Abstract
Contending with variability in drug exposure and effect in disease populations requires patient characterization for changes in drug metabolism and transport pathways and predictive modelling platforms within the framework of systems pharmacology. In this chapter, we explore current and emerging patient characterization approaches, the role of physiologically based pharmacokinetic modelling in stratified versus individualized predictions, the possibility of exploring the impact of permutations of comorbidities, and application of these elements in model-informed precision dosing.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Variability
- Drug metabolism and disposition
- In vitro–in vivo
- Extrapolation (IVIVE)
- Physiologically Based Pharmacokinetics (PBPK)
- Quantitative proteomics
- Disease perturbation
1 Introduction
“Variability is the law of life, and as no two faces are the same, so no two bodies are alike, and no individuals react alike and behave alike under the abnormal conditions which we know as disease” – Sir William Osler (1849–1919), Professor of Medicine, Oxford, England
Current drug development mainly focuses on ‘typical’ representation of patients and many of the subtypes involving other comorbidities are studied at later stages after regulatory approval. The latter does not provide any evidence for potential requirements of dosage adjustment that might be necessary for effective and safe medication of patients with certain comorbidities or combinations of these. Hence, in recent years, drug regulatory agencies, as well as professional associations related to drug development and pharmacotherapy, advocated widening the recruitment criteria during clinical studies to provide information on the fate of drugs beyond what is known in a ‘typical patient’. Two recent Guidance for the Industry documents issued by the US Food and Drug Administration (FDA) concerning “Enhancing the Diversity of Clinical Trial Populations — Eligibility Criteria, Enrollment Practices, and Trial Designs” [41] and “Diversity Plans to Improve Enrollment of Participants from Underrepresented Racial and Ethnic Populations in Clinical Trials” [42] are typical examples of such attempts. Although widening recruitment improves gathering of information, and subsequent data analysis can highlight some of the significant changes using sparse samples and non-linear mixed-effect models (so-called population pharmacokinetics or POP-PK), these are not a panacea for the huge lack of data in special populations that may suffer from more than one or two comorbidities. Requesting conduct of clinical studies for every given permutation of concurrent comorbidities places such an act on the verge of impossibility for any drug development entity. However, we cannot leave patients in such special groups without sound and scientific decisions on the best dosage regimen for a given drug and permit off-label use to become the norm for these patients.
The solution may lie within the so-called mechanistic models framing the fate of drugs. These are known as physiologically based pharmacokinetic (PBPK) models and they can accommodate and propagate the physiological and pathological changes in bodily systems to consequences for any given drug if the interplay of parameters with the drug is adequately characterized using in vitro systems. Even in the case of known comorbidities, such as organ impairment (mainly renal or hepatic), which are assessed much more often than other conditions regarding their impact on the fate of drugs, Jadhav et al. [49] reported in 2015 that over 50% of drugs released onto market did not have any information on the impact of severe impairment. These authors as well as others went on to say that PBPK models can fill the void in such conditions. The situation with the void of information on subpopulations has not improved since the report by Jadhav et al. in 2015 as evidenced by our internal unpublished data that demonstrate the case for renal impairment patients (Fig. 6.1). In this chapter, we explore the role of PBPK, in conjunction with existing and emerging patient characterization approaches, in addressing this lack of dosing information for special populations.
The number of FDA-approved drugs without explicit dosing recommendation for patients with renal impairment at the point of entry to the market. The plot shows data (for 2013 and 2014) from Jadhav et al. [49] and unpublished in-house data (for 2015–2019)
2 Accounting for Sources of Interindividual Variability in Pharmacokinetics
Numerous internal and external factors, with complex interplay, affect between-patient variability in drug kinetics (Fig. 6.2); these have an impact on patient physiology, biology, and expression of proteins involved in drug disposition. Other factors unrelated to patient biology, such as compliance, can also add to the apparent PK variability [68]. Quantitative assessment of variability in the fate of drugs in the body due to these factors follows mathematical formalism of pharmacokinetics. This can be in the form of simple equations that describe temporal changes of drug concentration after dosing. Finding the covariates that define interindividual differences in the various model parameters is an a posteriori activity within these models (e.g. using POP-PK methods for sparse samples that employ non-linear mixed-effect methods). However, predicting such individual variations in concentration–time profiles in advance of conducting clinical studies requires more mechanistic models in the form of physiologically based pharmacokinetics (PBPK) [19].
Factors intrinsic and extrinsic to the patient which affect variability in drug exposure and response. Factors are inter-dependent making accounting for their effect challenging. The use of modelling allows prediction of the effect of different permutations of such factors. Abbreviations: PD pharmacodynamics, PK pharmacokinetics
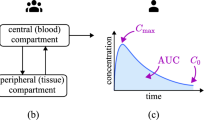
Adequate drug exposure, as defined by the area under the curve of the concentration-time profile (AUC) or either maximum or minimum exposure, respectively defined by the highest concentration (Cmax) or trough concentration (Ctrough), is an essential element of reaching therapeutic response. Together, absorption, distribution, metabolism, and excretion (ADME) of drugs determine the features of the concentration-time profile following drug administration. Data generated from in vitro studies are used to determine and understand ADME variations in different individuals by integration of data with PBPK models [80, 81]. However, the multi-scale nature of these mechanistic models [96] necessitates large efforts and wide expertise to create and verify the model elements, putting PBPK under the framework of systems pharmacology/biology [51] that requires drug-independent systems data, as enlisted below [52].
-
Physiological, anatomical, biological, and biochemical data for each individual (some are defined based on demography, such as ethnicity, sex, age, and environment of the population that an individual belongs to when the actual individual values are not known).
-
Trial design parameters, such as the conditions under which the drug is taken (e.g. fed versus fasted state) or any concomitant drugs interfering with the functions of the systems that handle the drug (e.g. perturbing enzyme expression or function).
