission spectrum is dependent on the polarity of its micro-environment [46]. In a non-polar medium, the fluorescence emission spectrum of pyrene exhibits five vibronic bands, namely, I1, I2, I3, I4 and I5 at 373 nm, 378 nm, 384 nm, 389 nm and 394 nm, respectively which reduces to three bands in a polar medium (I1, I3 and I5) [47]. A qualitative estimate of the polarity of the pyrene microenvironment can be obtained from the ratio of the fluorescence intensities at I3 and I1. Higher I3/I1 ratio indicates that the environment surrounding the pyrene molecules is more hydrophobic. Additionally, when two pyrene molecules come closer within a distance of ~5 Å, they stack together and form excited state dimer or excimer exhibiting an emission in the range of 450–480 nm. The fluorescence properties of pyrene have been successfully used to detect the formation of oligomers and aggregates whereby a high I3/I1 ratio of pyrene accompanied by a blue-shift in the excimer emission and an enhancement in the excimer intensity are observed [48–50]. Additionally, these covalently bound dyes have proven to be effective in accurate determination of the specific roles of different protein domains (either sequential or concurrent involvement) in the amyloid fibril assembly kinetics [51–53]. Very recently, a new fluorescent probe called MFC, based on 2-(2-furyl)-3-hydroxychromone (maleimide derivative), has been reported to be extremely sensitive towards the detection of oligomers during initial stages of protein aggregation [54]. The probe works on the principles of ESIPT (excited-state intramolecular proton transfer) whereby upon excitation, there is a rapid proton transfer within the molecule (keto-enol tautomerization) at subnanoseconds timescale generating a tautomer. Distinct dual emission profiles are observed from both the “normal” (N*) and the tautomeric form (T*); the latter being red-shifted. Since the probe is environmentally-sensitive, the ratio of emission intensities IT*/IN* is a more convenient way to express the changes in polarity of the microenvironment. An increase in IT*/IN* ratio suggests a reduction in polarity and hydrogen bonding capability of the environment. These properties of the probe have been successfully utilized to monitor the formation of oligomers from α-synuclein wherein a 15-fold rise in the T* emission was observed compared to a 2-fold rise in the N* emission and these changes were detected prior to that observed by a rise in ThT intensity [54]. Using these different fluorescent probes, it is now possible to detect and distinguish between the prefibrillar oligomers and mature amyloid fibrils unambiguously leading to a better understanding of the structural diversity of different species involved in the sequence of events leading to amyloid formation.
1.4.2 Fluorescence Polarization (Anisotropy)
The fluorescence anisotropy measurements offer information about the overall size and the rigidity of the probe attached to biomolecules [24], a marker of the rate of reorientation (tumbling) of the fluorophore(s) (Fig. 1.2). This implies that larger the size, slower is the tumbling, and higher is the anisotropy. The steady-state fluorescence anisotropy is related to the overall size of the protein assuming that the overall size changes are predominant compared to the changes in internal (local) dynamics. The concentration dependence of fluorescence anisotropy indicates oligomer formation prior to fibril formation (Fig. 1.4) [31]. In the fluorescence anisotropy methodology, the fluorophore is excited by a vertically polarized light from a continuous light source following which the emission is collected using parallel and perpendicular geometry of the polarizers at a constant emission wavelength either in L- or in T-format. The steady-state anisotropy is estimated by the ratio of difference between vertically and horizontally polarized light to the total light intensity using the following equation:
The change in fluorescence anisotropy of (a) intrinsic tryptophans and (b) extrinsic Nile red as a function of BSA concentration at pH 3 and 50 mM NaCl at room temperature. All the observations suggest that BSA forms soluble oligomeric aggregates under our experimental condition at which fibrillation experiments were carried out ([BSA] = 100 μM, pH = 3) (Adapted from Ref. [31] with permission from the American Chemical Society)
where III and I⊥ are parallel and perpendicular fluorescence intensities, respectively and the perpendicular components are corrected using corresponding G-factors. A continuous rise in fluorescence anisotropy of intrinsic and/or extrinsic (covalent and non-covalently bound) fluorophore(s) is expected as aggregation progresses suggesting the formation and growth of oligomeric and large-sized aggregates [25, 30, 31, 55]. As mentioned earlier, the steady-state anisotropy gives an average description of the system whereas time-resolved anisotropy can be particularly advantageous in differentiating between the local (residue-specific) and the global dynamics of the fluorophores (Fig. 1.5) [51]. In this technique, the time-dependent fluorescence anisotropy r(t) decay of the fluorescent species at a fixed emission wavelength is monitored upon excitation by vertically polarized laser pulse followed by fitting and analysis of the anisotropy decay in terms of the rotational correlation time φ. Quantitative analysis results in the determination of the rate of rotational motion of the fluorophore by using the following equation:
Time-resolved fluorescence anisotropy decay of IAEDANS-labeled barstar in native form (i), low-pH oligomeric form (ii) and amyloid fibrils (iii). The intrinsic anisotropy (r0) was measured by immobilizing the probe in glycerol. The inset shows the wobbling-in-cone model to illustrate the local and global rotational dynamics of fluorophores covalently attached to a protein (This figure was reproduced with permission from Elsevier (Ref. [51]))
where r0 is the intrinsic fluorescence anisotropy at time zero for a given fluorophore (usually close to 0.4). Incidentally, both local (fast rotation) and global (slow rotation) dynamics of a fluorophore attached to a protein molecule can contribute to the decay of intrinsic fluorescence anisotropy r0 of the fluorophore which can be characterized by the following equation (Fig. 1.5):
where φfast is related to the fluorophore which undergoes fast local rotation. On the other hand, φslow is associated with the global motion of the aggregate which is directly proportional to the overall aggregate size (V) as given by the Stokes–Einstein–Debye equation:
where η is the viscosity and \( \mathrm{V}=4/3{\uppi \mathrm{r}}_{\mathrm{h}}^3 \) where rh is the hydrodynamic radius. βfast denotes the amplitude of the local motion which is a measure of the rotational freedom (or the “degree of orientational constraint”) of the fluorescent probe. Larger the volume of the molecular species, slower will be the anisotropy decay and therefore longer will be the rotational correlation time (φ). The utility of time-resolved fluorescence anisotropy measurements as a tool to detect protein aggregation was demonstrated [56] whereby an external fluorescein label was employed to study the aggregation of amyloid-beta peptide. The time-resolved fluorescence anisotropy measurements have been effectively used to investigate protein aggregation using both intrinsic and extrinsic fluorescent probes in addition to studying the participation of different residues of a protein at various stages of aggregation [26, 51, 57, 58]. In these studies the local dynamics indicated the conformational flexibility/rigidity whereas the global dynamics allowed the estimation of aggregate size. However, the accurate estimation of the size of large aggregates is not possible using nanosecond lifetime probes. One would require to use long lifetime probes to determine the size of large aggregates and fibrils.
