Abstract
The development of cancer results from the evolutionary balance between the proliferating aptitude of cancer cells and the response of the host’s tissues. Some cancers are characterized by genetic instability dependent upon impaired DNA repair mechanisms that lead to the chaotic disruption of multiple cellular functions often in excess of the cancer survival needs and may exert broad effects on surrounding tissues, some beneficial and some detrimental to cancer growth. Among them, inflammatory processes that accompany wound healing may initiate a reaction of the host against the neo-formation. This is possibly triggered by the release by dying cancer cells of molecules known as Damage-Associated Molecular Patterns (DAMPs) following a process termed Immunogenic Cell Death (ICD) that initiates an immune response through innate and adaptive mechanisms. Indeed, analysis of large cancer data sets has shown that ICD is strictly associated with the activation of other immune effector or immune-regulatory pathways. Here, we will describe how immune activation and compensatory immune-regulatory mechanisms balance anti-cancer immune surveillance and the roles that innate and adaptive immunity play including the weight that neo-epitopes may exert as initiators and sculptors of high-affinity memory and effector immune responses against cancer. We will discuss the evolutionary basis for the existence of immune checkpoints and how several theories raised to explain cancer resistance to immunotherapy represent a facet of a similar evolutionary phenomenon that we described in the Theory of Everything. We will show how the biology of immunogenicity and counterbalancing immune regulation is widespread across cancers independent of their ontogenesis while subtle idiosyncratic differences are discernible among them. Finally, we will suggest that overcoming immune resistance implies distinct approaches relevant to the immune context of individual cancers.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Cancer development results from the evolutionary balance between the proliferating aptitude of cancer cells and the response of the host‘s tissues [1]. The latter is dependent upon paracrine factor released by cancer cells and upon cell-to-cell contact interactions that stimulate tissue remodeling to support the growth of the neo-formation. In the process, a balance can be stricken by two opposite strategies [2, 3]: in the simplest, cancer initiation, promotion, and progression follow a cumulative and coherent process of progressive genetic alterations that lead to cancer-initiating cells [4, 5] that involve critical mutations of genes regulating normal cell growth and differentiation with minimal disruption of surrounding tissues similar to the strategy adopted by non-cancer stem cells during normal development [6]. At the other extreme, genetic instability dependent upon disruption of several surveillance mechanisms such as DNA damage checkpoints, DNA repair machinery, and mitotic checkpoints results in the disorderly and non-sequential damage of multiple genes controlling cell division which have been associated with immune inflamed tumors and their responsiveness to Immune Oncology (IO) agents [7,8,9,10]. This instability results in the disruption of several cellular functions of which only a few are required for survival while several have broader effects on the surrounding tissues of which some benefit from and some are detrimental for cancer growth [1]. This phenomenon may amplify and exaggerate the tissue remodeling properties described originally by Virchow’s healing wound theory [11] that includes inflammatory processes that may complement wound healing but are not relevant to cancer growth [1, 12]. The latter may be amplified in immunogenic tumors by the presence of Damage-Associated Molecular Patterns (DAMPs) that induce an exaggerated immune response by the host [13, 14]. DAMPs are released by degenerate cancer cell death also referred to as Immunogenic Cell Death (ICD) that is strictly associated with the activation of other immune effector and immune-regulatory pathways [3]. In particular, we observed that the strength of ICD activation is highly proportional to the activation of immune effector responses and of immune-suppressive mechanisms that mold the balance of immune surveillance including the checkpoint cluster, regulatory T cells, the Interleukin (IL) 23-Th17 axis, myeloid suppressor cells, and metabolic suppressors [3].
Here, we will describe immune-active landscapes and the compensatory regulatory mechanisms balancing immune surveillance. We will discuss the roles that adaptive and innate immunity play in the conundrum of self-non-self-recognition including the weight that neo-epitopes may exert as initiators and sculptors of high-affinity memory and effector immune responses against cancer. We will discuss the evolutionary root for the existence of immune checkpoints and how several theories explaining cancer immune resistance to immunotherapy are rather a facet of a similar phenomenon that we assembled in the Theory of Everything [3]. We will discuss how the biology of immunogenicity and counterbalancing immune regulation is widespread across different cancer histologies at the same time indicating idiosyncratic differences among them. Finally, we will discuss how overcoming immune resistance implies different approaches relevant to the immune context of individual cancers.
2 The Immune Contexture of Cancer
It is broadly accepted that morphologically the Tumor Micro-environment (TME) can be portrayed according to three immune phenotypes: immune active, immune desert, or immune excluded [3, 15, 16]. This segregation is independent of individual cancer ontogenesis [2, 3]. The immune-active landscape is characterized by the homogenous infiltration of CD8 T cells intermixed with cancer cells within proliferating tumor nests. The immune-deserted landscape is devoid of immune infiltrates, in particular, T cells. The immune excluded is characterized by T cell lineups confined to the periphery of cancer nests reminiscent of the peri-insulitis that associates with experimental and clinical type 1 diabetes [17].
The morphological distinction is reduced to only two immune phenotypes when gene expression profiling is applied to bulk tumor tissues such as the samples collected by the The Cancer Genome Atlas (TCGA). In this case, no information about spatial distribution of various cell types is available. Thus, only a gradual progression in the level of expression of transcripts associated with immune infiltration and activation can be imputed for classification of immune landscapes based on studies where immune histochemical parameters were compared to transcriptional profiling [18,19,20,21,22]. Therefore, at the transcriptional level, tumors can be classified as immune active at one extreme and immune silent at the other [2, 3, 23,24,25,26]. It has been our unpublished observation that immune-excluded tumors behave most frequently as functionally immune-active tumors suggesting that immune exclusion is not due to immune ignorance but rather to the presence of functional and/or mechanical barrier that prevents progression of cognitive T cells. With the exception of immune exclusion, the functional distinction based on transcriptional signatures fairly represents the morphological classification as it has been shown that the immune infiltrated landscape is defined not only by the density and spatial distribution of Tumor-Infiltrating Lymphocytes (TILs) but also by their functional orientation toward a Th1 effector phenotype [18, 19].
Combined peri-tumoral and intra-tumor infiltration of CD8 T cells functionally oriented toward a Th1 phenotype has been shown to be associated with improved survival of patients with colorectal cancer [19, 27, 28]. This observation has been recently validated by a multi-national, multi-institutional consortium where a parameter called the immunoscore that includes the combined assessment of CD3 and CD8 expressing T cells [20, 22, 29, 30] accurately predicted prognosis in a multivariate analysis [31].
We identified a functional signature associated with the immune-active landscape that is an independent prognostic marker in several data sets of breast cancer [23, 25]. This signature can be applied to sub-classify cancers according to degree of immune activation [3]. Moreover, this transcriptional signature or some of its components is predictive of responsiveness to IO agents [32, 33] and defines the mechanism of immune-mediated tumor rejection [34,35,36,37,38]. We termed this signature the Immunologic Constant of Rejection (ICR) [12]. A survey of two open-access data sets comprising approximately 3,000 cases of breast cancer [23, 24, 39] allowed us to rank cancers according to the transcriptional expression of genes associated with immune-mediated tissue-specific destruction. This consists of a conserved mechanism responsible for destructive flares of autoimmunity, acute allograft rejection, and graft-versus-host disease, clearance of pathogen-infected cells and rejection of cancer [12, 40]. Expression of the ICR signature is unconditionally associated with this process [12].
