Abstract
Because of the variety of genetic components, plants are affected by complex genetic-environment-management interactions that drive phenotypic plasticity. Reliable, automatic, multifunctional, and high-throughput phenotypic technology are more taken into consideration as crucial tools for the speedy development of genetic gain in breeding schemes. With the rapid advancement in high-throughput phenotyping (HTP) technology, a study in this aspect is ingoing a novel era called “phenomics.” The crop phenotyping network now no longer most effective needs to construct a multi-domain, multilevel, and multiscale crop phenotyping large database however additionally to analyze technical structures for phenotypic developments identity and increase bioinformatics technology for statistics extraction from the overpowering quantities of omics data. HTP platforms can monitor phenotypic variation in plants using red, green, and blue (RGB) cameras, hyperspectral sensors, and computed tomography are examples of sensors that can be linked to environmental and genotypic data. Since whole-genome sequencing (WGS) and next-generation sequencing (NGS) of many crops has been achieved, functional genomic studies of crops have divided into the main data and the high-throughput era. However, obtaining large-scale phenotypic data has become one of the most significant roadblocks to crop breeding and functional genomics research. Phenomics can give vast amounts of phenotypic data at a minimal cost because it is an inherently big scale and high throughput. Obtaining high-quality digital phenotypic data with detailed information (e.g., descriptions of protocols, growth conditions, etc., in a standardized format) allows for the examination and quantitative modeling of molecular networks influencing complex features including development, stress tolerance, and metabolism as well as community relationships. In this chapter, we will discuss how image-based HTP can be used to bridge the phenotype–genotype gap for yield prediction, root phenotyping, climate-resilient crop development, pathogen and pest detection, and quantitative trait measurement for long-term food security. Finally, we will discuss some conceptual issues and provide suggestions for bridging the phenotype–genotype divide. There is little question that precise HTP will speed up plant genetic advancements and help to usher in the next crop breeding revolution.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction of Phenomics
Phenomics is the study of the phenome with the goal of characterizing phenotypes in a rigorous and formal fashion and linking them to the genes and gene variations that cause them (alleles). The study of plant development, performance, and composition is known as plant phenomics. Phenotyping technologies are used in forward phenomics to “sieve” collections of germplasm for important traits. The sieve or screen may be high-throughput, completely automated, and low-resolution, with higher-resolution, lower-throughput measurements following. Abiotic or biotic stress challenges may be used in screenings, which must be repeatable and physiologically relevant. Reverse phenomics is the detailed dissection of attributes that have been found to be valuable to disclose mechanistic insight and use this mechanism in new techniques. A physiological characteristic may need to be reduced to biochemical or biophysical processes and subsequently to a gene or genes.
High-Throughput Phenomics (HTP)
Phenomics is the study of multidimensional phenotypes with high throughput and correct achievement at different (cell, tissues, organs, individual plant, plots, and field) levels during the developmental stages of a crop. The phenotypic performance of crops is totally dependent upon the interaction between genotypes and environmental factors such as climatic factors, biotic and abiotic factors, and management methods of crops. In other words, phenomics can be defined as the whole study of high dimensional phenotypes. To improve the crop and understand the plant biology, its necessary to obtain data on all main features of phenotype in detail, as shown in Table 6.1.
Phenotypic Technologies
The phenotypic traits can measure exactly through the fast growth of harmless or nontoxic senses and advanced techniques of imaging in which visible, thermal infrared, fluorescence, 3D, and hyperspectral imaging, as well as tomographic imaging using magnetic resonance imaging (MRI) or X-ray computed tomography (CT) are involved. The number of high-throughput phenotyping (HTP) platforms can increase with the help of different technologies like sensing technologies, automatic controlled technologies, computers, robotics, and aeronautics for crop phenotypic traits inquiry. Numerous phenotypic platforms for the traits of the crop at numerous application scales are developed by scientists. In this chapter, there are three types of phenotypic platforms: microscopic, ground-based, and aerial phenotyping platforms split on the bases of imaging levels, which permit the representation of phenotypic traits at the different levels (tissue level, individual plant level, plot level, and field level). For the highly developing field of phenomics and giving rise to an increasing amount and diversity of data, high-throughput technologies are generally used. Turn off the extent data into the beneficial forecast and perception the artificial intelligence (AI) acts as a game changer. Although we need specialized programming skills and deep knowledge about machine learning, deep learning and ensemble learning algorithms are used to understand this artificial intelligence.
