Abstract
The integration of genomics with phenomics leads to efficient breeding and the development of climate-resilient and crop varieties adaptable to the needs of modern breeding. Next-generation high-throughput approaches and plant phenotyping platforms have enabled the efficient, precise, and robust assessment of multiple plant traits in the last two decades. These approaches also mediate the relationship between plant growth and development traits on one hand and reproduction under diverse environmental conditions on the other. Nevertheless, recent high-tech advances develop novel tools with potential solutions to explore large-scale phenotyping data acquisition and processing in the coming years. In this book chapter, we discuss the significant achievement and advancement in high-throughput and phenomics in controlled environmental conditions and its uses for microphenotyping. We also discuss the latest multitudinal genomics research aided with high-throughput phenotyping with plant genetic studies. Finally, we propose few conceptual challenges and provide our future perspectives on bridging the phenotype-genotype-envirotype gap.
Graphical Abstract

Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
10.1 Introduction
Since the new millennium, emerging next-generation sequencing (NGS) technology has assisted researchers in measuring intractable and complex traits in biological data acquisition (Furbank et al. 2019). Among these technologies, crop functional genomics and whole-genome sequencing (Li et al. 2018) allowing in the acquisition of genome-wide association studies (GWAS) and quantitative trait locus (QTL) mapping (Xiao et al. 2017) on the large-scale phenotypic and genetic architecture of complex traits (Shi et al. 2019; Wang et al. 2019) have stepped into the high-throughput and big-data era. For instance, plant genome annotation revealed 26500 loci for Arabidopsis to 41000 genes in rice. Recently, the poplar genome was revealed to encode 45000 genes, and more than 40000 genes regulate multidimensional physiological and biological processes in Medicago and Lotus (Sterck et al. 2007). However, a genome size of 950 MB encoding about 35,000 genes in the tomato revolution our understanding of tomato biology (Barone et al. 2008). In the last two decades, thousands of genes in tomato (Barone et al. 2008), rice (Yao et al. 2018), and Arabidopsis (Bouché and Bouchez 2001) were functionally characterized through various traditional phenotypic techniques. The challenge is to attach the functions of these enormous numbers of genes restricting functional genomics studies and crop breeding (Deery et al. 2016).
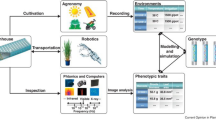
The term “phenome” was first characterized by Davis (1949): approach to systematically explore the comprehensive set of extragenic, non-autoreproductive portions of the cell and represented the set of phenotypes, either cytoplasmic or nuclear. Later on, phenomics was described as a complex interaction of an organism’s genotype and phenotype (Houle et al. 2010). Conventional crop phenotyping is laborious, tedious, intensive, and potentially injurious to plants (Chen et al. 2014). Recently, plant phenomics has been growing and advancing rapidly in the last decades includes the set of approaches used to precisely assess individual cells, tissue, leaf, or plant to the large scale, i.e., ecosystem (Fiorani and Schurr 2013). In addition, crop phenomics is the inter/intradisciplinary study of high-throughput phenotyping platforms for accurate acquisition and an organism-wide scale analysis of phenotypes in crop development (Fig. 10.1) (Zhao et al. 2019). In recent years, next-generation genotyping and phenotyping have been advantageous over traditional breeding approaches due to the accuracy of these methods and their robust capability to accelerate crop breeding (Pasala and Pandey 2020).
Advanced sophisticated sensors, vision-guided robotics, automation technology, and machine learning, with applications in harvesting, quality assessment, sorting, screening, and packaging, have been extensively implemented in the agri-production industry to promote efficiency (Ruiz-Garcia et al. 2009) and for breeders to have a breakthrough in making rapid genetic progress (Furbank et al. 2019). The integration of genomics and phenomics can accelerate genetics gain in breeding programs and identify new traits in diverse plant germplasms that help breed populations through the crossing and artificial genomics (Bortesi and Fischer 2015). However, these approaches should apply at the early stages of plant development. To search for how and why to measure the whole genome and whole-plant phenotypes has been extensively explored. Over the last few decades, the answer to the former has been examined in detail. Recent achievements in high-throughput technologies allow us to conclude how and why to measure organism-level phenotype in the coming decades (Houle et al. 2010). For instance, RICE 2020 has been initiated to systematically and functionally characterize all protein transcripts and gene transcripts in rice by 2020 (Zhang et al. 2008). This chapter tries to cover the significant advances in applying integrated high-throughput and phenomics approaches in genetics studies. Finally, we discuss the challenges in agri-phenomics and specify our standpoint on phenomics-related studies.
10.2 Phenomics
The plant being sessile organisms interact with multiple environmental stress across the life cycle and developed multiple avoidance strategies (Lymperopoulos et al. 2018). Conventional phenomics approaches focus on crop productivity and yield, while advanced sensing approaches enable plant scientists to record the environmental history of plants, together with their dynamic responses (Pratap et al. 2019). Robust phenomics is vital to plant breeding due to its fundamental basis for developing new varieties. However, advancement in phenomics including robotics, image processing, and deep learning enables non-destructive monitoring of plants development and function to extract valuable information (Hickey et al. 2019). The ongoing challenges with new generation phenomics are data handling and continuous contribution by computational technologies critical to maintaining rapid advancements in accelerated breeding programs (Tester and Langridge 2010).