1.4.3 Fluorescence Quenching
Fluorescence quenching is a phenomenon by which the fluorescence intensity of a sample decreases due to intermolecular interaction between a fluorophore (intrinsic and/or extrinsic) and a quencher [24]. The measurements on fluorescence quenching provide information about the accessibility of the fluorophore (buried or exposed) to the quencher such as oxygen, iodide, acrylamide etc. which in turn, provide a quantitative estimation of the level of exposure of the fluorophore on the polypeptide chain. Fluorescence quenching can be of two types namely, static and dynamic. In static quenching, the fluorophore and the quencher form a complex which does not fluoresce and the distance between the two is fixed. In dynamic or collisional quenching which is distance-dependent, the quencher diffuses and forms a complex with the fluorophore in the excited-state whereby the fluorophore does not emit any photon upon relaxation to the ground-state. Depending on the fluorescence lifetime of the fluorophore, diffusion can occur even at large distances. Also, by choosing an appropriate quencher, one can selectively quench a fluorophore of interest. Quenching analysis is generally performed by Stern-Volmer plots whereby the fluorescence intensities in the presence and absence of a quencher are collected independently and the ratio of the intensities is plotted as a function of varying quencher concentration from which Stern-Volmer quenching constants are estimated. A high quenching constant would indicate that the fluorophore is exposed to solvent whereas a low quenching constant would imply that the fluorophore is buried inside. There are few reports where fluorescence quenching, in addition to other fluorescence parameters, has been employed in protein aggregation studies [25, 29, 59]. As expected, low quenching efficiency was observed upon protein oligomer and aggregate formation wherein the fluorophore got buried inside the interior of the aggregates. Additionally, based on the quenching experiments, site-specific fluorescent protein mutants gave information about solvent accessibility of various segments of the protein during oligomerization and aggregation [25]. Additionally, internal (inbuilt) amino acid quenchers (Cys, Met, His etc) can also act as conformational reporters of fluorescently labeled proteins.
1.4.4 Fluorescence Resonance Energy Transfer (FRET )
FRET, also known as Förster resonance energy transfer, is a photophysical phenomenon providing a quantitative estimate of the biomolecular distance in the range of 1–10 nm [24]. The FRET technique is used as a reporter of alterations in biomolecular distance as a function of conformational changes and dynamics. It involves energy transfer from a donor (D) fluorophore in the excited state to an acceptor (A) fluorophore in the ground state Consequently, the acceptor emission intensity increases at the expense of donor emission (in steady-state mode) or the donor lifetime decreases (in the time-resolved mode). The FRET efficiency (E) is expressed as:
where R0 is the Förster distance (the distance at which energy transfer efficiency is 50 %) for a given donor-acceptor pair and r is the distance between donor and acceptor whereby the transfer efficiency is inversely proportional to the sixth power of interchromophore distance (r). The Förster distance (R0) is simplistically expressed as:
R0 shows dependence on the following factors: (i) extent of spectral overlap (J(λ)) of the donor emission spectrum with the acceptor absorption spectrum (ii) distance (r) between the donor and acceptor moieties (iii) the donor’s quantum yield (QD) in the absence of acceptor and (iv) relative orientation (κ2) of the donor and acceptor transition dipoles. n is the refractive index which is generally assumed to be 1.4 for biomolecules in aqueous solution and κ2 is assumed to be 2/3 since the rotational motion of the donor and acceptor can “randomize the relative orientation” prior to energy transfer [24]. The FRET technique has been extended to protein aggregation studies, especially to oligomer detection, as reported by several groups. By suitably labeling the protein with donor and acceptor fluorophores, FRET measurements are performed to investigate and monitor conformational fluctuations in the monomeric protein as a consequence of changes in solution conditions conducive to aggregation and oligomeric intermediate formation along with conformational rearrangement preceding amyloid fibrillation in combination with other fluorescence techniques [25, 60, 61]. Earlier efforts in α-synuclein aggregation studies using FRET from tyrosine to tryptophan indicated the presence of early oligomeric species during the lag phase which eventually formed amyloid fibrils. Interestingly, few “partly oligomeric” intermediates were still present in the solution even after the completion of fibril formation [25]. Another study using FRET in confocal microscopy during the investigation of polyglutamine (polyQ) oligomer cytotoxicity revealed that polyQ oligomers are soluble and assemble via “length-dependent manner” inside cells and permeabilize the cell membranes [62]. Very recently, in vivo FRET imaging studies have been carried out in a living multicellular organism C. Elegans to probe α-synuclein aggregation that have shed light into the nature of the aggregated species which were found to be less ordered than the amyloid fibrils and the aggregation kinetics was also monitored [63].