The above are combined with drug data (physicochemical properties, e.g. LogP and pKa, drug intrinsic clearance by certain enzymes, affinity to certain transporters) to help not only understand but also predict the behaviour of the drug in certain individuals or a subgroup of patients using a realistic compartmental structure defined as a set of differential equations. Critical considerations are listed below.
-
The factors affecting the variability of the absorption and bioavailability of orally administered drugs are described previously [53]. It is important to note that cytochrome P450 (CYP) 3A and multidrug resistance P-glycoprotein (P-gp), which have wide interindividual variability, are present at high levels in the villi tips of enterocytes in the small intestine [6] and they can cause variations in the bioavailability of drugs, as shown for tacrolimus controlled-release formulation in the case of Afro-Americans versus their Caucasian counterparts [60, 91]. Variations in these proteins as well as other CYP and non-CYP enzymes and transporters in the small intestine are demonstrated in disease states, such as Crohn’s disease [9] and can play a significant role in altering the fate of drugs in such patients [10].
-
Early screening tools can assess the relative importance of the routes of metabolism by various metabolic pathways. Hence, it is now possible to employ information on in vivo intrinsic clearance as well as transporter-mediated uptake to postulate about variability associated with hepatic clearance in human populations [80]. This is facilitated with knowledge of scaling factors [1, 2, 13, 67].
-
Aspects defining variations in renal excretion are also formulated under systems pharmacology [83, 84] and capture the role of urine flow and pH alongside the physical chemistry, lipophilicity, and ionization of the compound that define plasma protein and erythrocyte binding and add knowledge of drug affinity to efflux transporters [85] and abundance of such transporters in human kidney [7].
-
Variability in volume of distribution does not have an impact on overall exposure (as measured by AUC0). However, it defines the shape of the temporal changes of the concentration-time profiles (Cmax and Ctrough); hence, defining/predicting its variability is important. The physical volume of tissues and their blood flows are components of PBPK models that capture population variability related to these parameters. Nonetheless, there are other aspects of the volume of distribution which are more relevant to protein binding in the systemic circulation as well as tissues. Many of these can be measured in vitro and used for in vitro–in vivo extrapolation (IVIVE) purposes through PBPK models [16, 72, 75].
3 The Growing Role of Physiologically Based Pharmacokinetics (PBPK)
In a recent survey, El-Khateeb et al. [37] demonstrated that the first two decades of the twenty-first century have witnessed a more than 40-fold increase in the applications of PBPK (based on the number of publications in the literature). This was in contrast to the general discipline of pharmacokinetics which had a relatively modest increase of around fourfold, in line with an increase in the bulk of scientific publications by threefold. The fastest growing area of PBPK applications according to the survey was focused on addressing alterations to kinetics (or lack thereof) in special populations. Indeed, this was one of the areas that regulatory scientists advocated for the use of PBPK over a decade ago [101] by harnessing the natural compatibility between PBPK and assessment of internal/external factors affecting the kinetics of drugs in various patients.
So, what are the attributes of PBPK that make it so popular with determining the impact of patient variability? The essence of PBPK modelling was described by Rostami-Hodjegan [77] in relation to separation of the system parameters from those of drugs and formulations (Fig. 6.3). Therapeutic effects of a minority of drugs can be monitored relatively easily using established biomarkers (e.g. international normalized ratio, INR, for anticoagulants, blood pressure for antihypertensive agents, and blood glucose for antidiabetic agents) for dose adjustment. However, for most drugs, such effects are not readily measurable or finding out the outcome takes a long time (e.g. patient survival). On the other hand, accounting for drug exposure differences can minimize a large part of the variation in patient outcomes that is related to kinetics. One of the major sources of variability in kinetics is related to interindividual differences in metabolic and transporter-mediated clearance. Clearance and first pass gut and liver metabolism together define internal exposure of the bioavailable dose after entering the gut wall. Many drugs have an optimal therapeutic window for exposure. Whereas for renal clearance, creatinine can be used as a general marker for glomerular filtration as well as active secretion of the drugs into the urine, hepatic clearance does not have a single universal marker that can be applied to all drugs. Characterization of metabolism becomes very important in various groups of patients when we consider that >70% of 698 orally administered marketed drugs have high levels of metabolism as part of their clearance [15]. As shown by Rostami-Hodjegan and Tucker [80], PBPK models can readily incorporate the known variations in drug metabolism and propagate them to projected clearance values using IVIVE techniques. Availability of liver [93, 99], intestinal [9, 33], brain [5, 17, 87], kidney [7, 57], skin [29], and lung [40] tissue for conducting quantitative analysis of proteins related to drug ADME has contributed to advancing a priori understanding of likely differences in kinetics in special populations before conducting any clinical studies. Table 6.1 summarizes prominent examples of such applications. With this approach, and if the baseline in healthy adults (or other control cohorts) is established, it is possible to simulate kinetics in special populations, such as foetal exposure to medications taken by pregnant mothers, or in disease groups, such as patients with hepatic impairment [67]. While availability of human samples for these applications is certainly increasing, access is still restricted by ethical and logistic obstacles, with samples largely limited to post-mortem or surgical surplus tissue.
Separation of parameters related to the drug from parameters related to the population in PBPK models. This approach enables testing different permutations of factors, allowing assessment of changes in PK (or PD) in the target population a priori to conducting clinical studies. Abbreviations: ADME, absorption, distribution, metabolism and excretion; IVIVE, in vitro–in vivo extrapolation; PBPK, physiologically based pharmacokinetics; PD, pharmacodynamics; PK, pharmacokinetics
The principles of PBPK can also be extended to propagation of interindividual variability to drug pharmacodynamics (PD) within the framework of quantitative systems pharmacology [25]. PBPK-PD models are mainly used to predict drug effects in special populations (e.g. predicting dental analgesic effect of ibuprofen in children [30]) and PD effects of drug-drug interactions (DDI) (e.g. the impact of coadministration of domperidone and ketoconazole on QT prolongation in the electrocardiogram of patients [65]). Application of quantitative proteomics to monitoring changes in drug receptors and other PD targets, such as the insulin receptor (INSR) in the human blood–brain barrier [92] and receptor tyrosine kinases in human metastatic liver cancer from colon [95], is expected to facilitate modelling of drug concentration–effect relationships in special/disease populations.