HTP Methods
RGB Imaging
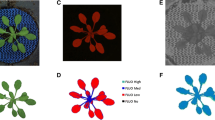
RGB camera or RGB imaging method is mostly used to measure the morphological effects of plants (caused by its cost efficiency and ease of insertion). RGB cameras consist of an infrared blocking filter (VIS camera) that can detect the light wavelength of 400–700 nm instead of customer cameras. For the measurement of the color of every pixel, this camera used different color sensors (red, green, and blue). Scientists concluded that the model’s prediction accuracy could be enhanced by adding other elements such as the growth date (Golzarian et al., 2011) or more anatomical or physiological characteristics in a more complex model (Chen et al., 2014). Because of its cost-effectiveness and ease of installation, the RGB camera approach is the most extensively used technology for measuring plant morphological features. RGB cameras (VIS cameras) include an infrared blocking filter that detects light wavelengths between (400 and 700 nm), unlike consumer cameras. The VIS camera measures through red, green, and blue color sensors as shown in Fig. 6.1.
Near-Infrared Imaging
In different topics, the highest reflectance is in the near-infrared wavelength range (700–1400 nm). This near-infrared imaging (NIR) attribute is employed to confirm plant transformations during drought stress.
Hyperspectral Imaging
Hundreds of thousands of bands per pixel are detected by hyperspectral sensors, which cover the visible (400–700 nm), NIR (700–1000 nm), and SWIR (1000–2500 nm) wavelength ranges. Even though the data’s physical complexity mandates the use of high-performance analytical processors, sensitive detectors are required, and also pictures may be recorded at high resolution with restricted spatial coverage to distinguish reactions to different pressures and large data storage capacity.
Fluorescence Imaging
With the fluorescence sensors, we can easily test the photosynthetic ability of the crop, and it can be tested by the estimation of chlorophyll fluorescence. For example, the excretion of unessential energy by the plant in the form of fluorescence.
Applications of High-Throughput (HTP) Phenotypic (Phenomics)
According to Soulé (1967), the word phenome implies an entire phenotype like genome manifestation in a certain location (Houle et al., 2010; Chen et al., 2014). As a result, a plant phenotype in an agricultural system must be viewed as the outcome of complicated G*E*M interactions (Houle et al., 2010). The word phenomics was coined in 1997 (schork, 1997) and was described as the methodical study of phenotypes at an organism-wide ranging like genomics and the various further omics technologies (Houle et al., 2010). The set of morphological, physiological, and recital-associated features of a genotype in the environment is known as the phenotype (Dhondt et al., 2013). In other disputes, phenomic is an inclusive wide-ranging study of high-dimensional phenotypes that is vital for the generation of meticulous data on all important aspects of phenotypes and for an improved understanding of plant biology and crop improvement. Therefore, the phenotyping system not only includes tools for performing phenotypes on its own but also plants in a specific environment, from the tightly controlled condition of the climatic chamber to the natural environment of the field it also means to grow (Dhont et al., 2013). The throughput of a plant phenotyping system relates to the number of individual units at a given organization. Steven A., a UC Berkeley and LBNL scientist, invented the term phenomics to describe the scientific study of phenotypes. As a result, it is a multidisciplinary field of study that includes biology, data science, engineering, and other disciplines.
Phenotypic Technologies
Multidisciplinary collaboration and some of the initial developments were targeted toward assessing genetically modified crops on a large scale (Reuzeau et al., 2005). Plant refurbishment approaches, Trait Mill, a suite of proprietary bioinformatics tools, a high-throughput gene engineering system, Crop Design (Belgium) established a HTP stage that was utilized to detect morphometric characteristics. (aboveground biomass, plant shape, and plant color) that might have an impact on yield. TraitMill’s details, as well as the trial methodology and results, were inappropriately kept confidential (Reuzeau et al., 2005).
High-Throughput Phenomic Methods
-
Scanalyzer 3D platform:
Hairmansis et al. (2014) and Neilson et al. (2015) reported that the Scanalyzer 3D platform was created by Lemna Tec in Germany, and it has been implemented in numerous countries. Computer-controlled conveyor systems are installed at the Plant Accelerator (Australian Plant Phenomics Facility, University of Adelaide, Australia), automated weighing–watering devices, imaging stations, Near-infrared (NIR), fluorescence (at near-infrared wavelengths between 700 and 1400 nm, the green portions of plants had the maximum reflectivity), and Hyperspectral imaging (hundreds of thousands of bands per pixel are detected using hyperspectral sensors, which cover the visible spectrum) (Mathieu et al., 2015).
RADIX Imaging Marié et al. (2016) and Jeudy et al. (2016) examined the RhizoTubes (an automated “plant-to-sensor” platform including 1200 rhizo tubes to acquire the RSA in about 6–8 weeks) and RhizoSlide (a rhizoslide platform used to screen the shoots and roots of 200 maize plants) (Le Marié et al., 2016).