1.4.5 Single-Molecule Fluorescence Studies
Single-molecule fluorescence methodologies are incredibly powerful tools to investigate the complex conformational behavior of aggregation prone amyloidogenic proteins [64, 65]. Using single molecule fluorescence, it is possible to detect, interrogate and analyze individual fluorescently labeled protein or protein complexes. One of us was involved in characterizing natively unfolded or intrinsically disordered yeast prion determinant of Sup35 protein [66]. Single-molecule FRET (SM-FRET) was used to monitor the conformational properties of dual-labeled protein. These experiments indicated that the protein adopts a collapsed conformer under native condition. Additionally, fluorescence correlation spectroscopy (FCS) measurements revealed the presence of nanosecond conformational fluctuations. These results established that the monomeric form of the yeast prion determinant of Sup35 protein adopts an ensemble of relatively unordered collapsed states with rapid conformational fluctuations [66]. Using single-molecule fluorescence, the mechanism of prion propagation has been suggested [67]. Recently, amyloidogenic oligomers have been elegantly detected and characterized by single-molecule fluorescence coincidence [68] and single-molecule photobleaching methods [69].
1.4.6 Fluorescence Correlation Spectroscopy (FCS)
FCS is an extremely sensitive technique which can be performed at nanomolar concentration and is sensitive down to single molecules. The basic principle of this methodology is as follows. As fluorescently-labeled molecules diffuse in-and-out through an illuminated confocal volume (≤1 femtoliter) defined by a focussed laser beam and pinhole, the fluorescence intensity fluctuates spontaneously as a result of the translational diffusion . Statistical analysis of these time-dependent fluctuations in fluorescence intensity using an autocorrelation function yields an autocorrelation fit/curve from which an average number of molecules in the observation volume and the average diffusion coefficient of the molecule are determined [70, 71]. The normalized autocorrelation function for the fluorescence intensity (F(t)) fluctuations is expressed as follows:
which can be further denoted by:
for diffusion-induced intensity fluctuations wherein Veff is the effective volume (illuminated area), 〈C〉 is the average concentration of molecules, Veff〈C〉 = 〈N〉 where N is the average number of molecules, τD is the translational diffusion time of the molecule and is given by:
where r0 is the detection volume and D: diffusion coefficient and r 20 /z 20 is the structure parameter related to the Gaussian observation volume. From the diffusion coefficient (D), the average size (or hydrodynamic radius; rh) of the molecule can be estimated using the following equation: [72, 73]
However, apart from diffusion, fast conformational dynamics and triplet-state dynamics also contribute to the fluctuations in the observed fluorescence intensity which typically occur at faster timescales compared to the diffusion and can be explained by a simple exponential decay function. Hence, in order to incorporate these factors, the autocorrelation function can be rewritten as:
where T is the number of molecules occupying the dark triplet state [74, 75]. As a consequence of fast timescales of these processes, a shoulder is commonly observed in the measured curve. FCS has been widely used to probe protein aggregates and their size distribution either solely or in conjunction with FRET both in solution [61, 65, 76–81] as well as in the cells [82–84]. In a recent report on probing the effect of lipid on tau protein aggregation by FCS at various pH, it was shown that anionic lipid vesicles induce aggregation of the K18 fragment of tau protein when the latter reaches a critical aggregation concentration (CAC) . The CAC was found to increase with an increase in pH of the solution. Both FCS and ThT-assay indicated the presence of tau-lipid co-aggregates and absence of the vesicles did not promote aggregation [78]. FCS has also been successfully applied to monitor inhibition of protein aggregation [85]. In another interesting study, FCS in combination with TEM has been utilized to investigate the size, molecular mass distribution and morphology of Aβ protein aggregates formed by two distinct routes. It was observed that at low pH, Aβ forms a dimer and possibly 12-mer species (spherical intermediates) that serve as precursors to amyloid fibrils whereas it forms trimers prior to the formation of toxic amylospheroids (ASPDs) which do not culminate in amyloid fibrils [79]. In another observation, it was pointed out from diffusion coefficient measurements that the Sup35 protein oligomers (diffused in the yeast cell cytoplasm) are actually fibrillar-shaped instead of being globular-shaped [84]. All the FCS measurements generally involve one-photon excitation of extrinsically-labeled fluorescent species, but recently, two-photon FCS using intrinsic tryptophan has been shown to be efficient in probing the formation of soluble oligomers pertinent to protein aggregation [86]. Additionally, the tryptophan lifetime as a function of aggregation could be monitored simultaneously with two-photon FCS technique. An extension of FCS is FCCS (fluorescence cross-correlation spectroscopy) whereby instead of a single-wavelength-labeled fluorescent species, the temporal fluctuations in fluorescence intensity of dual-wavelength-labeled fluorescent species are observed whereby the two colors are cross-correlated [72, 87] By this technique, one can observe the cross-correlation pattern arising out of “joint fluctuations ” when the dual-colored molecules diffuse through the observation volume simultaneously [88]. The working principles of both FCS and FCCS have been used in image correlation spectroscopy . Raster image correlation spectroscopy (RICS) [73, 82] is one such technique which allows one to generate spatio-temporal autocorrelation curves from the images of the fluorescent species by raster-scanning the laser beam across the surface to acquire the image.
1.5 Case Studies from our Laboratory at IISER Mohali
In this section, we will briefly discuss our efforts directed towards the elucidation of aggregation mechanisms of three classes of model proteins, namely lysozyme (α+β) and bovine serum albumin (all α-helical) and κ-casein (intrinsically disordered protein) using primarily fluorescence spectroscopy in combination with other biophysical techniques [30, 31, 89, 90]. In all these studies, steady-state fluorescence spectroscopy was employed to monitor the simultaneous changes in both conformation and size. Information about the monomeric as well as oligomeric intermediates and early stages of the aggregation process were extracted by monitoring the aggregation kinetics using a variety of fluorescence readouts.