The ICR bears both predictive and prognostic implications in various cancers within the continuum of anti-cancer immune surveillance [18]. Specifically, the ICR signature includes four functional categories summarized by 20 representative genes: CXCR3/CCR5 chemokines (CXCL9, CXCL10, CCL5), Th1 signaling (IFNG, IL12B, TBX21, CD8A, STAT1, IRF1, CD8B), effector (GNLY, PRF1, GZMA, GZMB, GZMH), and immune-regulatory (CD274, CTLA4, FOXP3, IDO1, PDCD1) functions. The expression of the 20 representative genes is highly correlated with the extended ICR signature that includes approximately five hundred genes [12, 35, 40]. The 20-gene ICR bears strong analogy with other signatures predictive of immune responsiveness to human recombinant IL-2-based therapy [32,33,34] and Checkpoint Inhibitor Therapy (CIT) [41]. For instance, Ayers et al. using RNA from baseline tumor samples of pembrolizumab-treated patients identified and validated a pan-tumor T cell-inflammation gene signature (tumor inflammation signature, TIS) that defines prognostic landscapes of cancer [42] and is predictive of cancer responsiveness to CIT. The TIS contains Interferon (IFN)-γ-responsive genes (CD27, STAT1,Footnote 1 IDO1, HLA-E, NKG2) related to antigen presentation (HLA-DQA1, HLA-DRB1, PSMB10, CMKLR1) chemokine and chemokine receptors (CCL5, CXCL9, CXCR6), cytotoxic activity (CD8A), and adaptive immune resistance (TIGIT, LAG3, CD274, CD276, PDCD1LG2). The expression of the TIS signature tightly correlates with the ICR signature and it has been developed into a clinical-grade assay currently being evaluated in ongoing pembrolizumab trials [43].
The expression of ICR genes is consistently accompanied by that of the ICD signature and of genes associated with immune-regulatory functions believed to determine immune resistance [3, 23]. These include regulatory T cells, IL-23-Th17 axis activation, myeloid suppressor cells, the PI3K-γ pathway, the checkpoint cluster, and the IDO/NOS signature [3].
3 The Paradox of Immune Exclusion
The phenomenon of immune exclusion represents a biological paradox. The presence of T cell at the periphery of tumor nests and their functional orientation toward a Th1 phenotype suggests that some immunogenic stimulus attracts them there. What stops them from infiltrating the tumor nests? Cancers are clonal, especially within a single tumor nest. Yet, repetitive patterns of heterogeneity suggest that from the “germinal center” of each tumor nest toward its periphery a transformation occurs in the biology of cancer cells and/or their by-products that affect migration and function of immune cells. The transition from an immune depleted germinal center toward an immunologically active periphery is discordant with the relatively homogeneous biology implied by a progeny of cells deriving from a single clone [44]. What determines this rapid change within a few cell divisions? Various hypotheses have been raised to explain immune exclusion but no conclusive understanding of the weight that each one plays in the context of human cancers has been reached [3, 45].
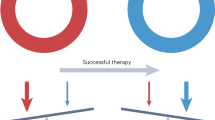
While immune ignorance may most likely shape the development of silent cancers, it is possible that immune exclusion is determined by functional or mechanical barriers that prevent a cognitive immune response. Thus, the biology of immune exclusion may be quite different from that of silent cancers. In fact, the reasons for immune exclusions may depend on two opposite vectors (Fig. 5.1). A centrifugal gradient of escalating immunogenicity responsible for attracting T cells at the periphery may depend upon a metamorphosis of tumor cell biology transition from cancer-initiating cells in the center to their progeny at the periphery [46] following a process reminiscent of epithelial–mesenchymal transition. This in turn has been shown to bear profound effects on the immune orientation of the TME [47]. Moreover, evolving patterns of cell death from the germinal center to the periphery may progressively increase immunogenicity. Perhaps, cancer cells may differentiate from a stem-cell-like core to a degenerate progeny at the periphery prone to ICD [46]. This could be tested by surface expression of calreticulin or other ICD markers or the analysis of expression of DAMPs [48]. Alternatively, cancer cell death occurs only at the periphery while the “germinal” centers continue to proliferate and this could be tested using proliferation markers such as Ki67, proliferating cell nuclear antigen, and mini-chromosome maintenance proteins [49]. This may be associated with decreasing chemoattraction from the periphery to the center with higher expression of T cell attracting cytokines such as the CXCR3 and CCR5-ligand chemokines tightly correlated with the immune-active TME and part of the ICR signature [12, 23, 40].
The paradox of immune exclusion—Example of distinct theories that may explain the confined presence of T cells at the periphery of immune-excluded cancer nests. Blue and red arrows describe, respectively, centrifugal or centripetal vectors of increasing or decreasing immune attraction of exclusion from the center of a cancer nest to the periphery. In addition, a functional and dynamic barrier is described of immune interactions at the first encounter of cancer and immune cells at the periphery of tumor nests
Alternatively, cancer cell immunogenic potential may remain constant while increasing intensity of immune exclusion mechanisms may follow a centripetal gradient (Fig. 5.1). Several mechanisms could be postulated such as an insufficient or abnormal expression of factors determining vascularization like Vascular Endothelial Growth Factor (VEGF)-α [50, 51] gradually increasing expression of Transforming Growth Factor (TGF)-β, production of immune-suppressive metabolites such as Nitric Oxide Synthase (NOS) proteins and their bio-products [52, 53], or Indoleamine 2,3-Dioxygenase (IDO) [54, 55], hypoxia gradients [56, 57], altered glycolytic, and electrolyte rates [58]. In addition, mechanical rather than functional barriers may be developed at the interface between tumor margins and the host stroma while proliferation continues undisturbed at the center of the tumor nests [51, 59,60,61,62].
Finally, it is possible that a dynamic functional barrier is established upon the first encounter of T cells with cancer cells at the periphery of tumors cells which triggers a cross talk among different immune cells and cancer cells leading to the production of inflammatory cytokines such as IL-12 and IFN-γ and consequently the activation of checkpoint suppressor functions [63, 64] that limits the migration and proliferation of T cells beyond the periphery of the tumor nests.
Clinical evidence show that the ability of cancers to induce adaptive memory responses may be required but not sufficient for the localization of T lymphocytes at the tumor site and that the TME is the ultimate modulator of Adoptive Cell Therapy (ACT) products homing by determining chemoattraction, function, persistence, and relevance of transferred T cells was provided by in vivo trafficking studies [65]. Specifically, 111Indium-labeled tumor antigen-specific TILs were studied for their ability to localize in melanoma metastases upon adoptive transfer. TILs migrate into the target lesions only in a fraction of patients, while in several other cases they were trapped in the lungs and in lymphoid organs. Responses were observed exclusively in patients in which TILs localized at the tumor site suggesting that in a large proportion of patients the lack of response to ACT is due to lack of infiltration of T cells in tumors from patients who had cancer-specific adaptive memory responses. Interestingly, among those cases in which the TILs were localized at the tumor site only a fraction of patients responded to therapy demonstrating observable regression of tumor masses [66]. Another example of dissociation between the development of memory responses against cancer and their deployment within the TME is the successful induction of circulating tumor antigen-specific immune responses by vaccines that, however, do not correspond to cancer regression mostly because of lack of localization of the vaccine-induced T cells at the tumor site [67, 68].
In summary, we suggest that the immune-excluded TME landscape is closer in its “phylogeny” to the immune-active cancers rather than the silent ones and it may be an optimal model system to understand mechanisms of immune modulation in humans by understanding the biology of functional and/or mechanical barriers affecting immune infiltration. These tumors with a heterogeneous architecture may represent ideal targets to address the relevance of the TME in linking tumor immunogenicity with immune infiltration and identifying distinct biological mechanisms and combination of them that may determine this phenotype and could be modulated pharmacologically. This will require the use of spatially resolved multiplexed, multi-analyte molecular assessment of the tumor heterogeneity using newly emerging technology [69] platforms such as the GeoMx DSP which allows for digital quantification of the distribution of RNAs or proteins from formalin-fixed, paraffin-embedded tissue sections in a spatially resolved and highly multiplexed manner. Consequently, digital spatial profiling has the potential to reveal actionable changes in biology that otherwise may be obscured by non-spatially resolved tissue profiling. This sensitive, robust, and rapid platform enables the users to efficiently assess immune cell response, T cell activation/proliferation, tumor reactivity, and heterogeneity in multiple types of cancer cells [70, 71].