Quantitative Plant Morphology Detection Through Phenomics
To maintain future food security, it is critical to developing crop tolerance to abiotic stresses and new pests brought on by climate change. The growing use of gene editing, as well as the continuous utilization of natural genetic diversity, present excellent prospects for producing novel alleles and selecting natural sources of genetic variation for crop development. This necessitates the examination of hundreds of lines growing in a variety of environments. At the same time, breakthroughs in DNA marker assays and sequencing technology have enabled genotyping to achieve this throughput at a reasonable cost, similar innovations provide an urgent demand for high-throughput and meaningful phenotypic data. The purpose of plant phenomics, which we describe as the study of the development of plants, is to achieve this and implementation of a set of tools and methodologies that are used to achieve three key objectives—gathering data on the structures, functions, and performances of huge groups of plants, as well as their surroundings; analyzing, organizing, and storing the generated datasets; and constructing models that can untangle and recreate plant activity in a variety of settings. Plant phenomics has advanced significantly in the last decade, with new sensors and imaging approaches being developed for a variety of features, organs, and conditions. When it comes to turning sensor data into knowledge, however, data handling and processing remain significant obstacles (Tardieu et al., 2017).
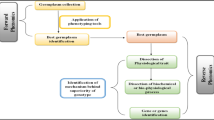
Plant phenotyping is the study of complex plant characteristics such as growth, development, tolerance, resistance, architecture, physiology, ecology, and yield, as well as the fundamental assessment of quantitative parameters that serve as the foundation for more sophisticated aspects. Photosynthetic efficiency, root shape, biomass, leaf features, fruit traits, and yield-related aspects direct measuring parameters in the plant phenotype, including biotic and abiotic stress response. To enable current genetic crop development, there is a necessity for more effective and reliable phenotyping data, given the fast development of high-throughput genotype screening for associated growth, yield, and resistance to various biotic and abiotic stresses in plant breeding and genomics. Currently, expert visual evaluation is used to assess phenotypic features for disease resistance or stress in breeding programs (Yang et al., 2020). This takes time and may result in prejudice between experts and experimental duplicates as shown in Fig. 6.2.
It will be a huge challenge for plant science and crop development to ensure that crop production is sufficient to meet the needs of a human population that is predicted to reach more than 9 billion by 2050. This aim is difficult to achieve because crop output increases at a 1.3% annual rate, which is insufficient to keep up with population growth. High-yielding, stress-tolerant plants can be selected significantly more quickly and efficiently than is now possible by connecting the genotype to the phenotype. Breeders can benefit from advances in technology such as next-generation DNA sequencing, which might potentially boost the rate of genetic improvement through molecular breeding (Jaradat, 2018). However, our capacity to unravel the genetics of quantitative variables relevant to growth, yield, and stress adaptability is limited because of a paucity of phenotyping skills. Long before the discovery of DNA and molecular markers, plant breeders and farmers made decisions based on phenotypes. The more crosses and habitats that are used for selection, the better the chance of finding a superior variety.
It is necessary to improve breeding efficiency to fulfill future demands. The establishment of huge mapping populations has been aided by high-throughput genotyping and phenotyping panels made up of hundreds of recombinant inbred lines and the creation of massive mapping populations and phenotyping diversity panels made up of hundreds of recombinant inbred lines. Although molecular breeding techniques place a greater focus on genotypic information, phenotypic data is still required. Phenotypes are used for selection and to train a prediction model in genomic selection. A single phenotyping cycle is utilized in maker-assisted recurrent selection phenotyping to develop markers for future selection through generations to find potential events in transgenic investigations (Chaerle & Straeten, 2001).
Breakthroughs in phenotyping are critical for capitalizing on advances in traditional, molecular, and transgenic breeding (Li et al., 2014). Plant phenomics is concerned with defining the plasticity of the plant phenome when subjected to a variety of environmental variables rather than just correlating a genotype with one phenotype in a specific state (e.g., in a controlled environment). In contrast to most animals, which maintain roughly the same structure regardless of their environment, plants can take on a variety of architectural forms depending on the circumstances. After being exposed to either short- or long-day circumstances, the same type of Arabidopsis thaliana can produce a huge 30-leaf plant or a small 8-leaf plant. Water deficiency, nitrogen deficiency, and poor light all have a significant impact on the quantity and size of plant organs. As a result, plant phenomics research concentrates on the study of variation in organism structure, whereas animal phenomics research is primarily concerned with metabolism (Tardieu et al., 2017).
Quantitative Plant Morphology Detection Through Phenomics
Analysis of plant development, production, and formation is called Phenomics. According to (Soulé, 1967), the word “phenome” implies an entire phenotype like; genome manifestation in a certain location; collection of expression in a body belongs to phenome. To get valuable characters from assembled germplasm, forward phenomics act as a tool. Complete analysis of traits exposes systematic understanding and permits the development of this mechanism in new methods. Measurable and qualitative characteristics of a fused at a certain phase of ontogenesis in certain living conditions of a living system. At a considerable range, a superior proportion of phenotypic data find through phenomics. Skelly, Lobos, Orgogozo, and some other scientists proved that phenomics is usually considered parallel to genomics.