4 Predicting Pharmacokinetics in Subgroups of Patients Versus Predictions in an Individual
Despite advances made in the prediction of changes that occur in pharmacokinetics in subgroups of patients, predicting the fate of drugs in a specific individual who may not be the average patient in his or her subgroup requires characterization of changes that happen in ADME proteins in that particular individual as opposed to the average person in the relevant subgroup. Figure 6.4 summarizes current and emerging characterization methods.
Methods used for the characterization of drug-metabolizing and transporting pathways. Traditionally used methods include genotyping of polymorphic enzymes/transporters, characterization with specific probes (in cocktails administered orally) or the use of endogenous biomarkers for enzyme and transporter activity. More recent methods assess the expression/activity of enzymes and transporters in human samples (either from surgical surplus or post-mortem), in tissue biopsies (from individual patients), or in liquid biopsies (tissue-shedded exosomes). The measurements require modelling platforms for prediction of drug exposure and response. Abbreviations: CB conjugated bilirubin, CPI/III coproporphyrin I and III, CYP cytochrome P450, GCDCA-S glycochenodeoxycholate-3-O-sulphate, MATE1/2 K multidrug and toxin extrusion protein 1 and 2 K, NAT2 N-acetyltransferase 2, NMN N1-methylnicotinamide, OATP1B1/3 organic anion transporting polypeptide 1B1 (gene name SLCO1B1) and 1B3, OAT1/3 organic anion transporter 1 and 3, OCT2 organic cation transporter 2, P-gp P-glycoprotein, TPMT thiopurine methyltransferase, UCB unconjugated bilirubin. Under phenotyping cocktails, superscript numbers indicate the pathways each cocktail can monitor
Genotyping can identify the bracket of the pharmacogenetic subgroup for an individual patient, which is then linked to a specific activity score, such as the case of CYP2D6 genotype [44]. The Clinical Pharmacogenetics Implementation Consortium (CPIC) [23] has published several guideline reports demonstrating the value of such tests in managing optimal dosing for many drugs, e.g. tacrolimus (CYP3A5 genotype), clopidogrel (CYP2C19 genotype), and efavirenz (CYP2B6 genotype) [18, 32, 86]. CPIC guidelines typically offer recommendation of dose adjustment, the use of therapeutic drug monitoring or consideration of alternative therapeutic agents for each genotype group. However, there are wide population variations in the activity of proteins encoded by the same gene, and indeed, some ADME proteins with large population variability in abundance and activity do not have known genotypes that correlate with changes in activity. Hence, endogenous biomarkers and exogenous probes have been used to characterize patients regarding sets of important ADME pathways (e.g. cocktails of drug substrates). Established probe cocktails include the Geneva cocktail (6 enzymes and 1 transporter [20]), the Cooperstown 5 + 1 cocktail (5 enzymes [24]), the Karolinska cocktail (5 enzymes [26]), and the Pittsburgh cocktail (5 enzymes [43]). The issue with these biomarkers is their limited scope which does not cover all relevant pathways of metabolism and transport for the range of clinically used drugs and the specificity of several substrates shows considerable overlap. Whereas tissue proteomics is able to address the quantitative nature of ADME/PD proteins for large sets of targets (a few thousand proteins in the same experiment), obtaining tissue from donors is fraught with ethical and logistic challenges. Hence, the recently introduced possibility of using liquid biopsy offers a more practical alternative for characterization of patients as an input compatible with PBPK models.
Liquid biopsies are biofluids sampled from a patient for diagnostic, companion diagnostic or therapeutic applications. Exosomes shedded by tissue into a biofluid offer a snapshot of the cellular biomolecular pool of macromolecules, which reflect the functional state of their tissue of origin (Fig. 6.5). The vesicles (30–150 nm in size) enclose DNA, (non-coding, messenger and micro) RNA, and (transmembrane and non-membrane) proteins, offering protection from degradation, and therefore longer half-lives of cargo molecules in systemic circulation [21]. ‘Omics’ analysis generates quantitative data for the cargo of extracted exosomes and the levels are linked to the abundance/activity of corresponding proteins in the liver or other organs. Several FDA-approved diagnostic oncology tests rely on liquid biopsy profiling with RNA or DNA sequencing to generate qualitative expression and mutation profiles of batteries of disease markers (e.g. the receptor tyrosine kinases, EGFR and ERBB2) [63]. Integration of quantitative transcriptomic and proteomic analyses into such assays is the next step in the development trajectory of current screening tests towards precision diagnostics and therapeutics. In addition to monitoring ADME proteins (PK variability), liquid biopsy can be used to define between-patient variability in receptors and other therapeutic targets (PD variability). Achour et al. [3, 4] demonstrated the possibility to monitor variability in the expression of over 500 ADME genes (171 enzymes, 362 transporters and the neonatal Fc receptor, FcRn) and over 80 FDA-approved drug targets after appropriate normalization for between-patient differences in the rate of shedding (defined based on expression of a set of tissue-specific stably expressed markers). Although not very well understood, exosome shedding is, in essence, a physiological process that is altered under pathological conditions, adding another parameter that modelling PK variability needs to contend with. Determination of such parameter becomes critical when the patient cohort includes a heterogeneous mix of diseases. The use of tissue-specific cell surface markers can help with purification, by immunoenrichment, of specific extracellular vesicles originating from the tissue of interest, e.g. asialoglycoprotein receptor 1, ASGR1, in the case of liver exosomes [76].
