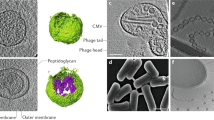
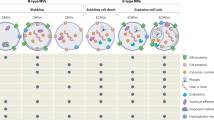
Abstract
Bacterial membrane vesicles represent a universal secretion mechanism enabling both Gram-negative and Gram-positive organisms to transfer cargo to eukaryotic cells, as well as to other bacterial cells. Bacterial vesicles can deliver to target cells an extremely wide range of virulence factors, including exotoxins, lipids, nucleic acids, and small molecules. Although there has been extensive research to decipher the mechanisms regulating cellular uptake of Gram-negative bacterial outer membrane vesicles (OMVs), much less is known about the cellular uptake of Gram-positive bacterial membrane vesicles (MVs). This chapter focuses on a selection of major bacterial pathogens and summarizes the present knowledge of OMV and MV-mediated virulence factor delivery, as well as mechanisms of bacterial vesicle–host cell interaction and uptake by mammalian cells.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
7.1 Bacterial Membrane Vesicle-Mediated Protein Delivery
Bacterial pathogenicity is enhanced by secretion systems that export virulence factors, either by secretion or injection, into the environment or adjacent host cells. Once delivered, these virulence factors then interfere with or stimulate host cellular processes. Eight bacterial secretion systems designated types I–VIII have been characterized to date (Green and Mecsas 2016). Both Gram-negative and Gram-positive bacteria of several different bacterial species release membrane vesicles to augment their pathogenic potential. Release of bacterial membrane vesicles, a very basic and relevant mode of protein transport, presumably also occurs during infection. Compared to other secretion mechanisms, membrane vesicle release has special implications since vesicles can deliver cargo, e.g., virulence factors, over much longer distances than secretory systems dependent upon direct bacterial contact. Effectively, these bacterial membrane vesicles become vehicles of multifunctional cargo, delivering a multitude of virulence factors, including metabolites, several protein toxins, nucleic acids, and immune modulators such as peptidoglycan (Berleman and Auer 2013).
7.1.1 Escherichia coli
Bacterial membrane vesicles were initially discovered as a product of Gram-negative bacterial outer membrane blebbing, and are therefore often referred to as outer membrane vesicles (OMVs). However, in 1976, Hoekstra et al. reported that membrane fragments, consisting of essentially unmodified outer membranes, were present in the culture supernatant of E. coli during normal growth (Hoekstra et al. 1976). Since then, a number of studies have described E. coli OMV biogenesis as well as the physiological cargo of these OMVs. When newly synthesized OMVs are released from E. coli, they contain active heat-labile enterotoxin (ELT) (Gankema et al. 1980; Wai et al. 2003). Because ELT is associated with lipopolysaccharide (LPS) on the OMV surface, host cell uptake of enterotoxigenic E. coli (ETEC) OMVs is allowed by binding to LT-receptor (GM1) (Horstman and Kuehn 2000, 2002; Kesty et al. 2004). The LeoA protein, a homolog of eukaryotic GTPase, secretes ELT from the periplasm of ETEC bacteria and it has been suggested that LeoA contributes to OMV formation and protein content (Brown and Hardwidge 2007).
Earlier studies demonstrating that a cytotoxic protein, cytolysin A (ClyA), in E. coli was exported through OMVs pointed to the potential physiological relevance of E. coli OMVs (Wai et al. 2003). In a process that involves redox-dependent oligomerization, ClyA is incorporated into OMVs, thus appearing to possess an intrinsic ability to translocate to the bacterial periplasm. ClyA incorporated into OMVs has considerably higher cytotoxicity toward mammalian cells compared to ClyA purified from the bacterial periplasm. Thus, protein localization in OMVs may play a direct role in activating and delivering virulence effector proteins (Wai et al. 2003).
Additional studies in E. coli examining vesicle-mediated export of bacterial virulence factors revealed the capacity of OMVs to deliver toxigenic cargo. For example, during infection, E. coli OMVs may represent an alternative pathway to deliver type I-secreted alpha-hemolysin from bacteria to host cells (Balsalobre et al. 2006). Another study reported that OMVs are a vehicle for bacteria to transfer cytotoxic necrotizing factor-1 (CNF1) to the environment and to infected tissue (Kouokam et al. 2006). Similarly, Enterohemorrhagic E. coli (EHEC) use OMVs to release hemolysin toxin (EHEC-Hly), a typical repeats-in-toxin protein (RTX) that lyses host cells through a mechanism of pore formation (Aldick et al. 2009). The toxin can exist as free EHEC-Hly and as EHEC-Hly associated with OMVs, and both forms are released during EHEC growth. Free EHEC-Hly is lytic toward human endothelial cells, whereas OMV-associated EHEC-Hly is not lytic toward microvascular endothelial cells (HBMEC) and the colon epithelial cell line Caco-2, although it can trigger apoptosis (Bielaszewska et al. 2013). Research into whether an MV-associated genotoxin from intestinal E. coli can promote cancer development revealed that E. coli-derived OMVs are readily internalized into target cells. Within these target cells, OMVs have the potential to induce oxidative stress, which would lead to DNA damage, replication, and aneuploidy in susceptible cells (Tyrer et al. 2014).
During hyper-biofilm formation, the kil gene, located in a three-gene cluster on the E. coli ColE1 plasmid, induces release of proteinous materials and aberrant OMVs into the extracellular environment (Nakao et al. 2018). A variety of pathogen-associated molecular pattern molecules are enriched in OMVs isolated from E. coli. These include LPS, lipoproteins, CpG DNA, flagellin, and peptidoglycan, most of which are Toll-like receptor (TLR) and nucleotide-binding and oligomerization domain (NOD) ligands (Ellis et al. 2010). Therefore, bacterial OMVs are capable of activating epithelial cells, endothelial cells, macrophages, and dendritic cells to release TNF-α, IL-1β, IL-6, and IL-8 (Bauman and Kuehn 2006; Bielaszewska et al. 2018; Canas et al. 2018; Lee et al. 2018). OMVs from pathogenic E. coli have also been known to cause sepsis-induced cardiac dysfunction, demonstrated both in vitro and in vivo (Svennerholm et al. 2017).
