Abstract
Rhizosphere harbors potential microbiomes which play a pivotal role in nutrient cycling, enhancing soil fertility, maintaining plant health and productivity. Specific microbiomes that are assembled near roots are considered to be some of the most complex ecosystems on the Earth. Heterogeneous microbial communities of rhizospheric microbiomes considerably vary by soil type, land use pattern, plant species, and host genotype. It is demonstrated that root exudates act as substrates and signaling molecules which are required for establishing plant–rhizobacterial interactions. The present chapter focused on the rhizosphere microbiomes of different agricultural crops, their functions, and possible biotechnological applications for increasing crop production in a sustainable manner. Further, the plant growth-promoting mechanisms of rhizobacteria were highlighted. Although much work has been done on the biocontrol characteristics of rhizospheric bacteria, it has to be considered that soil type, plant species, and the pathogen affect altogether influence the biocontrol efficiency of strain applied against a soil-borne pathogen.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1.1 Introduction
Soil microorganisms play a pivotal role in nutrient cycling, regulating soil fertility, maintaining plant health, and productivity (Wagg et al. 2014). Soil microbial communities are exceedingly complex and consist of various organisms such as bacteria, archaea, fungi, algae, and viruses. Most of these microorganisms largely utilize plant root-derived nutrients such as root exudates and secondary metabolites (Huang et al. 2014). Rhizosphere microorganisms are component of microbiomes that assemble near plant roots. Rhizospheric microbiomes are considered to be some of the most complex ecosystems on Earth. It is estimated that one gram of soil contains more than 50,000 different microbial species, but majority of them are uncultivable in nature (Roesch et al. 2007a, b). Beneficial free-living rhizospheric bacteria are generally referred to as plant growth-promoting rhizobacteria—“PGPR”. Conceptually, “PGPR” represents beneficial portion of rhizospheric microbiome and can have positive effect on both growth and development of plants by direct or indirect mechanisms.
Chemical compounds that are released by roots apparently modify physical and chemical characteristics of the soil (Mukherjee et al. 2018) and subsequently regulates the diversity and composition of soil microbial community in the rhizosphere (Huang et al. 2014). Moreover, plants may also influence composition of rhizosphere microbial communities by selectively stimulating microorganisms with beneficial traits that are needed for both plant growth and health (Chaparro et al. 2014). For example, Acidobacteria, Proteobacteria (mainly Alpha, Beta, and Deltaproteobacteria classes), Chloroflexi, and Actinobacteria are enriched in the rhizosphere of Oryza sativa, whereas soybean selected a specific microbial community consists of Bacteroidetes, Acidobacteria, Proteobacteria, and Actinobacteria (Lu et al. 2018; Ding et al. 2019; Yadav et al. 2016b). These microbial populations are found to colonize in the root rhizosphere because of their functional traits and also beneficial to plant nutrient absorption, growth, and disease suppression. In turn, the plant provides root exudates to the microbes which are used as substrates and signaling molecules (Mendes et al. 2013).
Studies revealed that root microbiomes considerably vary by soil type, habitat, land use pattern, plant species, and host genotype (Bouffaud et al. 2014; Fitzpatrick et al. 2018; Lu et al. 2018; Ding et al. 2019; Yadav et al. 2019f). In recent, the relationship between rhizosphere microbial communities and plant genotypes is well studied and the results may lead to increased plant productivity (Bouffaud et al. 2014; Bulgarelli et al. 2015; Pérez-Jaramillo et al. 2017; Leff et al. 2017; Ding et al. 2019). In this chapter, we summarize recent progress made in rhizosphere microbiomes of agriculture crops. We also discuss the importance of rhizosphere microbial communities particularly PGPR and their immense biotechnological values for sustainable production and productivity of agriculture crops.
1.2 Rhizosphere and Root Exudates
The narrow zone of soil surrounding the plant roots and influenced by roots, root hair, and plant-produced exudates is referred to as rhizosphere (Dessaux et al. 2009). There are three distinct interacting systems which are reported in the plant rhizosphere, viz., rhizoplane, rhizosphere, and the root itself. Rhizoplane is defined as the root surface including the strongly adhering soil particles. Group of bacteria which are inhabitants of rhizosphere and able to compete in colonizing the root system is known as “rhizobacteria” while the total microbial component (prokaryotes, eukaryotes, and viruses) of rhizosphere is termed as rhizo-microbiome or rhizosphere microbiome. The “rhizobacteria” term was first time introduced by Kloepper and Schroth (1978) to refer the soil bacterial population that competitively colonize the roots and stimulate plant growth, thereby reducing the incidence of diseases in a sustainable manner.
Specific microbiomes that are assembled near roots are proposed to be some of the most complex ecosystems on the Earth (Raaijmakers et al. 2009). Most of these microorganisms utilize diverse array of compounds/nutrients which are derived from plant roots in the rhizosphere (Lu et al. 2018; Yadav et al. 2017b). The chemicals that are released by roots in the soil are known as “root exudates.” It was suggested that chemicals secreted by plant roots act as signaling molecules and recruit wide variety of heterogeneous and metabolically active soil microbial populations (Ahemad and Kibert 2014) (Table 1.1).
Most importantly, the exudation of chemical compounds by roots apparently modifies the physical and chemical characteristics of the soil and subsequently regulates the structure and composition of rhizosphere microbial community (Doornbos et al. 2012). Impact of root exudates on bacterial communities in the rhizosphere was extensively reviewed by Doornbos et al. (2012). Further, it is estimated that around five to twenty-one percent of caron (photosynthetically fixed carbon) gets transported to the rhizosphere through the process of root exudation (Doornbos et al. 2012). Therefore, the rhizosphere is redefined by Dessaux et al. (2009) as “any volume of soil selectively influenced by plant roots, root hairs and plant-produced materials.”
The quality and quantity of the root exudates rely on type of plant species and distinct developmental/physiological status of plants (Kang et al. 2010). Furthermore, root exudates significantly enhance the plant-beneficial microbial–symbiotic interactions in the rhizosphere. These interactions, in turn, affect the rooting patterns, supply of available nutrients, thereby modifying the quantity and/or quality of root exudates. Microbial colonization in/on root tissues is known as root colonization, similarly microbial colonization of the adjoining volume of soil under the influence of the plant root system is defined as “rhizosphere colonization” (Ahemad and Kibert 2014). Compared with the bulk soil, microbial activity and biomass are relatively enhanced in the rhizosphere as a result of root exudation (Ahemad and Kibert 2014; Huang et al. 2014).
1.3 Rhizosphere Microbiome and Its Diversity
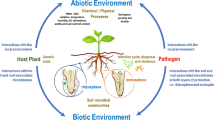
Most of the soils contain exceedingly high microbial diversity including bacteria, fungi, algae, viruses, and protozoa. It was reported that one gram of soil contains approximately 9 × 107 bacteria, 2 × 105 fungi, 4 × 106 actinomycetes, 5 × 103 protozoa, and 3 × 104 algae. The rhizosphere which is under influence of root exudates can harbor up to 10−11 microbial cells and around 30,000 different prokaryotic species per gram of root (Egamberdieva et al. 2008). Metagenomic analysis of tomato rhizosphere revealed that approximately 3,050 different bacterial species (OTUs at 3% distance cutoff) were associated in the rhizosphere (Tian et al. 2015). The rhizosphere microbiomes are very diverse and can actively interact with plants and mediate distinct agro-ecological process. The rhizosphere microbiome is considerably important in bridging the plant microbiomes and bulk soil and facilitates plant growth promotion by providing nutrition (Pathak et al. 2016) . The rhizobacterial microbiota also improves host plant’s health by protecting from phytopathogens and promotes plant growth and fitness in different physiochemical stresses by producing phytohormones (Fig. 1.1). It is imperative to elucidate the assembly, composition, and variation among the microbial communities present in the rhizosphere for understanding the diversity and metabolic functions of the rhizosphere microbiome. This information could be beneficial for sustainable management of plant health and the underlying mechanisms that drive microbiome assembly.
It has been revealed that the rhizosphere, rhizoplane (root surface), endosphere (root interior), and of host plants harbor a distinct microbiome (Edwards et al. 2015). Diversity, distribution, and the composition of the core rhizospheric microbiomes from several plant species such as Arabidopsis (Bulgarelli et al. 2012; Carvalhais et al. 2013; Chaparro et al. 2014), and economically important crops, viz., maize (Bouffaud et al. 2014), rice (Edwards et al. 2015; Malyan et al. 2016a, b; Lu et al. 2018; Moronta-Barrios et al. 2018; Ding et al. 2019), barley (Bulgarelli et al. 2015), citrus (Xu et al. 2018), sugar beet (Chapelle et al. 2016), sunflower (Leff et al. 2017), tomato (Tian et al. 2015), French bean (Pérez-Jaramillo et al. 2017), soybean (Mendes et al. 2011, 2014), wheat (Kour et al. 2019d; Verma et al. 2015a, b, 2016a, b; Yadav 2017a, 2019), and other tropical crop plants (Yadav 2017a, b; Yadav et al. 2019a; Yadav and Yadav 2018) have been established. All these studies have utilized 16S rRNA gene-based high-throughput sequencing analysis for understanding the microbial community dynamics. Although different methodologies have been suggested to explore soil microbial diversity and functions, culture-independent molecular methods are appropriate choice for deciphering diversity of microbiomes in high resolution (Fig. 1.2). Dominant microbial communities and their functions in core rhizospheric microbiomes of different agricultural crops have been extensively summarized in Table 1.2.
1.3.1 Diversity of Rhizospheric Microbiome in Wild Plants
Microorganisms represent the richest gamut of molecular and chemical diversity in nature, as they comprise the simplest yet dynamic forms of life (Yadav et al. 2015). Interest in the exploration of microbial diversity has been spurred by the fact that microbes are essential for life as they perform numerous functions integral to the sustenance of the biosphere, including nutrient cycling and environmental detoxification, which involve process such as augmentation, supplementation, and recycling of plant nutrients, so vital to sustainable agriculture (Kumar et al. 2019; Malyan et al. 2019; Rana et al. 2018; Yadav et al. 2017a, c, d). More recently, this largely unexplored reservoir of resources is the focus of investigations for innovative applications useful to mankind (Rastegari et al. 2019; Yadav et al. 2019c, d, e).
The distribution and diversity of bacterial community compositions in the rhizosphere microbiomes of six different wild plant species (Bidens biternata of the Asterales order, Ageratum conyzoides, Artemisia argyi, Euphorbia hirta, Viola japonica of the Malpighiales order, and Erigeron annuus) were evaluated by Lei et al. (2019). All the six different wild plant species were grown in the same experimental field. In this study, high-throughput sequencing of 16S rRNA gene targeting the hypervariable V3 and V4 regions was carried out with Illumina MiSeq platform. Comprehensive details for composition and distribution of rhizospheric microbiomes of wild plants have been shown in Fig. 1.2.
Approximately, 3000 OTUs for each rhizosphere sample were obtained. Rhizosphere microbiomes in the six wild plant species were dominated by bacterial phyla Proteobacteria (35%), Acidobacteria (12%), Actinobacteria (11%), Bacteroidetes (10%), Planctomycetes (8%), Chloroflexi (6%), and Verrucomicrobia (6%) and the details have been shown in Fig. 1.3a, b, c, d. Rhizobiales (8%) and Sphingomonadales (3.5%) orders of class Alphaproteobacteria (15%); Nitrosomonadales (4.28% ± 1.24%), and Burkholderiales (3%) orders of class Betaproteobacteria (9%); Myxococcales (5.5%) order of class Deltaproteobacteria (8%); and Xanthomonadales (4%) orders of class Gammaproteobacteria (7%) were found to be abundant in phylum Proteobacteria (Fig. 1.3a). Abundant members of phylum Actinobacteria were found to be Acidimicrobiales (4%). Similarly, Subgroup 4 (6%) and Subgroup 6 (4%) were abundant in Acidobacteria phylum.

Adapted with permission from Lie et al. (2019)
The composition and relative abundance of major bacterial taxa in a typical rhizosphere of six different wild plant species
The core rhizospheric microbiome of wild plant species showed a total of 1,109 operational taxonomic units (OTUs) affiliated to 113 bacterial genera accounting for more than 70% of the total sequencing data analyzed. The predominant bacterial genera of core OTUs are Variovorax, Acidibacter, Ferruginibacter, Bradyrhizobium, Blastocatella, Variibacter, Sphingomonas, and unclassified bacteria (Fig. 1.3e). The predominant bacterial orders were found to be composed of Xanthomonadales, Rhodospirillales, Rhizobiales, Burkholderiales, Sphingomonadales, Myxococcales, Nitrosomonadales of Proteobacteria; Acidimicrobiales of Actinobacteria; Subgroup 4 and Subgroup 6 of Acidobacteria.
Variations in microbial community compositions at the order level in the rhizosphere of six different plant species were also demonstrated (Lei et al. 2019). Predominant bacterial group in E. hirta rhizosphere is Proteobacteria, while the same group is least represent in V. japonica microbiome. Highly enriched Rhizobiales order of Proteobacteria was found in V. japonica and A. argyi. Predominant members of Myxococcales were noticed in V. japonica rhizosphere. Abundant members of Nitrosomonadales were observed in E. hirta. Similarly, higher abundance of Burkholderiales and Sphingomonadales was noticed in E. annuus. Members of Xanthomonadales were dominated in V. japonica rhizosphere.