Liquid biopsies, their nature, and attributes. (a) The anatomical origin and level of invasiveness of commonly sampled liquid biopsies (+, least invasive; ++++, most invasive, but all are less invasive than tissue biopsies). (b) Biofluids used to probe ADME/PD protein expression in liver, kidney, lung, and brain tissue as some of the main systems studied in PK/PD research. (c) Blood is the most widely used liquid biopsy with diagnostic, companion diagnostic and therapeutic applications. Tissue (liver) is perfused in blood and continuously sheds microvesicles (exosomes) into the systemic circulation. Molecules shedded include proteins and RNA (of PK and PD targets). The electron micrograph shows exosomes extracted from plasma (size range: 30–150 nm). Abbreviations: ADME absorption, distribution, metabolism and excretion, PD pharmacodynamics, PK pharmacokinetics
With sufficient validation and rapidly declining costs, the use of liquid biopsy will facilitate implementation of model-informed precision dosing owing to the inherent advantages of the technique; it is minimally invasive, quantitative (connecting exosomal profiles to tissue expression), and compatible with modelling platforms, such as Virtual Twins [31, 71]. Virtual Twins should incorporate detailed individual data, such as demographics, genotype, PK/PD expression grades (e.g. from liquid biopsy), and clinical scores (e.g. eGFR for renal function) into a generic PBPK model of the cohort that the individual patient belongs to (Fig. 6.6). The use of liquid biopsy data with such modelling platforms opens the possibility of a priori selection of the optimal initial dose in a treatment regimen for an individual patient and allows identification of patients most likely to experience adverse events or lack of efficacy (for closer therapeutic monitoring). Achour et al. [4] demonstrated correlation with activity in a cohort of patients with cardiovascular disease monitored with the Geneva cocktail (for CYP1A2, CYP2B6, CYP2C9, CYP3A and P-gp), in support of findings by Rowland et al. [82] for CYP3A4 in a set of healthy volunteers before and after induction. Early applications have focused on precision dosing and investigation of DDI potential [3, 76].
The use of a liquid biopsy with modelling platforms for precision therapeutics. (a) Liquid biopsy can be used as a test for grading patients based on quantitative measurement of PK/PD targets while traditional tests (in oncology diagnostics) rely on qualitative evidence of the presence/absence of disease markers and the mutation profiles of such markers. (b) Quantitative data for PK and PD targets from liquid biopsy can be used to generate Virtual Twin models for individualized therapeutics. Abbreviations: PD pharmacodynamics, PK pharmacokinetics
Despite its potential applications, liquid biopsy requires specialist expertise in isolation and purification of exosomes from biofluids, extraction of RNA and protein, and multi-omic analysis (genomics, RNAseq and proteomics). For this reason, the bulk of recent work has focused on assessment of enzymes and transporters in readily accessible systems, such as plasma exosomes [3, 27, 45, 56, 82], while measurements of ADME targets in more challenging biofluids, such as urine [28] and cerebrospinal fluid, are lacking.
5 Modelling the Impact of Permutations of Various Comorbidities
The added value of PBPK becomes paramount when we consider combinations of factors that influence the fate of the drug, which are very difficult, if not impossible, to study in advance of the drug becoming available on the market. We take the example of DDIs as the case here. In 1999, Krayenbühl et al. [55] proposed that interpretation of interaction studies should focus not only on mean DDI effect but also observed and theoretically conceivable extremes. This initiated some efforts within the PBPK community to conduct virtual clinical studies involving large groups of virtual patients where various scenarios could be tested (the platform later became known as the Simcyp Population Based PBPK Platform) [52]. One of the essential elements of the system is its ability to run “what if” scenarios such as those shown in Fig. 6.7. “What if” scenarios take into account factors that affect the outcome of the interaction, e.g. genetics, renal/hepatic impairment, age, or combinations of these elements [79]. It took almost another decade before such facilities were put to practical use by some scientists. In 2012, researchers at the Office of Clinical Pharmacology (OCP) at the US FDA published a PBPK study that verified a previously reported case study [88] on DDI in renal impairment for telithromycin [97]. They went on then to prospectively project on the level of DDI for rivaroxaban in renal impairment where no clinical data were available [46]. The study informed the label for the drug and was a guide to prescribers dealing with these rare occasions. Almost 10 years later, real-world data (RWD) analysis on retrospective information for rivaroxaban and associated side effects clearly demonstrated twofold higher incidence of bleeding in renally impaired patients who were receiving inhibitors of metabolic/transporter clearance (Grillo et al. [47]). The analysis was based on extracts from electronic health records (EHR) from HIPAA-compliant anonymized individual-patient-level data for 117 US institutions in the Cerner-Oracle RWD dataset for a 5-year period (2017–2021). One can postulate that such adverse effects could have been more frequent if the label did not contain the information on the combined impact of renal impairment and DDI. The example above is not unique and there are now many other cases where PBPK information has informed the drug label in the absence of clinical data. Table 6.2 shows a list of examples collated in an internal database by Certara. Similar but less comprehensive lists are published elsewhere [48, 50].
“What if” scenarios simulated with PBPK modelling for a DDI between a substrate (metabolized by CYP1A2 and CYP2D6) and an inhibitor of CYP1A2. Scenarios examined the magnitude of interaction in renal impairment, CYP2D6 poor metabolizer genotype and a combination of the two. Abbreviations: AUCpo area under the plasma concentration-time profile after oral administration, DDI drug-drug interaction. (The concept of the figure is adopted from Rostami-Hodjegan and Tucker [79])
6 Conclusions and Future Use of PBPK for Model-Informed Precision Dosing
While the debate on the nature of PBPK models (Open Source Code versus Closed Source Code) continues [78], the use of closed-source systems has certainly accelerated applications in drug development. Achieving a similar success in model-informed precision dosing faces many hurdles and not just the lack of a user-friendly interface for PBPK. These are discussed by Darwich et al. [31] in the lines of creating virtual twins of patients [71]. However, the first critical step of such efforts is the faithful characterization of patients’ phenotypes beyond genetic-based categorization. It appears that liquid biopsy, in conjunction of omics analyses, may just provide such capacity, if the technical aspects of such game-changing initiatives are addressed [4].