The procedure of development, transformation, and regulation of phenotypic expressions in living systems are examined in phenomics, then decreased to regularities (Skelly et al., 2013; Orgogozo et al., 2015; Lobos et al., 2017) though varies from genomics. Because of differences in phenotypic manifestation of characteristics upon the ecological circumstances, the entire classification of a genome is feasible in genomics, while the entire classification of phenome is not easy in phenomics, have been proved by Houle (Houle et al., 2010). Achievement of molecular genetics and breeding efforts particularly in field crops ever more defines through phenomics (Afonnikov et al., 2016). Plant phenotyping is considered an advanced and practically designed method of plant physiology (Furbank et al., 2011). Phenotypical modification that detect structural variation is quite easy to identify and examine. Stress response phenotypes depend on structural markers to compute stress responses in both ways. Phenotypes of numerous plants were recorded by Parent in 2015 because of the development of visual imagination and remote sensing skills and can easily be calculated at once mechanically and constantly (Parent et al., 2015).
Several sensors are utilized by extraordinary-data examining systems to access some structural characteristics like: plant height, canopy size, leaf area, green leaf pigment, shoot angle, virus spot size, and plant wilt degree. An individual can widely explain the significance of characters through this data like plant manner, nutritional value, drought acceptance, and virus resistance. A number of technologies are applied in various plants. By utilizing the Tomato Analyzer image, fruit shape traits were evaluated in 21 eggplant accessions from four varieties (Hurtado et al., 2013). A total of 23 fruit form parameters were calculated for agreement for fruit shape index, blockiness, homogeneity, proximal fruit end shape, asymmetry, internal eccentricity, and slenderness. (Hui et al., 2018). Three-dimensional (3D) canopy of cucumber, pepper, and eggplant based on multiview stereo (MVS) about plant canopy. By utilizing Crop Circle ACS-470 technique, Jaradat (2018) has isolated phenotypic information and some factors involving less heat tolerance during propagation, premature plant, etc. (Jaradat, 2018). Brassica napus verities that commonly grow in the Midwestern United States having extraordinary yields, and these can be measured through attributes. Tomato biomass and important linear association among expected shoot area, plant numeral biomass was analyzed by a scientist Laxman et al. (2018) using Scan analyzer 3D large scale imaging platform (Laxman et al., 2018). Bernotas et al. (2019) brief about photometer stereoscopic (PS) images, which consist of phenotype arrangement and its eudicot variety, including some vegetables like cabbage, tomato, and oilseed rapes (Bernotas et al., 2019).
Phenotyping of Roots in Plants Through Phenomics
Root system architecture (RSA) explains the spatial configuration of the root system rising from root morphology, topology, and distribution (Lynch, 1995). Root architecture, domestication environment, and techniques may be used to direct breeding programs to connect a root system with a life strategy and agroecology, increasing system adaptability (Bullock et al., 2017; Schmidt et al., 2016). Hypocotyl roots, as well as basal roots in some species, initiate epigeal germination between the radical and the cotyledons, which are elevated out of the soil. Above the basic root, there are three groups of epigeal germinators (Zobel, 2011). Basal roots of common beans can be divided into three types (Zobel & Waisel, 2010). Hypocotyl, main, and basal roots are among the types of roots. Basal roots emerge from the hypocotyl’s base. Root morphology (Zhu et al., 2011), leaf features (Micol, 2009), biomass (Tackenberg, 2007; Golzarian et al., 2011), yield-related traits (Duan et al., 2011), photosynthetic proficiency (Clark et al., 2011), and abiotic stress response (Rellan-Alvarez et al., 2015) are the most widely considered phenotypic traits Here, we will go through some of the most important plant phenotyping instruments, as well as some of the most promising photonics-based technologies.
As a result, despite notable breakthroughs, there are relatively few publically available root phenotyping datasets. Laboratory investigations benefit from improved levels of control, and at least in a few cases, loci with fundamental RSA in early root development have been identified. Nonetheless, the growth flasks utilized in these experiments, which were filled with actual or artificial soil (Rellan-Alvarez et al., 2015), limited geographic and temporal explanations for small or immature root systems (Judd et al., 2015; Lobet et al., 2013). Individual tools offer varying degrees of computational automation, ranging from manual to semiautomatic to fully automatic, making this software assembly an exciting prospect. None of these, however, give a combined stage that can (a) combine secondary root images with environmental and phenotypic metadata, (b) provide nontechnical users with continuous access to supercomputing resources, and (c) communicate content within a cooperative team and with the general public.
We formed DIRT in order to speak about these issues. The DIRT stage includes several key features that enable researchers to: (a) manage root picture collections and metadata, (b) interactively standardize dimension pipelines, (c) calculate crop root traits on available high-throughput compute platforms; and (d) analyze computation outcomes. DIRT enables researchers to process thousands of root images, complete the pipeline with routine parameters, and display and analyze calculated RSA output connected to the raw images. As a result, our stage allows researchers with few practical capabilities to access high-throughput computational stages. Thus, automation, remote control, and data (image) investigation pipelines agreeable to HTP stages acceptable showing of large plant populations, germplasm collections (core collections), breeding material, and mapping populations with increased accuracy and precision in phenotypic trait achievement attached with decreased labor input attained by high-throughput (Junker et al., 2015).