1.3.2 Diversity of Rhizospheric Microbiome in Agriculture Crops
1.3.2.1 Rhizospheric Microbiome of Rice
The structure of microbial communities present in the rice rhizosphere is very complex, dynamic, and diverse (Edwards et al. 2015; Lu et al. 2018; Moronta-Barrios et al. 2018; Ding et al. 2019). Recently, microbiome inhabiting rice roots and rhizosphere is extensively reviewed by Ding et al. (2019). A study taken by Edwards et al. (2015) revealed that endosphere (inside the root compartment), rhizoplane (surface of the root), and rhizosphere of rice had distinct microbiomes. Microbial communities from the rice rhizosphere are established by amplification of the 16S rRNA gene (variable regions V4-V5) followed by high-throughput sequencing using the Illumina MiSeq platform (Edwards et al. 2015). Results indicate that rice endosphere microbial communities had the lowest α-diversity, whereas rice rhizosphere had higher α-diversity. Furthermore, the mean α-diversity was found to be relatively high in the rhizosphere than in the bulk soil (Edwards et al. 2015).
The most dominant bacterial genera of rice rhizosphere is summarized in Fig. 1.4. Bacterial community profiles and their relative abundance are shown in Fig. 1.5 (Lu et al. 2018). Bacterial, archaeal, and fungal communities and their relative abundance in the rice rhizosphere have been studied (Ding et al. 2019). Bacterial populations were found to be abundant in the rice rhizosphere (Edwards et al. 2015). The abundance of rhizosphere microbial populations such as bacterial, fungal, and archaeal was twice those that of the bulk soil which is an indication of rhizospheric effect (Ding et al. 2019).

Adapted from Moronta-Barrios et al. (2018)
Dominant bacterial genera in the rhizosphere microbiome of rice

Adapted with permission from Lu et al. (2018)
Bacterial community profiling in the rhizospheric microbiomes of Hordeum vulgare (Barley), Triticum aestivum (Wheat), Oryza sativa Indica and Japonica (Rice)
Proteobacteria dominated the microbiome of rice rhizosphere accounting more than 71%. Among Proteobacteria, the most abundant class was Gammaproteobacteria followed by Betaproteobacteria, Alphaproteobacteria, Deltaproteobacteria, and Epsilonproteobacteria (Moronta-Barrios et al. 2018). Representatives of Epsilonproteobacteria and Deltaproteobacteria classes were not detected in the rice endorhizosphere. Phyla Verrucomicrobia and Bacteroidetes were abundant across the samples. Representative members of Nitrospirae and Acidobacteria were found only in rice-rhizospheric samples (Moronta-Barrios et al. 2018). Bacterial phyla Proteobacteria, Firmicutes, Chloroflexi, Nitrospirae, Spirochaetes, Fibrobacteres, Planctomycetes, Bacteroidetes, Proteobacteria, Actinobacteria, Verrucomicrobia, Cyanobacteria, and Acidobacteria are the most commonly found bacterial members of the rice rhizosphere (Edwards et al. 2015; Lu et al. 2018; Moronta-Barrios et al. 2018; Ding et al. 2019). Similarly, the most common bacterial genera of rice rhizosphere are as follows: Pseudomonas sp., Limnobacter, Devosia, Opitutus, Flavobacterium, Shewanella, Caulobacter, Agrobacterium, Pseudomonas veronii, Methylotenera mobilis, Microvirgula aerodenitrificans, Pedobacter, Rhodoferax, Variovorax, Mycoplana, Rheinheimera, Flavisolibacter, Fluviicola, Chryseobacterium, Asticcacaulis, Halothiobacillus, Pleomorphomonas, Sphingobium, Thiobacillus, Bacillus sp., Flavobacterium gelidilacus, Methylophaga, and Acidovorax (Moronta-Barrios et al. 2018). Further details on dominant microbial communities and their functions in rice-rhizospheric microbiomes have been summarized in Table 1.2.
Methanogenic archaea, viz., Methanobacterium, Methanosarcina, Methanocella, and Methanosaeta were also reported in the rice rhizosphere (Malyan et al. 2016a; Edwards et al. 2015). Firmicutes, Bacteroidetes, Betaproteobacteria, and Chloroflexi were found to be differentially enriched in rice rhizosphere. The Betaproteobacterial OTUs that are enriched in rice rhizosphere belong to mainly Comamonadaceae and Rhodocyclaceae families (Edwards et al. 2015). Total bacterial count of rice rhizosphere is approximately 5 × 10 9 cells g dw−1 soil, whereas as archeal members are found to be 2.5 × 10 8 cell g dwt−1 soil.
1.3.2.2 Rhizospheric Microbiome of Wheat and Barley
Rhizosphere community of wheat was analyzed by comparative metatranscriptomics approach (Hayden et al. 2018). The rhizosphere community of wheat was predominately bacteria. Classes Gammaproteobacteria, Alphaproteobacteria, and Actinobacteria were dominant in the rhizosphere of wheat and barley. Bacterial families such as Micrococcaceae, Enterobacteriaceae, and Pseudomonadaceae were abundant in the rhizosphere microbiomes (Hayden et al. 2018). Predominant archaeal members in the rhizosphere are affiliated to family Nitrososphaeraceae under phylum Thaumarchaeota. Ascomycota is the dominant fungal phylum found in the rhizosphere representing more than 72% of total fungal transcripts. Other fungal phyla in the rhizosphere of wheat and barley were affiliated to Basidiomycota (>10%), which includes the genus Rhizoctonia, and Glomeromycota (4%) form arbuscular mycorrhizae. Interestingly, fungal families represent a smaller proportion of the total microbial transcripts analyzed in the rhizosphere (Hayden et al. 2018). Rhizosphere community of barley was reported by Lu et al. (2018). Bacterial phyla Proteobacteria, Bacteroidetes, Acidobacteria, Planctomycetes, Nitrospirae, Actinobacteria, Verrucomicrobia, Firmicutes, Cyanobacteria, Chloroflexi, and Gemmatimonadetes were associated with barley rhizosphere. Further, fungi Ascomycota, Basidiomycota, Zygomycota, and Unidentified fungi were distributed in the barley rhizosphere (Hayden et al. 2018).
1.3.2.3 Rhizospheric Microbiome of Soybean
Shotgun metagenomics approach was used to study functional and taxonomic diversities of microbial communities in the rhizosphere of soybean, Glycine max (L.) (Mendes et al. 2014). Metagenomic libraries were dominated by bacteria (>95%) followed by eukaryotes (3%) and archaea and virus (1%). Proteobacteria was found to be the most abundant phylum in soybean rhizosphere and represented around 47% distribution. Other dominant bacterial phyla in the rhizosphere of soybean were found to be Actinobacteria (23%), Acidobacteria (5%), and Firmicutes (6%) (Mendes et al. 2014). In general, 28% of total sequences found in the soybean rhizosphere were novel and were not affiliated to known bacterial taxa. Results indicate clear differences in microbial community structure among rhizosphere and bulk soil. Overrepresentation of the phyla Acidobacteria, Actinobacteria, Chloroflexi, Chlamydiae, Cyanobacteria, Deferribacteres, Tenericutes, Chlorobi, Aquificae, and Verrucomicrobia was found in rhizosphere and the results were significant at P < 0.01 (Mendes et al. 2014). Similarly, abundance of class Mollicutes, Bacilli, Clostridia, Epsilonproteobacteria, Gammaproteobacteria, Thermomicrobia, and Chlamydiae was found in the rhizosphere of Glycine max (L.).
1.3.2.4 Rhizospheric Microbiome of French Bean
Microbiome of French bean was elucidated by amplification of 16S rRNA (V3–V4 region) followed by high-throughput sequencing performed at Illumina MiSeq platform (Pérez-Jaramillo et al. 2017). Phylum Proteobacteria was the dominant member, whereas lower abundance of Acidobacteria was noticed in wild bean rhizosphere. The phyla Verrucomicrobia and Bacteroidetes were predominant in the wild bean rhizosphere. Phylum Actinobacteria was found to be more abundant in the modern bean rhizosphere and these results were statistically significant.
Significant increase in the relative abundance of bacterial families Sphingomonadaceae and Rhizobiaceae was observed in the rhizosphere as compared to the bulk soil. Furthermore, it was noticed that there is a gradual decrease in the relative abundance of the Chitinophagaceae and Cytophagaceae of the Bacteroidetes phylum in the French bean rhizosphere. Gradual increases in relative abundance of families Streptomycetae and Nocardiodaceae of Actinobacteria and Rhizobiaceae of Proteobacteria.
1.3.2.5 Microbiome of Maize and Other Members of Poaceae Crops
Rhizospheric microbiome of Poaceae crops such as Zea mays L; Zea mays ssp. Parviglumis; Sorghum bicolor cv. Arprim; Triticum aestivum L. cv. Fiorina was established by Bouffaud et al. (2014). The dominant bacterial members of Microbiome of Poaceae crops were found to be Rhodospirillales such as Gluconacetobacter, Rhodospirillum, Azospirillum, and Sphingomonadaceae of class Burkholderiales; Actinomycetales such as Corynebacterium, Actinomyces, Propionibacterium, and Kocuria; Acidovorax of Alphaproteobacteria; Xanthomonas, Francisella, Pantoea, Moraxella, Pseudomonas, and Photorhabdus of class Gammaproteobacteria; Burkholderia, Hydrogenophaga, and Alcaligenes of class Betaproteobacteria; Myxococcales such as Anaeromyxobacter of class Deltaproteobacteria; Mogibacterium, Bacillales (Firmicutes) such as Bacillus and Paenibacillus; Megasphaera and Collinsella (Bouffaud et al. 2014).
1.4 Factors Influencing Rhizospheric Microbiome in Agriculture Crops
The rhizosphere microbiomes participate in very important functions suitable for plant growth promotion. The key functions mediated by rhizosphere microbiome include abiotic stress tolerance, nutrient acquisition, and protection against plant pathogen infection. Therefore, understanding the assembly of rhizosphere microbiome and their molecular mechanisms will provide us basic information. This information will be useful to develop soil management practices, designing of healthy rhizosphere microbiome, and introduction of biofertilizers and biological control agents to develop sustainable agricultural strategies. Different factors that are influencing structure, assembly, and function of rhizospheric microbiomes are depicted in Fig. 1.6.
Rhizobacterial community composition in Phaseolus vulgaris was influenced by specific root morphological traits and host plant genotype (Pérez-Jaramillo et al. 2017). Impact of host plant genotype on rhizosphere microbial community was mediated by qualitative and quantitative composition of root exudates (Huang et al. 2014; Ahemad and Kibert 2014). Host genotype had a tremendous effect on the composition of root-associated microbial communities in Hordeum vulgare (Bulgarelli et al. 2015). Bulgarelli et al. (2012) reported that host genotype and soil type define the diversity of root-inhabiting bacterial communities in Arabidopsis thaliana. Plant cell wall properties confer sufficient colonization (40%) of root-associated microbiota in Arabidopsis thaliana (Bulgarelli et al. 2012).
Invading fungal pathogens and plant stress response induces a shift in microbiome composition of sugar beet (Chapelle et al. 2016). Rhizosphere microbial community structure varied according to the Poaceae genotype (Bouffaud et al. 2014). Evolutionary divergence among host plants and type of plant species affects the assembly of the rhizosphere and endosphere and microbiome (Fitzpatrick et al. 2018). The root microbiome is also associated with drought tolerance of host plants (Fitzpatrick et al. 2018, Kour et al. 2019). Different developmental stages of plant also influence rhizosphere microbiome assemblages (Chaparro et al. 2014). Rhizosphere microbiome of Oryza sativa is shaped by plant and soil-related conditions such as soil type, geographic location, rice genotype, oxic–anoxic interface, agricultural management, and growth stages (Ding et al. 2019). Selection of the microbial community in the wheat rhizosphere depends on niche-based processes as a result of environmental factors and the selection power of the plant (Mendes et al. 2014). Further, agricultural management practices and growth stages of host plants exerted much influence on the rice rhizosphere microbiome (Edwards et al. 2015).
1.5 Plant Growth-Promoting Mechanisms of Rhizospheric Microbiome
Rhizobacteria plays a crucial role in growth promotion and immunity of the agricultural crops. These plant growth promoters follow certain mechanisms during the entire sequential process for nutrient mobilization, phytohormones for the growth and development, and chemical agents for defense-related issues of the crops (Suman et al. 2016; Verma et al. 2017; Yadav et al. 2018a, b). According to Mahanty et al. (2016), similar mechanisms are adopted by all bacterial genera during promotion of plant growth, although they are phylogenetically of different origins. The mechanisms behind the scene could mainly be divided into two types, direct mechanisms and indirect mechanisms. Comprehensive details of plant growth-promoting mechanisms of rhizobacteria in different agricultural crops have been summarized in Table 1.3.
1.5.1 Direct Mechanism
The direct mechanisms mainly involve the bacterial activities like phosphate solubilization, nitrogen fixation, secretion of plant hormones, ACC deaminase activities, and siderophore production.
1.5.1.1 Phosphate Solubilization
In spite of the large reservoir of phosphorus in soil, a very low amount of it is available to the plants (Ahemad and Kibret 2014). This is because plants utilize them in only two forms: (a) monobasic and (b) dibasic ions (Bhattacharyya and Jha 2012). It has also been reported that due to rapid conversion of phosphorus into insoluble complexes of different metal oxides most of the cultivable soils are deficit of available phosphate (Sandilya et al. 2016). Phosphate fertilizers are mostly applied to the agricultural soils in order to overcome the overall loss. But, continuous use of these chemical fertilizers is harmful to the soil and the environment in vivo. Hence, the importance of biofertilizers having plant growth-promoting traits was raised worldwide.
