Abstract
In many key applications of metabolomics, such as toxicology or nutrigenomics, it is of interest to profile and detect changes in metabolic processes, usually represented in the form of pathways. As an alternative, a broader point of view would enable investigators to better understand the relations between entities that exist in different processes. Therefore, relating a possible perturbation to several known processes represents a new approach to this field of study. We propose to use a network representation of metabolism in terms of reactants, enzymes and metabolites. To model these systems, it is possible to describe both reactions and relations among enzymes and metabolites. In this way, analysis of the impact of changes in some metabolites or enzymes on different processes are easier to understand, detect and predict.
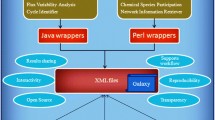
Results. We release the MetaboX library, an open source PHP framework for developing metabolic networks from a set of compounds. This library uses data stored in the Kyoto Encyclopedia for Genes and Genomes (KEGG) database using its RESTful Application Programming Interfaces (APIs), and methods to enhance manipulation of the information retrieved from the KEGG webservice. The MetaboX library includes methods to extract information about a resource of interest (e.g. metabolite, reaction and/or enzyme) and to build reactants network, bipartite enzyme-metabolite and unipartite enzyme networks. These networks can be exported in different formats for data visualization with standard tools. As a case study, the networks built from a subset of the Glycolysis pathway are described and discussed.
Conclusions. The advantages of using such a library imply the ability to model complex systems with few starting information represented by a collection of metabolites KEGG IDs.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
H.K., et al.: Metabolic network modeling and simulation for drug targeting and discovery. Biotechnol J., 30–42 (2011)
Cloots, L., et al.: Network-based functional modeling of genomics, transcriptomics and metabolism in bacteria. Curr Opin Microbiol, 599–607 (2011)
Raosaheb, K., et al.: Uncovering transcriptional regulation of metabolism by using metabolic network topology. PNAS, 2685–2689 (2005)
Krieger, C.J., et al.: MetaCyc: a multiorganism database of metabolic pathways and enzymes. Oxford Journals Nucl. Acids Res, 511–516 (2004)
Keseler, I.M., et al.: EcoCyc: fusing model organism databases with systems biology, Oxford Journals Nucl. Oxford Journals Nucl. Acids Res, D605-D612 (2013)
Wishart, D.S., et al.: HMDB: the Human Metabolome Database. Nucleic Acids Res, D521-D526 (2007)
Sud, M., et al.: LMSD: LIPID MAPS structure database. Oxford Journals Nucl. Acids Res, 527–532 (2007)
Karp, P.D., et al.: Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucl. Acids Res, 6083–6089 (2005)
Matthews, L., et al.: Reactome knowledgebase of human biological pathways and processes. Nucl. Acids Res, D619-D622 (2009)
Wang, Y., et al.: PubChem’s BioAssay Database, Nucl. Nucl. Acids Res, D400-D412 (2012)
Hastings, J., et al.: The ChEBI reference database and ontology for biologically relevant chemistry: enhancements for 2013. Oxford Journals Nucl. Acids Res, 456–463 (2013)
Pence, H.E., et al.: ChemSpider: An Online Chemical Information Resource. J. Chem. Educ., 1123–1124 (2010)
Smith, C.A., et al.: METLIN: A Metabolite Mass Spectral Database Therapeutic Drug Monitoring. In: Proc. of the 9th ICTDM, pp. 747–751 (2005)
Menikarachchi, L.C., et al.: Silico Enzymatic Synthesis of a 400.000 Compound Biochemical Database for Nontargeted Metabolomics. J. Chem. Inf. Model., 2483–2492 (2013)
Kanehisa, M., et al.: KEGG for integration and interpretation of large-scale molecular data sets. Nucl. Acid Res. 14, D109-D114 (2011)
Altman, T., et al.: A systematic comparison of the MetaCyc and KEGG pathway databases. BMC Bioinformatics (2013)
Jitao, D.Z., et al.: KEGGgraph: a graph approach to KEGG PATHWAY in R and bioconductor. Bioinformatics, 1470–1471 (2009)
Xia, J., et al.: MetaboAnalyst 2.0 - a comprehensive server for metabolomic data analysis. Nucl. Acids Res., 1–7 (2012)
Xia, J., et al.: INMEXa web-based tool for integrative meta-analysis of expression data. Nucl. Acids Res., W63-70 (2013)
Mak, T.D., et al.: MetaboLyzer: A Novel Statistical Workflow for Analyzing Postprocessed LCMS Metabolomics Data. Anal. Chem. Article ASAP, 506–513 (2013)
Posma, J.M., et al.: MetaboNetworks, an interactive Matlab-based toolbox for creating, customizing and exploring sub-networks from KEGG. Bioinformatics (2013)
Cline, M.S., et al.: Integration of biological networks and gene expression data using Cytoscape. Nat Protoc., 2366–2382 (2007)
Sharma, A., et al.: Rigidity and flexibility in protein-protein interaction networks: a case study on neuromuscular disorders, arXiv, arXiv:1402.2304v2 (2014)
Franceschini, A., Szklarczyk, D., et al.: STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41, D808–D815 (2013)
Hagberg, A.A., et al.: Exploring Network Structure, Dynamics, and Function using NetworkX. Proc. SciPy, 11-16 (2008)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this paper
Cite this paper
Maiorano, F., Ambrosino, L., Guarracino, M.R. (2015). The MetaboX Library: Building Metabolic Networks from KEGG Database. In: Ortuño, F., Rojas, I. (eds) Bioinformatics and Biomedical Engineering. IWBBIO 2015. Lecture Notes in Computer Science(), vol 9043. Springer, Cham. https://doi.org/10.1007/978-3-319-16483-0_55
Download citation
DOI: https://doi.org/10.1007/978-3-319-16483-0_55
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-16482-3
Online ISBN: 978-3-319-16483-0
eBook Packages: Computer ScienceComputer Science (R0)