More study is needed to fully utilize genomics and molecular breeding methods in crop improvement, which address the creation of phenotyping tools and technologies, phenomics for a specific trait, phenotyping requirements, ongoing initiatives, and obstacles (Furbank & Tester, 2011; Cobb et al., 2013; Lobet, 2017). This review study aims to: emphasize the importance of phenomics and phenotypic constraints in crop improvement in the genomics era, (i) review the current status of phenomics stages and accommodations worldwide, (ii) emphasize the use of high-throughput phenomics platforms for trait separation in different crop plants and detection of genes/QTLs for a variety of traits in different crop plants, and (iii) emphasize the need for phenomics files and phenotyping. Responsible root function in soil, as well as root structure and growth screening, has long been a fascinating area (Gregory et al., 2009). For cereal species growing on stored soil moisture, access to water at penetration is critical for drought tolerance, and a study using model species to identify genes relevant for root characteristics is now underway. Small, short-lifecycle crop models, which are better suitable for cereal species, have recently been produced and are great systems for phenomic display (Watt et al., 2009).
This topic denotes a few technologies ranging from imaging in thin layers of soil or reproduction media to MRI and X-ray CT-scanning (Faget et al., 2009; Nagel et al., 2009). Root crown phenotyping occurs at the apex of crop root systems and can be utilized for marker-assisted breeding, genetic mapping, and a more sympathetic understanding of how roots inspire soil resource acquisition. There are a number of imaging methodologies and picture series available, but none of them are optimized for high-throughput, reproducible, and vigorous root crown phenotyping. The RhizoVision Crown stage includes an imaging unit, picture detention software, and image analysis software that have been upgraded to remove measurements from huge numbers of root crowns in a uniform manner to identify that root crown shapes. The hardware platform uses a backlight and a monochrome machine vision camera. The RhizoVision Imager and RhizoVision Analyzer are free, open-source applications that improve picture capture and analysis by incorporating spontaneous graphical user boundaries.
Physical validation of the RhizoVision Analyzer was done using copper wire, and feature validation was done with 10,464 ground truth simulated images of dicot and monocot root systems. The soybean and wheat root crowns were then phenotyped using this platform. The researchers phenotyped 2799 soybean (Glycine max) root crowns from 187 lines and 1753 wheat (Triticum aestivum) root crowns from 186 lines in both species; principal component analysis revealed comparable connections between characteristics. The greatest heritability was 0.74 in soybean and 0.22 in wheat, demonstrating that species and population variations must be taken into account. The RhizoVision Crown platform enables HTP of crop root crowns and establishes a benchmark against which open plant phenotyping platforms can be measured.
The total volume of soil that roots can investigate is influenced by RSA, which is shaped by interactions between genetic and environmental components (Schmidt et al., 2016). The number, length, growth angle, elongation rate, diameter, and branching of axial and lateral root phenes (or elemental units of phenotypic) shape the final RSA. Weaver and colleagues (Weaver, 1925; Weaver & Bruner, 1926) pioneered root-digging, diagramming, and photography methods that have been widely utilized for almost half a century (Böhm, 2012). These classical methods were since changed by Stoeckeler and Kluender (1938) with the use of water to remove soil particles from the root systems on a large scale and the use of high-pressure air to enter soil pores while leaving roots complete (Kosola et al., 2007). Devised hydro pneumatic root elutriation to give a rapid and repeatable method for extracting roots from the soil of field-composed soil important samples with minimal harm (Smucker et al., 2009).
Traditional digging methods are best for trees and shrubs because the root systems of woody plants are often stronger and more resistant to breakage than grasses or annual crops. The existing inability to quantify root architecture in the field is a major hindrance to current “phenomic” technologies’ claims of marker-assisted selection for better root system features. Traditional field approaches for root phenotyping, such as digging up soil cores and using standard digging techniques to control root depth, root branching density, and root angle, are still considered the best (Trachsel et al., 2011; Nielsen et al., 1997). Such approaches, however, do not reveal the finer aspects of root architecture, anatomy (for example, root hair densities), or function (e.g., nutrient uptake as a result, identifying crops with increased root architectural traits, as well as developing appropriate instruments for studying root growth in the soil, largely under field settings, remains a major challenge for current plant biology. Under drought conditions, root architecture changes dramatically, favoring the formation of more long lateral roots and root hairs to increase total surface area for better water absorption (Osmont et al., 2007). The increase in root quantity, mostly deeper in the soil, results in an enhanced plant water status, which is required to promote biomass production and yield when combined with techniques that limit water loss, such as stomatal closure, leaf systematic, and leaf abscission.