Native rhizobacteria pays an immense attribute to solubilize the inorganic phosphate so as to make it available for the utilization of various crops or plants (Widawati 2011). Certain bacterial genera, viz., Bacillus, Pseudomonas, Azospirillum, Achromobacter, Acetobacter, Acinetobacter, Enterobacter, Klebsiella, and Serratia are able to solubilize the inorganic form of phosphate to the available form (Kumar et al. 2012; Rana et al. 2019a, b). Besides, the role of bacterial organic acids for cation uptake by the plants is also worth mentioning (Sandilya et al. 2016). Researchers further stated that the bacterial genera belonging to the Proteobacteria and some of the Firmicutes and Actinobacteria are the most capable of the abovementioned conversion process.
1.5.1.2 Nitrogen Fixation
Nitrogen being the most important limiting factors, its fixation in nature is an interesting phenomena led by the plant growth-promoting rhizobacteria both in the symbiotic and non-symbiotic or free-living forms (Fagodiya et al. 2017a, b). It has been believed that the free-living nitrogen fixers provide a very lower amount of available nitrogen to the plants in comparison to the symbiotic nitrogen fixers since time immemorial (James and Olivares 1997). The nif genes found in the nitrogen-fixing rhizobacteria complete the nitrogenase enzyme by the means of its structural and regulatory proteins responsible for activation of the Fe protein, iron molybdenum, cofactor biosynthesis, and electron donation in case of the former and synthesis and function of the enzyme in the later (Glick 2012). Numerous PGPR genera capable of converting nitrate into nitrite by the catalysis of the nitrate reductase enzyme have also been reported. The most common among them are Azospirillum, Azotobacter, Achromobacter, Bradyrhizobium, Beijerinckia, and Rhizobium (Kour et al. 2019b, c; Yadav et al. 2019b).
1.5.1.3 Phytohormones and ACC Deaminase Enzyme Activity
Major plant hormones such as IAA and GA3 (Marques et al. 2010; Ahmed and Hasnain 2010 and Khan et al. 2014) along with cytokinin secretion (Liu et al. 2013) by the PGPR’s have often being reported by various authors. The IAA secreted by the bacterial population associated with the roots of the agricultural crops could augment the root surface area and length that could pave an easier route for absorption of the soil nutrients by the plants (Ahemad and Khan 2012). Amino acid tryptophan being a major precursor of IAA boosts the level of IAA biosynthesis. Almost five different types of IAA pathways have been reported by Spaepen and Vanderleyden (2011).
The role of GA3 has also been explained by some authors in the context of plant growth-promoting rhizobacteria. The most important among them are the induction of seed germination and emergence and development of stem, leaf, flower, and fruits (Bottoni et al. 2004). The most common bacterial strains Bacillus cereus, Sphingomonas sp. LK11 were reported by them to enhance the growth and production of red pepper and tomato. Similarly, some other mechanisms of a plant body, viz., cytokinesis, sensitivity of vascular cambium, and their differentiation and root apical dominance are being conducted by the hormone cytokinin. Root-associated bacteria such as Azotobacter chroococcum, Bacillus megaterium, and B. subtilis were accounted to produce cytokinin thereby enhancing plant growth. On the other hand, synthesis of ethylene by the plant growth-promoting rhizobacteria induces ripening of fruits, opening of flowers, and leaf abscission.
Plants growing under stress are able to withstand the adverse effects of the environment with the due help of these phytohormones (de Garcia et al. 2006). Ethylenes produced in such conditions are called as “stress ethylene” that adds to the existing production of ethylene. However, excessive production of ethylene is a harmful phenomena for the longer development of the roots and in order to check such level of production, PGPR’s with the help of 1-aminocyclopropane-1-carboxylate (ACC) deaminase plays a vital role in the early stages of growth which modulates the level of ethylene by hydrolyzing ACC, a precursor of ethylene, in ammonia and a-ketobutyrate (Glick et al. 1998; Marques et al. 2010). Bacteria synthesizing IAA along with endogenous plant IAA could stimulate plant growth or accelerate the amalgamation of the enzyme ACC synthase translating the compound S-adenosyl methionine to ACC being the immediate precursor of ethylene in higher plants (Glick 2012). Different kinds of phytohormones and their plant growth-promoting activity in agriculture crops have been summarized in Table 1.4.
1.5.1.4 Siderophore Production
Iron being one of the most important nutrients for all forms of life is found to occur as Fe3+ that could most likely form insoluble hydroxides and oxyhydroxides making it nearly impossible for plants and microflora for easy access (Rajkumar et al. 2010). In order to overcome such situations, bacteria secretes siderophores which are iron chelating agents with low molecular mass. According to Glick (2012), siderophores are mostly water soluble and could be divided into extracellular and intracellular siderophores. Siderophore forming Fe3+-siderophore complex on the bacterial membranes gets reduced to Fe2+. These ionic forms of iron are released into the cell from the complex via another mechanism linking both the membrane systems (inner and outer) which may finally lead to the destruction or recycling of the left out siderophore (Rajkumar et al. 2010). Thus, the siderophores prove to be excellent iron solubilizing agents from minerals and other inorganic sources. Pseudomonads, the bacterial genera, are the best-known secretors of siderophores playing an important role in the overall plant growth promotion activities (Sandilya et al. 2017).
1.5.2 Indirect Mechanisms
Plant growth-promoting rhizobacteria has been implemented in various crop fields for their promising capability to work both as biocontrol agents and growth promoters since last two decades. Bacteria secretes various metabolites and chemical agents that makes them wonderful candidates for controlling different crop diseases most of them being originated from fungal sources. According to Bhattacharyya and Jha (2012), PGPRs are able to synthesize different antifungal secondary metabolites such as phenazines, HCN, pyrrolnitrin, 2, 4-diacetylphloroglucinol, viscosinamide, tensin, and pyoluteorin. Availability of bacterial antagonist in the rhizosphere soil may even adapt the plant for developing induced systemic resistance against broad-spectrum bacterial, fungal, and viral pathogens (Lugtenberg and Kamilova 2009). Cyanide is the most dangerous chemical known for its high toxic properties which can well inhibit the pathogens sensitizing agricultural crops. HCN being the secondary metabolite secreted by the PGPRs does not have any pessimistic effect on the host plants, and hence they are frequently used for controlling weeds (Zeller et al. 2007). According to various reports, HCN-producing PGPRs are very helpful in controlling dreaded phytopathogens such as Pythium ultimum, Fusarium oxysporum, and pathogenic Agrobacterium. The mode of action mechanisms involves lysis of fungal cell walls (Maksimov et al. 2011), root colonization (Kamilova et al. 2005), reduction of stress ethylene level (Van Loon 2007), siderophore and antibiotic production (Beneduzi et al. 2012).
Certain genera like Bacillus have been best studied for their ability to secrete antimicrobial traits with higher rate of agricultural applicability (Compant et al. 2005). The members of this group of bacteria hold a key role in biocontrol aspects as they could reluctantly replicate at a very faster rate and are mostly resistant to environmental stress (Shafi et al. 2017). They secrete bacillomycins, iturins, and mycosubtilin very much effective against fungal pathogens, particularly Aspergillus flavus (Gong et al. 2015). Similarly, Lee et al. (2015) reported almost 99.1% of the antagonistic success in crops fields inoculated with Bacillus amyloliquefaciens strain HK34 against Phytophthora cactorum in Lycopersicum esculentum, Sclerotium rolfsii, Capsicum annuum var. acuminatum, Colletotrichum gloeosporioides, and Cucumis sativus.
Apart from that, other bacterial genera like Pseudomonas and Paenibacillus have also been reported by various authors having antimicrobial properties in both in vitro and in vivo conditions. Although laboratory results may not always be relied under field conditions, PGPR has been reported to be effective in both the conditions in different agricultural cropping systems. That is why they may be termed as multifunctional agents by controlling a wide spectrum range of phytopathogens and a spectacular replacement for chemical fertilizers by enhancing plant growth and overall yield per hectares of cultivated soil further playing a vital role in maintaining ecological balance across the globe (Ahemad and Kibret 2014).
Although much work has been done on the biocontrol characteristics of rhizospheric bacteria, it has to be considered that soil type, plant species, and pathogen affect in rhizosphere competence and/or biocontrol efficiency of applied biocontrol strain against a soil-borne pathogen.
1.6 Biotechnological Applications of Rhizosphere Microbiomes
In the recent past, sustainable technologies have gained lot of momentum to improve quality and yield of agricultural crop production. Nevertheless, still there is uncertainty about success of chemical-based formulations in plant protection management. In general, pests and diseases are mainly controlled by chemical-based pesticides which pose major health risks as well as adverse negative impacts in the ecosystem and environment. In addition to this, indiscriminate use of chemical fertilizers resulted in negative impacts on biodiversity and function of biogeochemical cycles. Most importantly, agricultural practices require novel products according to the demand of farmers and consumers. Therefore, alternative management tools have to be developed on the basis of biological solutions.
The plant rhizosphere hosts a considerable amount of microbiome. Plant growth-promoting rhizobacteria (PGPR) is an integral component of rhizosphere microbiome and is competent to promote plant growth by direct and indirect mechanisms. PGPR also promotes defense against diseases causing organisms using diverse plant-beneficial functions. Therefore, it is anticipated that crop inoculation with suitable PGPR could reduce the use of pesticides and fertilizers in agrosystems. Biotechnological applications of various PGPR inoculants for enhancing crop production were summarized in Table 1.5. Since most of the research information on PGPR comes from rhizosphere microbiome, one can further explore and exploit biotechnical prospects of rhizosphere microbiomes for sustainable agricultural production. We have specially highlighted the production of extracellular lytic enzymes, bioactive metabolites, and volatile organic compounds (VOCs) of rhizosphere bacteria in this section and the details are given extensively in Table 1.6.
1.6.1 Production of Lytic Enzymes by Rhizospheric Bacteria
Rhizosphere bacteria can benefit plant growth indirectly through biocontrol mechanisms which can inhibit the growth and colonization of phytopathogens. This potential antagonism character of biocontrol agent might occur through different mechanisms which include production of extracellular lytic enzymes, secondary metabolites, siderophores, antibiotics, and induction of systemic responses (Saraf et al. 2014, Jadhav and Sayyed 2016; Kour et al. 2019a; Yadav et al. 2016a, 2019f). One of the important mechanisms for biocontrol agent is the production of lytic enzymes which are able to degrade the membrane constituents of phytopathogens, such as proteases (Felestrino et al. 2018), acylases, and lactonases (Combes-Meynet et al. 2011). These hydrolytic enzymes degrade the structural integrity of the pathogen cell wall. Their ability to inhibit phytopathogens makes them to be the preferable choice in biological control process. The application of these hydrolytic enzymes from rhizospheric origin is a viae solution as they are totally natural and are eco-friendly in nature (Mishra et al. 2019).
Lytic enzymes produced by various microorganisms can hydrolyze polymeric compounds like cellulose, hemicellulose, chitin, and protein of phytopathogens. Extracellular hydrolytic enzymes like chitinases, lipases, proteases, and glucanases are involved in the lysis of fungal cell wall (Neeraja et al. 2010). These enzymes either disintegrate or digest the molecular components of cell wall of fungal phytopathogens. Therefore, this process would be considered as eco-friendly control of soil-borne pathogens in agriculture crops. These enzymes further involve in nutrient cycling by decomposition of organic matter and plant residues in the rhizosphere. It is demonstrated that extracellular lytic enzymes produced by Myxobacteria sp. have the ability to suppress fungal plant pathogens (Bull et al. 2002). In an another study, glucanase-producing antagonistic bacteria Lysobacter sp. is capable of controlling diseases of Pythium sp. and Bipolaris sp. (Palumbo 2005). These hydrolytic enzymes rescue plants from biotic stresses and directly contribute in the parasitization of phytopathogens.
Hydrolytic enzymes of rhizospheric microbes were reviewed extensively by Jadhav and Sayyed 2016. Many rhizobacterial microbial species are capable of producing extracellular enzymes and effectively hydrolyze wide variety of polymeric substances like cellulose, hemicellulose, proteins, and chitin of phytopathogens (Jadhav and Sayyed 2016). Microbial strains like B. subtilis strains PCL1608 PCL1612, Streptomyces cyaneofuscatus B-49, Serratia marcescens strain ETR17, Pseudomonas fluorescens, Serratia marcescens strain ETR17, and many other antagonistic microbes have a potential to synthesize hydrolytic enzymes for the biocontrol of fungal phytopathogens like P. ultimum, F. oxysporum, R. solani, and S. rolfsii, (Cazorla et al. 2007; Kumar et al. 2012a, b; Purkayastha et al. 2018, El-Gamal et al. 2016). The mode of actions of extracellular enzymes is given in Table 1.7.
Chitinolytic microorganisms are heavily colonized in plant rhizosphere among which actinobacteria are the most abundant members (Yadav et al. 2018c). Actinobacteria such as Streptomyces flavotricini, Streptomyces kanamyceticu, Streptomyces cyaneofuscatus, and Streptomyces rochei produce chitinases and inhibit the growth of phytopathogen, viz, Verticillium dahlia in cotton rhizosphere (Xue et al. 2013). Chitinase-producing Bacillus thuringiensis spp. colmeri can inhibit the growth of plant pathogenic fungi, including Rhizoctonia solani, Penicillium chrysogenum, and Physalospora piricola (Liu et al. 2010). Biocontrol agent Bacillus subtilis inhibits the growth of pathogenic fungi Fusarium oxysporum through production of extracellular chitinase (Gajbhiye et al. 2010). Chitinases produced by Brevibacillus laterosporus effectively inhibit the growth of phytopathogenic fungi Fusarium equiseti (Prasanna et al. 2013). Lysobacter enzymogenes showed to inhibit Pythium aphanidermatum by producing extracellular protease and lipases (Folman et al. 2003)
Minimal use of chitinase-based fungicides in agriculture crops was associated with the perception that their efficacy will be slowly reduced in the soil environment. Nevertheless, Dahiya et al. (2006) extensively reviewed biotechnological prospects of chitinolytic enzymes and suggested that chitinases can be used as supplementary inputs along with other chemical-based fungicides to enhance their effectiveness against phytopathogenic fungi and reduce the required amount of chemical fungicides. In addition to this, it was shown that the application of mixed consortia containing two different chitinolytic bacteria is more effective in controlling the pathogen. Application of chitinase-producing Streptomyces sp. 385, Paenibacillus sp. 300, and both together is more effective in controlling cucumber wilt caused by F. oxysporum than individual strains applied (Singh et al. 1999). Similar kind of observation was reported by El-Tarabily et al. (2000) wherein growth of fungal pathogen Sclerotinia responsible for vegetable rot was effectively controlled by combination of S marcescens, Streptomyces viridodiasticus, and Micromonospora carbonacea strains. In recent, chitinase, protease, lipase, and cellulose-producing Serratia marcescens strain ETR17 showed in vitro antagonism toward nine different root and foliar pathogens of tea (Purkayastha et al. 2018).