1.5.1 Lysozyme Aggregation
We have recently studied sodium dodecyl sulfate (SDS)-induced aggregation of lysozyme [30]. In our study, we investigated the aggregation kinetics in real time by monitoring the changes in tryptophan, ANS, fluorescein and dansyl fluorescence intensity and anisotropy with an emphasis towards detection and characterization of the early stages involved in the fibril assembly. The negatively-charged SDS, at a concentration (150 μM) much below its critical micellar concentration (8.6 mM), initiates aggregation of lysozyme (at a very low protein concentration ~5 μM) at pH 9.2 at room temperature (Fig. 1.6). During aggregation kinetics experiments, an increase in both fluorescence intensity and anisotropy of all the fluorescent probes were observed as a function of time suggesting conformational- and size changes leading to aggregation. The rate of change in fluorescence intensity of the fluorophores could be fitted to a biexponential function indicating that the conformational change during lysozyme aggregation is a biphasic process whereas the rate of change in fluorescence anisotropy could be satisfactorily explained by a single exponential function indicating an overall size growth. Comparison of the apparent rate constants obtained during intensity and anisotropy kinetics suggested that conformational changes precede the formation of large-sized aggregates (Fig. 1.7). It was proposed that interaction between the positively-charged lysozyme and negatively charged SDS results in conformational expansion with increased hydrophobicity and such conformational expansion facilitates the formation of soluble oligomers.
The time course of ANS fluorescence intensity during lysozyme aggregation upon addition of SDS. A triggering experiment shows that SDS is essential for lysozyme aggregation at pH 9.2 (Adapted from Ref. [30])
(a) The observed rate constants obtained during SDS-induced lysozyme aggregation using various fluorescence readouts indicate that the conformational changes in lysozyme precedes overall size growth of the aggregates. (b) A plausible model of SDS-induced lysozyme aggregation (Adapted from Ref. [30])
1.5.2 Serum Albumin Aggregation
In another study on serum albumin aggregation, we first identified and characterized a ‘molten-globule-like’ intermediate of bovine serum albumin (BSA) at pH 3 that served as an amyloidogenic precursor. Using various fluorescence techniques involving tryptophan fluorescence intensity and anisotropy, ANS fluorescence intensity and anisotropy and pyrene I3/I1 measurements, we established that BSA adopts an expanded conformational state at pH 3 [89]. Figure 1.8 shows the plot of tryptophan fluorescence anisotropy for both bovine and human serum albumins as a function of pH indicating multiple conformational transitions. At pH 3 and elevated temperature, BSA fibrillation was observed [31] in the presence of salt whereby ThT fluorescence assay indicated a dramatic rise in the ThT fluorescence intensity as aggregation proceeded. The fibrillation occurred instantaneously without any lag-phase. A comparative analysis of BSA fibrillation kinetics (monitored by ThT) at various pH indicated that the rate of fibrillation was the fastest at pH 3. After establishing the formation of amyloid fibrils at low pH, attempts were made to extract detailed information about the alterations in conformation and size of the aggregates by observing the rate of changes in fluorescence intensity and anisotropy of both intrinsic (tryptophan) and extrinsic (ANS, IAEDANS) fluorophores as a function of time. Prior to heating, the presence of oligomers was detected by all the fluorescent probes whereby higher fluorescence anisotropy was observed compared to that of the monomeric protein indicating an increase in size (Fig. 1.4). After heating, as the aggregation progressed, the tryptophan fluorescence intensity decreased as a function of time but an increase in the ANS and AEDANS fluorescence intensity with a simultaneous blue-shifted emission were observed. It was suggested that as the hydrophobically associated oligomers accumulated and assembled into amyloid fibrils, the tryptophans were quenched due to proximal amino acid residues such as histidines, phenylalanines and disulfides whereas ANS and AEDANS served as fluorescent markers for enhanced hydrophobicity. The fluorescence anisotropy of all the probes showed a monotonic increase as a function of aggregation suggesting the formation of large-sized aggregates. Both the rate of change in fluorescence intensity and anisotropy of the fluorescent reporters could be fitted to a mono-exponential function to extract the apparent rate constants. Comparison of the rate constants obtained from various probes yielded a unified observation that the conformational conversion in the preformed molten oligomers occurs much faster than the overall size growth. Interestingly, the average rate constant obtained from ThT-fluorescence kinetics was similar to that observed for fluorescence anisotropy, thus, re-confirming the fact that ThT is more sensitive to mature β-sheet-rich amyloid fibrils. It was proposed that the ‘molten-globule-like’ conformer of BSA forms soluble oligomers readily at higher protein concentration assisted by the salt. At an increased temperature, conformational rearrangements occur in these molten oligomers that culminate in fibrils formation (Fig. 1.9).
pH-induced conformational isomerism using intrinsic protein fluorescence: Changes in tryptophan fluorescence anisotropy of conformational isomers of (a) BSA and (b) HSA at room temperature. A molten-globule-like expanded state of serum albumins at pH 3–3.5 is characterized by an enhancement in the tryptophan anisotropy (Adapted from Ref. [89])
(a) The rate constants of BSA fibrillation kinetics of multiple structural probes used to monitor the changes in conformation and size to delineate the steps involved in the fibrillation event. (b) A proposed model for fibril formation from serum albumins based on aggregation kinetics (Reproduced from Ref. [31] with permission from the American Chemical Society)
1.5.3 Conformational Property of κ-Casein – A Model Intrinsically Disordered Protein
Caseins are flexible milk proteins which belong to the class of intrinsically disordered proteins (IDPs) that lack specific secondary structure. They form protein micelles under native conditions. We have used bovine κ-casein as a model IDP to study conformational and aggregation behaviors. Using pyrene fluorescence vibronic band ratio (I3/I1), we show concentration dependent micellar aggregation of native κ-casein in water (Fig. 1.10a). We monomerized it by denaturing followed by disulfide reduction and subsequent carboxymethylation of free cysteines in order to prevent intermolecular disulfide formation. Upon renaturation, carboxymethylated κ-casein showed much lower tryptophan fluorescence anisotropy compared to native κ-casein micelles (Fig. 1.10b). We used monomeric κ-casein as a model protein to study conformational behavior of an IDP. Polypeptide chain collapse of IDPs is believed to play a key role in protein misfolding and amyloid aggregation. Using a variety of fluorescence spectroscopic tools, we have first established that monomeric κ-casein adopts a collapsed ‘pre-molten globule’ like conformers under native condition. We then took the advantage of two free cysteines that are separated by 77-amino acid residues and then covalently labeled the cysteines using thiol-reactive pyrene maleimide. This dual-labeled protein demonstrated a strong excimer formation upon renaturation from urea- and acid-denatured states both under equilibrium and kinetic conditions providing a compelling evidence of polypeptide chain collapse under physiological conditions (Fig. 1.10c) [90]. We believe that our pyrene excimer fluorescence-based methodology will be applied to other IDPs. Currently, investigations of aggregation behavior of IDPs are in progress in our laboratory.