10.2.1 Phenomics for Crop Microphenotypic Traits
Phenotyping at organ, tissue, and cellular levels requires complex procedures, and several automated high-throughput and large-scale phenotyping platforms have been developed for screening and assessing Arabidopsis, rice, and maize under controlled environmental conditions or in the field (Table 10.1) (Kaul, Koo et al. 2000). Phonoscope, a large-scale high-throughput phenotyping platform, monitors the plant growth rates of more than 700+ plants (Tisné et al. 2013). Renovator, accurate quantification for genotypic variation in natural genetic population using growth and photosynthesis as a phenotypic indicator of plant performance (Flood et al. 2016), utilizes a monochrome camera on the moving rail system platform with 1440 plants carrying capacity. The renovator can collect leaf area and light-use efficiency. Instead, PlantScreen with automatic weighing and watering moving conveyor belts transported plants from growing chamber to RGB imaging cabinets, chlorophyll fluorescence, and dark acclimation chamber (Awlia, Nigro et al. 2016). Plant root traits play a critical role in nutrient acquisition and the transport of water from the soil to the aerial parts of plants. Microimaging and sensing have improved our understanding of root anatomy and functions. Wu et al. (2011) developed a low-cost computer-aided 3-D visualization and quantitative analysis technique based on classical paraffin embedding serial sections and microtome techniques. In 2012, a high-throughput, high-resolution phenotyping platform, RootSlice, aided with laser and 3-D visualization was introduced to analyze (Burton, Williams et al. 2012) and quantify root anatomy with semi-auto RootScan (Chimungu, Loades et al. 2015). RootAnalyzer, automatic versatile root tissues, and root cells image phenotypic analysis tools. RootAnalyzer has more than 90% accuracy and improves image segmentation efficiency in quantifying the properties of tissues (Chopin, Laga et al. 2015). Walter et al. (2007) introduced an automated imaging pipeline GROWSCREEN to acquire the dynamics of seedling growth acclimation such as relative growth rate, total leaf area, and root area. Moreover, GROWSCREEN FLUORO allowed the simultaneous phenotyping of chlorophyll fluorescence in rosette plants (Jansen, Gilmer et al. 2009). The mapping power of GROWSCREEN for GWAS or QTL analyses reduced due to relatively limited carrying capacity in of platform in combination with micro-environmental heterogeneity.
TraitMill by CropDesign (Belgium) yield-related morphometric traits measure exclusive bioinformatics tools and a high-throughput phenotyping platform (Reuzeau et al. 2010). Computer-controlled Scanalyzer3D platform accelerating plant phenomics aided with automated watering and weighing, a conveyor with a capacity of 2400 plants, and RGB imaging stations to estimate plant biomass using RGB images (Virlet et al. 2016). In addition, the salinity tolerance of chickpea can be assessed by Scanalyzer3D (Hairmansis, Berger et al. 2014). In addition, rice (Hairmansis, Berger et al. 2014), and nutrient starvation in crops, in field diseases monitoring, and more physiological features including growth date. In recent years, an array of algorithms and tools has been developed (Table 10.1). However, there is still a need to simplify complex phenotypic procedures at cellular and tissue levels. The introduction of advanced imaging techniques will accelerate microscopic phenotyping and assist in advanced phenotyping studies, particularly on specific cell phenotypes and crop organ characteristics.
10.2.2 High-Throughput Plant Phenotyping Platforms in a Controllable Environment
In crops, the breeding and selection of desirable traits is vital for sustainable agriculture, global food security, and the growing global demand for fiber, feed, and fuel (Dungey et al. 2018). High-throughput plant phenotyping with non-destructively image approaches facilitate efficient screening of plants based on their morphological and physiological traits, may assist in increasing productivity, shorten the crop cycle, improve plant efficiency in the environment, and help in linking phenomics to genomics (Li et al. 2014). However, the application of high-throughput plant phenotyping in agriculture is still in its infancy. It needs to have the accuracy and efficiency to assess the growth and morphological traits of plants, such as growth patterns, development rate, plant aerial architecture, root architecture system, and plant biomass. These features are fundamental to understand function-structure of plant in assesing biotic and bitoic responses for sustainable management of crops.
Global warming has posed a risk to global food demand as it has impacted agricultural productivity in the past few decades. Despite the negative influences of climate change, however, the selection and breeding of environmental resistance crops are needed today, but they should be done without compromising the quality and quantity of crops (Langstroff et al. 2021). Controlled environment phenotyping (CEP) is a nondestructive approach for exploring plant behaviors, which enable breeders to search for genotypes capable of coping with future environments (Xue et al. 2019). The primary problems in upcoming plant breeding programs are the lack of infrastructure and the diversity of users (Carpentier et al. 2019). Currently, a practical approach that is being used for searching scientific opportunities is bibliometric science mapping, which is done by analyzing scientific publications (Van Raan 2004). This method has been applied for analyzing phenomics. Under such a scenario, there is a need to develop dedicated tools, infrastructures, and resources for phenotyping genomics resources. The production of such high-throughput plant phenotyping resources could only be possible through a public-private partnership.
Luckily few initiatives arose to integrate fully controlled environment facilities climate-specific locations between laboratory-based work and “real-world” scenarios (Carpentier et al. 2019; Costa et al. 2019). For instance, the projects EPPN (http://www.plant-phenotyping-network.eu/), the COST Action FA1306 (http://www.cost.eu/COST_Actions/fa/FA1306), and EPPN2020 (https://eppn2020.plant-phenotyping.eu/). Similarly, European Infrastructure for Pan-Phenomics and simulation for global food security, the ESFRI-project EMPHASIS, jointly launched synergistic pan-European excellence in phylogenomics for developing relevant approaches and shared infrastructures. German Plant Phenotyping Network (DPPN) provides a robust phenotypic portfolio and shares productive and efficient infrastructure. On a global scale, China, USA, and Canada national phenotyping efforts include Asia-Pacific Plant Phenotyping Conference (APPP, www.APPP-con.org), North American Plant Phenotyping Network (NAPPN, https://www.plantphenotyping.org/), and the International Plant Phenotyping Network (IPPN; https://www.plant-phenotyping.org/) develops integrated approaches beyond the national and regional perspectives.
High-throughput plant phenotyping can be integrated into greenhouses which allow crop phenotyping at the whole-population level and observe natural-variation in GWAS citrus (Minamikawa et al. 2017), rice (Crowell et al. 2016; Rebolledo et al. 2016; Yang et al. 2014), maize (Gage et al. 2018; Wang et al. 2019), wheat (Beyer et al. 2019; Rasheed et al. 2014), barley (Bergsträsser et al. 2015; Neumann et al. 2017), soybean (Bergsträsser et al. 2015). Besides acting as a shield from light, rain, and extreme temperatures, greenhouses provide a straightforward environment conducive for plant nutrient, salinity, and drought studies (Neumann et al. 2015). During drought, identification, and selection of precisely multitude drought heritable traits beneficial to characterize a phenotype (Chen et al. 2014). Global warming impacts agriculture productivity at global scale proxies drought indices to account and predict drought severity (Mukherjee et al. 2018) owing to few crop species spatiotemporal adaptations with varying productivity. Temperatures have drastic effects on crop yield and productivity (Zhao et al. 2017); for example, an elevated temperature decreases crop yields in maize up to 90% (Hatfield and Prueger 2015). Control environment integrated with high-throughput phenotyping enables identification of QTL and GWAS necessary for repeated phenotyping to ultimate phenotype (Muraya et al. 2017). The ecosphere is highly sensitive to temperature changes and differentially affects plant growth at different altitudes (Rosenzweig et al. 2014), influencing plant adaptation and productivity. Flowering in plant is controlled by daily temperature fluctuation, day length, light intensity, and seasonal cues to help understand the dynamic genetic components to plant adaptation (Li et al. 2010). In general, intercepted light can increase total plant biomass (60%) (Poorter et al. 2016). Dynamic environmental components are integrated with high throughput approaches to explore genetic variations in field or controlled environments.