Actinomycetes were considered to be strong biocontrol agents against fungal pathogens. This is mainly due to production of different types of antifungal compounds such as antibiotics and extracellular hydrolytic enzymes which includes chitinases and glucanases (Xue et al. 2013; Yadav et al. 2018c). Streptomyces halstedii, Streptomyces cavourensis SY224, and Streptomyces griseus are known to produce potential antifungal extracellular chitinases, which makes them to be used as biocontrol agents in crop protection strategies (Ki et al. 2012; Gherbawy et al. 2012). Lysobacter spp. was reported to be an effective biocontrol agent against soil-borne pathogens through production of extracellular enzymes and other metabolites (Folman et al. 2003). Lysobacter spp. was abundant in the soil which is suppressive to root pathogen, viz., Rhizoctonia solani. Certain antagonistic strains showed in vitro biocontrol activity against Xanthomonas campestris, R. solani, and other important phytopathogens such as Aspergillus niger, Fusarium oxysporum, and Pythium ultimum.
These natural microbial biofungicides will be used as integrated pest management supplement for reduction of negative impact of chemical pesticides on the environment and maintain the sustainable production of agriculture.
1.6.2 Production of Antibiotics
Rhizospheric bacteria produce distinct antimicrobial products to inhibit the growth and colonization of plant pathogens to compete the nutrients present in the rhizosphere. This has become a beneficial trait to the host plant as disease development is significantly reduced by PGPR. Rhizosphere harbors diverse actinomycetes species which have been further exploited for secondary metabolites (Yadav et al. 2018b; Geetanjali and Jain 2016). Actinobacteria is known to produce wide variety of natural antimicrobial products (approximately 10,000 secondary metabolites) (Passari et al. 2015, 2017; Yadav et al. 2018a, b). Production of antibiotics by Actinobacteria was extensively reviewed by Yadav et al. (2018b, c). Application of secondary metabolites producing rhizobacterial isolates against phytopathogens is increasing over the past decade (Yilmaz et al. 2008). A variety of antimicrobial agents such as 2,4-diacetylphloroglucinol (DAPG), pyoluteorin (PRN), phenazine, cyclic lipopeptides, tensin, and pyrrolnitrin (PLT) have been screened and identified from Pseudomonas sp., Arthrobacter sp., and Streptomyces sp., (Weller 2007; Gupta et al. 2015). Details of antibiotics/secondary metabolites producing organisms and their application in different crops have been summarized in Table 1.6.
Rhizospheric soil isolates Bacillus sp. S2 and Pseudomonas fluorescens S5 were found to exert good antimicrobial activity against multi-drug-resistant clinical pathogens such as Pseudomonas aeruginosa, Klebsiella pneumonia, Escherichia coli, and Staphylococcus aureus obtained from different samples (Dhore et al. 2014). Thirty Pseudomonas fluorescens strains isolated from rice rhizosphere against pathogenic fungi Sarocladium oryzae, Dreschelaria oryzae, Magnaporthe grisea, and Rhizoctonia solani. Among these, P. fluorescens Pf 003 effectively inhibited (62–85%) the mycelial growth in all the pathogenic fungi in dual culture. The antifungal compounds extracted with ethyl acetate from P. fluorescens at 5% completely inhibited the pathogens (Reddy et al. 2007). Walia et al. (2013) isolated the bacteria from the tomato rhizosphere for having broad-spectrum antifungal activity against Sclerotinia sclerotiorum, Rhizoctonia solani, and Fusarium oxysporum.
DAPG, phenazines, PLT, and PRN are considered to be potent antibiotics synthesized by Pseudomonas biocontrol agents affiliated to gammaproteobacteria (Table 1.6). In recent, antibiotics-producing Pseudomonas spp. has got much attention in biocontrol research, and corresponding genes involved in the expression and regulation of these metabolites are now fully understood (Weller 2007and there in references). For the last 30 years, developments on biocontrol applications of Pseudomonas sp. against soil-borne pathogens have been summarized by Weller (2007). P. fluorescens strain CHA0 was isolated from tobacco rhizosphere which is naturally suppressive to black root rot of tobacco caused by Thielaviopsis basicola (Stutz et al. 1986). P. fluorescens CHA0 produces siderophore (pseudobactin), PLT, DAPG, PRN, HCN, salicylic acid, pyoverdine, indoleacetic acid, pyochelin, and other secondary metabolites (Voisard et al. 1994). Antagonistic bacterium P. fluorescens F113 isolated from sugar beet was applied in the field for suppression of damping-off of sugar beet infection caused by a pathogen Pythium ultimum (Cronin et al. 1997a, b).
Antibiotics such as bacilysin- and iturin-producing Bacillus subtilis ME488 suppressed soil-borne pathogens in pepper and cucumber crops (Chung et al. 2008). Secondary metabolites, viz., Pyrrolnitrin and prodigiosin-producing Serratia marcescens strain ETR17 Serratia marcescens strain ETR17 showed significant level of in vitro antagonistic property against different root and foliar pathogens of tea (Purkayastha et al. 2018). Antifungal lipopeptides such as surfactin-, fengycin-, and iturin-producing B. subtilis strains PCL1608 and PCL1612 have shown biocontrol mechanism toward soil-borne pathogen Fusarium oxysporum (Cazorla et al. 2007). Paenibacillus sp. strain B2 isolated sorghum mycorrhizosphere showed production of antibiotic polymyxin B1 and significantly inhibited the growth of fungal pathogens (Selim et al. 2005). Antifungal peptides-producing Bacillus sp. KM 5 isolated from rice rhizosphere showed antagonist activity toward pathogenic fungi Gibberella fujikuroi, Sclerotium rolfsii Saccardo, Fusarium udum, Helminthosporium oryzae, and Rhizoctonia solani Nees (Majumdar et al. 2011).
1.6.3 Production of Volatile Organic Compounds (VOCs)
Volatile organic compounds are lipophilic low molecular weight (<300 g mol − 1) compounds emitted from microbial metabolic pathways with high vapor pressure and low boiling point. VOCs can act as signal molecules in rhizosphere over short and long distances (Fincheira and Quiroz 2018). It is evidenced that VOCs released from diverse rhizospheric microorganisms, e.g., Arthrobacter sp., Proteus sp., Bacillus sp., Fusarium sp., Pseudomonas sp., Alternaria sp. and Laccaria sp., can promote plant growth on a specific “target”. Detailed description about chemical nature of VOCs and their functions have been summarized in Table 1.6. Ryu et al. (2003) reported for the first time about the mechanism mediated by volatile organic compounds released by Bacillus subtilis GB03 which induced growth on Arabidopsis thaliana. This study evidenced that VOCs can modulate stress, growth, nutrition, and health processes in host plants. Some identified VOCs compounds, such as acetoin, β-Caryophyllene 2,3-butanediol, Sesquiterpenes, 2-pentylfuran, and dimethylhexadecylamine, have shown their ability to elicit plant growth at above and below ground biomass (Fincheira and Quiroz 2018; Chung et al. 2016) (Table 1.6).
Few studies indicate that VOCs act as signals and chemical messengers to regulate phytohormone synthesis, metabolic pathways, and nutrition levels. Effects of VOCs for induction of resistance and tolerance in plants are documented, wherein compounds such as 3-pentanol, dimethyl disulfide, 6-pentyl-α-pyrone, and acetoin were reported. VOCs derived by rhizospheric bacteria showed antagonistic activity toward plant pathogen Rhizoctonia solani and inhibit mycelial growth (Kai et al. 2007). Certain plant volatiles are proven to induce plant growth promotion through biochemical signals, eliciting local defence reactions known as induced systemic resistance (Chung et al. 2016; Kai et al. 2007). Long-chain VOCs signaling molecules, acetoin 2,3-butanediol, ethanethiol, isoprene, and acetic acid-butyl ester, and tridecane are found to be involved in induced resistance in Arabidopsis (Lee et al. 2012a, b). Yi et al. (2016) reported that 2,3-butanediol is produced by a Bacillus subtilis isolate involved in plant defense mechanisms. Root exudates of pepper inoculated with the B. subtilis were used to challenge various phytopathogens. For example, growth of Trichoderma sp (saprophytic fungus) and Ralstonia solanacearum (soil-borne pathogen) was inhibited by VOCs. This indicates that VOCs triggered the secretion of root exudates and subsequently acted as a plant defence inducer toward soil-borne fungal and bacterial pathogens.
Volatile organic compounds such as dehydroaromadendrene, α-pinene, tetrahydro-2,2,5,5-tetramethylfuran, (-)-trans-caryophyllene, and (+)-sativene-producing Cladosporium cladosporioides strain CL-1 showed increased growth parameters in Tobacco crop (Paul and Park 2013). In an another study, rhizospheric isolates such as Bacillus subtilis GB03, Bacillus amyloliquefaciens IN937, Pseudomonas fluorescens 89B-61, and Paenibacillus polymyxa E681 produced Brassinosteroid a long-chain VOC and signaling molecules such as acetoin 2,3-butanediol, ethanethiol, acetic acid-butyl ester, and isoprene. These VOCs are involved in induced systemic resistance in Arabidopsis (Lee et al. 2012a, b). Fresh weight, shoot length, chlorophyll concentration, and lateral root numbers of Sorghum were significantly increased by dimethylhexadecylamine produced by Arthrobacter agilis UMCV2. Salt tolerance, increased shoot and root length, fresh weight, and leaf surface area were increased in soybean by VOCs, 4-nitroguaiacol, and quinoline produced by Vaishnav et al. (2016). VOCs of fungal origin also showed increased growth parameters in host plants like lettuce, Arabidopsis, and tobacco. Fusarium oxysporum MSA 35 showed production of β-Caryophyllene and increased fresh weight of tobacco in field experiment (Minerdi et al. 2011). Sesquiterpenes synthesized by ectomycorrhizal fungi Laccaria bicolour increased the lateral root of Arabidopsis (Ditengou et al. 2015). In the same study, it was demonstrated that other ectomycorrhizal ascomycote, Cenococcum geophilum, which cannot synthesize Sesquiterpenes does not promote lateral root of Arabidopsis. These studies indicate that volatile organic compounds emitted by microorganisms in the rhizosphere are cheaper, effective, efficient, and eco-friendly alternatives for controlling phytopathogens.
Environmentally friendly biotechnological approaches offer the development of PGPR inoculants and their potential application in metal-contaminated systems. Plant growth promotion by PGPR is a result from improved nutrient acquisition or phytohormonal stimulation (Table 1.3). Different mechanisms involved in plant growth promotion were shown in Fig. 1.6. PGPR inoculants were widely used in agriculture, forestry, horticulture, and in environmental restoration/phytoremediation sectors.
1.7 Conclusion and Future Prospects
Although studies have focused on plant microbiome structure and its function under natural and agricultural environments, there have been no significant coordinated efforts to combine and translate research results into practical solutions for farmers. According to Busby et al. (2017), integration of beneficial plant microbiome into agricultural production is one of the ways to assist in achieving these goals. However, this requires large-scale efforts from academic and industry researchers, farmers, and policy-makers to understand and manage complex plant–microbiome interactions under current challenges of the agriculture production.
For achieving this goal, five key research priorities have been identified by Busby et al. (2017). Few research priorities include development of host–microbiome model systems with associated microbial culture collections and reference genomes; characterization and refinement of a model “plant genotype–environment stress–microbiome–management interactions”; elucidation of the role core microbiome and determine functional mechanisms of plant–microbiome interactions. These research priorities may enable us to manipulate agricultural microbiomes and thereby to develop management strategies for increased production and productivity of global agriculture in a sustainable manner. One of the challenges for future research work includes protection and conservation of rhizosphere biodiversity and their potential application in agricultural soils. Sustainable agriculture production may not be possible unless integration of plant germplasm and beneficial microbial species in the current agricultural practices globally.
Exploitation and production of natural drug formulations from microbial species have gained a significant leap during last three decades. Therapeutic applications of anticancerous compounds extracted from actinobacteria have been well addressed (Busi and Pattnaik 2018). The research priority is now shifted toward rhizosphere microbial communities for developing new drugs through high-throughput screening and fermentation techniques. Exploitation of bioprospecting potential of rhizosphere microbiomes is an upcoming new avenue.
