Abstract
Mortality in elderly population having type II diabetes (T2D) can be prevented sometimes through intervention. For that risk assessment can be performed through predictive modeling. This study is part of a collaboration with Maccabi Healthcare Services’ Electronic Health Records (EHR) data, that consists on up to 10 years of 18,000 elderly T2D patients. EHR data is typically heterogeneous and sparse, and for that the use of temporal abstraction and time intervals mining to discover frequent time-interval related patterns (TIRPs) are employed, which then are used as features for a predictive model. However, while the temporal relations between symbolic time intervals in a TIRP are discovered, the temporal relations between TIRPs are not represented. In this paper we introduce a novel TIRPs based patient data representation called Integer-TIRP (iTirp), in which the TIRPs become channels represented by values representing the number of TIRP’s instances that were detected. Then, the iTirps representation is fed into a Deep Learning Architecture, which can learn this kind of sequential relations, using a Recurrent Neural Network (RNN) or a Convolutional Neural Network (CNN). Finally, we introduce a predictive model that consists of a committee, in which two inputs were concatenated, a raw data and iTirps data. Our results indicate that iTirps based models, showed superior performance compared to raw data representation and the committee showed even better results, this by taking advantage of each representations.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Diabetes is a major chronic disease in the western society and its prevalence is on the rise worldwide. Type 2 diabetes (T2D) patients often suffer from heart disease and the prevalence of coronary artery disease and heart failure is also much higher among diabetic patients [4]. Moreover, cardiac related in-hospital mortality is also much higher among patients with diabetes [12].
Israel’s HMOs had implemented disease management programs to improve quality of care for diabetes and prevent those complications through risk reduction [10]. These programs aim at achieving centrally controlled documented multifactorial risk reduction that are implemented mainly by primary care givers. To date the effect of these programs on cardiac morbidity and mortality were not assessed. A potential deficiency in these plans may be the lack of targeted case management for high-risk patients. Such identification may lead to on time intensive intervention that may reduce morbidity and mortality. Namely, prevent hospitalization for cardiac disease and lower cardiac mortality. To this end it is desirable to develop a predictive model that will help to identify the patients that are more prone to cardiac deterioration. This will form the basis for intervention aimed at prevention of costly and lethal consequences. For that purpose, in this paper the focus is on prediction of All-Cause Mortality in T2D patients.
In this paper we introduce for the first time iTirps, which are temporal patterns based representation that can be later fed into temporal architectures of Artificial Neural Networks (ANNs), which we use to learn predictive models for outcomes, which in our study is all cause mortality in T2D patients. To have the iTirps representation, first temporal abstraction is used [19] and time-intervals mining to discover TIRPs [19]. Then these are transformed into a new representation, called integer-TIRPs, which are described in greater details later. The contributions of the paper are the following: 1. iTirps, a novel representation for temporal data consisting on frequent TIRPs instances, which enable to represent a time period according to the relations among the temporal variables along time, and their appearances, which are hard to represent by TIRPs, nor by temporal ANNs. 2. A rigorous evaluation on a large real-life data of T2D patients, using iTirps for the prediction of all-cause mortality.
2 Related Work
We start with a review of the use of data science in diabetic patients’ data. Then we proceed with discussing time intervals related patterns mining in heterogeneous multivariate temporal data and their use for classification, and then we go over approaches in the field of ANN for time series classification.
2.1 Outcomes Prediction in Diabetes
The use of data mining and machine learning methods in diabetes related research is constantly increasing [11]. There is a relatively small number of studies that intend to predict mortality in T2D patient. For example, prediction of ICU mortality of diabetic patient by applying several classifiers on aggregated data and showed good results on predict risk of mortality [1]. Most of current research that assesses mortality risk in diabetic patients, are using Cox proportional hazards model, in [16] used Cox model to create risk equations for all cause, cardiovascular, and non-cardiovascular mortality diagnosed of type 2 diabetes patients. In [5] a Cox model was used specifically for prediction of mortality in adults population. The use of ANN in diabetes related research was not very extensive, and most of the work use feed forward (FF) network on a temporal data [9].