10.3 Application of Machine Learning in Phenomics
Integration of artificial intelligence in interdisciplinary fields has been grown exponentially in the last decade. Artificial intelligence applications such as deep learning, sensors, and machine learning successfully enable high-throughput phenotyping of plant traits into non-invasive imaging approaches (Nabwire et al. 2021). The accuracy and efficiency of data collection and analysis Improve through deep learning and machine learning for vigorous image analysis and influential study of phenotypes. Conventional breeding approaches of phenotyping are destructive with sufficient resolution and require crop harvesting at specific plant growth stages (Furbank and Tester 2011). The plant breeding programs are significantly lagging behind genomics, slow, time consuming, and require repeated experiments to validate certain traits pivotal for crop improvement (Fahlgren et al. 2015). The non-invasive high-throughput imaging approaches enable phenotype visualization at a cellular scale. The imaging techniques such as chlorophyll fluorescence (Zarco-Tejada et al. 2009), thermography (Oerke et al. 2006), spectroscopic imaging (Montes et al. 2006), and digital imaging (Jensen et al. 2007) carry a large amount of extractable data to support biological interpretations of plant growth (Walter et al. 2010). Hight-throughput artificial intelligence architecture applied in phenotyping are listed in Table 10.2.
Currently, high-throughput approaches like growth chambers (Bai et al. 2016), imaging sensors (Chaerle and Van Der Straeten 2000), data acquisition, and statistical software are employed for data collection, management, and interpretation at laboratory and field levels. Integration of these techniques into artificial intelligence in the form of machine learning (Kruse et al. 2014) and computer vision (Casanova et al. 2014) attribute to the non-invasion aspect of phenomics (Montes et al. 2007). Artificial intelligence applications are expanding with a public-private partnership in developing and disseminating these phenomic approaches that address the challenges of costly infrastructure and proprietary data formats. Thus far, computer vision, deep learning, and machine learning have been applied in phenomics. Since 1970, various machine learning models, such as Bayesian networks, support vector machines, and perceptron, have been developed, but none have proved to be the best as they all have certain limitations (Roscher et al. 2020). Later on, neural networks integrated with data collection and information processing infrastructure enable machine learning to determine best-fitting models (Roscher et al. 2020). One of the advantages of phytogenomic machine learning is their ability to simultaneously processing massive amounts of data in combination with other related features (Roscher et al. 2020) assist in the identification and classification of plant traits, including disease or pest detection (Wetterich et al.), floral transition (Wetterich et al.), and seeds classification (Sabanci et al. 2017).
Deep learning is a subset of machine learning instead of several complex high throughput sensors with a wide range of phenotype applications (LeCun et al. 2015). However, the data collected through this versatile tool contained high variability making its application more complicated but providing more reliable prediction (Singh et al. 2018). Deep learning in computer vision-based phenomics ensures the more reliable processing of phenotypic images involving multilayered approaches network, each performing its operation in succession, improving prediction and discrimination ability (Pound et al. 2017) by a process called transfer learning. Table 10.2 lists some deep learning phenomics approaches that have been used for plant morphology and stress identification. However, applying machine learning and deep learning subsets of artificial intelligence enables plant scientists robust identification, classification, and detection of environmental variability influences on plant growth, development, and other related physiological parameters.
10.4 High-Throughput Phenomics Enhances Phytogenetics
High-throughput phenomics has been widely used in remote sensing, root phenomics, deep learning for plant stress, and vision sensing technologies in disease and pest detection (Atkinson et al. 2019; Mahlein et al. 2018). However, applying phenotyping technologies and genome sequencing is still limited in genetics and crop breeding studies both in the field and within the lab. A plethora of studies have identified QTL in many plant species, but still sveral issues need to be resolved. Among them is how to charctarizes dymanic QTLs for complex traits at mutiple grwoth stage or at different species level or districtive tarit measurements including root architechture, biomass allocation, and nutrient assessment. Identification and efficient functional characterization of potential QTLs. Integrating various genomics approaches with systemic and synthetic molecular biology approaches will significantly facilitate future breeding programs.
The regulation of the size of maize shoot apical meristems (SAMs) is correlated with flowering. High throughput analysis enables integration of SAM morphological traits with GWAS and QTL, demonstrated their contribution to SAM development (Leiboff et al. 2015). High-throughput non-destructively micro-CT-RGB phenotyping and genomics enable large-scale assessment of rice tiller traits, tiller growth, and plant traits nine growth stages. Among these traits, 402 significantly influence grain yield, vigor-related traits and yield (Wu et al. 2019). However, such integration of genomics and phenomics is beneficial in crop breeding programs required for high yields and compact planting. Genome selection is another robust genotype-phenotype approach that involves statistical modeling and genome-wide markers. It allows efficient and accurate markers to be identified, but associated phenotype prediction is still a bottleneck in crop breeding (Taylor 2014). The process of photosynthesis and transpiration in leaves (Wang et al. 2015) depends on the number of leaves inm plants as well as the leaf size, shape, and greenness (Wang et al. 2011). The genetics study of rice and maize leaves by high-throughput leaf scoring revealed nine loci associated with leaf traits in 533 rice accessions at thee growth stages. In maize, QTL mapping of 22 leaf traits of a RIL population at 16 growth stages predited leaf tarits (leaf angle and length) being an indicator of yield (Yang et al. 2015).
Integration of high throughput phenomics with large-scale GWAS or QTLs expanded our understanding of crop developmental dynamics and emerged as a tool for plant genomics, gene expression, and characterization. The root system architecture is a promising trait for nutrients and water acquisition from the soil. Dissecting the root genetic will be helpful in increased nutrient and water acquisition from the soil. Two genetic studies on root traits were performed on rice and Brassica (Courtois et al. 2013; Shi et al. 2013). They conducted a GWAS of 15 root traits using vision sensors and detected associations between deep root number and mass. In Brassica napus, 38 QTLs were predictive indicators under phosphorus variability. Shi et al. (2013) used an agar-based high-throughput root phenotyping system to identify QTLs associated with phosphate variability correlated with Brassica napus root architectural traits.