A new machine vision-based facility for the automated evaluation of yield-related characteristics in rice has been developed. This work resulted in the creation of an integrated facility for completely automated yield trait scoring. The facility can thresh rice panicles, evaluate rice production attributes, and pack loaded spikelets automatically. The accuracy (mean absolute percentage error is less than 5%) and efficiency of this unique machine vision-based facility were demonstrated in tests (1440 plants per continuous 24 h workday) (Table 6.2).
Future Perspective
Phenotypic–genotypic integrated breeding as the sequencing technology of crops has advanced. With the emergence of phenomics, breeding has entered a new age. It allows breeders to accurately phenotype many samples. Breeders may be able to correlate a lot more traits with accordant genotypes if it is combined with NGS technology. In current years, even more innovative attempts in the phenomics field have been made; computational approaches like machine learning (ML), deep learning (DL), and artificial intelligence (AI) have been integrated with HTP analyses to anticipate the population of many crops.
References
Afonnikov, D. A., Genaev, M. A., Doroshkov, A. V., Komyshev, E. G., & Pshenichnikova, T. A. (2016). Methods of high-throughput plant phenotyping for large-scale breeding and genetic experiments. Russian Journal of Genetics, 52, 688. https://doi.org/10.1134/S1022795416070024
Bernotas, G., Scorza, L. C. T., Hansen, M. F., Hales, I. J., Halliday, K. J., Smith, L. N., Smith, M. L., & Cormick, A. J. (2019). A photometric stereo-based 3D imaging system using computer vision and deep learning for tracking plant growth. Gigascience, 8, giz056.
Böhm, W. (2012). Methods of studying root systems (Vol. 33). Springer.
Bullock, J. M., Dhanjal-Adams, K. L., Milne, A., Oliver, T. H., Todman, L. C., Whitmore, A. P., & Pywell, R. F. (2017). Resilience and food security: Rethinking an ecological concept. Journal of Ecology, 105, 880–884. https://doi.org/10.1111/1365-2745.12791
Chaerle, L., & Van Der Straeten, D. (2001). Seeing is believing: Imaging techniques to monitor plant health. Biochimica et Biophysica Acta – Gene Structure and Expression, 1519, 153–166.
Chen, D., Neumann, K., Friedel, S., Kilian, B., Chen, M., Altmann, T., et al. (2014). Dissecting the phenotypic components of crop plant growth and drought responses based on high-throughput image analysis. Plant Cell, 26, 4636–4655.
Clark, R. T., et al. (2011). Three-dimensional root phenotyping with a novel imaging and software platform. Plant Physiology, 156(2), 455–465.
Cobb, J. N., et al. (2013). Next-generation phenotyping: Requirements and strategies for enhancing our understanding of genotype–phenotype relationships and its relevance to crop improvement. Theoretical and Applied Genetics, 126(4), 867–887.
Duan, L. F., Yang, W. N., Huang, C. L., & Liu, Q. (2011). A novel machine-vision based facility for the automatic evaluation of yield-related traits in rice. Plant Methods, 7, 44.
Dhondt, S., Wuyts, N., & Inzé, D. (2013). Cell to whole-plant phenotyping: the best is yet to come. Trends in plant science, 18(8), 428–439.
Faget, M., Herrera, J. M., Stamp, P., Aulinger-Leipner, I., Frossard, E., & Liedgens, M. (2009). The use of green fluorescent protein as a tool to identify roots in mixed plant stands. Functional Plant Biology, 36, 930–937. https://doi.org/10.1071/FP09125
Furbank, R. T., & Tester, M. (2011). Phenomics—Technologies to relieve the phenotyping bottleneck. Trends in Plant Science, 16, 635–644. https://doi.org/10.1016/j.tplants.2011.09.005
Furbank, R. T., Jimenez-Berni, J. A., George-Jaeggli, B., Potgieter, A. B., and Deery, D. M. (2019). Field crop phenomics: enabling breeding for radiation use efficiency and biomass in cereal crops. New Phytol. 223, 1714–1727.
Golzarian, M. R., Frick, R. A., Rajendran, K., Berger, B., Roy, S., Tester, M., & Lun, D. S. (2011). Accurate inference of shoot biomass from high throughput images of cereal plants. Plant Methods, 7, 2.
Gregory, P. J., Bengough, A. G., Grinev, D., Schmidt, S., Thomas, W. T. B., Wojciechowski, T., & Young, I. M. (2009). Root phenomics of crops: Opportunities and challenges. Functional Plant Biology, 36, 922–929.
Hairmansis, A., Berger, B., Tester, M., & Roy, S. J. (2014). Image-based phenotyping for non-destructive screening of different salinity tolerance traits in rice. Rice, 7(1), 1–10.
Houle, D., Govindaraju, D. R., & Omholt, S. (2010). Phenomics: The next challenge. Nature Reviews. Genetics, 11, 855–866.
Hui, F., Zhu, J., Hu, P., Meng, L., Zhu, B., Guo, Y., Li, B. G., & Ma, Y. T. (2018). Image-based dynamic quantification and high-accuracy 3D evaluation of canopy structure of plant populations. Annals of Botany, 121, 1079–1088.