2.2 Temporal Abstraction, TIRPs Discovery and TIRPs Based Classification
A major challenge in analyzing EHR data is the heterogeneity of the sampling forms of the data. Additionally, challenges may include sparsity, and exploiting the temporal information. Therefore, increase usage of temporal abstraction (TA) and time intervals mining is being reported [19]. In order to transform the heterogeneous temporal variables into a uniform representation, state TA is used, in which the time point series are transformed into symbolic time intervals (STIs), given a set of cutoffs. The cutoffs can be knowledge based [20], or data driven, based on discretization methods, such as Symbolic Aggregate approXimation [14] or the Temporal Discretization for Classification (TD4C) [19, 21]. Another type of temporal abstraction is gradient abstraction, which segments the data based on the first derivative into periods of time, in which the variable is increasing or decreasing [20]. Once symbolic time intervals series are created, frequent Time Intervals Related Patterns (TIRPs) can be discovered. Several methods for TIRPs discovery were proposed in the past [19, 21], mostly consisting on Allen’s temporal relations [17] which include seven relations such as before, meet, overlap, and more, and their inverse. Beyond temporal knowledge discovery, frequent TIRPs were shown to be effective for classification and prediction in electronic health records [3, 17, 18, 22]. However, incorporating the use of TIRPs to represent the temporal relations between heterogeneous temporal variables in ANNs based architectures is still a challenge, which we explore in this paper.
2.3 Artificial Neural Networks for Temporal Data
ANNs designed for temporal data were successfully used in several domains and tasks. For example, RNN can store information about previous inputs in internal memory (hidden states), that abstract and carry information from earlier time stamps and CNN are achieving state-of-the-art results in a high variety of tasks including computer vision tasks and more [13]. These methods are increasingly employed also in clinical data. RNN based methods showed superior results than the use of classical algorithm like logistic regression (LR) and multilayer perceptron with hand-engineered features in predicting diagnosis codes [8, 15]. RNN based method for missing values imputation in temporal data, called GRU-D [6], showed better performance than traditional methods, such as mean-imputation, imputation with k-nearest neighbor and other. Modified CNN, for capturing temporal relations, was trained CNN on temporal matrix representation of medical codes for outcome prediction and showed better results than LR using aggregated clinical features on real world EHR data [7].
3 Methods
We present here the framework for the development of the iTirps and the iTirpsMap representation which is used later with temporal ANNs for classification and specifically prediction in this study. With iTirps representation we preserve more temporal information then the regular TIRP representation. iTirps hold the information about the starting and ending time of each TIRP, and through that the duration of each TIRP instance is captured, including its relative location in the time series. This information can be learned by ANN and improve the classification performance.
3.1 iTirps and iTirpsMap
Figure 1 presents the steps in the creation of iTirps and a corresponding iTirpsMap. First the multivariate temporal data abstracted and transformed into a uniform representation of symbolic time intervals [19]. Then, frequent TIRPs are discovered by mining the symbolic time intervals. For the mining process, we use the KarmaLego algorithm [21]. The result is a bag of frequent TIRPs. Next, the TIRPs are detected and transformed to iTirp, that are passed in the form of iTirpsMap as input to a CNN/RNN.
Temporal Abstraction. In this study we perform state abstraction using Symbolic Aggregate approXimation (SAX) [14], in which the states are derived from the Gaussian distribution of the values, and Temporal Discretization for Classification (TD4C) [19] that determines the cutoffs in a supervised manner, so that the states distribution are most different among the classes. The result of the Temporal Abstraction process is a uniform representation of the temporal variables as symbolic time intervals. A symbolic time interval, \(I = {<}s, e, sym{>}\), is an ordered pair of time points, start-time (s) and end-time (e), and a symbol (sym) that represents one of the domain’s symbolic concepts, which in our study can be laboratory results that went through abstraction, conditions or procedures. As mentioned in the background, once the data is transformed into a uniform representation of symbolic time intervals, TIRPs can be discovered.
TIRPs Discovery. To discover TIRPs, the KarmaLego algorithm [19] is used, which uses Allen’s temporal relations, such as starts, meets, overlap, contains, and more, and their inverse [20] to represent the temporal relation among a pair of symbolic time intervals. In this study a set of generalized temporal relations, which are the disjunction of part of Allen’s seven relations were used.
These include: BEFORE based on before||meets; OVERLAP based on overlaps; and CONTAIN based on \(\{starts\) || contains || \(finish-by\) || \(equal\}\). In addition, a maximum allowed gap duration is set for the before relation [20]. A non-ambiguous TIRP P is defined as \(P = {I ,R}\), where \(I = {I1, I2,.., Ik}\) is a set of k ordered symbolic time intervals and the conjunction of all their pairwise temporal relations among each of the (k2 − k)/2 pairs of the symbolic time intervals in I, \( R= {U}_{i=1}^{k-1}{U}_{j=i+1}^{k}r(I^{i},I^{j}){r_{1,2} (I^1,I^2 ),..,r_{1,k} (I^1,I^k ),...,r_{k-1,k}(I^{k-1}I^k )}\). Thus, given a database of entities (i.e., patients), the vertical support of a TIRP P (frequency in the database) is denoted by the cardinality of the distinct entities having P, relative to the size of the database. However, in this study we propose a novel use of the TIRPs, which become channels, and called iTirps.