Environmental factors, including both abiotic and biotic factors, can produce a variety of phenotypic effects. The rapid development of non-destructive high-throughput plant phenotyping approaches has been popularized in a plethora of crop populations to reveal the genetics of complex quantitative traits to various environmental factors (Yang et al. 2020), such as phosphate deficiency tolerance of Brassica napus (Shi et al. 2013), drought response of wheat (Parent et al. 2015), salinity tolerance of rice (Al-Tamimi et al. 2016), and drought resistance of rice (Guo et al. 2018). Most of these studies have focused on external responses, such as the morphology, biomass, and greenness-related traits. The internal response of plants to drought is mainly unknown. Wu et al. (2021) develop a non-destructively image-based traits (i-traits) approach to plant responses to drought. The i-traits are high-throughput image analysis pipelines aided by RGB optical sensors, X-ray computed tomography and hyperspectral imaging. In maize, i-traits identified 4322 drought-responsive loci encoding 1529 QTLs, including 15 QTLs containing potential markers for drought tolerance breeding in maize. Combining crop genetics information with genotype-phenotype approaches revolutionizes researchers’ understanding of complex traits and reinforces the new era of crop breeding.
10.5 Conclusion and Future Perspective
Crop breeding evolved from conventional approaches to phenotypes-genotype aided breeding through advancement in next-generation sequencing of crops. Plant breeding enters the next era of phenomics which enables breeders accurate sampling to phenotype various traits. In the last few decades, phenomics has entered a new period of advancements as it integrated machine learning, deep learning, and artificial intelligence to predict the phenotypic characteristics of different crop populations. These approaches inherent data from various sources tend to accelerate crop breeding programs.
Considerable efforts have been placed in agriculture and global food security to maintain sustainable crop growth and productivity. Sustainable agriculture and crop production by integrating genetic tools lead to the advancement in phenomics, but demand for financial investments diminishes the development of agriculture. Phenotypic studies focus on aerial plant parts and underground plant parts; however, the integrated role of aerial-phenomics to underground phenomics influenced, but various biotic and abiotic stresses need to consider. For instance, the primary root of plant roots determines the plant’s capacity to store water or other essential plant nutrients in the stem. Integrated phenotype, the ratio of root length to stem height, could be related to crop yield. For such integrated phenotypes, algorithms to compute plant imagery are required to be developed.
Furthermore, for early detection of environmental stresses, including water, drought, salinity, temperature, etc, effort should be placed to investigate the phenotypes for characterizing the propagating stress and classified them into different stages such as moderate, extreme, or exceptional. Similarly, controlling the root growth angel may contribute to the speed of recovery.
The advancement of phenomics in “big data” enables the plant science community to establish new theories in plant phenotypic approaches to integrate artificial and collaborative research at global levels. Crop phenotypic information should focus on developing tools that comprehensively integrate multi-tudinal scale emphasize on pheno-envir-genotype and physiological parameters to systematically and complete phenomic information. The critical problem of functional phenomics is its development and application in phenotyping. The introduction of new methodologies integrated with artificial intelligence and machine learning help minimize environmental challenges. These throughput approaches collect digital features efficiently. These features’ precise and robust interpretation dig out critical quantitative and qualitative phenotypic traits for functional genomics. High throughput approaches also facilitate the integration of multi-tudinal phenotypic information for big-data development, management, shareability, and globality in crop geno-pheno-envirotype analysis and utilization. In short, for the future of phenomics, we urgently need synergism at the global level. Search for novel tools and methodologies offer powerful tools to dissect the processes in plant growth, development, and producing high-yielding and climate-resilient crops.
References
Al-Tamimi N, Brien C, Oakey H, Berger B, Saade S, Ho YS, Schmöckel SM, Tester M, Negrão S (2016) Salinity tolerance loci revealed in rice using high-throughput non-invasive phenotyping. Nat Commun 7:1–11
Amal TC, Thottathil AT, Veerakumari KP, Rakkiyappan R, Vasanth K (2020) Morphological traits of drought tolerant horse gram germplasm: classification through machine learning. J Sci Food Agric 100:4959–4967
Atkinson JA, Pound MP, Bennett MJ, Wells DM (2019) Uncovering the hidden half of plants using new advances in root phenotyping. Curr Opin Biotechnol 55:1–8
Awlia M, Nigro A, Fajkus J, Schmoeckel SM, Negrão S, Santelia D, Trtílek M, Tester M, Julkowska MM, Panzarová K (2016) High-throughput non-destructive phenotyping of traits that contribute to salinity tolerance in Arabidopsis thaliana. Front in Plant Sci. 7:1414
Bai G, Ge Y, Hussain W, Baenziger PS, Graef G (2016) A multi-sensor system for high throughput field phenotyping in soybean and wheat breeding. Comput Electron Agric 128:181–192
Barone A, Chiusano ML, Ercolano MR, Giuliano G, Grandillo S, Frusciante L (2008) Structural and functional genomics of tomato. Int J Plant Genomics 2008:820274–820274
Bergsträsser S, Fanourakis D, Schmittgen S, Cendrero-Mateo MP, Jansen M, Scharr H, Rascher U (2015) HyperART: non-invasive quantification of leaf traits using hyperspectral absorption-reflectance-transmittance imaging. Plant Methods 11:1–17