References
Abd El-Fattah DA, Ewedab WE, Zayed MS, Hassaneina MK (2013) Effect of carrier materials, sterilization method, and storage temperature on survival and biological activities of Azotobacter chroococcum inoculants. Ann Agric Sci 58:111–118
Abd El-Lattief EA (2016) Use of azospirillum and azobacter bacteria as biofertilizers in cereal crops: a review. Int J Res Eng Appl Sci 6(7):36–44
Ahemad M, Khan MS (2009) Effect of insecticide-tolerant and plant growth promoting Mesorhizobium on the performance of chick pea grown in insecticide stressed alluvial soils. J Crop Sci Biotechnol 12:213–222
Ahemad M, Khan MS (2010a) Ameliorative effects of Mesorhizobium sp. MRC4 on chickpea yield and yield components under different doses of herbicide stress. Pestic Biochem Physiol 98:183–190
Ahemad M, Khan MS (2010b) Improvement in the growth and symbiotic attributes of fungicide-stressed chickpea plants following plant growth promoting fungicide-tolerant Mesorhizobium inoculation. Afr J Basic Appl Sci 2:111–116
Ahemad M, Khan MS (2012) Effect of fungicides on plant growth promoting activities of phosphate solubilizing Pseudomonas putida isolated from mustard (Brassica campestris) rhizosphere. Chemosphere 86(9):945–950
Ahemad M, Kibret M (2014) Mechanisms and application of plant growth promoting rhizobacteria: Current perspective. J King Saud Univ Sci 26(1):1–20
Ahmed A, Hasnain S (2010) Auxin-producing Bacillus sp.: auxin quantification and effect on the growth of Solanum tuberosum. Pure Appl Chem 82(1):313–319
Ahsan T, Chen J, Zhao X, Irfan M, Wu Y (2017) Extraction and identification of bioactive compounds (eicosane and dibutyl phthalate) produced by Streptomyces strain KX852460 for the biological control of Rhizoctonia solani AG-3 strain KX852461 to control target spot disease in tobacco leaf. AMB Exp 7:54
Babalola OO, Osir EO, Sanni AI, Odhaimbo GD, Bulimo WD (2003) Amplification of 1-aminocyclopropane-1-carboxylic (ACC) deaminase from plant growth promoting rhizobacteria in Striga-infested soils. Afr J Biotechnol 2(6):157–160
Bal HB, Das S, Dangar TK, Adhya TK (2013) ACC deaminase and IAA producing growth promoting bacteria from the rhizosphere soil of tropical rice plants. J Basic Microbio 53(12):972–984
Belimov AA, Hontzeas N, Safronova VI, Demchinskaya SV, Piluzza G, Bullitta S, Glick BR (2005) Cadmium-tolerant plant growth-promoting bacteria associated with the roots of Indian mustard (Brassica juncea L. Czern.). Soil Biol Biochem 37:241–250
Beneduzi A, Ambrosini A, Passaglia LMP (2012) Plant growth-promoting rhizobacteria (PGPR): their potential as antagonists and biocontrol agents. Gen Mole Biol 35(4):1044–1051
Bent E, Tuzun S, Chanway CP, Enebak S (2001) Alterations in plant growth and in root hormone levels of lodgepole pines inoculated with rhizobacteria. Can J Microbiol 47(9):793–800
Bhardwaj D, Ansari MW, Sahoo RK, Tuteja N (2014) Biofertilizers function as key player in sustainable agriculture by improving soil fertility, plant tolerance and crop productivity. Micro Cell Fact 13:66
Bhattacharyya D, Garladinne M, Lee Y (2015) Volatile indole produced by rhizobacteriumProteus vulgaris JBLS202 stimulates growth of Arabidopsis thaliana through auxin, cytokinin, and brassinosteroid pathways. J Plant Growth Regul 34:158–168
Bhattacharyya PN, Jha DK (2012) Plant growth-promoting rhizobacteria (PGPR): emergence in agriculture. World J Microbiol Biotechnol 28:1327–1350
Bouffaud ML, Poirier MA, Muller D, Moënne-Loccoz Y (2014) Root microbiome relates to plant host evolution in maize and other P oaceae. Env Microbiol 16(9):2804–2814
Bulgarelli D, Garrido-Oter R, Münch PC, Weiman A, Dröge J, Pan Y, McHardy AC, Schulze-Lefert P (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17(3):392–403
Bulgarelli D, Rott M, Schlaeppi K, van Themaat EVL, Ahmadinejad N, Assenza F, Rau P, Huettel B, Reinhardt R, Schmelzer E, Peplies J (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488(7409):91
Bull CT, Shetty KG, Subbarao KV (2002) Interactions between Myxobacteria, plant pathogenic fungi, and biocontrol agents. Plant Dis 86:889–896
Busby PE, Soman C, Wagner MR, Friesen ML, Kremer J, Bennett A, Morsy M, Eisen JA, Leach JE, Dangl JL (2017) Research priorities for harnessing plant microbiomes in sustainable agriculture. PLoS Biol 15(3):2001793
Busi S, Pattnaik SS (2018) Current status and applications of actinobacteria in the production of anticancerous compounds. In: New and future developments in microbial biotechnology and bioengineering, Elsevier, pp 137–153
CarvalhaisLC Dennis PG, Badri DV, Tyson GW, Vivanco JM, Schenk PM (2013) Activation of the jasmonic acid plant defence pathway alters the composition of rhizosphere bacterial communities. PLoS ONE 8(2):56457
Cassan F, Maiale S, Masciarellia O, Vidal A, Luna V, Ruiz O (2009) Cadaverine production by Azospirillum brasilense and its possible role in plant growth promotion and osmotic stress mitigation. Eur J Soil Biol 45:12–19
Castulo-Rubio DY, Alejandre-Ramírez NA, Orozco-Mosqueda MC, Santoyo G, Macías-Rodríguez L, Valencia-Cantero E (2015) Volatile organic compounds produced by the rhizobacterium Arthrobacter agilis UMCV2 modulate Sorghum bicolor (Strategy II Plant) morphogenesis and SbFRO1 transcription in vitro. J Plant Growth Regul 34:611–623
Cazorla FM, Romero D, Pérez-García A, Lugtenberg BJJ, Vicente AD, Bloemberg G (2007) Isolation and characterization of antagonistic Bacillus subtilis strains from the avocado rhizoplane displaying biocontrol activity. J Appl Microbiol 103(5):1950–1959
Chaparro JM, Badri DV, Vivanco JM (2014) Rhizospheremicrobiome assemblage is affected by plant development. ISME J 8(4):790
Chapelle E, Mendes R, Bakker PAH, Raaijmakers JM (2016) Fungal invasion of the rhizospheremicrobiome. ISME J 10(1):265
Chin-A-Woeng TF, Bloemberg GV, Lugtenberg BJ (2003) Phenazines and their role in biocontrol by Pseudomonas bacteria. New Phytol 157(3):503–523
Chung JH, Song GC, Ryu CM (2016) Sweet scents from good bacteria: case studies on bacterial volatile compounds for plant growth and immunity. Plant Mol Biol 90:677–687
Chung S, Kong H, Buyer JS, Lakshman DK, Lydon J, Kim SD, Roberts DP (2008) Isolation and partial characterization of Bacillus subtilis ME488 for suppression of soil borne pathogens of cucumber and pepper. Appl Microbiol Biotechnol 80(1):115–123
Combes-Meynet E, Pothier JF, Moenne-Loccoz Y, Prigent-Combaret C (2011) The Pseudomonas secondary metabolite 2,4-diacetylphloroglucinol is a signal inducing rhizoplane expression of Azospirillumgenes involved in plant-growth promotion. Mol Plant Microbe Interact 24:271–284
Compant S, Brion D, Jerzy N, Christophe C, Essaid AB (2005) Use of Plant Growth-Promoting Bacteria for biocontrol of plant diseases: principles, mechanisms of action and future prospects. Appl Environ Microbiol 71(9):4951–4959
Creus CM, Sueldo RJ, Barassi CA (2004) Water relations and yield in Azospirillum inoculated wheat exposed to drought in the field. Can J Bot 82:273–281
Cronin D, Moënne-Loccoz Y, Fenton A, Dunne C, Dowling DN, O’Gara F (1997a) Ecological interaction of a biocontrol Pseudomonas fluorescens strain producing 2,4-diacetylphloroglucinol with the soft rot potato pathogen Erwinia carotovora subsp. atroseptica. FEMS Microbiol Ecol 23:95–106
Cronin D, Moënne-Loccoz Y, Fenton A, Dunne C, Dowling DN, O’Gara F (1997b) Role of 2,4-diacetylphloroglucinol in the interactions of the biocontrol pseudomonad strain F113 with the potato cyst nematode Globodera rostochiensis. Appl Environ Microbiol 63:1357–1361
Dahiya N, Tewari R, Hoondal GS (2006) Biotechnological aspects of chitinolytic enzymes: a review. Appl Microbiol Biotech 25:1–10
de Garcia, Salamone IE, Hynes RK, Nelson LM (2006) Role of cytokinins in plant growth promotion by rhizosphere bacteria. PGPR: biocontrol and biofertilization. Springer, Netherlands, Amsterdam, pp 173–195
de Salamone IEG, Hynes RK, Nelson LM (2001) Cytokinin production by plant growth promoting rhizobacteria and selected mutants. Can J Microbiol 47(5):404–411
Dessaux Y, Hinsinger P, Lemanceau P (2009) Rhizosphere: so many achievements and even more challenges. Plant Soil 321:1–3
Dhore M, Barate D, Musaddiq M (2014) Studies on in-vitro anti microbial potential of rhizospheric soil bacteria against multi drug resistant clinical isolates. Ind J Appl Res 4(7):446–449
Ding LJ, Cui HL, Nie SA, Long XE, Duan GL, Zhu YG (2019) Microbiomes inhabiting rice roots and rhizosphere. FEMS Microbiol Ecol. https://doi.org/10.1093/femsec/fiz040
Ditengou FA, Muller A, Rosenkranz M, Felten J, Lasok H, Van Doorn MM, Legué V, Palme K, Schnitzler JP, Polle A (2015) Volatile signalling by sesquiterpenes from ectomycorrhizal fungi reprogrammes root architecture. Nat Commun 6:6279
Doornbos R, Loon L, Bakker PHM (2012) Impact of root exudates and plant defence signaling on bacterial communities in the rhizosphere: a review. Agron Sust Dev 32(1):227–243
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Eisen JA, Sundaresan V (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. P Natl Acad Sci 112(8):911–920
Egamberdieva D, Kamilova F, Validov S, Gafurova L, Kucharova Z, Lugtenberg B (2008) High incidence of plant growth-stimulating bacteria associated with the rhizosphere of wheat grown on salinated soil in Uzbekistan. Environ Microbiol 10:1–9
El-Gamal NG, Shehata AN, Hamed ER, Shehata HS (2016) Improvement of lytic enzymes producing Pseudomonas fluorescens and Bacillus subtilis isolates for enhancing their biocontrol potential against root rot disease in tomato plants. Res J Pharm Biol Chem Sci 7(1):1393–1400
El-Tarabily KA (2006) Rhizosphere-competent isolates of streptomycete and non-streptomycete actinomycetes capable of producing cell-wall-degrading enzymes to control Pythium aphanidermatum damping-off disease of cucumber. Bot 84(2):211–222
El-Tarabily KA, Soliman MH, Nassar AH, Al-Hassani HA, Sivasithamparam K, McKenna F, Hardy GESTJ (2000) Biological control of Sclerotinia minor using a chitinolytic bacterium and actinomycetes. Plant Pathol 49(5):573–583
Fagodiya RK, Pathak H, Bhatia A, Kumar A, Singh SD, Jain N (2017a) Simulation of Maize (Zea Mays L.) yield under alternative nitrogen fertilization using infocrop-maize model. Biochem Cell Arch 17:65–71
Fagodiya RK, Pathak H, Kumar A, Bhatia A, Jain N (2017b) Global temperature change potential of nitrogen use in agriculture: a 50-year assessment. Sci Rep 7:44928
Felestrino ÉB, Vieira IT, Caneschi WL, Cordeiro IF, Assis RDAB, de CarvalhoLemes CG, Fonseca NP, Sanchez AB, Cepeda JCC, Ferro JA, Garcia CCM (2018) Biotechnological potential of plant growth-promoting bacteria from the roots and rhizospheres of endemic plants in ironstone vegetation in southeastern Brazil. World J Microbiol Biotechn 34(10):156
Fierro-Coronado RA, Quiroz-Figueroa FR, García-Pérez LM, Ramírez-Chávez E, Molina-Torres J, Maldonado-Mendoza IE (2014) IAA-producing rhizobacteria from chickpea (Cicerarietinum L.) induce changes in root architecture and increase root biomass. Can J Microbiol 60(10): 639–648
Fincheira P, Quiroz A (2018) Microbial volatiles as plant growth inducers. Microbiol Res 208:63–75
Fitzpatrick CR, Copeland J, Wang PW, Guttman DS, Kotanen PM, Johnson MT (2018) Assembly and ecological function of the root microbiome across angiosperm plant species. Proc Natl Acad Sci USA 115(6):E1157–E1165
Folman LB, Postma J, Van Veen JA (2003) Characterisation of Lysobacter enzymogenes (Christensen and Cook 1978) strain 3.1 T8, a powerful antagonist of fungal diseases of cucumber. Microbiol Res 158:107–115
Gajbhiye A, Rai AR, Meshram SU, Dongre AB (2010) Isolation, evaluation and characterization of Bacillus subtilis from cotton rhizospheric soil with biocontrol activity against Fusariumoxysporum. World J Microbiol Biotechnol 26(7):1187–1194
Geetanjali, Jain P (2016) Antibiotic production by rhizospheric soil microflora-a review. Int J Pharm Sci Res 7(11):4304–4314
Gherbawy Y, Elhariry H, Altalhi A, El-Deeb B, Khiralla G (2012) Molecular screening of Streptomycesisolates for antifungal activity and family 19 chitinase enzymes. J Microbiol 50(3):459–468