iTirp and iTirpsMap Creation. We introduce here iTirps, a new temporal representation of multivariate temporal data through TIRPs’ instances that results in a numeric matrix representation of the appearance of the TIRPs along time, which can be later fed to various methods, such as RNN/CNN as happens in this study. In previous studies TIRPs were used as features for classifiers [3, 19], however, in order to represent them explicitly along time, we present iTirpsMap. Figure 2 illustrates the process of the iTirps and iTirpsMap creation. The description starts at the bottom and goes up. The x-axis is the time by months along 12 months. Starting with the symbolic time intervals at the bottom, which can be raw concepts, such as drug exposers, conditions, or a state abstraction of time point series, such as lab tests.
In Fig. 2 there are three STIs at the bottom, HbA1c_High, which represents their measurements abstracted along three months (which a HbA1c test is valid for), Dipeptidyl peptidase-4 (DPP4) Inhibitors, which are a class of medications that decrease high blood glucose, and Doctor Visit events. Two TIRPs examples are presented above: TIRP HH b Do (HH before Do) appears three times in the periods of 16, 1–8, and 10–12; and the TIRP HH b D c Do (HbA1c before DPP4 Inhibitors, HbA1c High before Doctor Visit, and DPP4 Inhibitors contains Doctor Visit) which appears twice (since there two Doctor Visits) during 1–9. In fact, the HH_b_Do TIRP is shown in the bottom illustration of the STIs surrounding the relevant STIs which include each HH_b_Do instance – there are two. To create iTirps that construct the iTirpsMap, we have two steps, in the first step each TIRP instance becomes a time series of one and zero values (one values are placed from TIRP starting point to ending point). In the second step, we aggregate the TIRPs to create iTIRPs. Thus, for example, iTirp HH b Do value is 2 in time stamps 1–6, since there are two instances of the TIRP during this time stamp. Thus, an iTirp represents the number of the TIRP appearances in each time stamp. Eventually, the entire set of iTirps are combined into an iTirpsMap 3-dimensional matrix (of the Entities, the time axis, and the TIRPs’ channels) representation.
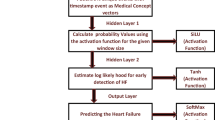
Artificial Neural Network. In this paper the purpose of iTirps and iTirpsMap is to enable to combine the advantages of TIRPs in capturing temporal relations between heterogeneous temporal variables and the advantages of neural learning, specifically when using temporal versions of ANNs, such as the RNN, CNN and their ensemble. RNN-ALSTM. Our RNN architecture is an Attention block followed by a LSTM (ALSTM). The attention mechanism enables the network to better learn long-term dependencies for the prediction task proposed by [2]. Long Short-Term Memory (LSTM) [13] is variation of RNN, that can overcome RNN’s limitations like vanishing and exploding gradient by a gating mechanism that regulates the information flow. Encoder-CNN. For the CNN architecture the Encoder is used. In Encoder the first three layers are CNN that followed by attention mechanism, that summarize the temporal dimension, proposed in [23]. To map the network output to a probability distribution the last layer is SoftMax, for both networks. Committee. We experiment also with a committee of two classifiers, in which the first classifier is based on the raw data, and the second classifier is based on the iTirpsMap input. First, we train each model separately, with the different inputs, one with raw data and the second with the iTirpsMap, based on some type of TA. Then the SoftMax layer is removed from both models, while the last layers of the network are concatenated, and a new SoftMax added as the last layer.
4 Evaluation and Results
We first state our research questions, and then we describe the data, and the experiments that were designed to answer the questions, and the results.
4.1 Research Questions
1. What type of temporal abstraction is best for classification? 2. What are the best prediction time periods? 3. Which ANN with iTirps performs best, in comparison to the use of raw data? 4. What ‘Committee’ of ANNs is the best for outcome prediction?
4.2 Dataset
The diabetes dataset of Maccabi Healthcare Services contains data of up to 10 years on 18,000 elderly patients with T2D. The dataset includes 9,000 cases, which are T2D patients who experienced an outcome, defined as all-cause mortality. The data collected from the years 2008–2018. Cohort Inclusion criteria: (1) Patients with diabetes according to the diabetes registry (and not defined as type 1). (2) Experiences an outcome from 2011–2018. 9,000 controls, which are patients without the outcome that were matched according 2 parameters Age and Gender to control patients. Control (Matched patients) will be defined as followed: (1) Patients with diabetes according to the diabetes registry (and not defined as type 1). (2) Being Maccabi Health Services members and without recorded outcomes during the outcome period. Patients included in cancer registry prior to outcome are excluded from the dataset. Control patients, outcome date will be defined as January 1st. The variables include Demographic data, Therapies (medication), Co-morbidities indicators, Lab results, hospitalizations and inpatients and outpatient visits.