(a) Pyrene I3/I1 as a function of κ-casien concentration. (b) Tryptophan fluorescence anisotropy of micellar and monomeric κ-casien. (c) A schematic of excimer formation upon polypeptide chain collapse of monomeric κ-casien dually labeled with pyrene maleimide (Adapted with permission from Ref. [90])
1.6 Conclusions and Future Directions
In this review, we have provided an overview of protein misfolding and aggregation studies using fluorescence spectroscopy at various levels of sophistication from steady-state fluorimeter measurements to time-resolved and single molecule fluorescence measurements. It is expected that this review will serve as a benchmark to the practitioners in the field of protein misfolding and aggregation. Monitoring different fluorescence readouts during the course of aggregation will allow us to delineate the steps that are involved in the aggregation process. In addition to fluorescence spectroscopic experiments, high-resolution fluorescence microscopy will provide a handle to directly visualize aggregate formation and maturation. Such imaging experiments using a variety of fluorescence microscopy techniques have illuminated intricate structural and mechanistic details of amyloid fibrils. For instance, total internal reflection fluorescence microscopy has been employed to watch fibril growth in real time [67, 91]. Recently, amyloid fibrils have been imaged using near-field scanning optical microscopy technique that is capable of imaging with the nanoscale resolution (beyond the diffraction limit of conventional microscope) [92]. Another exciting development in the field will be to directly watch aggregation and fibril formation in living cells. A recent work describes the methodology to probe fibrillation using FRET imaging in living cells [93]. We anticipate that in the near future, developments in fluorescent probe chemistry, protein engineering and (unnatural) fluorescent amino acid incorporation, cell biology and fluorescence microscopy techniques will further push the boundaries to drive the amyloid biology field in new and exciting directions.
References
Dill KA, Chan HS (1997) From Levinthal to pathways to funnels. Nat Struct Biol 4:10–19
Jahn TR, Radford SE (2008) Folding vs aggregation: polypeptide conformations on competing pathways. Arch Biochem Biophys 469:100–117
Dobson CM (2006) Protein aggregation and its consequences for human disease. Protein Pept Lett 13:219–227
Uversky VN, Fink AL (2004) Conformational constraints for amyloid fibrillation: the importance of being unfolded. Biochim Biophys Acta 1698:131–153
Morel B, Casares S, Conejero-Lara FA (2006) Single mutation induces amyloid aggregation in the α-spectrin SH3 domain: analysis of the early stages of fibril formation. J Mol Biol 356:453–468
Kumar S, Udgaonkar JB (2010) Mechanisms of amyloid fibril formation by proteins. Curr Sci 98:639–656
Guijarro JI, Sunde M, Jones JA, Campbell ID, Dobson CM (1998) Amyloid fibril formation by an SH3 domain. Proc Natl Acad Sci U S A 95:4224–4228
Chiti F, Webster P, Taddei N, Clark A, Stefani M, Ramponi G, Dobson CM (1999) Designing conditions for in vitro formation of amyloid protofilaments and fibrils. Proc Natl Acad Sci U S A 96:3590–3594
Chiti F, Stefani M, Taddei N, Ramponi G, Dobson CM (2003) Rationalization of the effects of mutations on peptide and protein aggregation rates. Nature 424:805–808
Bucciantini M, Giannoni E, Chiti F, Baroni F, Formigli L, Zurdo J, Taddei N, Ramponi G, Dobson CM, Stefani M (2002) Inherent toxicity of aggregates implies a common mechanism for protein misfolding diseases. Nature 416:507–511
Glabe CG (2008) Structural classification of toxic amyloid oligomers. J Biol Chem 283:29639–29643
Tycko R (2006) Molecular structure of amyloid fibrils: insights from solid state NMR. Q Rev Biophys 39:1–55
Sivanandam VN, Jayaraman M, Hoop CL, Kodali R, Wetzel R, van der Wel PCA (2011) The aggregation-enhancing Huntingtin N-terminus is helical in amyloid fibrils. J Am Chem Soc 133:4558–4566
Carulla N, Zhou M, Arimon M, Gairí M, Giralte E, Robinson CV, Dobson CM (2009) Experimental characterization of disordered and ordered aggregates populated during the process of amyloid fibril formation. Proc Natl Acad Sci U S A 106:7828–7833
Fabian H, Gast K, Laue M, Misselwitz R, Uchanska-Ziegler B, Ziegler A, Naumann D (2008) Early stages of misfolding and association of β 2−microglobulin: insights from infrared spectroscopy and dynamic light scattering. Biochemistry 47:6895–6906
Ma S, Cao X, Mak M, Sadik A, Walkner C, Freedman TB, Lednev IK, Dukor RK, Nafie LA (2007) Vibrational circular dichroism shows unusual sensitivity to protein fibril formation and development in solution. J Am Chem Soc 129:12364–12365
Measey TJ, Schweitzer-Stenner R (2011) Vibrational circular dichroism as a probe of fibrillogenesis: the origin of the anomalous intensity enhancement of amyloid-like fibrils. J Am Chem Soc 133:1066–1076