4 What’s the Buzz with Neo-epitopes?
The explanations for the development of an active immune landscape revolve around two non-mutually exclusive interpretations of the mechanisms leading to immune surveillance. One suggests that the primary stimulus that induces anti-cancer immune responses is the recognition of non-self-proteins by the adaptive immune responses. Thus, self-non-self-discrimination promotes priming of adaptive immune responses as the primary mechanism leading to cancer rejection through recognition against non-self-antigens (neo-antigens) and respective neo-epitopes generated by the translation of missense mutations into novel protein domains. This hypothesis is based on several experimental [72, 73] and clinical observations. For instance, cancers with high mutational burden are more frequently observed within the immune-active landscape and a large number of studies reported various degrees of association with prognosis and/or immune responsiveness to IO agents [72,73,74,75,76,77,78,79,80,81,82,83]. The weight that adaptive immune responses play in the development of cancer landscapes bears important implications for the development of IO agents. For instance, the development of T cell responses against neo-epitope suggests that these may depend on the activation of T cells bearing high-affinity TCRs that have not gone through negative thymus selection and, therefore, could be more effective anti-tumor agents [73, 84, 85]. Such postulate, however, has not been conclusively demonstrated in humans where high-affinity TCRs against non-mutated antigens can also be detected [86, 87]. Another important concept is that the development of adaptive immune responses against mutated antigens gives confidence that a therapeutic agent will not have an on-target, off-tumor effect that could cause major adverse events [88, 89].
Recently, Thomas et al. investigated the genomic, transcriptomic, and clinical database from TCGA and from the Molecular Taxonomy of Breast Cancer International Consortium cohorts to examine associations between Tumor Mutational Burden (TMB) and survival in patients bearing tumors characterized by distinct signatures associated with [24] prognostic connotation reflecting favorable, weak, or poor immune-infiltrate dispositions. A high density of tumor-infiltrating lymphocytes, as estimated by the expression of monocytes, DC, and T, NK, and B cells transcripts, was associated with improved survival in patients with high TMB but not in those with low TMB independently of tumor stage, molecular subtype, age, and treatment. Interestingly, in the same study, preferential amplification of chromosome 1q immune-regulatory genes in tumor with poor immune infiltrate disposition was noted [75]. Similarly, in the same dataset, the prognostic role of immune-related signatures was restricted to tumor with high proliferative capacity [24, 25]. This data suggests that, in tumor with high mutational burden and/or high proliferative capacity, the immune favorable disposition is elicited by specific features of cancer cells, while in the ones with low proliferative capacity/low mutational burden such transcriptional signature is the result of a bystander immune infiltration with no protective effect. Therefore, it is possible to speculate that a proportion of phenotypically “immune-active” tumors are functionally “immune silent.” A combined analysis of tumor cell intrinsic features and immune disposition might facilitate the identification of true “immune-active” tumor and enhance the predictive values of transcriptomic signatures in the setting of cancer immunotherapy (Roelands et al., in preparation).
Although the value of self-non-self-discrimination by adaptive immune response bears an unquestionable therapeutic impact, the initiating role of adaptive self-non-self-recognition in determining cancer immune landscapes has been questioned by recent observations by us [23, 42] and by others [90].
Basic understanding of immunologic processes confutes the primary role that adaptive immunity plays in the rejection of cancer emphasizing the need for a first signal [13, 14, 91,92,93,94,95,96]. The conditionality of adaptive immune responses is also suggested by experimental evidence that they are not an essential requirement for the rejection of cancer as exemplified by the transferrable anti-cancer innate immunity model [97,98,99,100] and by oncotropic virus-mediated rejection of xenografts in immune-deficient mice [36, 37]. From the clinical standpoint, the secondary role played by adaptive immunity could also explain the paradoxical observation that vaccines aimed at priming adaptive immune responses can consistently elicit systemic immunity, which, however, does not correlate with tumor rejection [67, 68]. Thus, immune ignorance is not a sole explanation for cancer immune resistance to surveillance mechanisms. It should also be pointed out that seminal studies done on the effectiveness of tumor-infiltrating lymphocytes demonstrated that their homing at the tumor site is necessary, though not sufficient, to induce tumor regression pointing to the critical role that chemoattraction may play in immune responsiveness [66, 96].
5 The Chicken and Egg Conundrum of Self-non-Self-recognition
As for all immunologic phenomena, the TME landscapes are likely shaped by a cascade of innate mechanisms as a first signal secondarily followed by adaptive immune responses as originally suggested by Charles A. Janeway Jr. and Polly Matzinger’s danger model [13, 14, 91,92,93,94,95,96, 101,102,103]. These models align with the central role subsequently proposed by Kroemer et al. [104] for ICD in initiating anti-cancer immune responses [96, 104,105,106]. This concept is not in antithesis with the role played by self-non-self-recognition by the adaptive immune system but rather proposes a sequential path that requires initiation through innate mechanisms and may not necessarily require self-non-self-discrimination as demonstrated by the natural immune responses against non-mutated melanoma-associated melanocyte lineage-specific antigens [107, 108]. Self-non-self-discrimination by innate immunity mechanisms postulates that activation of innate and adaptive immunity is dependent upon recognition of foreign material such as pathogen-associated molecular patterns sensed by pattern recognition receptors excluding endogenous sources as mediators of immune activation. In contrast, the danger model proposes that non-physiological, sterile, cell death can activate the innate immune system by releasing DAMPs [14, 103]. Garg et al. [96] described a pathogen response-like recruitment and activation of neutrophils by sterile immunogenic dying cells that drove primarily neutrophil-mediated killing of cancer cells awakening at the same time secondary mechanisms of innate and adaptive immune recognition.
As previously discussed, the ICD gene signature was positively associated with the inflamed immune landscapes suggesting that it is a requirement for the development of the immune-active phenotypes [3]. Conversely, absence of ICD in immune-silent tumors leads to immune ignorance. Some chemotherapies such as anthracyclines and oxaliplatin can induce ICD [109], which is characterized by overexpression by dying cells of calreticulin, the ER-associated protein disulfide isomerase 57 [110] that promotes the uptake of dying cells by dendritic cells and/or the release of High Mobility Group Box 1 (HMGB1) and ATP [48], which in turn activate downstream pathways of innate sensing of danger [104]. Recently, it has been shown that other anti-cancer modalities of treatments can induce ICD in a tumor cell-specific manner. For instance, an epidermal growth factor receptor-neutralizing antibody affects cancer growth primarily by blocking receptor–ligand interactions. However, its anti-cancer activity is amplified when the antibody is administered in combination with chemotherapeutic agents [111].
Previous studies on ICD focus on the activation of antigen presentation by dendritic cell to prime and sustain adaptive immune responses. However, ICD activates also innate effector mechanisms through the recruitment of neutrophils, macrophages, and NK cells. These innate mechanisms recapitulate the immune response to pathogens and are evolutionarily conserved across vertebrates [96, 112]. Furthermore, both tumor cells undergoing ICD and innate mechanisms activated by ICD induce the secretion of chemokines such as CXCL1, CCL2, CXCL9, and CXCL10 that secondarily recruit adaptive immune cells [96, 104]. Thus, we propose that the TME is primarily molded by the way cancer cells die which defines the primary signal for immune surveillance.
6 Cancer as an Evolutionary Process and the Theory of Everything
In the theory of everything, we suggested that cancer immune phenotypes are shaped by a binary evolutionary choice [3]. In the majority of cases, genes associated with immune-regulatory functions correlate in expression with the ICR genes. Thus, we propose that the natural history of cancer is shaped at the crossroad of two biological evolutionary paths as a “Two-Option Choice” (TOC) or Hobson’s predicament (Fig. 5.2): (1) immunogenic tumors can only survive in the host when immune-suppressive mechanisms balance the reaction of the host and (2) silent tumors can grow undisturbed [2].
The two-option choice or Hobson’s predicament in cancer survival—Reproduced from Turan et al. [3]
The segregation between the biology of immune-active compared to immune-silent tumors is corroborated by the observation that immune-silent tumors are characterized by a distinctive mutational profile characterized by low prevalence of mutations within oncogenes, which suggests in turn a more orderly growth process [23]. It is, therefore, reasonable to suppose that clean tumor growth is dependent on an orderly and sequential accumulation of relevant genetic alterations likely to be centered in a good proportion of cases on the PI3Kγ/SFK/pGSK3/β-catenin axis [113,114,115]. This process is evolutionarily fit to achieve growth efficiency without unnecessarily interfering with other physiological processes that regulate the host’s homeostasis.