Gholami A, Shahsavani S, Nezarat S (2009) The effect of plant growth promoting rhizobacteria (PGPR) on germination seedling growth and yield of maize. Int J Biol Life Sci 5:1
Glick BR (2012) Plant growth promoting bacteria: mechanisms and applications. Scientifica 1–5
Glick BR, Penrose DM, Li J (1998) A model for the lowering of plant ethylene concentrations by plant growth promoting bacteria. J Theoretical Biol 190:63–68
Gong AD, Li HP, Yuan QS, Song XS, Yao W, He WJ, Zhang JB, Liao YC (2015) Antagonistic mechanism of Iturin A and Plipastatin A from Bacillus amyloliquefaciens S76-3 from wheat spikes against Fusarium graminearum. PLoS ONE 10(2):e0116871
Grossman JM, Schipanski ME, Sooksanguan T, Drinkwater LE (2011) Diversity of rhizobia nodulating soybean Glycine max (Vinton) varies under organicand conventional management. Appl Soil Ecol 50:14–20
Gutiérrez‐Mañero FJ, Ramos‐Solano B, Probanza AN, Mehouachi J, Tadeo FR, Talon M (2001) The plant‐growth‐promoting rhizobacteria Bacillus pumilus and Bacillus licheniformis produce high amounts of physiologically active gibberellins. Physiol Plant 111(2):206–211
Harrison LA, Letendre L, Kovacevich P, Pierson E, Weller D (1993) Purification of an antibiotic effective against Gaeumannomyces graminis var. tritici produced by a biocontrol agent, Pseudomonas aureofaciens. Soil Biol Biochem 25:215–221
Hashem A, Abd Allah EF, Alqarawi A, Al-Huqail AA, Wirth S, Egamberdieva D (2016) The interaction between arbuscularmy corrhizal fungi and endophytic bacteria enhances plant growth of Acacia gerrardii under salt stress. Front Plant Sci 7:1089
Hayden HL, Savin K, Wadeson J, Gupta V, Mele PM (2018) Comparative metatranscriptomics of wheat rhizosphere microbiomes in disease suppressive and non-suppressive soils for Rhizoctonia solani AG8. Front Microbiol 9:859
Hidri R, Barea JM, Mahmoud OM, Abdelly C, Azcón R (2016) Impact of microbial inoculation on biomass accumulation by Sulla carnosa provenances, and in regulating nutrition, physiological and antioxidant activities of this species under non-saline and saline conditions. J Plant Physiol 201:28–41
Howie WJ, Suslow T (1991) Role of antibiotic synthesis in the inhibition of Pythium ultimum in the cotton spermosphere and rhizosphere by Pseudomonas fluorescens. Mol Plant-Microbe Interact 4:393–399
Huang XF, Chaparro JM, Reardon KF, Zhang R, Shen Q, Vivanco JM (2014) Rhizosphere interactions: root exudates, microbes, and microbial communities. Botany 92(4):267–275
Hussain N, Mujeeb F, Tahir M, Khan GD, Hassan NM, Bari A (2002) Effectiveness of Rhizobium under salinity stress. Asian J Plant Sci 1:12–14
Jadhav HP, Sayyed RZ (2016) Hydrolytic enzymes of rhizospheric microbes in crop protection. MOJ Cell Sci Rep 3(5):00070
James EK (2000) Nitrogen fixation in endophytic and associative symbiosis. Field Crops Res 65:197–209
James EK, Olivares FL (1997) Infection and colonization of sugar cane and other graminaceous plants by endophytic diazotrophs. Crit Rev Plant Sci 17(1):77–119
Jimtha JC, Jishma P, Arathy GB, C Anisha, Radhakrishnan EK (2016) Identification of plant growth promoting Rhizosphere Bacillus sp. WG4 antagonistic to Pythium myriotylum and its enhanced antifungal effect in association with Trichoderma. J Soil Sci Plant Nutr 16(3):578–590
Kai M, Effmert U, Berg G, Piechulla B (2007) Volatiles of bacterial antagonists inhibit mycelial growth of the plant pathogen Rhizoctonia solani. Arch Microbiol 187:351–360
Kamilova F, Validov S, Azarova T, Mulders I, Lugtenberg B (2005) Enrichment for enhanced competitive plant root tip colonizers selects for a new class of biocontrol bacteria. Environ Microbiol 7:1809–1817
Kang BG, Kim WT, Yun HS, Chang SC (2010) Use of plant growth-promoting rhizobacteria to control stress responses of plant roots. Plant Biotechnol Rep 4:179–183
Kaushik R, Saxena AK, Tilak KVBR (2000) World J Microbiol Biotechnol 16(6):567–570
Khan AL, Hamayun M, Kim YH, Kang SM, Lee JH, Lee IN (2011) Gibberellins producing endophytic Aspergillus fumigatus sp. LH02 influenced endogenous phytohormonal levels, isoflavonoids production and plant growth in salinity stress. Process Biochem 46:440–447
Khan AL, Waqas M, Kang SM (2014) Bacterial endophytes Sphingomonas sp LK11 produces gibberellins and IAA and promotes tomato plant growth. J Microbiol 52:689–695
Kloepper JW, Schroth MN (1978) Plant growth-promoting rhizobacteria on radishes. In: Proceedings of the 4th international conference on plant pathogenic bacteria. Gilbert-Clarey, Tours, pp 879–882
Kour D, Rana KL, Kumar A, Rastegari AA, Yadav N, Yadav AN, Gupta VK (2019a) Extremophiles for hydrolytic enzymes productions: biodiversity and potential biotechnological applications. In: Molina G, Gupta VK, Singh BN, Gathergood N (eds) Bioprocessing for biomolecules production. Wiley, USA, pp 321–372
Kour D, Rana KL, Yadav AN, Yadav N, Kumar V, Kumar A, Sayyed RZ, Hesham AEL, Dhaliwal HS, Saxena, AK (2019). Drought-tolerant phosphorus-solubilizing microbes: biodiversity and biotechnological applications for alleviation of drought stress in plants. In: Plant growth promoting rhizobacteria for sustainable stress management. Springer, Singapore, pp 255–308
Kour D, Rana KL, Yadav N, Yadav AN (2019b) Bioprospecting of phosphorus solubilizing bacteria from Renuka Lake Ecosystems, Lesser Himalayas. J Appl Biol Biotechnol 7:1–6
Kour D, Rana KL, Yadav N, Yadav AN, Kumar A, Meena VS, Singh B, Chauhan VS, Dhaliwal HS, Saxena AK (2019c) Rhizospheric microbiomes: biodiversity, mechanisms of plant growth promotion, and biotechnological applications for sustainable agriculture. In: Kumar A, Meena VS (eds) Plant growth promoting rhizobacteria for agricultural sustainability: from theory to practices. Springer Singapore, Singapore, pp 19–65. https://doi.org/10.1007/978-981-13-7553-8_2
Kour D, Rana KL, Yadav N, Yadav AN, Singh J, Rastegari AA, Saxena AK (2019d) Agriculturally and industrially important fungi: current developments and potential biotechnological applications. In: Yadav AN, Singh S, Mishra S, Gupta A (eds) Recent advancement in white biotechnology through fungi: Volume 2: perspective for value-added products and environments. Springer International Publishing, Cham, pp 1–64. https://doi.org/10.1007/978-3-030-14846-1_1
Kumar A, Chaturvedi AK, Yadav K, Arunkumar KP, Malyan SK, Raja P, Kumar R, Khan SA, Yadav KK, Rana KL, Kour D, Yadav N, Yadav AN (2019) Fungal Phytoremediation of Heavy Metal-Contaminated Resources: Current Scenario and Future Prospects. In: Yadav A, Singh S, Mishra S, Gupta A (eds) Recent Advancement in White Biotechnology Through Fungi. Fungal Biology. Springer, Cham
Kumar A, Gupta DK, Kumar M (2013) Green manure crops: a boon for agricultural soil. Int J Agri Environ Biotechnol 6:193
Kumar A, Kumar A, Devi S, Patil S, Chandani P, Nagi S (2012) Isolation, screening and characterization of bacteria from rhizospheric soils from different plant growth promotion activities: as in vitro study. Recent Res Sci Technol 4(1):1–5
Kumar P, Dubey RC, Maheshwari DK (2012) Bacillus strains isolated from rhizosphere showed plant growth promoting and antagonistic activity against phytopathogens. Microbiol Res 167(8):493–499
Lee B, Farag MA, Park HB, Kloepper JW, Lee SH, Ryu CM (2012a) Induced resistance by a long-chain bacterial volatile: elicitation of plant systemic defense by a C13 volatile produced by Paenibacilluspolymyxa. PLoS One 7:48744
Lee BD, Dutta S, Ryu H, Yoo SJ, Suh DS, Park K (2015) Induction of systemic resistance in Panax ginseng against Phytophthora cactorum by native Bacillus amyloliquefaciens HK34. J Ginseng Res 39(3):213–220
Lee SY, Tindwa H, Lee YS, Naing KW, Hong SH, Nam Y, Kim KY (2012b) Biocontrol of anthracnose in pepper using chitinase, beta-1,3 glucanase, and 2-furancarboxaldehyde produced by Streptomyces cavourensis SY224. J Microbiol Biotechnol 2(10):1359–1366
Leff JW, Lynch RC, Kane NC, Fierer N (2017) Plant domestication and the assembly of bacterial and fungal communities associated with strains of the common sunflower, Helianthus annuus. New Phytol 214(1):412–423
Lei S, Xu X, Cheng Z, Xiong J, Ma R, Zhang L, Yang X, Zhu Y, Zhang B, Tian B (2019) Analysis of the community composition and bacterial diversity of the rhizosphere microbiome across different plant taxa. Microbiol Open. https://doi.org/10.1002/mbo3.762
Liu D, Cai J, XieCh-Ch Liu Ch, Chen Y-H (2010) Purification and partial characterization of a 36-kDa chitinase from Bacillus thuringiensis spp. colmeri, and its biocontrol potential. Enzyme Microb Technol 46:252–256
Liu F, Xing S, Ma H, Du Z, Ma B (2013) Cytokinin-producing, plant growth-promoting rhizobacteria that confer resistance to drought stress in Platycladus orientalis container seedlings. Appl Microbiol Biotechnol 97:9155–9164
Lu T, Ke M, Peijnenburg WJGM, Zhu Y, Zhang M, Sun L, Fu Z, Qian H (2018) Investigation of rhizospheric microbial communities in wheat, barley, and two rice varieties at the seedling stage. J Agri Food Chem 66(11):2645–2653
Lucas JA, Solano BR, Montes F, Ojeda J, Megias M, Gutierrez Ma-nero FJ (2009) Use of two PGPR strains in the integrated management of blast disease in rice (Oryza sativa) in Southern Spain. Field Crop Res 114:404–410
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting rhizo-bacteria. Annu Rev Microbiol 63:541–556
Ma W, Guinel FC, Glick BR (2003) Rhizobium leguminosarum biovar viciae 1-aminocyclopropane-1-carboxylate deaminase promotes nodulation of pea plants. Appl Environ Microbiol 69(8):4396–4402
Madhaiyan M, Poonguzhali S, Ryu J, Sa T (2006) Regulation of ethylene levels in canola (Brassica campestris) by 1-aminocyclopropane-1-carboxylate deaminase-containing Methylobacterium fujisawaense. Planta 224:268–278
Mahanty T, Bhattacharjee S, Madhurankhi Goswami, Bhattacharyya P, Bannhi Das, Ghosh A, Tribedi P (2016) Biofertilizers: a potential approach for sustainable agriculture development. Environ Sci Pollut Res 24(4):3315–3335
Majumdar K, Razdan M, Aggarwal N, Murali KK, Bhattacharya RC, Dureja P (2011) Isolation and characterization of a potential biocontrol agent Bacillus KM5 from rhizosphere soil of a rice plant. Arch Phytopathol PFL 44(12):1196–1212
Maksimov IV, Abizgil’dina RR, Pusenkova LI (2011) Plant growth promoting rhizobacteria as alternative to chemical crop protectors from pathogens (Review). Appl Biochem Microbiol 47:333–345
Mali GV, Bodhankar MG (2009) Antifungal and phytohormone production potential of Azotobacterchroococcum isolates from Groundnut (Arachis hypogea L.) rhizosphere. Asian J Exp Sci 23:293–297
Malyan SK, Bhatia A, Kumar A, Gupta DK, Singh R, Kumar SK, Tomer R, Kumar O, Jain N, (2016a) Methane production, oxidation and mitigation: a mechanistic understanding and comprehensive evaluation of influencing factors. Science of The Total Environment 572:874–896
Malyan SK, Kumar A, Baram S, Kumar J, Singh S, Kumar SS, Yadav AN (2019) Role of Fungi in Climate Change Abatement Through Carbon Sequestration. In: Yadav A., Singh S., Mishra S., Gupta A. (eds) Recent Advancement in White Biotechnology Through Fungi. Fungal Biology. Springer, Cham
Malyan SK, Kumar SS, Kumar A, Kumar J (2016b) Water management tool in rice to combat two major environmental issues: global warming and water scarcity. In: Kumar S, Beg MA (eds) Environmental concerns of 21st century: Indian and global context, pp 43–58. (ISBN: 978-93-83281-65-7)
Marques APGC, Pires C, Moreira H, Rangel AOSS, Castro PML (2010) Assessment of the plant growth promotion abilities using Zea mays as indicator plant. Soil Biol Biochem 42:1229–1235
Mehnaz S, Baig DN, Lazarovits G (2010) Genetic and phenotypic diversity of plant growth promoting rhizobacteria isolated from sugarcane plants growing in Pakistan. J Microbiol Biotechnol 20:1614–1623
Mehnaz S, Lazarovits G (2006) Inoculation effects of Pseudomonas putida, Gluconabacter azotocaptans and Azospirilum lipoferum on corn plant growth under greenhouse conditions. Microbial Ecol 51:326–335
