Abstract
Nowadays a widely production and low cost of sequencing has allowed the extension of metagenomics in order to explore the genomic information from diverse environments. This offers the opportunity to examine new approaches for sequence binning and functional assignment. Driving of metagenomic studies using high throughput sequencing, usually follows the same pipeline used to analyze single genomes: sequence assembling, gene prediction, functional annotation and phylogenetic classification of reads, contigs or scaffolds, nevertheless, the accuracy of this approach is limited by the length of the reads or resulting contigs.
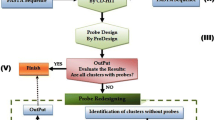
In Silico Hybridization System is an approach of functional and taxonomical assignment. It has default assignment at gender taxonomic level binning. This tool works with two general pipelines: Probes Creator (CrSo), ordered to design DNA probes (fingerprints) to gender taxonomic level and Sequential In silico Hybridator (HISS) which use the probes to make the hybridization with the community reads.
This bioinformatics tool allows characterization of the microbial metabolism in charge of biogeochemical cycles, tracking their key stages using debugged reference information. This strategy resulted in an increasing of binning accuracy. The simulated and real scenarios were better described using one probe and selection threshold fitted to a logarithmic distribution, with mean sensitivity of 85% and mean specificity of 83%.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
References
Altschul, S.F., et al.: Basic local alignment search tool. J. Mol. Biol. 215(3), 403–410 (1990)
Amann, R.I., et al.: Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microb. Rev. 59(1), 143 (1995)
Angly, F.E., et al.: Grinder: a versatile amplicon and shotgun sequence simulator. Nucleic Acids Res. 40(12), 94 (2012)
Darling, A.E., et al.: The design, implementation, and evaluation of mpiBLAST. Presented at the Conference on Linux Clusters: The HPC Revolution 2003 in ClusterWorld Conference & Expo and the 4th International (2003)
Eddy, S.R.: Profile hidden Markov models. Bioinfor. 14(9), 755–763 (1998)
Edgar, R.C.: Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26(19), 2460–2461 (2010)
Foerstner, K.U., et al.: Comparative Analysis of Environmental Sequences: Potential and Challenges. Philosophical Transactions: Biological Sciences 361(1467), 519–523 (2006)
Haque, M., et al.: SOrt-ITEMS: Sequence orthology based approach for improved taxonomic estimation of metagenomic sequences. Bioinformatics 25(14), 1722–1730 (2009)
Heshan Lin et al.: Efficient Data Access for Parallel BLAST. Presented at the 19th IEEE Internat. Parallel and Distributed Processing Symposium (2005)
Hugenholtz, P., Tyson, G.W.: Microbiology: metagenomics (2008)
Huson, D.H., et al.: MEGAN analysis of metagenomic data. Genome Res. 17(3), 377–386 (2007)
Huson, D.H., et al.: Methods for comparative metagenomics. BMC Bioinformatics 10(suppl. 1), S12 (2009)
Kane, M.D., et al.: Assessment of the sensitivity and specificity of oligonucleotide (50mer) microarrays. Nucleic Acids Res. 28(22), 4552–4557 (2000)
Liebich, J., et al.: Improvement of oligonucleotide probe design criteria for functional gene microarrays in environmental applications. Appl. Environ. Microbiol. 72(2), 1688–1691 (2006)
Mavromatis, K., et al.: Use of simulated data sets to evaluate the fidelity of metagenomic processing methods. Nat. Methods 4(6), 495–500 (2007)
Mende, D.R., et al.: Assessment of metagenomic assembly using simulated next generation sequencing data. PLoS ONE 7(2), e31386 (2012)
Moran, M.: Metatranscriptomics: eavesdropping on complex microbial communities. Microbe (2009)
Mouillot, D., et al.: Functional regularity: a neglected aspect of functional diversity. Oecologia 142(3), 353–359 (2004)
Noguchi, H., et al.: MetaGene: prokaryotic gene finding from environmental genome shotgun sequences. Nucleic Acids Res. (2006)
Rho, M., et al.: FragGeneScan: predicting genes in short and error-prone reads. Nucleic Acids Res. 38(20), e191 (2010)
Rotthauwe, J.H., et al.: The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl. Environ. Microbiol. 63(12), 4704–4712 (1997)
Sandberg, R.R., et al.: Capturing whole-genome characteristics in short sequences using a naïve Bayesian classifier. Genes Dev. 11(8), 1404 (2001)
Schloss, P.D., Handelsman, J.: Introducing SONS, a Tool for Operational Taxonomic Unit-Based Comparisons of Microbial Community Memberships and Structures. Appl. Environ. Microbiol. 72(10), 6773–6779 (2006)
Tamura, K., et al.: MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol. Biol. E 28(10), 2731–2739 (2011)
Tringe, S.G., et al.: Comparative metagenomics of microbial communities. Science 308(5721), 554–557 (2005)
Urich, T., et al.: Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome. PLoS ONE 3(6), e2527 (2008)
Vila-Costa, M., et al.: Transcriptomic analysis of a marine bacterial community enriched with dimethylsulfoniopropionate. ISME J., 1–11 (2010)
Wernersson, R., Nielsen, H.B.: OligoWiz 2.0–integrating sequence feature annotation into the design of microarray probes. Nucleic Acids Res. 33(Web Server issue), W611–W615 (2005)
Ye, J., et al.: Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13(1), 134 (2012)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this paper
Cite this paper
Torres-Estupiñan, G.G., Barreto-Hernández, E. (2014). In Silico Hybridization System for Mapping Functional Genes of Soil Microorganism Using Next Generation Sequencing. In: Castillo, L., Cristancho, M., Isaza, G., Pinzón, A., Rodríguez, J. (eds) Advances in Computational Biology. Advances in Intelligent Systems and Computing, vol 232. Springer, Cham. https://doi.org/10.1007/978-3-319-01568-2_48
Download citation
DOI: https://doi.org/10.1007/978-3-319-01568-2_48
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-01567-5
Online ISBN: 978-3-319-01568-2
eBook Packages: EngineeringEngineering (R0)