7 The Duck Soup of Immune Checkpoints
As previously described, analyses of large data sets such as the TCGA repository suggested that immune checkpoints are predominantly co-expressed and the intensity of expression is directly proportional to the status of activation of ICD and immune effector mechanisms [2, 3]. This seems to be the case for most immune-regulatory mechanism and, in particular, for the checkpoint cluster that includes several inhibitory ligand–surface receptor interactions mostly controlled by immune activator signals such as IFN-dependent signaling [116, 117]. This can be partly explained by a modular expression of several checkpoints in the condition of T cell exhaustion as part of a broad co-inhibitory gene program driven by immune-regulatory cytokines such as IL-27 [118] or more generally the level of differentiation of T cells in the continuum from stem-cell-like to terminally differentiated [119,120,121,122].
However, some exceptions were noted particularly in non-immunogenic cancers. Thus, variety in the distribution of expression of immune checkpoints may be intrinsic to their biology independently of the level of immunogenicity of individual cancers such as, for example, melanomas. In this case, most co-inhibitory and co-stimulatory immune checkpoints are expressed in association with the immune-active landscape. Even in this case exceptions are noticed and some co-inhibitory checkpoints (CD276, VTCN1, TNFRSF14, CECAM1, SIRPA, PVR) seem to display an opposite distribution of expression divergent by the common checkpoint cluster pattern (Fig. 5.3).
Independent of their pattern of distribution, the checkpoints in cutaneous melanoma correlate in expression among themselves (Fig. 5.4). This is not the case in non-immunogenic tumors such as glioblastoma. In this case, immune checkpoints follow a less coordinated pattern of distribution suggesting that their activation is individual cancer specific. As a possible explanation, the immunogenic tumors may provide higher level of stimulation for the activation of immune checkpoint that overcomes different thresholds of activation therefore harmonizing their expression level. In the case of less immunogenic tumors, the lower intensity of stimulatory mechanisms may affect differently the expression of checkpoints requiring distinct threshold of activation and it may be more dependent on context-specific cofactors. However, to our knowledge, this, however, has not been tested in human cancers.
Indeed, the discordant distribution of expression of immune checkpoints among less immunogenic tumors has been described for different histologies including prostate and pancreatic cancer where the expression of V-domain Immunoglobulin Suppressor of T cell Activation (VISTA) seem to play a preeminent role compared with other tumors such as melanoma [123, 124].
Thus, for instance, the effectiveness of CIT is dependent upon two biological vectors: the first is the level of immunogenicity of a given tumor. More immunogenic tumors may stimulate a larger number of immune checkpoints that generate a complex network of stimulatory and inhibitory interactions. In this case, lack of responsiveness to a specific therapy such as anti-PD1 monotherapy may be dictated by the limited weight that one checkpoint mechanism plays in such a complex TME. The second vector is the specificity of expression of distinct checkpoints in the case particularly of less immunogenic tumors. Here lack of responsiveness may be due to the absence of expression of the targeted mechanism. For instance, as shown in Fig. 5.4, the expression of PD1 and another commonly targeted checkpoint such as T cell Immunoglobulin and Mucin-domain containing-3/Hepatitis A Virus Cellular Receptor 2 (TIM-3/HAVCR2) correlates strictly in melanoma but minimally in glioblastoma suggesting that in the former case a combination therapy against both inhibitory checkpoints may enhance chances of success by increasing the level of necessary suppression to allow immune rejection of cancer. In the latter, a more accurate, patient-specific approach may produce better results sparing the costs and added toxicities of combination therapy.
8 Mechanisms of Immune Resistance to Immunotherapy
While the usefulness of CIT may be limited to a low percentage of cancers endowed with a preexisting immune-active TME, resistance to CIT will affect the large majority of immune-deserted cancers where the most commonly targeted checkpoints are absent. This distinction among the mechanisms leading to resistance to CIT and more generally IO agents is critical since the modulation of ancillary cellular functions around antigen-specific signaling is completely different in different situations. For instance, modulating T cells suppression by hypoxia [56, 57] or TAM receptor tyrosine kinases [125, 126] or the ubiquitous serine/threonine phosphatase protein phosphatase-2A [127] may provide broader effectiveness by overcoming resistance in silent tumors as these mechanisms are more likely to be relevant to different immune landscapes [3].
A commonly adapted semantic to categorize lack of responsiveness to CIT (and more broadly to IO agents) is “primary” versus “secondary” immune resistance. The former appellation refers to lack of effectiveness of a given therapy at the first exposure and the latter refers to an acquired resistance stemming from immune escape mechanisms that determine relapse after an original response. Both of them are practical, clinically driven definitions that, particularly in the case of primary immune resistance, could be best sub-categorized according to a more accurate biological interpretation [3]. We suggest that several reasons may underline “primary” immune resistance that may be better defined as (a) authentic immune resistance when the mechanism of action of a given therapeutic does not match the biology of the targeted entity (such as CIT in immune-deserted tumors) since there is no reason for the regression of cancer to occur (128); (b) compensatory immune resistance, when the mechanism of action of a given therapeutic is relevant to the biology of the targeted TME and yet tumors do not regress. This could be the case for CIT or metabolic inhibitor therapy failures in an immune-active cancer landscape. Compensatory immune resistance is, therefore, most likely to occur in an immune-active cancer with a dense network of immune checkpoint mechanisms working in synchrony. In this case, targeting one at the time with monotherapy may not suffice to induce cancer regression; (c) pseudo-immune resistance, when factors extrinsic to the biology of individual tumors interfere with a potentially positive outcome. A good example is variations in the potency of the therapeutic agents tested. In the case of CIT that utilizes known entities such as well-characterized monoclonal antibodies, this maybe irrelevant, but in the case of ACT, variability in product potency may represent a critical extrinsic factor determining clinical outcome extraneous to the biology of the targeted cancer [129]. Other examples of pseudo-immune resistance are limiting toxicities and associated immune-suppressive mechanisms that may not allow a fair assessment of treatment efficacy [76]. In addition, environmental factors such as those affecting the microbiome [130, 131], or behavioral factors that affect the nutritional status [132], or comorbidities, associated therapies, and other hidden anamnestic and epidemiological factors [133]. All of these factors and the genetic background of the host may affect the molding of the TME which in turn may modulate the responsiveness of patients to IO agents [134]. Knowledge of the influence of these factors over the biology of the TME in the human settings, however, is still in its infancy.
However, with few exceptions, the majority of data regarding tumor–host interaction in humans has been derived from the analysis of a single tumor lesion, either a primary tumor or a metastatic site. A recent analysis of 31 colorectal cancer metastases from two patients over a 11-year-long follow-up has demonstrated that the tumor evolution is shaped by immunologic pressure [135]. Tumor lesions with genetic evidence of immunoediting (i.e., a lower than expected rate of neo-antigens) display a Th1/cytotoxic functional polarization. However, tumor clones that are immune-edited tend to be eliminated while progressive clones are immune privileged [135]. Studying the dynamic of tumor evolution in the context of parallel clonal expansion under immune selection might help to refine prognostic and predictive algorithms that take into account the “immunogenicity” of different tumor clones within and across different lesions. By applying reductionist approaches, the comparative analyses of multiple tumor sites corrected for clonal prevalence may lead to the identification of an aggregate “onco-immunogenomic score” that displays the lowest heterogeneity across metastases of the same individual. Since the sampling of every metastasis is unfeasible in the clinical setting, the definition of a relatively stable (i.e., relatively homogenous) multi-omics analyte might be used to better infer, from a single lesion, the chance to respond to therapy and/or to relapse.
9 How to Overcome Cancer-Specific Immune Resistance?
The big hurdle in IO development is addressing specifically the biology of individual cancers to prevent different types of immune resistance that are pre-determined and avoidable if appropriate selections are applied [136]. A good example is the recent failure of the large phase 3 trial evaluating the efficacy of the combined inhibition of IDO and PD1 in advanced melanoma [137]. It could be argued that a better study design, taking into account a refined understanding of the biology of the individual cancer targeted, may have produced better results and “better rationalized compounds and better rationalized trial designs will be important in the future to accurately gauge medical impact” [137].
In particular, disruption of the lean cancer cell biology of silent cancers should be considered to switch immune latency into an immunogenic outburst and open a window of opportunity for IO agents [16]. A good example is the effect on the TME of MAPK-induced alterations of cancer cell intrinsic and extrinsic pathways during pathway inhibitor therapy [138].