Hurtado, M., Vilanova, S., Plazas, M., Gramazio, P., Herraiz, F. J., & Andújar, I. (2013). Phenomics of fruit shape in eggplant (Solanum melongena L.) using tomato analyzer software. Scientia Horticulturae Amsterdam, 164, 625–632.
Jaradat, A. A. (2018). Integrating plant ontogeny and structure in Brassica napus L. I. forward phenomics. Euphytica, 214, 141.
Jin, X., Zarco-Tejada, P., Schmidhalter, U., Reynolds, M. P., Hawkesford, M. J., Varshney, R. K., Yang, T., Nie, C., Li, Z., Ming, B., Xiao, Y., Xie, Y., & Li, S. (2021). Li High-throughput estimation of crop traits: A review of ground and aerial phenotyping platforms. IEEE Geoscience and Remote Sensing Magazine, 9, 200–231.
Jeudy, C., Adrian, M., Baussard, C., Bernard, C., Bernaud, E., Bourion, V., ... & Salon, C. (2016). RhizoTubes as a new tool for high throughput imaging of plant root development and architecture: test, comparison with pot grown plants and validation. Plant methods, 12(1), 1–18.
Judd, L. A., Jackson, B. E., & Fonteno, W. C. (2015). Advancements in root growth measurement technologies and observation capabilities for container-grown plants. Plants., 4(3), 369–392.
Junker, A., et al. (2015). Optimizing experimental procedures for quantitative evaluation of crop plant performance in high throughput phenotyping systems. Frontiers in Plant Science, 5, 1–21.
Kosola, K. R., Workmaster, B. A. A., Busse, J. S., & Gilman, J. H. (2007). Sampling damage to tree fine roots: Comparing air excavation and hydropneumatic elutriation. HortScience, 42(3), 728–731.
Laxman, R. H., Hemamalini, P., Bhatt, R. M., & Sadashiva, A. T. (2018). Non-invasive quantification of tomato (Solanum lycopersicum L.) plant biomass through digital imaging using phenomics platform. Indian Journal of Plant Physiology, 23, 369–375.
Li, L., Zhang, Q., & Huang, D. (2014). A review of imaging techniques for plant phenotyping. Sensors (Switzerland), 14(11), 20078–20111. https://doi.org/10.3390/s141120078
Lobet, G. (2017). Image analysis in plant sciences: Publish then perish. Trends in Plant Science, 22(7), 559–566.
Lobet, G., Draye, X., & Perilleux, C. (2013). An online database for plant image analysis software tools. Plant Methods, 9, 38.
Lobos, G. A., Camargo, A. V., del Pozo, A., Araus, J. L., Ortiz, R., & Doonan, J. H. (2017). Editorial: Plant phenotyping and phenomics for plant breeding. Frontiers in Plant Science, 8, 2181. https://doi.org/10.3389/fpls.2017.02181
Lynch, J. (1995). Root architecture and plant productivity. Plant Physiology, 109, 7–13. https://doi.org/10.1104/pp.109.1.7
Mathieu, L., Lobet, G., Tocquin, P., & Perilleux, C. (2015). “Rhizoponics”: A novel hydroponic rhizotron for root system analyses on mature Arabidopsis thaliana plants. Plant Methods, 11, 3.
Marié, C., Kirchgessner, N., Flütsch, P., Pfeifer, J., Walter, A., & Hund, A. (2016). RADIX: rhizoslide platform allowing high throughput digital image analysis of root system expansion. Plant methods, 12(1), 1–15.
Micol, J. L. (2009). Leaf development: Time to turn over a new leaf? Current Opinion in Plant Biology, 12, 9.
Murchie, E. H., & Lawson, T. (2013). Chlorophyll fluorescence analysis: A guide to good practice and understanding some new applications. Journal of Experimental Botany, 64, 3983–3998.
Nagel, K. A., Kastenholz, B., Jahnke, S., van Dusschoten, D., Aach, T., et al. (2009). Temperature responses of roots: Impact on growth, root system architecture and implications for phenotyping. Functional Plant Biology, 36, 947–959. https://doi.org/10.1071/FP09184
Neilson, E. H., Edwards, A. M., Blomstedt, C. K., Berger, B., Moller, B. L., & Gleadow, R. M. (2015). Utilization of a high-throughput shoot imaging system to examine the dynamic phenotypic responses of a C4 cereal crop plant to nitrogen and water deficiency over time. Journal of Experimental Botany, 66, 1817–1832.
Nielsen, K. L., Lynch, J. P., & Weiss, H. N. (1997). Fractal geometry of bean root systems: Correlations between spatial and fractal dimension. American Journal of Botany, 84, 26–33.