References
Achour B, Barber J, Rostami-Hodjegan A (2014) Expression of hepatic drug-metabolizing cytochrome p450 enzymes and their intercorrelations: a meta-analysis. Drug Metab Dispos 42:1349–1356
Achour B, Rostami-Hodjegan A, Barber J (2014) Protein expression of various hepatic uridine 5′-diphosphate glucuronosyltransferase (UGT) enzymes and their inter-correlations: a meta-analysis. Biopharm Drug Dispos 35:353–361
Achour B, Al-Majdoub ZM, Grybos-Gajnak A et al (2021) Liquid biopsy enables quantification of the abundance and interindividual variability of hepatic enzymes and transporters. Clin Pharmacol Ther 109:222–232
Achour B, Gosselin P, Terrier J et al (2022) Liquid biopsy for patient characterization in cardiovascular disease: verification against markers of cytochrome P450 and P-glycoprotein activities. Clin Pharmacol Ther 111:1268–1277
Al-Majdoub ZM, Al Feteisi H, Achour B et al (2019) Proteomic quantification of human blood-brain barrier SLC and ABC transporters in healthy individuals and dementia patients. Mol Pharm 16:1220–1233
Al-Majdoub ZM, Couto N, Achour B et al (2021) Quantification of proteins involved in intestinal epithelial handling of xenobiotics. Clin Pharmacol Ther 109:1136–1146
Al-Majdoub ZM, Scotcher D, Achour B et al (2021) Quantitative proteomic map of enzymes and transporters in the human kidney: stepping closer to mechanistic kidney models to define local kinetics. Clin Pharmacol Ther 110:1389–1400
Al-Majdoub ZM, Achour B, Couto N et al (2021) Mass spectrometry-based abundance atlas of ABC transporters in human liver, gut, kidney, brain and skin. FEBS Lett 594:4134–4150
Alrubia S, Al-Majdoub ZM, Achour B et al (2022) Quantitative assessment of the impact of Crohn’s disease on protein abundance of human intestinal drug-metabolising enzymes and transporters. J Pharm Sci 111:2917–2929
Alrubia S et al (2022) Altered bioavailability and pharmacokinetics in Crohn’s disease: capturing systems parameters for PBPK to assist with predicting the fate of orally administered drugs. Clin Pharmacokinet 61:1365–1392
Anoshchenko O, Prasad B, Neradugomma NK et al (2020) Gestational age-dependent abundance of human placental transporters as determined by quantitative targeted proteomics. Drug Metab Dispos 48:735–741
Anoshchenko O, Storelli F, Unadkat JD (2021) Successful prediction of human fetal exposure to P-glycoprotein substrate drugs using the proteomics-informed relative expression factor approach and PBPK modeling and simulation. Drug Metab Dispos 49:919–928
Badée J, Achour B, Rostami-Hodjegan A et al (2015) Meta-analysis of expression of hepatic organic anion–transporting polypeptide (OATP) transporters in cellular systems relative to human liver tissue. Drug Metab Dispos 43:424–432
Bao X, Wu J, Xie Y et al (2020) Protein expression and functional relevance of efflux and uptake drug transporters at the blood-brain barrier of human brain and glioblastoma. Clin Pharmacol Ther 107:1116–1127
Benet LZ, Broccatelli F, Oprea TI (2011) BDDCS applied to over 900 drugs. AAPS J 13:519–547
Berezhkovskiy LM (2004) Volume of distribution at steady state for a linear pharmacokinetic system with peripheral elimination. J Pharm Sci 93(6):1628–1640
Billington S, Ray AS, Salphati L et al (2018) Transporter expression in noncancerous and cancerous liver tissue from donors with hepatocellular carcinoma and chronic hepatitis c infection quantified by LC-MS/MS proteomics. Drug Metab Dispos 46:189–196
Birdwell KA, Decker B, Barbarino JM et al (2015) Clinical pharmacogenetics implementation consortium (CPIC) guidelines for CYP3A5 genotype and tacrolimus dosing. Clin Pharmacol Ther 98:19–24
Bois FY, Jamei M, Clewell H (2010) PBPK modelling of inter-individual variability in the pharmacokinetics of environmental chemicals. Toxicology 278(3):256–267
Bosilkovska M, Samer CF, Déglon J et al (2014) Geneva cocktail for cytochrome P450 and P-glycoprotein activity assessment using dried blood spots. Clin Pharmacol Ther 96:349–359
Boukouris S, Mathivanan S (2015) Exosomes in bodily fluids are a highly stable resource of disease biomarkers. Proteomics Clin Appl 9:358–367
Brukner AM, Billington S, Benifla M et al (2021) Abundance of P-glycoprotein and breast cancer resistance protein measured by targeted proteomics in human epileptogenic brain tissue. Mol Pharm 18:2263–2273
Caudle KE, Klein TE, Hoffman JM et al (2014) Incorporation of pharmacogenomics into routine clinical practice: the Clinical Pharmacogenetics Implementation Consortium (CPIC) guideline development process. Curr Drug Metab 15:209–217
Chainuvati S, Nafziger AN, Leeder JS et al (2003) Combined phenotypic assessment of cytochrome P450 1A2, 2C9, 2C19, 2D6, and 3A, N-acetyltransferase-2, and xanthine oxidase activities with the “Cooperstown 5+1 cocktail”. Clin Pharmacol Ther 74:437–447
Chetty M, Rose RH, Abduljalil K et al (2014) Applications of linking PBPK and PD models to predict the impact of genotypic variability, formulation differences, differences in target binding capacity and target site drug concentrations on drug responses and variability. Front Pharmacol 5:258
Christensen M, Andersson K, Dalén P et al (2003) The Karolinska cocktail for phenotyping of five human cytochrome P450 enzymes. Clin Pharmacol Ther 73:517–528