Mehnaz S, Mirza MS, Haurat J, Bally R, Normand P, Bano A, Malik KA (2001) Isolation and 16S rRNA sequence analysis of the beneficial bacteria from the rhizosphere of rice. Can J Microbiol 47(2):110–117
Mehnaz S, Weselowski B, Lazarovits G (2007) Sphingobacterium canadense sp. nov., an isolate from corn roots. Syst Appl Microbiol 30:519–524
Mendes LW, Kuramae EE, Navarrete AA, Van Veen JA, Tsai SM (2014) Taxonomical and functional microbial community selection in soybean rhizosphere. ISME J 8(8):1577
Mendes R, Garbeva P, Raaijmakers JM (2013) The rhizosphere microbiome: significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol Rev 37(5):634–663
Mendes R, Kruijt M, De Bruijn I, Dekkers E, van der Voort M, Schneider JH, Piceno YM, DeSantis TZ, Andersen GL, Bakker PA, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Minerdi D, Bossi S, Maffei M, Gullino M, Garibaldi A (2011) Fusarium oxysporum and its bacterial consortium promote lettuce growth and expansin A5 gene expression through microbial volatile organic compounds (MVOC) emission. FEMS Microbiol Ecol 76:342–351
Mirza MS, Ahmad W, Latif F, Haurat J, Bally R, Normand P, Malik KA (2001) Isolation, partial characterization, and the effect of plant growth-promoting bacteria (PGPB) on micro-propagated sugarcane in vitro. Plant and Soil 237(1):47–54
Mishra S., Goyal D., Kumar A., Dantu P.K. (2019) Biotechnological Applications of β-Glucosidases in Biomass Degradation. In: Yadav A., Singh S., Mishra S., Gupta A. (eds) Recent Advancement in White Biotechnology Through Fungi. Fungal Biology. Springer, Cham
Mishra SK, Khan MH, Misra S, Dixit KV, Khare P, Srivastava S, Chauhan PS (2017) Characterisation of Pseudomonas spp. and Ochrobactrum sp. isolated from volcanic soil. Antonie Van Leeuwenhoek 110:253–270
Mohammadi K (2010) Ecophysiological response ofcanola (Brassica napus L.) to different fertility systems incrop rotation. PhD thesis. Agronomy Department. Tarbiat Modares University, Tehran, Iran, p 354
Mohammadi K, Ghalavand A, Aghaalikhani M, Heidari GR, Sohrabi Y (2011) Introducing the sustainable soilfertility system for chickpea (Cicer arietinum L.). Afr J Biotechnol 10(32):6011–6020
Moronta-Barrios F, Gionechetti F, Pallavicini A, Marys E, Venturi V (2018) Bacterial microbiota of rice roots: 16S-based taxonomic profiling of endophytic and rhizospheric diversity, endophytes isolation and simplified endophytic community. Microorgani 6(1):14
Mukherjee J, Mridha N, Mondal S, Chakraborty D, Kumar A (2018) Identifying suitable soil health indicators under variable climate scenarios: a ready reckoner for soil management. In: Bal S, Mukherjee J, Choudhury B, Dhawan A (eds) Advances in crop environment interaction. Springer, Singapore
Naz I, Bano A, Ul-Hassan T (2009) Isolation of phytohormones producing plant growth promoting rhizobacteria from weeds growing in Khewra salt range, Pakistan and their implication in providing salt tolerance to Glycine max L. Afr J Biotechnol 8:5762–5766
Neeraja C, Anil K, Purushotham P, Suma K, Sarma P et al (2010) Biotechnological approaches to develop bacterial chitinases as a bioshield against fungal diseases. Crit Rev Biotechnol 30:231–241
Nielsen TH, Christophersen C, Anthoni U, Sorensen J (1999) Viscosinamide, a new cyclic depsipeptide with surfactant and antifungal properties produced by Pseudomonas fluorescensDR54. J Appl Microbiol 87:80–90
Nowak-Thompson B, Gould SJ, Kraus J, Loper JE (1994) Production of 2,4-diacetylphloroglucinol by the biocontrol agent Pseudomonas fluorescens Pf-5. Can J Microbiol 40:1064–1066
Oliveira CA, Alves VMC, Marriel IE, Gomes EA, Scotti MR, Carneiro NP, Guimaraes CT, Schaffert RE, So NMH (2009) Phosphate solubilizing microorganisms isolated from rhizosphere of maize cultivated in an oxisol of the Brazilian Cerrado Biome. Soil Biol Biochem 41:1782–1787
Palumbo JD, Yuen GY, Jochum CC, Tatum K, Kobayashi DY (2005) Mutagenesis of beta-1,3-glucanase genes in Lysobacter enzymogenes strain C3 results in reduced biological control activity toward bipolaris leaf spot of tall fescue and Pythium damping-off of sugar beet. Phytopathol 95:701–707
Park Y, Dutta S, Ann M, Raaijmakers J, Park K (2015) Promotion of plant growth by Pseudomonas fluorescens strain SS101via novel volatile organic compounds. Biochem Biophys Res Commun 461:361–365
Pathak H, Jain N, Bhatia A, Kumar A, Chatterjee D (2016) Improved nitrogen management: a key to climate change adaptation and mitigation. Indian J Fertil 12(11):151–162
Patil PL, Medhane NS (1974) Seed inoculation studies in gram (Cicer arietinum) with different strains of Rhizobium sp. Plant Soil 40:221–223
Paul D, Park KS (2013) Identification of volatiles produced by Cladosporium cladosporioides CL-1, a fungal biocontrol agent that promotes plant growth. Sensors (Basel) 13:13969–13977
Pérez-Jaramillo JE, Carrión VJ, Bosse M, Ferrão LF, de Hollander M, Garcia AA, Ramírez CA, Mendes R, Raaijmakers JM (2017) Linking rhizospheremicrobiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J 11(10):2244
Perrig D, Boiero ML, Masciarelli OA, Penna C, Ruiz OA, Cassan FD, Luna MV (2007) Plant-growth promoting compounds produced by two agronomically important strains of Azospirillum brasilense, and implications forinoculant formulation. Appl Microbiol Biotechnol 75:1143–1150
Pierson LS III, Pierson EA (1996) Phenazine antibiotic production in Pseudomonas aureofaciens: role in rhizosphere ecology and pathogen suppression. FEMS Microbiol Lett 136:101–108
Prasanna L, Eijsink VGH, Meadow R, Gåseidnes S (2013) A novel strain of Brevibacillus laterosporus produces chitinases that contribute to its biocontrol potential. Appl Microbiol Biot 97(4):1601–1611
Purkayastha GD, Mangar P, Saha A, Saha D (2018) Evaluation of the biocontrol efficacy of a Serratia marcescens strain indigenous to tea rhizosphere for the management of root rot disease in tea. PLoS ONE 13(2):0191761
Raaijmakers JM, Paulitz TC, Steinberg C, Alabouvette C, Moenne-Loccoz Y (2009) The rhizosphere: a playground and battlefield for soil borne pathogens and beneficial microorganisms. Pl Soil 321:341–361
Rajkumar M, Ae N, Prasad MNV, Freitas H (2010) Potential of siderophore-producing bacteria for improving heavy metal phytoextraction. Trends Biotechnol 28:142–149
Ramachandran VK, East AK, Karunakaran R, Downie JA, Poole SP (2011) Adaptationof Rhizobium leguminosarum to pea, alfalfa and sugar beet rhizosphere investigated by comparative transcriptomics. Genome Biol 12:106–109
Rana KL, Kour D, Sheikh I, Dhiman A, Yadav N, Yadav AN, Rastegari AA, Singh K, Saxena AK (2019a) Endophytic fungi: biodiversity, ecological significance and potential industrial applications. In: Yadav AN, Mishra S, Singh S, Gupta A (eds) Recent Advancement in White Biotechnology through Fungi, vol 1. Diversity and Enzymes Perspectives. Springer, Switzerland, pp 1–62
Rana KL, Kour D, Sheikh I, Yadav N, Yadav AN, Kumar V, Singh BP, Dhaliwal HS, Saxena AK (2019b) Biodiversity of endophytic fungi from diverse niches and their biotechnological applications. In: Singh BP (ed) Advances in endophytic fungal research: present status and future challenges. Springer International Publishing, Cham, pp 105–144. https://doi.org/10.1007/978-3-030-03589-1_6
Rana KL, Kour D, Yadav AN (2018) Endophytic microbiomes: biodiversity, ecological significance and biotechnological applications. Res J Biotechnol 14:1–30
Rashid MH, Schafer H, Gonzalez J, Wink M (2012) Genetic diversity of rhizobia nodulating lentil (Lens culinaris) in Bangladesh. Syst Appl Microbiol 35:98–109
Rastegari AA, Yadav AN, Gupta A (2019) Prospects of renewable bioprocessing in future energy systems. Springer International Publishing, Cham
Raza A, Faisal M (2013) Growth promotion of maize by desiccation tolerant Micrococcus luteus-chp37 isolated from Cholistan desert, Pakistan. Aust J Crop Sci 7:1693–1698
Reddy KRN, Choudary KA, Reddy MS (2007) Antifungal metabolites of Pseudomonas fluorescens isolated from rhizosphere of rice crop. J Mycol Plant Pathol 37(2):280–284
Revillas JJ, Rodelas B, Pozo C, Martinez-Toledo MV, Gonzalez LJ (2000) Productionof B-Group vitamins by two Azotobacter strains with phenolic compoundsas sole carbon source under diazotrophic and adiazotrophic conditions. J Appl Microbiol 89:486–493
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Camargo FAO, Farmerie WG, Triplett EW (2007a) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Camargo FA, Farmerie WG, Triplett EW (2007b) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1:283–290
Ryu C, Farag M, Hu C, Reddy M, Wei H, Paré PW, Kloepper JW (2003) Bacterial volatiles promote growth in Arabidopsis. Proc Natl Acad Sci 100:4927–4932
Sandilya SP, Bhuyan PM, Gogoi DK, Kardong D (2016) Phosphorus solubilization and plant growth promotion ability of rhizobacteria of R. communis L. growing in Assam, India. Proc Natl Acad Sci India Sect B Biol Sci 88(3):959–966
Sandilya SP, Bhuyan PM, Vijay N, Gogoi DK, Kardong D (2017) Impact of Pseudomonas aeruginosa strain MAJ PIA03 affecting the growth and phytonutrient production of castor, a primary host-plant of Samia ricini. J Soil Sci Plant Nutr 17(2):499–515
Saleh SS, Glick BR (2001) Involvement of gacS and rpoS in enhancement of the plant growth-promoting capabilities of Enterobacter cloacae CAL2 and UW4. Can J Microbiol 47(8):698–705
Saraf M, Pandya U, Thakkar A (2014) Role of allelochemicals in plant growth promoting rhizobacteria for biocontrol of phytopathogens. Microbiol Res 169:18–29
Selim S, Negrel J, Govaerts C, Gianinazzi S, Van Tuinen D (2005) Isolation and partial characterization of antagonistic peptides produced by Paenibacillus sp. strain B2 isolated from the sorghum mycorrhizosphere. Appl Environ Microbiol 71(11):6501–6507
Shafi J, Tian H, Ji M (2017) Bacillus species as versatile weapons for plant pathogens: a review. Biotechnol Biotechnol Equipment. https://doi.org/10.1080/13102818.2017.1286950
Shaharoona B, Arshad M, Zahir ZA (2006) Effect of plant growth promoting rhizobacteria containing ACC-deaminase on maize (Zea mays L.) growth under axenic conditions and on nodulation in mung bean (Vigna radiata L.). Lett Appl Microbiol 42(2):155–159
Shanmugam PM, Veeraputhran R (2000) Effect oforganic manure, biofertilizers, inorganic nitrogen and zincon growth and yield of rabi rice. Madras Agric J 2:87–90
Sharma A, Kumar A, Dhaka TS (2012a) Impact on sugar factory effluent on chlorophyll and protein contents of Cicer arietinum and Tigonella foenum-gracecum. Curr Adv Agri Sci 4(1):62–63
Sharma A, Kumar A, Dhaka TS (2012b) Impact of sugar factory effluent on seed germination, seedling growth of Cicer arietinum and Trigonella foenum-graecum. Bioinfolet 9(2):220–221
Sharma P, Sardana V, Kandola SS (2011) Response of groundnut (Arachis hypogaea L.) to Rhizobium Inoculation. Libyan Agric Res Centre J Int 2:101–104
Sheng XF, Xia JJ (2006) Improvement of rape (Brassica napus) plant growth and cadmium uptake by cadmium-resistant bacteria. Chemosphere 64:1036–1042
Shih HD, Liu YC, Hsu FL, Mulabagal V, Dodda R, Huang JW (2003) Fungichromin: a substance from Streptomyces padanus with inhibitory effects on Rhizoctoniasolani. J Agric Food Chem 51(1):95–99
Shridhar BS (2012) Review: nitrogen fixing microorganisms. Int J Microbiol Res 3(1):46–52
Singh PP, Shin YC, Park CS, Chung YR (1999) Biological control of Fusarium wilt of cucumber by chitinolytic bacteria. Phytopathology 89:92–99
Singh R, Pandey DK, Kumar A et al (2017) PGPR isolates from the rhizosphere of vegetable crop Momordica charantia: characterization and application as biofertilizer. Int J Curr Microbiol App Sci 6(3):1789–1802
Singh RP, Jha PN (2016) A halotolerant bacterium Bacillus licheniformis HSW-16 augments induced systemic tolerance to salt stress in wheat plant (Triticumaestivum). Front Plant Sci 7:1890
Sirohi MH, Jackson J, Edwards M, Ollerton J (2015) Diversity and abundance of solitary and primitively eusocial bees in an urban centre: a case study from Northampton (England). J Insect Consev 123–136