Orgogozo, V., Morizot, B., & Martin, A. (2015). The differential view of genotype–phenotype relationships. Frontiers in Genetics, 6, 179. https://doi.org/10.3389/fgene.2015.00179
Osmont, K. S., Sibout, R., & Hardtke, C. S. (2007). Hidden branches: Developments in root system architecture. Annual Review of Plant Biology, 58, 93–113.
Parent, B., Shahinnia, F., Maphosa, L., Berger, B., Rabie, H., Chalmers, K., Kovalchuk, A., Langridge, P., & Fleury, D. (2015). Combining field performance with controlled environment plant imaging to identify the genetic control of growth and transpiration underlying yield response to water-deficit stress in wheat. Journal of Experimental Botany, 66, 5481–5492.
Rellan-Alvarez, R., et al. (2015). GLO-Roots: An imaging platform enabling multidimensional characterization of soil-grown root systems. eLife, 4, 016931.
Reuzeau, C., Pen, J., Frankard, V., de Wolf, J., Peerbolte, R., Broekaert, W., & van Camp, W. (2005). TraitMill: a discovery engine for identifying yield-enhancement genes in cereals. Molecular Plant Breeding, 3, 753–759.
Schmidt, J. E., Bowles, T. M., & Gaudin, A. C. M. (2016). Using ancient traits to convert soil health into crop yield: Impact of selection on maize root and rhizosphere function. Frontiers in Plant Science, 7, 373.
Skelly, D. A., Merrihew, G. E., Riffle, M., Connelly, C. F., Kerr, E. O., Johansson, M., Jaschob, D., Graczyk, B., Shulman, N. J., Wakefield, J., Cooper, S. J., Fields, S., Noble, W. S., Muller, E. G. D., Davis, T. N., et al. (2013). Integrative phenomics reveals insight into the structure of phenotypic diversity in budding yeast. Genome Research, 23, 1496. https://doi.org/10.1101/gr.155762.113
Smucker, A. J. M., McBurney, S. L., Srivastava, A. K., et al. (2009). Agronomy Journal, 74, 500.
Soulé, M. (1967). Phenetics of natural populations I. Phenetic relationships of insular populations of the side-blotched lizard. Evolution, 21, 584–591.
Stoeckeler, J. H., & Kluender, W. A. (1938). The hydraulic method of excavating the root systems of plants. Ecology, 19(3), 355–369.
Tackenberg, O. (2007). A new method for non-destructive measurement of biomass, growth rates, vertical biomass distribution and dry matter content based on digital image analysis. Annals of Botany, 99, 777–783.
Tardieu, F., Cabrera-Bosquet, L., Pridmore, T., & Bennett, M. (2017). Plant phenomics, from sensors to knowledge. Current Biology, 27(15), R770–R783. https://doi.org/10.1016/j.cub.2017.05.055
Trachsel, S., Kaeppler, S. M., Brown, K. M., & Lynch, J. P. (2011). Shovelomics: High throughput phenotyping of maize root architecture in the field. Plant and Soil, 341, 75–87.
Watt, M., Schneebeli, K., Dong, P., & Wilson, I. W. (2009). The shoot and root growth of Brachypodium and its potential as a model for wheat and other cereal crops. Functional Plant Biology, 36, 960–969. https://doi.org/10.1071/FP09214
Weaver, E. (1925). Investigations on the root habits of plants. American Journal of Botany, 12(8), 502–509.
Weaver, J. E., & Bruner, W. E. (1926). Root development of field crops. McGraw-Hill Book Company.
Yang, W., Feng, H., Zhang, X., Zhang, J., Doonan, J. H., Batchelor, W. D., et al. (2020). Crop phenomics and high-throughput phenotyping: Past decades, current challenges, and future perspectives. Molecular Plant, 13(2), 187–214.
Zhu, J. M., Ingram, P. A., Benfey, P. N., & Elich, T. (2011). From lab to field, new approaches to phenotyping root system architecture. Current Opinion in Plant Biology, 14, 310–317. This review summarizes the available approaches for root system architecture (RSA) phenotyping in the laboratory and field.
Zobel, R. W. (2011). A developmental genetic basis for defining root classes. Crop Science, 51, 1410–1413. https://doi.org/10.2135/cropsci2010.11.0652
Zobel, R., & Waisel, Y. (2010). A plant root system architectural taxonomy: A framework for root nomenclature. Plant Biosystems, 144, 507–512. https://doi.org/10.1080/1126350100376448
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ahmed, H.G.MD., Zeng, Y., Fiaz, S., Rashid, A.R. (2023). Applications of High-Throughput Phenotypic Phenomics. In: Prakash, C.S., Fiaz, S., Nadeem, M.A., Baloch, F.S., Qayyum, A. (eds) Sustainable Agriculture in the Era of the OMICs Revolution. Springer, Cham. https://doi.org/10.1007/978-3-031-15568-0_6
Download citation
DOI: https://doi.org/10.1007/978-3-031-15568-0_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-15567-3
Online ISBN: 978-3-031-15568-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)