Conde-Vancells J, Rodriguez-Suarez E, Embade N et al (2008) Characterization and comprehensive proteome profiling of exosomes secreted by hepatocytes. J Proteome Res 7:5157–5166
Console L, Scalise M, Tonazzi A et al (2018) Characterization of exosomal SLC22A5 (OCTN2) carnitine transporter. Sci Rep 8:3758–3767
Couto N, Newton JRA, Russo C et al (2021) Label-free quantitative proteomics and substrate based mass spectrometry imaging of xenobiotic metabolizing enzymes in ex vivo human skin and a human living skin equivalent model. Drug Metab Dispos 49:39–52
Cristofoletti R, Dressman JB (2016) Bridging the gap between in vitro dissolution and the time course of ibuprofen-mediating pain relief. J Pharm Sci 105:3658–3667
Darwich AS, Polasek TM, Aronson JK et al (2021) Model-informed precision dosing: background, requirements, validation, implementation, and forward trajectory of individualizing drug therapy. Annu Rev Pharmacol Toxicol 61:225–245
Desta Z, Gammal RS, Gong L et al (2019) Clinical pharmacogenetics implementation consortium (CPIC) guideline for CYP2B6 and efavirenz-containing antiretroviral therapy. Clin Pharmacol Ther 106:726–733
Drozdzik M, Gröer C, Penski J et al (2014) Protein abundance of clinically relevant multidrug transporters along the entire length of the human intestine. Mol Pharm 11:3547–3555
Drozdzik M, Szelag-Pieniek S, Post M et al (2020) Protein abundance of hepatic drug transporters in patients with different forms of liver damage. Clin Pharmacol Ther 107:1138–1148
Drozdzik M, Lapczuk-Romanska J, Wenzel C et al (2021) Gene expression and protein abundance of hepatic drug metabolizing enzymes in liver pathology. Pharmaceutics 13:1334
Drozdzik M, Lapczuk-Romanska J, Wenzel C et al (2022) Protein abundance of drug transporters in human hepatitis C livers. Internat J Mol Sci 23:7947
El-Khateeb E, Burkhill S, Murby S et al (2021) Physiological-based pharmacokinetic modeling trends in pharmaceutical drug development over the last 20-years; in-depth analysis of applications, organizations, and platforms. Biopharm Drug Dispos 42:107–117
El-Khateeb E, Achour B, Al-Majdoub ZM et al (2021) Non-uniformity of changes in drug-metabolizing enzymes and transporters in liver cirrhosis: implications for drug dosage adjustment. Mol Pharm 18:3563–3577
Erdmann P, Bruckmueller H, Martin P et al (2019) Dysregulation of mucosal membrane transporters and drug-metabolizing enzymes in ulcerative colitis. J Pharm Sci 108:1035–1046
Fallon JK, Houvig N, Booth-Genthe CL et al (2018) Quantification of membrane transporter proteins in human lung and immortalized cell lines using targeted quantitative proteomic analysis by isotope dilution nanoLC–MS/MS. J Pharm Biomed Anal 154:150–157
FDA (2020) Enhancing the diversity of clinical trial populations—eligibility criteria, enrollment practices, and trial designs, guidance for industry. Available from: https://www.fda.gov/regulatory-information/search-fda-guidance-documents/enhancing-diversity-clinical-trial-populations-eligibility-criteria-enrollment-practices-and-trial
FDA (2022) Diversity plans to improve enrollment of participants from underrepresented racial and ethnic populations in clinical trials; Draft Guidance for Industry. Available from: https://www.fda.gov/regulatory-information/search-fda-guidance-documents/diversity-plans-improve-enrollment-participants-underrepresented-racial-and-ethnic-populations
Frye RF, Matzke GR, Adedoyin A et al (1997) Validation of the five-drug “Pittsburgh cocktail” approach for assessment of selective regulation of drug-metabolizing enzymes. Clin Pharmacol Ther 62:365–376
Gaedigk A, Simon SD, Pearce RE et al (2008) The CYP2D6 activity score: translating genotype information into a qualitative measure of phenotype. Clin Pharmacol Ther 83:234–242
Gotanda K, Hirota T, Saito J et al (2016) Circulating intestine-derived exosomal miR-328 in plasma, a possible biomarker for estimating BCRP function in the human intestines. Sci Rep 6:32299–32307
Grillo JA, Zhao P, Bullock J et al (2012) Utility of a physiologically-based pharmacokinetic (PBPK) modeling approach to quantitatively predict a complex drug-drug-disease interaction scenario for rivaroxaban during the drug review process: implications for clinical practice. Biopharm Drug Dispos 33:99–110
Grillo J, Zhao P, McNair D (Unpublished) Analysis of Cerner-Oracle RWD dataset for 2017–2021
Huang S-M, Abernethy DR, Wang Y et al (2013) The utility of modeling and simulation in drug development and regulatory review. J Pharm Sci 102:2912–2923
Jadhav PR, Cook J, Sinha V et al (2015) A proposal for scientific framework enabling specific population drug dosing recommendations. J Clin Pharmacol 55:1073–1078
Jamei M (2016) Recent advances in development and application of physiologically-based pharmacokinetic (PBPK) models: a transition from academic curiosity to regulatory acceptance. Curr Pharmacol Rep 2:161–169
Jamei M, Dickinson GL, Rostami-Hodjegan A et al (2009) A framework for assessing inter-individual variability in pharmacokinetics using virtual human populations and integrating general knowledge of physical chemistry, biology, anatomy, physiology and genetics: a tale of ‘bottom-up’ vs ‘top-down’ recognition of covariates. Drug Metab Pharmacokinet 24:53–75
Jamei M, Marciniak S, Feng K et al (2009) The Simcyp® population-based ADME simulator. Expert Opin Drug Metab Toxicol 5:211–223