Sneha S, Anitha B, Sahair RA, Raghu N, Gopenath TS, Chandrashekrappa GK, Basalingappa KM (2018) Biofertilizer forcrop production and soil fertility. Acad J Agric Res 6(8):299–306
Sofia IA, Paula P, Castro ML (2014) Phosphate solubilizing rhizobacteria enhance Zea mays growth in agricultural P-deficient soils. Ecol Eng 73:526–535
Spaepen S, Vanderleyden J (2011) Auxin and plant-microbe interactions cold spring harbor. Perspect Biol 3(4):a001438
Stutz EG, Défago G, Kern H (1986) Naturally occurring fluorescent pseudomonads involved in suppression of black root rot of tobacco. Phytopathol 76:181–185
Subrahmanyam G, Archana G (2011) Plant growth promoting activity of Enterobacter sp. C1D in heavy metal contaminated soils. In: Plant growth-promoting rhizobacteria (PGPR) for sustainable agriculture, pp 440
Subrahmanyam G, Sharma RK, Kumar GN, Archana G (2018). Vigna radiata var. GM4 plant growth enhancement and root colonization by a multi-metal-resistant plant growth-promoting bacterium Enterobacter sp. C1D in Cr (VI)-amended soils. Pedosphere 28(1):144–156
Suman A, Yadav AN, Verma P (2016) Endophytic microbes in crops: diversity and beneficial impact for sustainable agriculture. In: Singh D, Abhilash P, Prabha R (eds) Microbial inoculants in sustainable agricultural productivity, research perspectives. Springer, India, pp 117–143. https://doi.org/10.1007/978-81-322-2647-5_7
Tian BY, Cao Y, Zhang KQ (2015) Metagenomic insights into communities, functions of endophytes, and their associates with infection by root-knot nematode, Meloidogyne incognita, in tomato roots. Sci Rep 5:17087
Timmusk S, Nicander B, Granhall U, Tillberg E (1999) Cytokinin production by Paenibacillus polymyxa. Soil Biol Biochem 31(13):1847–1852
Vaishnav A, Kumari S, Jain S, Varma A, Tuteja N, Choudhary DK (2016) PGPR mediated expression of salt tolerance gene in soybean through volatiles under sodium nitroprusside. J Basic Microbiol 56:1274–1288
Van Loon LC (2007) Plant responses to plant growth-promoting rhizobacteria. Eur J Plant Pathol 119:243–254
Venkataraman GS, Neelakantan S (1967) Effect of cellular constituents of the nitrogen fixing blue-green algae. Cylindrospermum musciola on the rootgrowth of rice seedlings. J General Appl Microbiol 13:53–61
Verma P, Yadav AN, Khannam KS, Kumar S, Saxena AK, Suman A (2016a) Molecular diversity and multifarious plant growth promoting attributes of Bacilli associated with wheat (Triticum aestivum L.) rhizosphere from six diverse agro-ecological zones of India. J Basic Microbiol 56:44–58
Verma P, Yadav AN, Khannam KS, Mishra S, Kumar S, Saxena AK, Suman A (2016b) Appraisal of diversity and functional attributes of thermotolerant wheat associated bacteria from the peninsular zone of India. Saudi J Biol Sci. https://doi.org/10.1016/j.sjbs.2016.01.042
Verma P, Yadav AN, Khannam KS, Panjiar N, Kumar S, Saxena AK, Suman A (2015a) Assessment of genetic diversity and plant growth promoting attributes of psychrotolerant bacteria allied with wheat (Triticum aestivum) from the northern hills zone of India. Ann Microbiol 65:1885–1899
Verma P, Yadav AN, Kumar V, Singh DP, Saxena AK (2017) Beneficial plant-microbes interactions: biodiversity of microbes from diverse extreme environments and its impact for crop improvement. In: Singh DP, Singh HB, Prabha R (eds) Plant-microbe interactions in agro-ecological perspectives: volume 2: microbial interactions and agro-ecological impacts. Springer Singapore, Singapore, pp 543–580. https://doi.org/10.1007/978-981-10-6593-4_22
Verma P, Yadav AN, Shukla L, Saxena AK, Suman A (2015b) Alleviation of cold stress in wheat seedlings by Bacillus amyloliquefaciens IARI-HHS2-30, an endophytic psychrotolerant K-solubilizing bacterium from NW Indian Himalayas. Natl J Life Sci 12:105–110
Voisard C, Bull CT, Keel C, Laville J, Maurhofer M, Schnider U, Défago G, Haas D (1994) Biocontrol of root diseases by Pseudomonas fluorescens CHA0: current concepts and experimental approaches. In: O’Gara F, Dowling DN, Boesten B (eds) Molecular ecology of rhizosphere microorganisms. VCH, Weinheim, Germany, pp 67–89
Wagg C, Bender SF, Widmer F, van der Heijden MGA (2014) Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci 111:5266–5270
Walia A, Mehta P, Chauhan A, Shirkot CK (2013) Antagonistic activity of plant growth promoting rhizobacteria isolated from tomato rhizosphere against soil borne fungal plant pathogens. Inte J Agri Environ Biotechnol 6(4):571–580
Wani PA, Khan MS (2010) Bacillus species enhance growth parameters of chickpea (Cicer arietinum L.) in chromium stressed soils. Food Chem Toxicol 48:3262–3267
Wani SA, Chand S, Ali T (2013) Potential use of Azotobacter chroococcum incrop production: an overview. Curr Agric Res J 1:35–38
Waqas M, Khan AL, Kamran M, Hamayun M, Kang SM, Kim YH, Lee IJ (2012) Endophytic fungi produce gibberellins and indoleacetic acid and promotes host-plant growth during stress. Mole 17:10754–10773
Weller DM (2007) Pseudomonas biocontrol agents of soilborne pathogens: looking back over 30 years. Phytopathology 97(2):250–256
Widawati S (2011) Diversity and phosphate solubilization by bacteria isolated from laki island coastal ecosystem. Biodiversitas 12(1):17–21
Xie X, Zhang H, Paré P (2009) Sustained growth promotion in Arabidopsis with longterm exposure to the beneficial soil bacterium Bacillus subtilis (GB03). Plant Signal Behav 4:948–953
Xu J, Zhang Y, Zhang P, Trivedi P, Riera N, Wang Y Liu X, Fan G, Tang J, Coletta-Filho HD, Cubero J (2018) The structure and function of the global citrus rhizosphere microbiome. Nat Commun 9(1):4894
Xue L, Xue Q, Chen Q, Lin C, Shen G, Zhao J (2013) Isolation and evaluation of rhizosphere actinomycetes with potential application for biocontrol of Verticillium wilt of cotton. Crop Prot 43:231–240
Yadav AN (2017a) Agriculturally important microbiomes: biodiversity and multifarious PGP Attributes for Amelioration of Diverse Abiotic Stresses in Crops for Sustainable Agriculture. Biomed J Sci Tech Res 1:1–4
Yadav AN (2017b) Beneficial role of extremophilic microbes for plant health and soil fertility. J Agric Sci 1:1–4
Yadav AN (2019) Microbiomes of wheat (Triticum aestivum L.) endowed with multifunctional plant growth promoting attributes. EC Microbiol 15:1–6
Yadav AN, Gulati S, Sharma D, Singh RN, Rajawat MVS, Kumar R, Dey R, Pal KK, Kaushik R, Saxena AK (2019a) Seasonal variations in culturable archaea and their plant growth promoting attributes to predict their role in establishment of vegetation in Rann of Kutch. Biologia 74:1031–1043. https://doi.org/10.2478/s11756-019-00259-2
Yadav AN, Kour D, Sharma S, Sachan SG, Singh B, Chauhan VS, Sayyed RZ, Kaushik R, Saxena AK (2019b) Psychrotrophic microbes: biodiversity, mechanisms of adaptation, and biotechnological implications in alleviation of cold stress in plants. In: Sayyed RZ, Arora NK, Reddy MS (eds) Plant growth promoting rhizobacteria for sustainable stress management: volume 1: rhizobacteria in abiotic stress management. Springer Singapore, Singapore, pp 219–253. https://doi.org/10.1007/978-981-13-6536-2_12
Yadav AN, Kumar R, Kumar S, Kumar V, Sugitha T, Singh B, Chauhan VS, Dhaliwal HS, Saxena AK (2017a) Beneficial microbiomes: Biodiversity and potential biotechnological applications for sustainable agriculture and human health. J Appl Biol Biotechnol 5:1–13
Yadav AN, Kumar V, Prasad R, Saxena AK, Dhaliwal HS (2018a) Microbiome in crops: diversity, distribution and potential role in crops improvements. In: Prasad R, Gill SS, Tuteja N (eds) Crop improvement through microbial biotechnology. Elsevier, USA, pp 305–332
Yadav AN, Mishra S, Singh S, Gupta A (2019c) Recent advancement in white biotechnology through fungi: volume 1: diversity and enzymes perspectives. Springer International Publishing, Cham
Yadav AN, Sachan SG, Verma P, Kaushik R, Saxena AK (2016a) Cold active hydrolytic enzymes production by psychrotrophic Bacilli isolated from three sub-glacial lakes of NW Indian Himalayas. J Basic Microbiol 56:294–307
Yadav AN, Sachan SG, Verma P, Saxena AK (2016b) Bioprospecting of plant growth promoting psychrotrophic Bacilli from cold desert of north western Indian Himalayas. Indian J Exp Biol 54:142–150
Yadav AN, Singh S, Mishra S, Gupta A (2019d) Recent advancement in white biotechnology through fungi: volume 2: perspective for value-added products and environments. Springer International Publishing, Cham
Yadav AN, Singh S, Mishra S, Gupta A (2019e) Recent advancement in white biotechnology through fungi: volume 3: perspective for sustainable environments. Springer International Publishing, Cham
Yadav AN, Verma P, Kaushik R, Dhaliwal HS, Saxena AK (2017b) Archaea endowed with plant growth promoting attributes. EC Microbiol 8:294–298
Yadav AN, Verma P, Kour D, Rana KL, Kumar V, Singh B, Chauahan VS, Sugitha T, Saxena AK, Dhaliwal HS (2017c) Plant microbiomes and its beneficial multifunctional plant growth promoting attributes. Int J Environ Sci Nat Resour 3:1–8 https://doi.org/10.19080/ijesnr.2017.03.555601
Yadav AN, Verma P, Kumar M, Pal KK, Dey R, Gupta A, Padaria JC, Gujar GT, Kumar S, Suman A, Prasanna R, Saxena AK (2015) Diversity and phylogenetic profiling of niche-specific Bacilli from extreme environments of India. Ann Microbiol 65:611–629
Yadav AN, Verma P, Kumar S, Kumar V, Kumar M, Singh BP, Saxena AK, Dhaliwal HS (2018b) Actinobacteria from rhizosphere: molecular diversity, distributions and potential biotechnological applications. In: Singh B, Gupta V, Passari A (eds) New and future developments in microbial biotechnology and bioengineering. USA, pp 13–41. https://doi.org/10.1016/b978-0-444-63994-3.00002-3
Yadav AN, Verma P, Kumar S, Kumar V, Kumar M, Sugitha TCK, Singh BP, Saxena AK, Dhaliwal HS (2018c) Actinobacteria from rhizosphere: molecular diversity, distributions, and potential biotechnological applications. In: New and future developments in microbial biotechnology and bioengineering, pp. 13–41
Yadav AN, Verma P, Singh B, Chauhan VS, Suman A, Saxena AK (2017c) Plant growth promoting bacteria: biodiversity and multifunctional attributes for sustainable agriculture. Adv Biotechnol Microbiol 5:1–16
Yadav AN, Yadav N (2018) Stress-adaptive microbes for plant growth promotion and alleviation of drought stress in plants. Acta Sci Agr 2:85–88
Yadav AN, Yadav N, Kour D, Kumar A, Yadav K, Kumar A, Rastegari AA, Sachan SG, Singh B, Chauhan V, Saxena AK (2019). Bacterial community composition in lakes. In: Freshwater microbiology. Academic Press, pp 1–71
Yadav AN, Yadav N, Sachan SG, Saxena AK (2019f) Biodiversity of psychrotrophic microbes and their biotechnological applications. J Appl Biol Biotechnol 7:99-108
Yi HS, Ahn YR, Song GC, Ghim SY, Lee S, Lee G, Ryu CM (2016) Impact of a bacterial volatile 2, 3-butanediol on Bacillus subtilis rhizosphere robustness. Front Microbiol 7:993
Zeller SL, Brand H, Schmid B (2007) Host-plant selectivity of rhizobacteria in a crop/weed model system. PLoS ONE 2(9):846
Zhang H, Sun Y, Xie X, Kim M, Dowd S, Paré P (2009) A soil bacterium regulates plant acquisition of iron via deficiency-inducible mechanisms. Plant J 58:568–577
Zou C, Li Z, Yu D (2010) Bacillus megaterium strain XTBG34 promotes plant growth by producing 2-pentylfuran. J Microbiol 48:460–466
Acknowledgements
Authors are thankful to Director, CMER&TI, Central Silk Board, Lahdoigarh for valuable guidance and constant encouragement.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Subrahmanyam, G., Kumar, A., Sandilya, S.P., Chutia, M., Yadav, A.N. (2020). Diversity, Plant Growth Promoting Attributes, and Agricultural Applications of Rhizospheric Microbes. In: Yadav, A., Singh, J., Rastegari, A., Yadav, N. (eds) Plant Microbiomes for Sustainable Agriculture. Sustainable Development and Biodiversity, vol 25. Springer, Cham. https://doi.org/10.1007/978-3-030-38453-1_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-38453-1_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-38452-4
Online ISBN: 978-3-030-38453-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)