Sunde M, Blake CCF (1998) From the globular to the fibrous state: protein structure and structural conversion in amyloid formation. Q Rev Biophys 31:1–39
Makin OS, Atkins E, Sikorski P, Johansson J, Serpell LC (2005) Molecular basis for amyloid fibril formation and stability. Proc Natl Acad Sci U S A 102:315–320
Larson JL, Ko E, Miranker AD (2000) Direct measurement of islet amyloid polypeptide fibrillogenesis by mass spectrometry. Protein Sci 9:427–431
Smith AM, Jahn TR, Ashcroft AE, Radford SE (2006) Direct observation of oligomeric species formed in the early stages of amyloid fibril formation using electrospray ionisation mass spectrometry. J Mol Biol 364:9–19
Goldsbury C, Baxa U, Simon MN, Steven AC, Engel A, Wall JS, Aebi U, Müller SA (2011) Amyloid structure and assembly: insights from scanning transmission electron microscopy. J Struct Biol 173:1–13
Gosal WS, Myers SL, Radford SE, Thomson NH (2006) Amyloid under the atomic force microscope. Protein Pept Lett 13:261–270
Lakowicz JR (2006) Principles of fluorescence spectroscopy, 3rd edn. Springer, New York
Dusa A, Kaylor J, Edridge S, Bodner N, Hong D-P, Fink AL (2006) Characterization of oligomers during α-synuclein aggregation using intrinsic tryptophan fluorescence. Biochemistry 45:2752–2760
Padrick SB, Miranker AD (2002) Islet amyloid: phase partitioning and secondary nucleation are central to the mechanism of fibrillogenesis. Biochemistry 41:4694–4703
Rolinski OJ, Amaro M, Birch DJS (2010) Early detection of amyloid aggregation using intrinsic fluorescence. Biosens Bioelectron 25:2249–2252
Maji SK, Amsden JJ, Rothschild KJ, Condron MM, Teplow DB (2005) Conformational dynamics of Amyloid β-protein assembly probed using intrinsic fluorescence. Biochemistry 44:13365–13376
van Rooijen BD, van Leijenhorst-Groener KA, Claessens MMAE, Subramaniam V (2009) Tryptophan fluorescence reveals structural features of α-synuclein oligomers. J Mol Biol 394:826–833
Jain N, Bhattacharya M, Mukhopadhyay S (2011) Kinetics of surfactant-induced aggregation of lysozyme studied by fluorescence spectroscopy. J Fluoresc 21:615–625
Bhattacharya M, Jain N, Mukhopadhyay S (2011) Insights into the mechanism of aggregation and fibril formation from bovine serum albumin. J Phys Chem B 115:4195–4205
Shukla A, Mukherjee S, Sharma S, Agrawal V, Radhakishan KV, Guptasarma P (2004) A novel UV laser-induced visible blue radiation from protein crystals and aggregates: scattering artifacts or fluorescence transitions of peptide electrons delocalized through hydrogen bonding? Arch Biochem Biophys 428:144–153
del Mercato LL, Pompa PP, Maruccio G, Torre AD, Sabella S, Tamburro AM, Cingolani R, Rinaldi R (2007) Charge transport and intrinsic fluorescence in amyloid-like fibrils. Proc Natl Acad Sci U S A 104:18019–18024
Hawe A, Sutter M, Jiskoot W (2008) Extrinsic fluorescent dyes as tools for protein characterization. Pharm Res 25:1487–1499
Levine H III (1999) Quantification of β-sheet amyloid fibril structures with thioflavin T. Methods Enzymol 309:274–284
Biancalana M, Koide S (2010) Molecular mechanism of thioflavin-T binding to amyloid fibrils. Biochim Biophys Acta 1804:1405–1412
Lindgren M, Sörgjerd K, Hammarström P (2005) Detection and characterization of aggregates, prefibrillar amyloidogenic oligomers, and protofibrils using fluorescence spectroscopy. Biophys J 88:4200–4212
Platt GW, Radford SE (2009) Glimpses of the molecular mechanisms of β2-microglobulin fibril formation in vitro: aggregation on a complex energy landscape. FEBS Lett 583:2623–2629
Mishra R, Sörgjerd K, Nyström S, Nordigården A, Yu Y-C, Hammarström P (2007) Lysozyme amyloidogenesis is accelerated by specific nicking and fragmentation but decelerated by intact protein binding and conversion. J Mol Biol 366:1029–1044
Hammarström P, Simon R, Nyström S, Konradsson P, Aslund A, Nilsson KPR (2010) A fluorescent pentameric thiophene derivative detects in vitro-formed prefibrillar protein aggregates. Biochemistry 49:6838–6845
Lindgren M, Hammarström P (2010) Amyloid oligomers: spectroscopic characterization of amyloidogenic protein states. FEBS J 277:1380–1388
Bjorndahl TC, Zhou G-P, Liu X, Perez-Pineiro R, Semenchenko V, Saleem F, Acharya S, Bujold A, Sobsey CA, Wishart DS (2011) Detailed biophysical characterization of the acid-induced PrPc to PrPβ conversion process. Biochemistry 50:1162–1173
Giurleo JT, He X, Talaga DS (2008) β-lactoglobulin assembles into amyloid through sequential aggregated intermediates. J Mol Biol 381:1332–1348
Bolognesi B, Kumita JR, Barros TP, Esbjorner EK, Luheshi LM, Crowther DC, Wilson MR, Dobson CM, Favrin G, Yerbury JJ (2010) ANS binding reveals common features of cytotoxic amyloid species. ACS Chem Biol 5:735–740
Daniel E, Weber G (1966) Cooperative effects in binding by bovine serum albumin I: the binding of 1-anilino-8-naphthalenesulfonate fluorimetric titrations. Biochemistry 5:1893–1900
Glushko V, Thaler MSR, Karp CD (1981) Pyrene fluorescence fine structure as a polarity probe of hydrophobic regions: behavior in model solvents. Arch Biochem Biophys 210:33–42