Jamei M, Turner D, Yang J et al (2009) Population-based mechanistic prediction of oral drug absorption. AAPS J 11:225–237
Jamwal R, de la Monte SM, Ogasawara K et al (2018) Nonalcoholic fatty liver disease and diabetes are associated with decreased cyp3a4 protein expression and activity in human liver. Mol Pharm 15:2621–2632
Krayenbühl JC, Vozeh S, Kondo-Oestreicher M et al (1999) Drug-drug interactions of new active substances: mibefradil example. Eur J Clin Pharmacol 55:559–565
Kumar S, Sinha N, Gerth KA et al (2017) Specific packaging and circulation of cytochromes P450, especially 2E1 isozyme, in human plasma exosomes and their implications in cellular communications. Biochem Biophys Res Commun 491:675–680
Kumar V, Yin J, Billington S et al (2018) The importance of incorporating OCT2 plasma membrane expression and membrane potential in IVIVE of metformin renal secretory clearance. Drug Metab Dispos 46:1441–1445
Kumar V, Yin M, Ishida K et al (2021) Prediction of transporter-mediated rosuvastatin hepatic uptake clearance and drug interaction in humans using proteomics-informed REF approach. Drug Metab Dispos 49:159–168
Kurzawski M, Szelag-Pieniek S, Łapczuk-Romańska J et al (2022) The reference liver-CYP450 and UGT enzymes in healthy donor and metastatic livers: the impact of genotype. Pharmacol Rep 74:204–215
Kuypers DRJ (2018) Tacrolimus formulations and African American kidney transplant recipients: when do details matter? Am J Kidney Dis 71:302–305
Kvitne KE, Hole K, Krogstad V et al (2022) Correlations between 4β-hydroxycholesterol and hepatic and intestinal CYP3A4: protein expression, microsomal ex vivo activity, and in vivo activity in patients with a wide body weight range. Eur J Clin Pharmacol 78:1289–1299
Ladumor MK, Thakur A, Sharma S et al (2019) A repository of protein abundance data of drug metabolizing enzymes and transporters for applications in physiologically based pharmacokinetic (PBPK) modelling and simulation. Sci Rep 9:9709
Lanman RB, Mortimer SA, Zill OA et al (2015) Analytical and clinical validation of a digital sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PLoS One 10:e0140712
Lloret-Linares C, Miyauchi E, Luo H et al (2016) Oral morphine pharmacokinetic in obesity: the role of P-glycoprotein, MRP2, MRP3, UGT2B7, and CYP3A4 jejunal contents and obesity-associated biomarkers. Mol Pharm 13:766–773
Mishra H, Polak S, Jamei M et al (2014) Interaction between domperidone and ketoconazole: toward prediction of consequent QTC prolongation using purely in vitro information. CPT Pharmacometrics Syst Pharmacol 3:e130
Miyauchi E, Tachikawa M, Declèves X et al (2016) Quantitative atlas of cytochrome P450, UDP-glucuronosyltransferase, and transporter proteins in jejunum of morbidly obese subjects. Mol Pharm 13:2631–2640
Neuhoff S, Harwood M, Rostami-Hodjegan A et al (2021) Application of proteomic data in the translation of in vitro observations to associated clinical outcomes. Drug Discov Today Technol 39:13–22
Ogna VF, Bassi I, Menetrey I et al (2017) Comparative long-term effect of three anti-P2Y12 drugs after percutaneous angioplasty: an observational study based on electronic drug adherence monitoring. Front Pharmacol 8:738
Oswald S, Müller J, Neugebauer U et al (2019) Protein abundance of clinically relevant drug transporters in the human kidneys. Int J Mol Sci 20:5303
Peng J, Ladumor MK, Unadkat JD (2022) Estimation of fetal-to-maternal unbound steady-state plasma concentration ratio of p-glycoprotein and/or breast cancer resistance protein substrate drugs using a maternal-fetal physiologically based pharmacokinetic model. Drug Metab Dispos 50:613–623
Polasek TM, Rostami-Hodjegan A (2020) Virtual twins: understanding the data required for model-informed precision dosing. Clin Pharmacol Ther 107:742–745
Poulin P, Theil FP (2002) Prediction of pharmacokinetics prior to in vivo studies. 1. Mechanism-based prediction of volume of distribution. J Pharm Sci 91:129–156
Prasad B, Johnson K, Billington S et al (2016) Abundance of drug transporters in the human kidney cortex as quantified by quantitative targeted proteomics. Drug Metab Dispos 44:1920–1924
Prasad B, Bhatt DK, Johnson K et al (2018) Abundance of phase 1 and 2 drug-metabolizing enzymes in alcoholic and hepatitis C cirrhotic livers: a quantitative targeted proteomics study. Drug Metab Dispos 46:943–952
Rodgers T, Rowland M (2007) Mechanistic approaches to volume of distribution predictions: understanding the processes. Pharm Res 24:918–933
Rodrigues AD, van Dyk M, Sorich MJ et al (2021) Exploring the use of serum-derived small extracellular vesicles as liquid biopsy to study the induction of hepatic cytochromes P450 and organic anion transporting polypeptides. Clin Pharmacol Ther 110:248–258
Rostami-Hodjegan A (2012) Physiologically based pharmacokinetics joined with in vitro-in vivo extrapolation of ADME: a marriage under the arch of systems pharmacology. Clin Pharmacol Ther 92:50–61
Rostami-Hodjegan A, Bois FY (2021) Opening a debate on open-source modeling tools: pouring fuel on fire versus extinguishing the flare of a healthy debate. CPT Pharmacometrics Syst Pharmacol 10:420–427
Rostami-Hodjegan A, Tucker GT (2004) ‘In silico’ simulations to assess the ‘in vivo’ consequences of ‘in vitro’ metabolic drug-drug interactions. Drug Discov Today Technol 1:441–448