The focus of this chapter is about immune-active cancers and strategies to overcome compensatory immune resistance that focused on the expression of transcriptional signatures associated with immune-regulatory properties [139] such as other immune checkpoints [116], regulatory T cells [140], IL-23/IL17 axis [141], myeloid suppressor cells [55], IDO [54, 136, 142], ICD [105], TAM tyrosine kinase receptors [143], hypoxia [144], cancer-associated fibroblasts [145], and barrier molecules [59]. Yet, it is likely that several of these mechanisms may overlap in different TME and a segregation among cancers should not be seen as a static division but should rather be taken as a starting point to design better clinical trials with the intent of altering individual cancer biology by combining agents that may affect ICD and activation of innate mechanisms in silent tumors turning them into suitable targets for IO agents. Defining better tools to understand the pharmacodynamics of individual treatments in the context of an evolving and unstable cancer TME prone to pharmacological and immunological manipulation should be a primary goal of future clinical trials.
Notes
- 1.
In bold are transcripts in common between the ICR and the TIS signature.
Abbreviations
- ACT :
-
Adoptive Cell Therapy
- CIR :
-
Cancer Immune Responsiveness
- CIT :
-
Checkpoint Inhibitor Therapy
- DAMPs :
-
Damage-Associated Molecular Patterns
- DSP :
-
Digital Spatial Profiling
- HMGB1 :
-
High Mobility Group Box 1
- ICD :
-
Immunogenic Cell Death
- ICR :
-
Immunologic Constant of Rejection
- IDO :
-
Indoleamine 2,3-Dioxygenase
- IFN :
-
Interferon
- IL :
-
Interleukin
- IO :
-
Immune Oncology
- NOS :
-
Nitric Oxide Synthase
- PDL1 :
-
Programmed Cell Death Protein Ligand 1
- TCGA :
-
The Cancer Genome Atlas
- TGF :
-
Transforming Growth Factor
- TILs :
-
Tumor-Infiltrating Lymphocytes
- TIM-3/HAVCR2 :
-
T cell Immunoglobulin and Mucin-domain containing-3/Hepatitis A Virus Cellular Receptor 2
- TIS :
-
Tumor Inflammation Signature
- TMB :
-
Tumor Mutational Burden
- TME :
-
Tumor Micro-environment
- TOC :
-
Two Option Choice
- TOE :
-
Theory Of Everything
- VEGF :
-
Vascular Endothelial Growth Factor
- VISTA :
-
V-domain Immunoglobulin Suppressor of T cell Activation
References
Mantovani A, Romero P, Palucka AK, Marincola FM (2008) Tumour immunity: effector response to tumour and role of the microenvironment. Lancet 371(9614):771–783
Lu R, Turan T, Samayoa J, Marincola FM (2017) Cancer immune resistance: can theories converge? Emerg Top Life Sci 1(5):411–419
Turan T, Kannan D, Patel M, Matthew Barnes J, Tanlimco SG, Lu R et al (2018) Immune oncology, immune responsiveness and the theory of everything. J Immunother Cancer 6(1):50
Tomasetti C, Vogelstein B, Parmigiani G (2013) Half or more of the somatic mutations in cancers of self-renewing tissues originate prior to tumor initiation. Proc Natl Acad Sci USA 110(6):1999–2004
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW (2013) Cancer genome landscapes. Science 339(6127):1546–1558
Tysnes BB, Bjerkvig R (2007) Cancer initiation and progression: involvement of stem cells and the microenvironment. Biochim Biophys Acta 1775(2):283–297
Yao Y, Dai W (2014) Genomic Instability and Cancer. J Carcinog Mutagen 5
Le DT, Durham JN, Smith KN, Wang H, Bartlett BR, Aulakh LK et al (2017) Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357(6349):409–413
Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD et al (2015) PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 372(26):2509–2520
Cahill DP, Kinzler KW, Vogelstein B, Lengauer C (1999) Genetic instability and darwinian selection in tumours. Trends Cell Biol 9(12):M57–M60
Balkwill F, Mantovani A (2001) Inflammation and cancer: back to Virchow? Lancet 357(9255):539–545
Wang E, Worschech A, Marincola FM (2008) The immunologic constant of rejection. Trends Immunol 29(6):256–262
Fuchs EJ, Matzinger P (1996) Is cancer dangerous to the immune system? Semin Immunol 8(5):271–280
Matzinger P (2002) An innate sense of danger. Ann N Y Acad Sci 961:341–342
Chen DS, Mellman I (2017) Elements of cancer immunity and the cancer-immune set point. Nature 541(7637):321–330
Gajewski TF (2015) The next hurdle in cancer immunotherapy: overcoming the non-t-cell-inflamed tumor microenvironment. Semin Oncol 42(4):663–671
Morgan NG, Richardson SJ (2018) Fifty years of pancreatic islet pathology in human type 1 diabetes: insights gained and progress made. Diabetologia
Galon J, Angell HK, Bedognetti D, Marincola FM (2013) The continuum of cancer immunosurveillance: prognostic, predictive, and mechanistic signatures. Immunity 39(1):11–26
Galon J, Costes A, Sanchez-Cabo F, Kirilovsky A, Mlecnik B, Lagorce-Pages C et al (2006) Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313(5795):1960–1964
Galon J, Fox BA, Bifulco CB, Masucci G, Rau T, Botti G et al (2016) Immunoscore and Immunoprofiling in cancer: an update from the melanoma and immunotherapy bridge 2015. J Transl Med 14:273
Galon J, Fridman WH, Pages F (2007) The adaptive immunologic microenvironment in colorectal cancer: a novel perspective. Cancer Res 67(5):1883–1886
Galon J, Pages F, Marincola FM, Angell HK, Thurin M, Lugli A et al (2012) Cancer classification using the immunoscore: a worldwide task force. J Transl Med 10:205
Hendrickx W, Simeone I, Anjum S, Mokrab Y, Bertucci F, Finetti P et al (2017) Identification of genetic determinants of breast cancer immune phenotypes by integrative genome-scale analysis. Oncoimmunology 6(2):e1253654
Miller LD, Chou JA, Black MA, Print C, Chifman J, Alistar A et al (2016) Immunogenic subtypes of breast cancer delineated by gene classifiers of immune responsiveness. Cancer Immunol Res 4(7):600–610
Bertucci F, Finetti P, Simeone I, Hendrickx W, Wang E, Marincola FM et al (2018) The immunologic constant of rejection classification refines the prognostic value of conventional prognostic signatures in breast cancer. Br J Cancer 119(11):1383–1391
Thorsson V, Gibbs DL, Brown SD, Wolf D, Bortone DS, Ou Yang TH et al (2018) The immune landscape of cancer. Immunity 48(4):812–830.e14
Pages F, Galon J, Dieu-Nosjean MC, Tartour E, Sautes-Fridman C, Fridman WH (2010) Immune infiltration in human tumors: a prognostic factor that should not be ignored. Oncogene 29(8):1093–1102
Pages F, Kirilovsky A, Mlecnik B, Asslaber M, Tosolini M, Bindea G et al (2009) In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J Clin Oncol 27(35):5944–5951
Galon J, Lugli A, Bifulco C, Pages F, Masucci G, Marincola FM et al (2017) World-wide immunoscore task force: meeting report from the “Melanoma Bridge”, Napoli, November 30th–December 3rd, 2016. J Transl Med. 15(1):212
Galon J, Pages F, Marincola FM, Thurin M, Trinchieri G, Fox BA et al (2012) The immune score as a new possible approach for the classification of cancer. J Transl Med 10:1
Pages F, Mlecnik B, Marliot F, Bindea G, Ou FS, Bifulco C et al (2018) International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet 391(10135):2128–2139
Weiss GR, Grosh WW, Chianese-Bullock KA, Zhao Y, Liu H, Slingluff CL Jr et al (2011) Molecular insights on the peripheral and intratumoral effects of systemic high-dose rIL-2 (aldesleukin) administration for the treatment of metastatic melanoma. Clin Cancer Res 17(23):7440–7450