Dong DC, Winnik MA (1982) The Py scale of solvent polarities. Solvent effects on the vibronic fine structure of pyrene fluorescence and empirical correlations with the E T and Y values. Photochem Photobiol 35:17–21
Christensen PA, Pedersen JS, Christiansen G, Otzen DE (2008) Spectroscopic evidence for the existence of an obligate pre-fibrillar oligomer during glucagon fibrillation. FEBS Lett 582:1341–1345
Krishnan R, Lindquist SL (2005) Structural insights into a yeast prion illuminate nucleation and strain diversity. Nature 435:765–772
Thirunavukkuarasu S, Jares-Erijman EA, Jovin TM (2008) Multiparametric fluorescence detection of early stages in the amyloid protein aggregation of pyrene-labeled α-synuclein. J Mol Biol 378:1064–1073
Mukhopadhyay S, Nayak PK, Udgaonkar JB, Krishnamoorthy G (2006) Characterization of the formation of amyloid protofibrils from barstar by mapping residue-specific fluorescence dynamics. J Mol Biol 358:935–942
Sun Y, Breydo L, Makarava N, Yang Q, Bocharova OV, Baskakov IV (2007) Site-specific conformational studies of prion protein (PrP) amyloid fibrils revealed two cooperative folding domains within amyloid structure. J Biol Chem 282:9090–9097
Yap TL, Pfefferkorn CM, Lee JC (2011) Residue-specific fluorescent probes of α-synuclein: detection of early events at the N- and C-termini during fibril assembly. Biochemistry 50:1963–1965
Yushchenko DA, Fauerbach JA, Thirunavukkuarasu S, Jares-Erijman EA, Jovin TM (2011) Fluorescent ratiometric MFC probe sensitive to early stages of α-synuclein aggregation. J Am Chem Soc 132:7860–7861
Koo BW, Miranker AD (2005) Contribution of the intrinsic disulfide to the assembly mechanism of islet amyloid. Protein Sci 14:231–239
Allsop D, Swanson L, Moore S, Davies Y, York A, El-Agnaf OMA, Soutar I (2001) Fluorescence anisotropy: a method for early detection of Alzheimer β-peptide (A-β) aggregation. Biochem Biophys Res Commun 285:58–63
Luk KC, Hyde EG, Trojanowski JQ, Lee VM-Y (2007) Sensitive fluorescence polarization technique for rapid screening of α-synuclein oligomerization/fibrillization inhibitors. Biochemistry 46:12522–12529
Jha A, Udgaonkar JB, Krishnamoorthy G (2009) Characterization of the heterogeneity and specificity of interpolypeptide interactions in amyloid protofibrils by measurement of site-specific fluorescence anisotropy decay kinetics. J Mol Biol 393:735–752
Kayser V, Turton DA, Aggeli A, Beevers A, Reid GD, Beddard GS (2004) Energy migration in novel pH-triggered self-assembled β-sheet ribbons. J Am Chem Soc 126:336–343
Teoh CL, Bekard IB, Asimakis P, Griffin MDW, Ryan TM, Dunstan DE, Howlett GJ (2011) Shear flow induced changes in apolipoprotein C-II conformation and amyloid fibril formation. Biochemistry 50:4046–4057
Nath S, Meuvis J, Hendrix J, Carl SA, Engelborghs Y (2010) Early aggregation steps in α-synuclein as measured by FCS and FRET: evidence for a contagious conformational change. Biophys J 98:1302–1311
Takahashi T, Kikuchi S, Katada S, Nagai Y, Nishizawa M, Onodera O (2008) Soluble polyglutamine oligomers formed prior to inclusion body formation are cytotoxic. Hum Mol Genet 17:345–356
Schierle GSK, Bertoncini CW, Chan FTS, van der Goot AT, Schwedler S, Skepper J, Schlachter S, van Ham T, Esposito A, Kumita JR, Nollen EAA, Dobson CM, Kaminski CF (2011) A FRET sensor for non-invasive imaging of amyloid formation in vivo. Chem Phys Chem 12:673–680
Mukhopadhyay S, Deniz AA (2007) Biomolecular structure and dynamics using fluorescence from single diffusing molecules. J Fluoresc 17:775–783
Hillger F, Nettels D, Dorsch S, Schuler B (2007) Detection and analysis of protein aggregation with confocal single molecule fluorescence spectroscopy. J Fluoresc 17:759–765
Mukhopadhyay S, Krishnan R, Lemke EA, Lindquist S, Deniz AA (2007) A natively unfolded yeast prion monomer adopts an ensemble of collapsed and rapidly fluctuating structures. Proc Natl Acad Sci U S A 104:2649–2654
Collins SR, Douglass A, Vale RD, Weissman JS (2004) Mechanism of prion propagation: amyloid growth occurs by monomer addition. PLoS Biol 2(e321):1582–1590
Orte A, Birkett NR, Clarke RW, Devlin GL, Dobson CM, Klenerman D (2008) Direct characterization of amyloidogenic oligomers by single-molecule fluorescence. Proc Natl Acad Sci U S A 105:14424–14429
Ding H, Wong PT, Lee EL, Gafni A, Steel DG (2009) Determination of the oligomer size of amyloidogenic protein β-Amyloid(1-40) by single-molecule spectroscopy. Biophys J 97:912–921
Magde D, Webb WW, Elson E (1972) Thermodynamic fluctuations in a reacting system-measurement by fluorescence correlation spectroscopy. Phys Rev Lett 29:705–708
Schwille P, Haustein E (2002) Fluorescence correlation spectroscopy: an introduction to its concepts and applications. Biophysics textbook online
Sahoo H, Schwille P (2011) FRET and FCS-friends or foes? Chem Phys Chem 12:532–541
Kolin DL, Wiseman PW (2007) Advances in image correlation spectroscopy: measuring number densities, aggregation states, and dynamics of fluorescently labeled macromolecules in cells. Cell Biochem Biophys 49:141–164