Beyer S, Daba S, Tyagi P, Bockelman H, Brown-Guedira G, Mohammadi M (2019) Loci and candidate genes controlling root traits in wheat seedlings—a wheat root GWAS. Funct Integr Genomics 19:91–107
Bortesi L, Fischer R (2015) The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol Adv 33:41–52
Bouché N, Bouchez D (2001) Arabidopsis gene knockout: phenotypes wanted. Curr Opin Plant Biol 4:111–117
Bucksch A, Burridge J, York LM, Das A, Nord E, Weitz JS, Lynch JP (2014) Image-based high-throughput field phenotyping of crop roots. Plant Physiol 166:470–486
Burridge J, Jochua CN, Bucksch A, Lynch JP (2016) Legume shovelomics: high—throughput phenotyping of common bean (Phaseolus vulgaris L.) and cowpea (Vigna unguiculata subsp, unguiculata) root architecture in the field. Field Crop Res 192:21–32
Busemeyer L, Mentrup D, Möller K, Wunder E, Alheit K, Hahn V, Maurer HP, Reif JC, Würschum T, Müller J (2013) BreedVision—A multi-sensor platform for non-destructive field-based phenotyping in plant breeding. Sensors 13:2830–2847
Bylesjö M, Segura V, Soolanayakanahally RY, Rae AM, Trygg J, Gustafsson P, Jansson S, Street NR (2008) LAMINA: a tool for rapid quantification of leaf size and shape parameters. BMC Plant Biol 8:1–9
Burton AL, Williams M, Lynch JP, Brown KM (2012) RootScan: software for high-throughput analysis of root anatomical traits. Plant and Soil 357(1):189–203
Chimungu JG, Loades KW, Lynch JP (2015) Root anatomical phenes predict root penetration ability and biomechanical properties in maize (Zea mays). J Experi Bot 66(11):3151–3162
Chopin J, Laga H, Huang CY, Heuer S, Miklavcic SJ (2015) Rootanalyzer: a cross-section image analysis tool for automated characterization of root cells and tissues. PloS one 10(9):e0137655
Carpentier S, Costa C, Schurr U, Loreto F, Menesatti P (2019) Plant phenotyping research trends, a science mapping approach. Front Plant Sci 9
Casanova JJ, O’Shaughnessy SA, Evett SR, Rush CM (2014) Development of a wireless computer vision instrument to detect biotic stress in wheat. Sensors 14:17753–17769
Chaerle L, Van Der Straeten D (2000) Imaging techniques and the early detection of plant stress. Trends Plant Sci 5:495–501
Chen D, Neumann K, Friedel S, Kilian B, Chen M, Altmann T, Klukas C (2014) Dissecting the phenotypic components of crop plant growth and drought responses based on high-throughput image analysis. Plant Cell 26:4636–4655
Clauw P, Coppens F, De Beuf K, Dhondt S, Van Daele T, Maleux K, Storme V, Clement L, Gonzalez N, Inzé D (2015) Leaf responses to mild drought stress in natural variants of Arabidopsis. Plant Physiol 167:800–816
Costa JM, Marques da Silva J, Pinheiro C, Barón M, Mylona P, Centritto M, Haworth M, Loreto F, Uzilday B, Turkan I (2019) Opportunities and limitations of crop phenotyping in southern European countries. Front Plant Sci 10:1125
Courtois B, Audebert A, Dardou A, Roques S, Ghneim-Herrera T, Droc G, Frouin J, Rouan L, Gozé E, Kilian A (2013) Genome-wide association mapping of root traits in a japonica rice panel. PLoS One 8:e78037
Crowell S, Korniliev P, Falcao A, Ismail A, Gregorio G, Mezey J, McCouch S (2016) Genome-wide association and high-resolution phenotyping link Oryza sativa panicle traits to numerous trait-specific QTL clusters. Nat Commun 7:1–14
Davis BD (1949) The isolation of biochemically deficient mutants of bacteria by means of penicillin. Proc Natl Acad Sci U S A 35:1
De Bei R, Cozzolino D, Sullivan W, Cynkar W, Fuentes S, Dambergs R, Pech J, Tyerman S (2011) Non-destructive measurement of grapevine water potential using near infrared spectroscopy. Aust J Grape Wine Res 17:62–71
De Diego N, Fürst T, Humplík JF, Ugena L, Podlešáková K, Spíchal L (2017) An automated method for high-throughput screening of Arabidopsis rosette growth in multi-well plates and its validation in stress conditions. Front Plant Sci 8:1702
De Vylder J, Vandenbussche F, Hu Y, Philips W, Van Der Straeten D (2012) Rosette tracker: an open source image analysis tool for automatic quantification of genotype effects. Plant Physiol 160:1149–1159
Deery DM, Rebetzke GJ, Jimenez-Berni JA, James RA, Condon AG, Bovill WD, Hutchinson P, Scarrow J, Davy R, Furbank RT (2016) Methodology for high-throughput field phenotyping of canopy temperature using airborne thermography. Front Plant Sci 7:1808–1808
Dungey HS, Dash JP, Pont D, Clinton PW, Watt MS, Telfer EJ (2018) Phenotyping whole forests will help to track genetic performance. Trends Plant Sci 23:854–864
Dutta P, Bandopadhyay P, Bera AK (2016) Identification of leaf based physiological markers for drought susceptibility during early seedling development of mungbean. Am J Plant Sci 7:1921
Fahlgren N, Gehan MA, Baxter I (2015) Lights, camera, action: high-throughput plant phenotyping is ready for a close-up. Curr Opin Plant Biol 24:93–99
Fiorani F, Schurr U (2013) Future scenarios for plant phenotyping. Annu Rev Plant Biol 64:267–291
Flood PJ, Kruijer W, Schnabel SK, van der Schoor R, Jalink H, Snel JFH, Harbinson J, Aarts MGM (2016) Phenomics for photosynthesis, growth and reflectance in Arabidopsis thaliana reveals circadian and long-term fluctuations in heritability. Plant Methods 12:1–14
Furbank RT, Tester M (2011) Phenomics–technologies to relieve the phenotyping bottleneck. Trends Plant Sci 16:635–644
Furbank RT, Jimenez-Berni JA, George-Jaeggli B, Potgieter AB, Deery DM (2019) Field crop phenomics: enabling breeding for radiation use efficiency and biomass in cereal crops. New Phytol 223:1714–1727
Gage JL, White MR, Edwards JW, Kaeppler S, de Leon N (2018) Selection signatures underlying dramatic male inflorescence transformation during modern hybrid maize breeding. Genetics 210:1125–1138