Bedognetti D, Spivey TL, Zhao Y, Uccellini L, Tomei S, Dudley ME et al (2013) CXCR3/CCR5 pathways in metastatic melanoma patients treated with adoptive therapy and interleukin-2. Br J Cancer 109(9):2412–2423
Wang E, Miller LD, Ohnmacht GA, Mocellin S, Perez-Diez A, Petersen D et al (2002) Prospective molecular profiling of melanoma metastases suggests classifiers of immune responsiveness. Cancer Res 62(13):3581–3586
Panelli MC, Stashower ME, Slade HB, Smith K, Norwood C, Abati A et al (2007) Sequential gene profiling of basal cell carcinomas treated with imiquimod in a placebo-controlled study defines the requirements for tissue rejection. Genome Biol 8(1):R8
Worschech A, Chen N, Yu YA, Zhang Q, Pos Z, Weibel S et al (2009) Systemic treatment of xenografts with vaccinia virus GLV-1h68 reveals the immunologic facet of oncolytic therapy. BMC Genom 10:301
Worschech A, Haddad D, Stroncek DF, Wang E, Marincola FM, Szalay AA (2009) The immunologic aspects of poxvirus oncolytic therapy. Cancer Immunol Immunother 58(9):1355–1362
Worschech A, Kmieciak M, Knutson KL, Bear HD, Szalay AA, Wang E et al (2008) Signatures associated with rejection or recurrence in HER-2/neu-positive mammary tumors. Cancer Res 68(7):2436–2446
Turan T, Kannan D, Patel M, Barnes MJ, Tanlimco SG, Lu R et al (2017) Immune oncology, immune responsiveness and the theory of everything. J Immunother Cancer (in press)
Spivey TL, Uccellini L, Ascierto ML, Zoppoli G, De Giorgi V, Delogu LG et al (2011) Gene expression profiling in acute allograft rejection: challenging the immunologic constant of rejection hypothesis. J Transl Med 9:174
Bedognetti D, Hendrickx W, Marincola FM, Miller LD (2015) Prognostic and predictive immune gene signatures in breast cancer. Curr Opin Oncol 27(6):433–444
Danaher P, Warren S, Lu R, Samayoa J, Sullivan A, Pekker I et al (2018) Pan-cancer adaptive immune resistance as defined by the Tumor Inflammation Signature (TIS): results from The Cancer Genome Atlas (TCGA). J Immunother Cancer 6(1):63
Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR et al (2017) IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest
Messerschmidt JL, Bhattacharya P, Messerschmidt GL (2017) Cancer clonal theory, immune escape, and their evolving roles in cancer multi-agent therapeutics. Curr Oncol Rep 19(10):66
Peranzoni E, Lemoine J, Vimeux L, Feuillet V, Barrin S, Kantari-Mimoun C et al (2018) Macrophages impede CD8 T cells from reaching tumor cells and limit the efficacy of anti-PD-1 treatment. Proc Natl Acad Sci USA. 115(17):E4041–E4050
Maccalli C, Parmiani G, Ferrone S (2017) Immunomodulating and immunoresistance properties of cancer-initiating cells: implications for the clinical success of immunotherapy. Immunol Invest 46(3):221–238
Chockley PJ, Keshamouni VG (2016) Immunological consequences of epithelial-mesenchymal transition in tumor progression. J Immunol 197(3):691–698
Kepp O, Senovilla L, Vitale I, Vacchelli E, Adjemian S, Agostinis P et al (2014) Consensus guidelines for the detection of immunogenic cell death. Oncoimmunology 3(9):e955691
Jurikova M, Danihel L, Polak S, Varga I (2016) Ki67, PCNA, and MCM proteins: markers of proliferation in the diagnosis of breast cancer. Acta Histochem 118(5):544–552
Costache MI, Ioana M, Iordache S, Ene D, Costache CA, Saftoiu A (2015) VEGF expression in pancreatic cancer and other malignancies: a review of the literature. Rom J Intern Med 53(3):199–208
Rahbari NN, Kedrin D, Incio J, Liu H, Ho WW, Nia HT et al (2016) Anti-VEGF therapy induces ECM remodeling and mechanical barriers to therapy in colorectal cancer liver metastases. Sci Transl Med 8(360):360ra135
Liu Q, Tomei S, Ascierto ML, De Giorgi V, Bedognetti D, Dai C et al (2014) Melanoma NOS1 expression promotes dysfunctional IFN signaling. J Clin Invest 124(5):2147–2159
Thomas DD, Wink DA (2017) NOS2 as an emergent player in progression of cancer. Antioxid Redox Signal 26(17):963–965
Mondanelli G, Ugel S, Grohmann U, Bronte V (2017) The immune regulation in cancer by the amino acid metabolizing enzymes ARG and IDO. Curr Opin Pharmacol 35:30–39
Munn DH, Bronte V (2016) Immune suppressive mechanisms in the tumor microenvironment. Curr Opin Immunol 39:1–6
Hatfield SM, Kjaergaard J, Lukashev D, Belikoff B, Schreiber TH, Sethumadhavan S et al (2014) Systemic oxygenation weakens the hypoxia and hypoxia inducible factor 1alpha-dependent and extracellular adenosine-mediated tumor protection. J Mol Med (Berl). 92(12):1283–1292
Hatfield SM, Kjaergaard J, Lukashev D, Schreiber TH, Belikoff B, Abbott R et al (2015) Immunological mechanisms of the antitumor effects of supplemental oxygenation. Sci Transl Med 7(277):277ra30
Cascone T, McKenzie JA, Mbofung RM, Punt S, Wang Z, Xu C et al (2018) Increased tumor glycolysis characterizes immune resistance to adoptive T cell therapy. Cell Metab 27(5):977–987.e4
Salerno EP, Bedognetti D, Mauldin IS, Deacon DH, Shea SM, Pinczewski J et al (2016) Human melanomas and ovarian cancers overexpressing mechanical barrier molecule genes lack immune signatures and have increased patient mortality risk. Oncoimmunology 5(12):e1240857
Kandalaft LE, Facciabene A, Buckanovich RJ, Coukos G (2009) Endothelin B receptor, a new target in cancer immune therapy. Clin Cancer Res 15(14):4521–4528
Johnson MO, Wolf MM, Madden MZ, Andrejeva G, Sugiura A, Contreras DC et al (2018) Distinct regulation of Th17 and Th1 cell differentiation by glutaminase-dependent metabolism. Cell 175(7):1780–1795.e19
Eil R, Vodnala SK, Clever D, Klebanoff CA, Sukumar M, Pan JH et al (2016) Ionic immune suppression within the tumour microenvironment limits T cell effector function. Nature 537(7621):539–543
Garris CS, Arlauckas SP, Kohler RH, Trefny MP, Garren S, Piot C et al (2018) Successful anti-PD-1 cancer immunotherapy requires T cell-dendritic cell crosstalk involving the cytokines IFN-gamma and IL-12. Immunity 49(6):1148–1161.e7
Mattox AK, Lee J, Westra WH, Pierce RH, Ghossein R, Faquin WC et al (2017) PD-1 expression in head and neck squamous cell carcinomas derives primarily from functionally anergic CD4(+) TILs in the presence of PD-L1(+) TAMs. Cancer Res 77(22):6365–6374
Turan T, Kannan D, Patel M, Barnes JM, Tanlimco SG, Lu R et al (2018) Immune oncology, immune responsiveness and the theory of everything. J Immunother Cancer (in press)
Pockaj BA, Sherry RM, Wei JP, Yannelli JR, Carter CS, Leitman SF et al (1994) Localization of 111indium-labeled tumor infiltrating lymphocytes to tumor in patients receiving adoptive immunotherapy. Augmentation with cyclophosphamide and correlation with response. Cancer 73(6):1731–1737
Lee KH, Panelli MC, Kim CJ, Riker AI, Bettinotti MP, Roden MM et al (1998) Functional dissociation between local and systemic immune response during anti-melanoma peptide vaccination. J Immunol 161(8):4183–4194
Lee KH, Wang E, Nielsen MB, Wunderlich J, Migueles S, Connors M et al (1999) Increased vaccine-specific T cell frequency after peptide-based vaccination correlates with increased susceptibility to in vitro stimulation but does not lead to tumor regression. J Immunol 163(11):6292–6300