Rostami-Hodjegan A, Tucker GT (2007) Simulation and prediction of in vivo drug metabolism in human populations from in vitro data. Nat Rev Drug Discov 6:140–148
Rowland M, Peck C, Tucker G (2011) Physiologically-based pharmacokinetics in drug development and regulatory science. Annu Rev Pharmacol Toxicol 51:45–73
Rowland A, Ruanglertboon W, van Dyk M et al (2019) Plasma extracellular nanovesicle (exosome)-derived biomarkers for drug metabolism pathways: a novel approach to characterize variability in drug exposure. Br J Clin Pharmacol 85:216–226
Scotcher D, Jones C, Posada M et al (2016) Key to opening kidney for in vitro–in vivo extrapolation entrance in health and disease: part I: in vitro systems and physiological data. AAPS J 18:1067–1081
Scotcher D, Jones C, Posada M et al (2016) Key to opening kidney for in vitro-in vivo extrapolation entrance in health and disease: part II: mechanistic models and in vitro-in vivo extrapolation. AAPS J 18:1082–1094
Scotcher D, Jones CR, Galetin A et al (2017) Delineating the role of various factors in renal disposition of digoxin through application of physiologically based kidney model to renal impairment populations. J Pharmacol Exper Ther 360:484–495
Scott SA, Sangkuhl K, Gardner EE et al (2011) Clinical pharmacogenetics implementation consortium guidelines for cytochrome P450-2C19 (CYP2C19) genotype and clopidogrel therapy. Clin Pharmacol Ther 90:328–332
Shawahna R, Uchida Y, Declèves X et al (2011) Transcriptomic and quantitative proteomic analysis of transporters and drug metabolizing enzymes in freshly isolated human brain microvessels. Mol Pharm 8:1332–1341
Shi J, Chapel S, Montay G et al (2005) Effect of ketoconazole on the pharmacokinetics and safety of telithromycin and clarithromycin in older subjects with renal impairment. Int J Clin Pharmacol Ther 43:123–133
Storelli F, Billington S, Kumar AR et al (2021) Abundance of P-glycoprotein and other drug transporters at the human blood-brain barrier in Alzheimer’s disease: a quantitative targeted proteomic study. Clin Pharmacol Ther 109:667–675
Szeląg-Pieniek S, Oswald S, Post M et al (2021) Hepatic drug-metabolizing enzymes and drug transporters in Wilson’s disease patients with liver failure. Pharmacol Rep 73:1427–1438
Trofe-Clark J, Brennan DC, West-Thielke P et al (2018) Results of ASERTAA, a randomized prospective crossover pharmacogenetic study of immediate-release versus extended-release tacrolimus in African American kidney transplant recipients. Am J Kidney Dis 71(3):315–326
Uchida Y, Ohtsuki S, Katsukura Y et al (2011) Quantitative targeted absolute proteomics of human blood–brain barrier transporters and receptors. J Neurochem 117:333–345
Vasilogianni A-M, El-Khateeb E, Al-Majdoub ZM et al (2022) Proteomic quantification of perturbation to pharmacokinetic target proteins in liver disease. J Proteomics 263:104601
Vasilogianni A-M, Al-Majdoub ZM, Achour B et al (2022) Quantitative proteomics of hepatic drug-metabolizing enzymes and transporters in patients with colorectal cancer metastasis. Clin Pharmacol Ther 112:699–710
Vasilogianni A-M, Al-Majdoub ZM, Achour B et al (2022) Proteomics of colorectal cancer liver metastasis: a quantitative focus on drug elimination and pharmacodynamics effects. Br J Clin Pharmacol 88:1811–1823
Vicini P (2010) Multiscale modeling in drug discovery and development: future opportunities and present challenges. Clin Pharmacol Ther 88(1):126–129
Vieira MLT, Zhao P, Berglund EG et al (2012) Predicting drug interaction potential with a physiologically based pharmacokinetic model: a case study of telithromycin, a time-dependent CYP3A inhibitor. Clin Pharmacol Ther 91:700–708
Wang L, Collins C, Kelly EJ et al (2016) Transporter expression in liver tissue from subjects with alcoholic or hepatitis C cirrhosis quantified by targeted quantitative proteomics. Drug Metab Dispos 44:1752–1758
Wegler C, Garcia LP, Klinting S et al (2021) Proteomics-informed prediction of rosuvastatin plasma profiles in patients with a wide range of body weight. Clin Pharmacol Ther 109:762–771
Wegler C, Wiśniewski JR, Robertson I et al (2022) Drug disposition protein quantification in matched human jejunum and liver from donors with obesity. Clin Pharmacol Ther 111:1142–1154
Zhao P, Rowland M, Huang S-M (2012) Best practice in the use of physiologically based pharmacokinetic modeling and simulation to address clinical pharmacology regulatory questions. Clin Pharmacol Ther 92:17–20
Acknowledgements
The authors would like to thank Eleanor Savill for assistance in the preparation and formatting of the chapter and Ellen Leinfuss for providing access to the Certara database of label claims.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Rostami-Hodjegan, A., Achour, B. (2023). On the Verge of Impossibility: Accounting for Variability Arising from Permutations of Comorbidities that Affect the Fate of Drugs in the Human Body. In: Macheras, P. (eds) Advances in Pharmacokinetics and Pharmacodynamics. AAPS Introductions in the Pharmaceutical Sciences, vol 9. Springer, Cham. https://doi.org/10.1007/978-3-031-29541-6_6
Download citation
DOI: https://doi.org/10.1007/978-3-031-29541-6_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-29540-9
Online ISBN: 978-3-031-29541-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)