Granier C, Aguirrezabal L, Chenu K, Cookson SJ, Dauzat M, Hamard P, Thioux JJ, Rolland G, Bouchier-Combaud S, Lebaudy A (2006) PHENOPSIS, an automated platform for reproducible phenotyping of plant responses to soil water deficit in Arabidopsis thaliana permitted the identification of an accession with low sensitivity to soil water deficit. New Phytol 169:623–635
Guo D, Juan J, Chang L, Zhang J, Huang D (2017) Discrimination of plant root zone water status in greenhouse production based on phenotyping and machine learning techniques. Sci Rep 7:1–12
Guo Z, Yang W, Chang Y, Ma X, Tu H, Xiong F, Jiang N, Feng H, Huang C, Yang P (2018) Genome-wide association studies of image traits reveal genetic architecture of drought resistance in rice. Mol Plant 11:789–805
Hatfield JL, Prueger JH (2015) Temperature extremes: effect on plant growth and development. Weather Clim Extremes 10:4–10
Hickey LT, Hafeez AN, Robinson H, Jackson SA, Leal-Bertioli SCM, Tester M, Gao C, Godwin ID, Hayes BJ, Wulff BBH (2019) Breeding crops to feed 10 billion. Nat Biotechnol 37:744–754
Houle D, Govindaraju DR, Omholt S (2010) Phenomics: the next challenge. Nat Rev Genet 11:855–866
Howarth CJ, Gay AP, Draper J, Powell W (2011) Development of high throughput plant phenotyping facilities at Aberystwyth. In: Proceedings of the phenomics workshop, San Diego
Humplík JF, Lazár D, Fürst T, Husičková A, Hýbl M, Spíchal L (2015) Automated integrative high-throughput phenotyping of plant shoots: a case study of the cold-tolerance of pea (Pisum sativum L.). Plant Methods 11:1–11
Hairmansis, A, Berger B, Tester M, Roy SJ (2014) Image-based phenotyping for non-destructive screening of different salinity tolerance traits in rice. Rice 7(1):1-10
Jansen M, Gilmer F, Biskup B, Nagel KA, Rascher U, Fischbach A, Briem S, Dreissen G, Tittmann S, Braun S (2009) Simultaneous phenotyping of leaf growth and chlorophyll fluorescence via GROWSCREEN FLUORO allows detection of stress tolerance in Arabidopsis thaliana and other rosette plants. Functional Plant Biology 36(11):902–914
Jensen T, Apan A, Young F, Zeller L (2007) Detecting the attributes of a wheat crop using digital imagery acquired from a low-altitude platform. Comput Electron Agric 59:66–77
Kaul S, Koo HL, Jenkins J, Rizzo M, Rooney T, Tallon LJ, Feldblyum T, Nierman W, Benito M-I, Lin X (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Natu 408(6814):796–815
Klukas C, Chen D, Pape J-M (2014) Integrated analysis platform: an open-source information system for high-throughput plant phenotyping. Plant Physiol 165:506–518
Kruse OMO, Prats-Montalbán JM, Indahl UG, Kvaal K, Ferrer A, Futsaether CM (2014) Pixel classification methods for identifying and quantifying leaf surface injury from digital images. Comput Electron Agric 108:155–165
Langstroff A, Heuermann MC, Stahl A, Junker A (2021) Opportunities and limits of controlled-environment plant phenotyping for climate response traits. Theor Appl Genet 135(1):1–16
LeCun Y, Bengio Y, Hinton G (2015) Deep learning. Nature 521:436–444
Leiboff S, Li X, Hu HC, Todt N, Yang J, Li X, Yu X, Muehlbauer GJ, Timmermans MC, Yu J, Schnable PS, Scanlon MJ (2015) Genetic control of morphometric diversity in the maize shoot apical meristem. Nat Commun 6:8974
Leport L, Musse M, Cambert M, De Franscesci L, Le Caherec F, Burel A, Mariette F, Bouchereau A (2011) Canola leaf senescence phenotyping and identification of subcellular changes using NMR tool. In: 2nd international plant phenotyping symposium toward plant phenotyping science: challenges and perspectives, Forschungszentrum Jülich, Germany, pp 13
Li L, Zhang Q, Huang D (2014) A review of imaging techniques for plant phenotyping. Sensors 14:20078–20111
Li Y, Huang Y, Bergelson J, Nordborg M, Borevitz JO (2010) Association mapping of local climate-sensitive quantitative trait loci in Arabidopsis thaliana. Proc Natl Acad Sci 107:21199–21204
Li Y, Xiao J, Chen L, Huang X, Cheng Z, Han B, Zhang Q, Wu C (2018) Rice functional genomics research: past decade and future. Mol Plant 11:359–380
Lymperopoulos P, Msanne J, Rabara R (2018) Phytochrome and phytohormones: working in Tandem for plant growth and development. Front Plant Sci 9:1037–1037
Mahlein AK, Kuska MT, Behmann J, Polder G, Walter A (2018) Hyperspectral sensors and imaging technologies in phytopathology: state of the art. Annu Rev Phytopathol 56:535–558
Minamikawa MF, Nonaka K, Kaminuma E, Kajiya-Kanegae H, Onogi A, Goto S, Yoshioka T, Imai A, Hamada H, Hayashi T (2017) Genome-wide association study and genomic prediction in citrus: potential of genomics-assisted breeding for fruit quality traits. Sci Rep 7:1–13
Montes JM, Utz HF, Schipprack W, Kusterer B, Muminovic J, Paul C, Melchinger AE (2006) Near-infrared spectroscopy on combine harvesters to measure maize grain dry matter content and quality parameters. Plant Breed 125:591–595
Montes JM, Melchinger AE, Reif JC (2007) Novel throughput phenotyping platforms in plant genetic studies. Trends Plant Sci 12:433–436
Mukherjee S, Mishra A, Trenberth KE (2018) Climate change and drought: a perspective on drought indices. Curr Clim Change Rep 4:145–163
Mula MG, Patil SB, Aden J, Rathore A, Anil Kumar V, Kumar RV (2016) Screening of pigeonpea genotypes for drought stress at early vegetative phase in Alfisol and Vertisol. Green Farming 7:507–511
Muraya MM, Chu J, Zhao Y, Junker A, Klukas C, Reif JC, Altmann T (2017) Genetic variation of growth dynamics in maize (Zea mays L.) revealed through automated non-invasive phenotyping. Plant J 89:366–380
Nabwire S, Suh H-K, Kim MS, Baek I, Cho B-K (2021) Application of artificial intelligence in phenomics. Sensors 21:4363
Nagel KA, Putz A, Gilmer F, Heinz K, Fischbach A, Pfeifer J, Faget M, Blossfeld S, Ernst M, Dimaki C (2012) GROWSCREEN-Rhizo is a novel phenotyping robot enabling simultaneous measurements of root and shoot growth for plants grown in soil-filled rhizotrons. Funct Plant Biol 39:891–904