4.3 Experimental Setup
To answer the research questions, while reflecting a real application conditions for continuous prediction using a sliding window, the most suitable study design is case-crossover-control. Thus, observation time windows are extracted from the cases, and the matched controls. In the cases, the latest observation time window, which is located a prediction time period prior to the outcome is labeled as positive, while the earlier observation time windows in the cases are labeled as negative, as well as the observation time windows from the controls’ data (taken randomly, since there are no outcomes), which enables to evaluate the method both on cases’, or controls’ time windows. We report quantitative results using the Receiver Operating Characteristic AUC (ROC-AUC), based on 10-fold grouped cross-validation (CV). Thus, time windows of a specific patient were either at the training or in the testing set. To answer the research questions, two experiments were designed. We used an observation time period of 12 months, and to discover TIRPs, KarmaLego was applied with 55% minimal vertical support, and unlimited maximal gap.
Experiment 1. The goal was to evaluate the iTirps based prediction, using an observation time window of 12 months. For this experiment we have two possible inputs for the ANN architectures: raw data, or the iTirpsMap based on the two types of abstraction (research question 1) using the SAX and TD4C-Cos with 2 states. All inputs with Encoder-CNN and RNN-ALSTM were evaluated on two prediction time periods of 90 and 180 days (research question 2, 3). Figure 3 presents the mean results of the iTirps based on the SAX or TD4C-Cos, in comparison to the use of raw data, with two prediction time periods of 90 and 180 days. Generally predicting 90 days performed better than within 180 days, which makes sense. Using the Encoder-CNN the use of iTirps with SAX performed significantly better than the other. Using the RNN-ALSTM the results were quite similar, and the iTirp with SAX performed best.
Experiment 2. To evaluate what is the best committee (research question 4) the performance of the ‘Committee’ merged network was evaluated. The committees included a combination of the raw data as input to one classifier, and the second with the iTirpsMap based on SAX or TD4C-Cos TA. Figure 4 presents the performance of the two committees, including for comparison the use of only raw data (as performed in the first experiment, as a baseline). The committee using iTirp with SAX and raw data, was significantly better with Encoder-CNN when predicting 90 days ahead, and also with the other options. Overall, the best performance was 84.5% AUC.
5 Discussion and Conclusions
Clinically, we focused on the prediction of all-cause mortality in T2D patients, to enable ideally prevention through intervention. In this paper, we introduced a new method for multivariate temporal data representation, called iTirps, which is consists on the discovery of frequent TIRPs, which enables to be fed to RNN or CNN. The use of temporal abstraction and TIRPs as features for classification, is very effective for the analysis of heterogeneous multivariate temporal data, such as often happens in Electronic Health Records. However, in order to employ the advantages of the temporal versions of ANNs, such as CNN or RNN, we propose the iTirps representation. Additionally, we proposed and experimented with the Committee network that combines both raw data and the iTirpsMap. The results of Experiment 1 show that using the iTirp with SAX with the Encoder-CNN had the best performance, and generally the use of the Encoder-CNN and RNN-ALSTM were comparable. In experiment 2, the performance of the committee network was evaluated, in which the committee of the Encoder-CNN with iTirp SAX and the raw data as input, performed best and was significantly higher compared to the raw data. The directions for future research include evaluation of additional discretization methods and number of bins, on a bigger number of SOTA ANN architectures.