Ascierto PA, Agarwala S, Botti G, Cesano A, Ciliberto G, Davies MA et al (2016) Future perspectives in melanoma research: meeting report from the “Melanoma Bridge”. Napoli, December 1st–4th 2015. J Transl Med 14(1):313
Amaria RN, Reddy SM, Tawbi HA, Davies MA, Ross MI, Glitza IC et al (2018) Neoadjuvant immune checkpoint blockade in high-risk resectable melanoma. Nat Med 24(11):1649–1654
Blank CU, Rozeman EA, Fanchi LF, Sikorska K, van de Wiel B, Kvistborg P et al (2018) Neoadjuvant versus adjuvant ipilimumab plus nivolumab in macroscopic stage III melanoma. Nat Med 24(11):1655–1661
Gubin MM, Zhang X, Schuster H, Caron E, Ward JP, Noguchi T et al (2014) Checkpoint blockade cancer immunotherapy targets tumour-specific mutant antigens. Nature 515(7528):577–581
Ward JP, Gubin MM, Schreiber RD (2016) The role of neoantigens in naturally occurring and therapeutically induced immune responses to cancer. Adv Immunol 130:25–74
Blankenstein T, Leisegang M, Uckert W, Schreiber H (2015) Targeting cancer-specific mutations by T cell receptor gene therapy. Curr Opin Immunol 33:112–119
Thomas A, Routh ED, Pullikuth A, Jin G, Su J, Chou JW et al (2018) Tumor mutational burden is a determinant of immune-mediated survival in breast cancer. Oncoimmunology 7(10):e1490854
Snyder A, Nathanson T, Funt SA, Ahuja A, Buros Novik J, Hellmann MD et al (2017) Contribution of systemic and somatic factors to clinical response and resistance to PD-L1 blockade in urothelial cancer: An exploratory multi-omic analysis. PLoS Med. 14(5):e1002309
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A et al (2014) Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 371(23):2189–2199
Goodman AM, Kato S, Bazhenova L, Patel SP, Frampton GM, Miller V et al (2017) Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol Cancer Ther 16(11):2598–2608
Johnson DB, Frampton GM, Rioth MJ, Yusko E, Xu Y, Guo X et al (2016) Targeted next generation sequencing identifies markers of response to PD-1 blockade. Cancer Immunol Res 4(11):959–967
McGranahan N, Furness AJ, Rosenthal R, Ramskov S, Lyngaa R, Saini SK et al (2016) Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351(6280):1463–1469
Rizvi H, Sanchez-Vega F, La K, Chatila W, Jonsson P, Halpenny D et al (2018) Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J Clin Oncol 36(7):633–641
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ et al (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348(6230):124–128
Saeterdal I, Bjorheim J, Lislerud K, Gjertsen MK, Bukholm IK, Olsen OC et al (2001) Frameshift-mutation-derived peptides as tumor-specific antigens in inherited and spontaneous colorectal cancer. Proc Natl Acad Sci USA 98(23):13255–13260
Brainstorm C, Anttila V, Bulik-Sullivan B, Finucane HK, Walters RK, Bras J et al (2018) Analysis of shared heritability in common disorders of the brain. Science. 360(6395)
Johanns TM, Ward JP, Miller CA, Wilson C, Kobayashi DK, Bender D et al (2016) Endogenous neoantigen-specific CD8 T cells identified in two glioblastoma models using a cancer immunogenomics approach. Cancer Immunol Res 4(12):1007–1015
Moore TV, Lyons GE, Brasic N, Roszkowski JJ, Voelkl S, Mackensen A et al (2009) Relationship between CD8-dependent antigen recognition, T cell functional avidity, and tumor cell recognition. Cancer Immunol Immunother 58(5):719–728
Voelkl S, Moore TV, Rehli M, Nishimura MI, Mackensen A, Fischer K (2009) Characterization of MHC class-I restricted TCRalphabeta+ CD4− CD8− double negative T cells recognizing the gp100 antigen from a melanoma patient after gp100 vaccination. Cancer Immunol Immunother 58(5):709–718
Zacharakis N, Chinnasamy H, Black M, Xu H, Lu YC, Zheng Z et al (2018) Immune recognition of somatic mutations leading to complete durable regression in metastatic breast cancer. Nat Med 24(6):724–730
Deniger DC, Pasetto A, Robbins PF, Gartner JJ, Prickett TD, Paria BC et al (2018) T-cell responses to TP53 “Hotspot” mutations and unique neoantigens expressed by human ovarian cancers. Clin Cancer Res 24(22):5562–5573
Spranger S, Luke JJ, Bao R, Zha Y, Hernandez KM, Li Y et al (2016) Density of immunogenic antigens does not explain the presence or absence of the T-cell-inflamed tumor microenvironment in melanoma. Proc Natl Acad Sci USA 113(48):E7759–E7768
Janeway CA, Jr (2013) Pillars article: approaching the asymptote? Evolution and revolution in immunology. Cold Spring Harb Symp Quant Biol 1989, 54:1–13. J Immunol 2013, 191(9):4475–4487
Janeway CA Jr (1989) Approaching the asymptote? Evolution and revolution in immunology. Cold Spring Harb Symp Quant Biol 54(Pt 1):1–13
Janeway CA Jr, Medzhitov R (1998) Introduction: the role of innate immunity in the adaptive immune response. Semin Immunol 10(5):349–350
Medzhitov R, Janeway CA Jr (1997) Innate immunity: the virtues of a nonclonal system of recognition. Cell 91(3):295–298
Medzhitov R, Janeway CA Jr (1997) Innate immunity: impact on the adaptive immune response. Curr Opin Immunol 9(1):4–9
Garg AD, Vandenberk L, Fang S, Fasche T, Van Eygen S, Maes J et al (2017) Pathogen response-like recruitment and activation of neutrophils by sterile immunogenic dying cells drives neutrophil-mediated residual cell killing. Cell Death Differ 24(5):832–843
Cui Z, Willingham MC, Hicks AM, Alexander-Miller MA, Howard TD, Hawkins GA et al (2003) Spontaneous regression of advanced cancer: identification of a unique genetically determined, age-dependent trait in mice. Proc Natl Acad Sci USA 100(11):6682–6687
Hicks AM, Riedlinger G, Willingham MC, Alexander-Miller MA, Von Kap-Herr C, Pettenati MJ et al (2006) Transferable anticancer innate immunity in spontaneous regression/complete resistance mice. Proc Natl Acad Sci USA 103(20):7753–7758
Hicks AM, Willingham MC, Du W, Pang CS, Old LJ, Cui Z (2006) Effector mechanisms of the anti-cancer immune responses of macrophages in SR/CR mice. Cancer Immun 6:11
Riedlinger G, Adams J, Stehle JR Jr, Blanks MJ, Sanders AM, Hicks AM et al (2010) The spectrum of resistance in SR/CR mice: the critical role of chemoattraction in the cancer/leukocyte interaction. BMC Cancer 10:179
Fuchs EJ, Matzinger P (1992) B cells turn off virgin but not memory T cells. Science 258(5085):1156–1159
Fuchs EJ, Ridge JP, Matzinger P (1996) Response: immunological tolerance. Science 272(5267):1406–1408
Matzinger P (1998) An innate sense of danger. Semin Immunol 10(5):399–415
Kroemer G, Galluzzi L, Kepp O, Zitvogel L (2013) Immunogenic cell death in cancer therapy. Annu Rev Immunol 31:51–72
Galluzzi L, Buque A, Kepp O, Zitvogel L, Kroemer G (2017) Immunogenic cell death in cancer and infectious disease. Nat Rev Immunol 17(2):97–111
Galluzzi L, Vitale I, Aaronson SA, Abrams JM, Adam D, Agostinis P et al (2018) Molecular mechanisms of cell death: recommendations of the nomenclature committee on cell death 2018. Cell Death Differ 25(3):486–541
Kawakami Y, Eliyahu S, Delgado CH, Robbins PF, Rivoltini L, Topalian SL et al (1994) Cloning of the gene coding for a shared human melanoma antigen recognized by autologous T cells infiltrating into tumor. Proc Natl Acad Sci USA 91(9):3515–3519
Kawakami Y, Eliyahu S, Delgado CH, Robbins PF, Sakaguchi K, Appella E et al (1994) Identification of a human melanoma antigen recognized by tumor-infiltrating lymphocytes associated with in vivo tumor rejection. Proc Natl Acad Sci USA 91(14):6458–6462