Neumann K, Klukas C, Friedel S, Rischbeck P, Chen D, Entzian A, Stein N, Graner A, Kilian B (2015) Dissecting spatiotemporal biomass accumulation in barley under different water regimes using high-throughput image analysis. Plant Cell Environ 38:1980–1996
Neumann K, Zhao Y, Chu J, Keilwagen J, Reif JC, Kilian B, Graner A (2017) Genetic architecture and temporal patterns of biomass accumulation in spring barley revealed by image analysis. BMC Plant Biol 17:1–12
Obidiegwu JE, Bryan GJ, Jones HG, Prashar A (2015) Coping with drought: stress and adaptive responses in potato and perspectives for improvement. Front Plant Sci 6:542
Oerke E-C, Steiner U, Dehne H-W, Lindenthal M (2006) Thermal imaging of cucumber leaves affected by downy mildew and environmental conditions. J Exp Bot 57:2121–2132
Parent B, Shahinnia F, Maphosa L, Berger B, Rabie H, Chalmers K, Kovalchuk A, Langridge P, Fleury D (2015) Combining field performance with controlled environment plant imaging to identify the genetic control of growth and transpiration underlying yield response to water-deficit stress in wheat. J Exp Bot 66:5481–5492
Pasala R, Pandey BB (2020) Plant phenomics: high-throughput technology for accelerating genomics. J Biosci 45:1–6
Patanè C, Scordia D, Testa G, Cosentino SL (2016) Physiological screening for drought tolerance in Mediterranean long-storage tomato. Plant Sci 249:25–34
Pereyra-Irujo GA, Gasco ED, Peirone LS, Aguirrezábal LAN (2012) GlyPh: a low-cost platform for phenotyping plant growth and water use. Funct Plant Biol 39:905–913
Poorter H, Fiorani F, Pieruschka R, Wojciechowski T, van der Putten WH, Kleyer M, Schurr U, Postma J (2016) Pampered inside, pestered outside? Differences and similarities between plants growing in controlled conditions and in the field. New Phytol 212:838–855
Pound MP, Atkinson JA, Townsend AJ, Wilson MH, Griffiths M, Jackson AS, Bulat A, Tzimiropoulos G, Wells DM, Murchie EH (2017) Deep machine learning provides state-of-the-art performance in image-based plant phenotyping. Gigascience 6:gix083
Pratap A, Gupta S, Nair RM, Gupta SK, Schafleitner R, Basu PS, Singh CM, Prajapati U, Gupta AK, Nayyar H, Mishra AK, Baek K-H (2019) Using plant phenomics to exploit the gains of genomics. Agronomy 9
Rasheed A, Xia X, Ogbonnaya F, Mahmood T, Zhang Z, Mujeeb-Kazi A, He Z (2014) Genome-wide association for grain morphology in synthetic hexaploid wheats using digital imaging analysis. BMC Plant Biol 14:1–21
Rebolledo MC, Peña AL, Duitama J, Cruz DF, Dingkuhn M, Grenier C, Tohme J (2016) Combining image analysis, genome wide association studies and different field trials to reveal stable genetic regions related to panicle architecture and the number of spikelets per panicle in rice. Front Plant Sci 7:1384
Reuzeau C, Pen J, Frankard V, de Wolf J, Peerbolte R, Broekaert WF, van Wim C (2010) TraitMill: a discovery engine for identifying yield-enhancement genes in cereals. Plant Gene Trait
Roscher R, Bohn B, Duarte MF, Garcke J (2020) Explainable machine learning for scientific insights and discoveries. Ieee Access 8:42200–42216
Rosenzweig C, Elliott J, Deryng D, Ruane AC, Müller C, Arneth A, Boote KJ, Folberth C, Glotter M, Khabarov N (2014) Assessing agricultural risks of climate change in the 21st century in a global gridded crop model intercomparison. Proc Natl Acad Sci 111:3268–3273
Ruiz-Garcia L, Lunadei L, Barreiro P, Robla I (2009) A review of wireless sensor technologies and applications in agriculture and food industry: state of the art and current trends. Sensors 9:4728–4750
Sabanci K, Kayabasi A, Toktas A (2017) Computer vision-based method for classification of wheat grains using artificial neural network. J Sci Food Agric 97:2588–2593
Sadok W, Naudin P, Boussuge B, Muller B, Welcker C, Tardieu F (2007) Leaf growth rate per unit thermal time follows QTL-dependent daily patterns in hundreds of maize lines under naturally fluctuating conditions. Plant Cell Environ 30:135–146
Shi C, Zhao L, Zhang X, Lv G, Pan Y, Chen F (2019) Gene regulatory network and abundant genetic variation play critical roles in heading stage of polyploidy wheat. BMC Plant Biol 19:1–16
Shi L, Shi T, Broadley MR, White PJ, Long Y, Meng J, Xu F, Hammond JP (2013) High-throughput root phenotyping screens identify genetic loci associated with root architectural traits in Brassica napus under contrasting phosphate availabilities. Ann Bot 112:381–389
Singh AK, Ganapathysubramanian B, Sarkar S, Singh A (2018) Deep learning for plant stress phenotyping: trends and future perspectives. Trends Plant Sci 23:883–898
Sirault X, Fripp J, Paproki A, Kuffner P, Nguyen C, Li R, Daily H, Guo J, Furbank R (2013) PlantScan: a three-dimensional phenotyping platform for capturing the structural dynamic of plant development and growth. In: Proceedings of the 7th international conference on functionalstructural plant models, Saariselk, Finland, pp 45–48
Sterck L, Rombauts S, Vandepoele K, Rouzé P, Van de Peer Y (2007) How many genes are there in plants (… and why are they there)? Curr Opin Plant Biol 10:199–203
Taylor JF (2014) Implementation and accuracy of genomic selection. Aquaculture 420:S8–S14
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822
Tisné S, Serrand Y, Bach L, Gilbault E, Ben Ameur R, Balasse H, Voisin R, Bouchez D, Durand-Tardif M, Guerche P, Chareyron G, Da Rugna J, Camilleri C, Loudet O (2013) Phenoscope: an automated large-scale phenotyping platform offering high spatial homogeneity. Plant J 74:534–544
Vadez V, Kholová J, Hummel G, Zhokhavets U, Gupta SK, Hash CT (2015) LeasyScan: a novel concept combining 3D imaging and lysimetry for high-throughput phenotyping of traits controlling plant water budget. J Exp Bot 66:5581–5593
Van Raan AFJ (2004) Measuring science, Handbook of quantitative science and technology research. Springer, pp 19–50
Virlet N, Sabermanesh K, Sadeghi-Tehran P, Hawkesford MJ (2016) Field Scanalyzer: an automated robotic field phenotyping platform for detailed crop monitoring. Functional Plant Bio 44(1):143–153.