Widengren J, Mets U, Rigler R (1995) Fluorescence correlation spectroscopy of triplet states in solution: a theoretical and experimental study. J Phys Chem 99:13368–13379
Mütze J, Ohrt T, Schwille P (2011) Fluorescence correlation spectroscopy in vivo. Laser Photonics Rev 5:52–67
Tjernberg LO, Pramanik A, Bjijrling S, Thyberg P, Thyberg J, Nordstedt C, Berndt KD, Terenius L, Rigler R (1999) Amyloid β-peptide polymerization studied using fluorescence correlation spectroscopy. Chem Biol 6:53–62
Kitamura A, Kubota H, Pack C-G, Matsumoto G, Hirayama S, Takahashi Y, Kimura H, Kinjo M, Morimoto RI, Nagata K (2006) Cytosolic chaperonin prevents polyglutamine toxicity with altering the aggregation state. Nat Cell Biol 8:1163–1169
Elbaum-Garfinkle S, Ramlall T, Rhoades E (2010) The role of the lipid bilayer in tau aggregation. Biophys J 98:2722–2730
Matsumura S, Shinoda K, Yamada M, Yokojima S, Inoue M, Ohnishi T, Shimada T, Kikuchi K, Masui D, Hashimoto S, Sato M, Ito A, Akioka M, Takagi S, Nakamura Y, Nemoto K, Hasegawa Y, Takamoto H, Inoue H, Nakamura S, Nabeshima Y, Teplow DB, Kinjo M, Hoshi M (2011) Two distinct amyloid β Protein (Aβ) assembly pathways leading to oligomers and fibrils identified by combined fluorescence correlation spectroscopy, morphology, and toxicity analyses. J Biol Chem 286:11555–11562
Sutter M, Oliveira S, Sanders NN, Lucas B, van Hoek A, Hink MA, Visser AJWG, De Smedt SC, Hennink WE, Jiskoot W (2007) Sensitive spectroscopic detection of large and denatured protein aggregates in solution by use of the fluorescent dye Nile red. J Fluoresc 17:181–192
Sengupta P, Garai K, Balaji J, Periasamy N, Maiti S (2003) Measuring size distribution in highly heterogeneous systems with fluorescence correlation spectroscopy. Biophys J 84:1977–1984
Vetri V, Ossato G, Militello V, Digman MA, Leone M, Gratton E (2011) Fluctuation methods to study protein aggregation in live cells: concanavalin A oligomers formation. Biophys J 100:774–783
Takahashi Y, Okamoto Y, Popiel HA, Fujikake N, Toda T, Kinjo M, Nagai Y (2007) Detection of polyglutamine protein oligomers in cells by fluorescence correlation spectroscopy. J Biol Chem 282:24039–24048
Kawai-Noma S, Pack C-G, Kojidani T, Asakawa H, Hiraoka Y, Kinjo M, Haraguchi T, Taguchi H, Hirata A (2010) In vivo evidence for the fibrillar structures of Sup35 prions in yeast cells. J Cell Biol 190:223–231
Ghosh R, Sharma S, Chattopadhyay K (2009) Effect of arginine on protein aggregation studied by fluorescence correlation spectroscopy and other biophysical methods. Biochemistry 48:1135–1143
Sahoo B, Balaji J, Nag S, Kaushalya SK, Maiti S (2008) Protein aggregation probed by two-photon fluorescence correlation spectroscopy of native tryptophan. J Chem Phys 129:0751031–0751035
Hwang LC, Wohland T (2007) Recent advances in fluorescence cross-correlation spectroscopy. Cell Biochem Biophys 49:1–13
Bieschke J, Schwille P (1998) Aggregation of prion protein investigated by dual-color fluorescence cross-correlation spectroscopy. In: Slavik J (ed) Fluorescence microscopy and fluorescent probes, vol 2. Plenum Press, New York
Bhattacharya M, Jain N, Bhasne K, Kumari V, Mukhopadhyay S (2011) pH-induced conformational isomerization of bovine serum albumin studied by extrinsic and intrinsic protein fluorescence. J Fluoresc 21:1083–1090
Jain N, Bhattacharya M, Mukhopadhyay S (2011) Pyrene excimer formation during polypeptide chain collapse of an amyloidogenic intrinsically disordered protein. Biophys J 101:1720–1729
Ban T, Yamaguchi K, Goto Y (2006) Direct observation of amyloid fibril growth, propagation, and adaptation. Acc Chem Res 39:663–670
Kitts CC, Bout DAV (2009) Near-field scanning optical microscopy measurements of fluorescent molecular probes binding to insulin amyloid fibrils. J Phys Chem B 113:12090–12095
Roberti MJ, Bertoncini CW, Klement R, Jares-Erijman EA, Jovin TM (2007) Fluorescence imaging of amyloid formation in living cells by a functional, tetracysteine-tagged α-synuclein. Nat Methods 4:345–351
Acknowledgments
We thank IISER Mohali for financial support and the past and present members of the Mukhopadhyay laboratory for their studies on protein aggregation using fluorescence spectroscopy. M.B. thanks the Department of Science and Technology (DST) Women Scientists’ Scheme and S.M. thanks the Council of Scientific and Industrial Research (CSIR), India for research grants.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Bhattacharya, M., Mukhopadhyay, S. (2016). Studying Protein Misfolding and Aggregation by Fluorescence Spectroscopy. In: Geddes, C. (eds) Reviews in Fluorescence 2015. Reviews in Fluorescence, vol 8. Springer, Cham. https://doi.org/10.1007/978-3-319-24609-3_1
Download citation
DOI: https://doi.org/10.1007/978-3-319-24609-3_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-24607-9
Online ISBN: 978-3-319-24609-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)