Galluzzi L, Senovilla L, Zitvogel L, Kroemer G (2012) The secret ally: immunostimulation by anticancer drugs. Nat Rev Drug Discov 11(3):215–233
Obeid M (2008) ERP57 membrane translocation dictates the immunogenicity of tumor cell death by controlling the membrane translocation of calreticulin. J Immunol 181(4):2533–2543
Pozzi C, Cuomo A, Spadoni I, Magni E, Silvola A, Conte A et al (2016) The EGFR-specific antibody cetuximab combined with chemotherapy triggers immunogenic cell death. Nat Med 22(6):624–631
Garg AD, Galluzzi L, Apetoh L, Baert T, Birge RB, Bravo-San Pedro JM et al (2015) Molecular and translational classifications of DAMPs in immunogenic cell death. Front Immunol 6:588
Spranger S, Gajewski TF (2015) A new paradigm for tumor immune escape: beta-catenin-driven immune exclusion. J Immunother Cancer 3:43
Spranger S, Sivan A, Corrales L, Gajewski TF (2016) Tumor and host factors controlling antitumor immunity and efficacy of cancer immunotherapy. Adv Immunol 130:75–93
Sweis RF, Spranger S, Bao R, Paner GP, Stadler WM, Steinberg G et al (2016) Molecular drivers of the non-T-cell-inflamed tumor microenvironment in urothelial bladder cancer. Cancer Immunol Res 4(7):563–568
Koyama S, Akbay EA, Li YY, Herter-Sprie GS, Buczkowski KA, Richards WG et al (2016) Adaptive resistance to therapeutic PD-1 blockade is associated with upregulation of alternative immune checkpoints. Nat Commun 7:10501
Benci JL, Xu B, Qiu Y, Wu TJ, Dada H, Twyman-Saint Victor C et al (2016) Tumor interferon signaling regulates a multigenic resistance program to immune checkpoint blockade. Cell 167(6):1540–1554.e12
Chihara N, Madi A, Kondo T, Zhang H, Acharya N, Singer M et al (2018) Induction and transcriptional regulation of the co-inhibitory gene module in T cells. Nature 558(7710):454–459
Gattinoni L, Lugli E, Ji Y, Pos Z, Paulos CM, Quigley MF et al (2011) A human memory T cell subset with stem cell-like properties. Nat Med 17(10):1290–1297
Gattinoni L, Klebanoff CA, Restifo NP (2012) Paths to stemness: building the ultimate antitumour T cell. Nat Rev Cancer 12(10):671–684
Crompton JG, Narayanan M, Cuddapah S, Roychoudhuri R, Ji Y, Yang W et al (2016) Lineage relationship of CD8(+) T cell subsets is revealed by progressive changes in the epigenetic landscape. Cell Mol Immunol 13(4):502–513
Restifo NP, Gattinoni L (2013) Lineage relationship of effector and memory T cells. Curr Opin Immunol 25(5):556–563
Blando J, Sharma A, Higa MG, Zhao H, Vence L, Yadav SS et al (2019) Comparison of immune infiltrates in melanoma and pancreatic cancer highlights VISTA as a potential target in pancreatic cancer. Proc Natl Acad Sci USA 116(5):1692–1697
Gao J, Ward JF, Pettaway CA, Shi LZ, Subudhi SK, Vence LM et al (2017) VISTA is an inhibitory immune checkpoint that is increased after ipilimumab therapy in patients with prostate cancer. Nat Med 23(5):551–555
Akalu YT, Rothlin CV, Ghosh S (2017) TAM receptor tyrosine kinases as emerging targets of innate immune checkpoint blockade for cancer therapy. Immunol Rev 276(1):165–177
Zhang B, Fang L, Wu HM, Ding PS, Xu K, Liu RY (2016) Mer receptor tyrosine kinase negatively regulates lipoteichoic acid-induced inflammatory response via PI3K/Akt and SOCS3. Mol Immunol 76:98–107
Ho WS, Wang H, Maggio D, Kovach JS, Zhang Q, Song Q et al (2018) Pharmacologic inhibition of protein phosphatase-2A achieves durable immune-mediated antitumor activity when combined with PD-1 blockade. Nat Commun 9(1):2126
Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR et al (2017) IFN-gamma-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest 127(8):2930–2940
Rossi J, Paczkowski P, Shen YW, Morse K, Flynn B, Kaiser A et al (2018) Preinfusion polyfunctional anti-CD19 chimeric antigen receptor T cells are associated with clinical outcomes in NHL. Blood 132(8):804–814
Iida N, Dzutsev A, Stewart CA, Smith L, Bouladoux N, Weingarten RA et al (2013) Commensal bacteria control cancer response to therapy by modulating the tumor microenvironment. Science 342(6161):967–970
Roy S, Trinchieri G (2017) Microbiota: a key orchestrator of cancer therapy. Nat Rev Cancer 17(5):271–285
Soldati L, Di Renzo L, Jirillo E, Ascierto PA, Marincola FM, De Lorenzo A (2018) The influence of diet on anti-cancer immune responsiveness. J Transl Med 16(1):75
Liebman MN, Molinaro S (2012) Computational modeling and epidemiologic approaches: a new section of the Journal of Translational Medicine. J Transl Med 10:210
Wang E, Uccellini L, Marincola FM (2012) A genetic inference on cancer immune responsiveness. Oncoimmunology 1(4):520–525
Angelova M, Mlecnik B, Vasaturo A, Bindea G, Fredriksen T, Lafontaine L et al (2018) Evolution of metastases in space and time under immune selection. Cell 175(3):751–765.e16
Botticelli A, Cerbelli B, Lionetto L, Zizzari I, Salati M, Pisano A et al (2018) Can IDO activity predict primary resistance to anti-PD-1 treatment in NSCLC? J Transl Med 16(1):219
Muller AJ, Manfredi MG, Zakharia Y, Prendergast GC (2019) Inhibiting IDO pathways to treat cancer: lessons from the ECHO-301 trial and beyond. Semin Immunopathol 41(1):41–48
Song C, Piva M, Sun L, Hong A, Moriceau G, Kong X et al (2017) Recurrent tumor cell-intrinsic and -extrinsic alterations during MAPKI-induced melanoma regression and early adaptation. Cancer Discov 7(11):1248–1265
Turan T, Kannan D, Patel M, Barnes MJ, Tanlimco SG, Lu R et al (2018) Immune oncology, immune responsiveness and the theory of everything. J Immunother Cancer (in press)
Abd Al Samid M, Chaudhary B, Khaled YS, Ammori BJ, Elkord E (2016) Combining FoxP3 and Helios with GARP/LAP markers can identify expanded Treg subsets in cancer patients. Oncotarget 7(12):14083–14094
Alinejad V, Dolati S, Motallebnezhad M, Yousefi M (2017) The role of IL17B-IL17RB signaling pathway in breast cancer. Biomed Pharmacother = Biomed & pharmacother 88:795–803
Moon PK, Tran S, Minhas PS (2019) Revisiting IDO and its value as a predictive marker for anti-PD-1 resistance. J Transl Med 17(1):31
Crittenden MR, Baird J, Friedman D, Savage T, Uhde L, Alice A et al (2016) Mertk on tumor macrophages is a therapeutic target to prevent tumor recurrence following radiation therapy. Oncotarget 7(48):78653–78666
Hatfield SM, Sitkovsky M (2016) A2A adenosine receptor antagonists to weaken the hypoxia-HIF-1alpha driven immunosuppression and improve immunotherapies of cancer. Curr Opin Pharmacol 29:90–96
Ohlund D, Handly-Santana A, Biffi G, Elyada E, Almeida AS, Ponz-Sarvise M et al (2017) Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J Exp Med 214(3):579–596
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bedognetti, D., Cesano, A., Marincola, F.M., Wang, E. (2020). The Biology of Immune-Active Cancers and Their Regulatory Mechanisms. In: Lee, P., Marincola, F. (eds) Tumor Microenvironment. Cancer Treatment and Research, vol 180. Springer, Cham. https://doi.org/10.1007/978-3-030-38862-1_5
Download citation
DOI: https://doi.org/10.1007/978-3-030-38862-1_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-38861-4
Online ISBN: 978-3-030-38862-1
eBook Packages: MedicineMedicine (R0)