Walter T, Shattuck DW, Baldock R, Bastin ME, Carpenter AE, Duce S, Ellenberg J, Fraser A, Hamilton N, Pieper S (2010) Visualization of image data from cells to organisms. Nat Methods 7:S26–S41
Walter A, Scharr H, Gilmer F, Zierer R, Nagel KA, Ernst M, Wiese A, Virnich O, Christ MM, Uhlig B (2007) Dynamics of seedling growth acclimation towards altered light conditions can be quantified via GROWSCREEN: a setup and procedure designed for rapid optical phenotyping of different plant species. New Phytologist 174(2):447–455
Wang P, Zhou G, Yu H, Yu S (2011) Fine mapping a major QTL for flag leaf size and yield-related traits in rice. Theor Appl Genet 123:1319–1330
Wang Q, Xie W, Xing H, Yan J, Meng X, Li X, Fu X, Xu J, Lian X, Yu S (2015) Genetic architecture of natural variation in rice chlorophyll content revealed by a genome-wide association study. Mol Plant 8:946–957
Wang X, Zhang R, Song W, Han L, Liu X, Sun X, Luo M, Chen K, Zhang Y, Yang H (2019) Dynamic plant height QTL revealed in maize through remote sensing phenotyping using a high-throughput unmanned aerial vehicle (UAV). Sci Rep 9:1–10
Waring RH, Cleary BD (1967) Plant moisture stress: evaluation by pressure bomb. Science 155:1248–1254
Weight C, Parnham D, Waites R (2008) TECHNICAL ADVANCE: LeafAnalyser: a computational method for rapid and large-scale analyses of leaf shape variation. Plant J 53:578–586
Wetterich CB, Kumar R, Sankaran S, Junior JB, Ehsani R, Marcassa LG A comparative study on application of computer vision and fluorescence imaging spectroscopy for detection of citrus huanglongbing disease in USA and Brazil. Opt Soc Am: JW3A–JW26A
Wu D, Guo Z, Ye J, Feng H, Liu J, Chen G, Zheng J, Yan D, Yang X, Xiong X, Liu Q, Niu Z, Gay AP, Doonan JH, Xiong L, Yang W (2019) Combining high-throughput micro-CT-RGB phenotyping and genome-wide association study to dissect the genetic architecture of tiller growth in rice. J Exp Bot 70:545–561
Wu X et al (2021) Using high-throughput multiple optical phenotyping to decipher the genetic architecture of maize drought tolerance. Genome Biol 22:185
Wu H, Jaeger M, Wang M, Li B, Zhang BG (2011) Three-dimensional distribution of vessels, passage cells and lateral roots along the root axis of winter wheat (Triticum aestivum). Ann of Bot 107(5):843–853
Xiao Y, Liu H, Wu L, Warburton M, Yan J (2017) Genome-wide association studies in maize: praise and stargaze. Mol Plant 10:359–374
Xue B, Sartori P, Leibler S (2019) Environment-to-phenotype mapping and adaptation strategies in varying environments. Proc Natl Acad Sci 116:13847–13855
Yang W, Guo Z, Huang C, Duan L, Chen G, Jiang N, Fang W, Feng H, Xie W, Lian X (2014) Combining high-throughput phenotyping and genome-wide association studies to reveal natural genetic variation in rice. Nat Commun 5:1–9
Yang W, Guo Z, Huang C, Wang K, Jiang N, Feng H, Chen G, Liu Q, Xiong L (2015) Genome-wide association study of rice (Oryza sativa L.) leaf traits with a high-throughput leaf scorer. J Exp Bot 66:5605–5615
Yang W, Feng H, Zhang X, Zhang J, Doonan JH, Batchelor WD, Xiong L, Yan J (2020) Crop phenomics and high-throughput phenotyping: past decades, current challenges, and future perspectives. Mol Plant 13:187–214
Yang Y, Chai R, He Y (2012) Early detection of rice blast (Pyricularia) at seedling stage in Nipponbare rice variety using near-infrared hyper-spectral image. Afr J Biotechnol 11:6809–6817
Yao W, Li G, Yu Y, Ouyang Y (2018) funRiceGenes dataset for comprehensive understanding and application of rice functional genes. Gigascience 7:1–9
Zakaluk R, Ranjan R (2008) Predicting the leaf water potential of potato plants using RGB reflectance. Can Biosyst Eng 50
Zarco-Tejada PJ, Berni JAJ, Suárez L, Sepulcre-Cantó G, Morales F, Miller JR (2009) Imaging chlorophyll fluorescence with an airborne narrow-band multispectral camera for vegetation stress detection. Remote Sens Environ 113:1262–1275
Zhang C, Pumphrey M, Zhou J, Gao H, Zhang Q, Sankaran S (2017) Development of automated highthroughput phenotyping system for controlled environment studies, Paper No. 1700581, 2017 American Society of Agricultural and Biological Engineers (ASABE) Annual International Meeting (AIM), Spokane, WA
Zhang Q, Li J, Xue Y, Han B, Deng XW (2008) Rice 2020: a call for an international coordinated effort in rice functional genomics. Mol Plant 1:715–719
Zhao C, Liu B, Piao S, Wang X, Lobell DB, Huang Y, Huang M, Yao Y, Bassu S, Ciais P (2017) Temperature increase reduces global yields of major crops in four independent estimates. Proc Natl Acad Sci 114:9326–9331
Zhao C, Zhang Y, Du J, Guo X, Wen W, Gu S, Wang J, Fan J (2019) Crop phenomics: current status and perspectives. Front Plant Sci:10
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Waseem, M., Shaheen, I., Aslam, M.M. (2022). Advances in Integrated High-Throughput and Phenomics Application in Plants and Agriculture. In: Prakash, C.S., Fiaz, S., Fahad, S. (eds) Principles and Practices of OMICS and Genome Editing for Crop Improvement. Springer, Cham. https://doi.org/10.1007/978-3-030-96925-7_10
Download citation
DOI: https://doi.org/10.1007/978-3-030-96925-7_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-96924-0
Online ISBN: 978-3-030-96925-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)