References
Anand, R.S., et al.: Predicting mortality in diabetic ICU patients using machine learning and severity indices. AMIA Summits Transl. Sci. Proc. 2018, 310 (2018)
Bahdanau, D., Cho, K., Bengio, Y.: Neural machine translation by jointly learning to align and translate. arXiv preprint arXiv:1409.0473 (2014)
Batal, I., Fradkin, D., Harrison, J., Moerchen, F., Hauskrecht, M.: Mining recent temporal patterns for event detection in multivariate time series data. In: Proceedings of the 18th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, pp. 280–288 (2012)
Bo, S., et al.: Patients with type 2 diabetes had higher rates of hospitalization than the general population. J. Clin. Epidemiol. 57(11), 1196–1201 (2004)
Chang, Y., et al.: A point-based mortality prediction system for older adults with diabetes. Sci. Rep. 7(1), 1–10 (2017)
Che, Z., Purushotham, S., Cho, K., Sontag, D., Liu, Y.: Recurrent neural networks for multivariate time series with missing values. Sci. Rep. 8(1), 1–12 (2018)
Cheng, Y., Wang, F., Zhang, P., Hu, J.: Risk prediction with electronic health records: a deep learning approach. In: Proceedings of the 2016 SIAM International Conference on Data Mining, pp. 432–440. SIAM (2016)
Choi, E., Bahadori, M.T., Schuetz, A., Stewart, W.F., Sun, J.: Doctor AI: predicting clinical events via recurrent neural networks. In: Machine Learning for Healthcare Conference, pp. 301–318 (2016)
El\(\_\)Jerjawi, N.S., Abu-Naser, S.S.: Diabetes prediction using artificial neural network (2018)
Heymann, A.D., et al.: The implementation of managed care for diabetes using medical informatics in a large Preferred Provider Organization. Diab. Res. Clin. Pract. 71(3), 290–298 (2006)
Kavakiotis, I., Tsave, O., Salifoglou, A., Maglaveras, N., Vlahavas, I., Chouvarda, I.: Machine learning and data mining methods in diabetes research. Comput. Struct. Biotechnol. J. 15, 104–116 (2017)
Khalid, J., Raluy-Callado, M., Curtis, B., Boye, K., Maguire, A., Reaney, M.: Rates and risk of hospitalisation among patients with type 2 diabetes: retrospective cohort study using the UK General Practice Research Database linked to English hospital episode statistics. Int. J. Clin. Pract. 68(1), 40–48 (2014)
LeCun, Y., Bengio, Y., Hinton, G.: Deep learning. Nature 521(7553), 436–444 (2015)
Lin, J., Keogh, E., Wei, L., Lonardi, S.: Experiencing SAX: a novel symbolic representation of time series. Data Min. Knowl. Discov. 15(2), 107–144 (2007)
Lipton, Z.C., Kale, D.C., Elkan, C., Wetzel, R.: Learning to diagnose with LSTM recurrent neural networks. arXiv preprint arXiv:1511.03677 (2015)
McEwen, L.N., et al.: Predictors of mortality over 8 years in type 2 diabetic patients: Translating Research Into Action for Diabetes (triad). Diabetes Care 35(6), 1301–1309 (2012)
Moskovitch, R., Choi, H., Hripcsak, G., Tatonetti, N.P.: Prognosis of clinical outcomes with temporal patterns and experiences with one class feature selection. IEEE/ACM Trans. Comput. Biol. Bioinform. 14(3), 555–563 (2016)
Moskovitch, R., Polubriaginof, F., Weiss, A., Ryan, P., Tatonetti, N.: Procedure prediction from symbolic electronic health records via time intervals analytics. J. Biomed. Inform. 75(C), 70–82 (2017). https://doi.org/10.1016/j.jbi.2017.07.018
Moskovitch, R., Shahar, Y.: Classification-driven temporal discretization of multivariate time series. Data Min. Knowl. Discov. 29(4), 871–913 (2014). https://doi.org/10.1007/s10618-014-0380-z
Moskovitch, R., Shahar, Y.: Classification of multivariate time series via temporal abstraction and time intervals mining. Knowl. Inf. Syst. 45(1), 35–74 (2015)
Moskovitch, R., Walsh, C., Wang, F., Hripcsak, G., Tatonetti, N.: Outcomes prediction via time intervals related patterns. In: 2015 IEEE International Conference on Data Mining, pp. 919–924. IEEE (2015)
Sacchi, L., Larizza, C., Combi, C., Bellazzi, R.: Data mining with temporal abstractions: learning rules from time series. Data Min. Knowl. Discov. 15(2), 217–247 (2007)
Serrà, J., Pascual, S., Karatzoglou, A.: Towards a universal neural network encoder for time series. In: CCIA, pp. 120–129 (2018)
Acknowledgements
The authors wish to thank the Israeli Ministry of Science and Technology, who assisted in funding this project with grant 8760521.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this paper
Cite this paper
Novitski, P., Cohen, C.M., Karasik, A., Shalev, V., Hodik, G., Moskovitch, R. (2020). All-Cause Mortality Prediction in T2D Patients. In: Michalowski, M., Moskovitch, R. (eds) Artificial Intelligence in Medicine. AIME 2020. Lecture Notes in Computer Science(), vol 12299. Springer, Cham. https://doi.org/10.1007/978-3-030-59137-3_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-59137-3_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-59136-6
Online ISBN: 978-3-030-59137-3
eBook Packages: Computer ScienceComputer Science (R0)