OMVs secreted by clinical isolates of EHEC O157 cause cell death by delivering into host cells a cocktail of virulence factors, such as Shiga toxin 2a (Stx2a), cytolethal distending toxin V (CdtV), EHEC hemolysin, and flagellin (Bielaszewska et al. 2017). Interestingly, OMVs from the nonpathogenic E. coli strain Nissle 1917 (EcN) can cause anti-inflammatory responses by reinforcing epithelial barrier integrity, thus affecting intestinal homeostasis (Alvarez et al. 2016; Behrouzi et al. 2018; Fabrega et al. 2017). In contrast, OMVs from the Nissle 1917 strain can also cause eukaryotic DNA double-stranded breaks (Canas et al. 2016). This strain harbors a cluster of genes that encode for proteins involved in the biosynthesis of hybrid non-ribosomal peptide-polyketide(s). It has been suggested that polyketides may be involved in inducing these eukaryotic DNA double-stranded breaks (Olier et al. 2012). In addition to host inflammatory responses, in colon cancer cells, OMVs from nonpathogenic commensal E. coli can induce epigenetic modifications (Vdovikova et al. 2018). Furthermore, OMVs from nonpathogenic E. coli can suppress the growth of established tumors as well as prevent tumor metastasis. These activities occur via an interferon-γ-mediated antitumor response, whereby OMVs deliver trypsin-sensitive surface proteins to the target cancer cells (Kim et al. 2017). Taken together, these studies demonstrate the enormous potential of bacterial OMVs from nonpathogenic E. coli as novel therapeutic agents against various cancers.
7.1.2 Vibrio cholerae
The formation of OMVs by Vibrio cholerae and Vibrio parahaemolyticus was first observed by researchers analyzing the cell structure of V. cholerae and V. parahaemolyticus using electron microscopy with freeze-substitution (Kondo et al. 1993). V. cholerae, the causal agent of the diarrheal disease cholera, possesses cholera toxin (CT) as its major virulence factor. In addition to CT, many other secreted protein toxins and enzymes that are important to V. cholerae pathogenesis have been reported to be associated with OMVs. These include Vibrio cytolysin (VCC) (Olivier et al. 2007), metalloprotease of Vibrio (PrtV) (Vaitkevicius et al. 2006), Zn-dependent hemagglutinin protease (HAP) (Ghosh et al. 2006; Hase and Finkelstein 1991), accessory cholera enterotoxin (Ace) (Kaper et al. 1995), and trypsin-like serine protease (VesC) (Syngkon et al. 2010).
OMVs from the V. cholerae strain O395 secrete biologically active CT (Chatterjee and Chaudhuri 2011). Using a GM1-independent mechanism, CT-containing OMVs are trafficked to host cells. This GMI-independent mechanism represents a secondary mechanism for CT secretion, in addition to the well-studied type II secretion system (Chatterjee and Chaudhuri 2011). Future studies to elucidate the functional details of this secondary mechanism of CT delivery are important to fully understand V. cholerae pathogenesis (Rasti et al. 2018).
Non-O1 and non-O139 V. cholerae (NOVC) serogroups are the causal agents of gastroenteritis and extraintestinal infections in humans; however, the virulence of NOVC strains is not well understood. OMVs from NOVC strains elicit NOD1- and NOD2-mediated immune responses in mammalian hosts. Quorum-sensing machinery attenuates OMVs’ inflammatory potential and thereby influences the immune responses (Bielig et al. 2011a, b).
Biologically active VCC, a pore-forming toxin, is released from the V. cholerae NOVC strain V:5/04 together with OMVs. OMV-associated VCC induces target cell autophagy, demonstrating that autophagy may play a role in cellular defense against an OMV-associated virulence factor (Elluri et al. 2014). The metalloprotease PrtV, a type II secretion system substrate protein, is also secreted from the V. cholerae strain C6706 together with OMVs (Rompikuntal et al. 2015). The biological activity of OMV-associated PrtV has been demonstrated in human colon carcinoma cells. Furthermore, the OMV-associated PrtV protease facilitates bacterial resistance toward the antimicrobial peptide LL-37 (Rompikuntal et al. 2015). HAP and VesC proteases are also released along with V. cholerae OMVs and the biologically active form of these proteases are delivered into human intestinal epithelial cells, causing cytotoxic and inflammatory responses (Mondal et al. 2016). Taken together, these studies demonstrate that the ability of V. cholerae to deliver virulence factors into host cells via OMV-mediated secretion is, therefore, a seemingly widespread feature among different Vibrio strains.
7.1.3 Pseudomonas aeruginosa
During normal growth, the Gram-negative bacterium Pseudomonas aeruginosa also releases OMVs. Pseudomonas OMVs carry and release several toxins and enzymes, including hemolysin, phospholipase C, alkaline phosphatase, protease, and elastase, which contribute to the organism’s pathogenicity (Kadurugamuwa and Beveridge 1995). Peptidoglycan hydrolases associated with OMVs that are naturally released by several Gram-negative bacterial strains, including Enterobacter, Citrobacter, Salmonella, Shigella, Escherichia, Klebsiella, Morganella, Pseudomonas, and Proteus, enable the lysis of both Gram-negative and Gram-positive bacteria (Kadurugamuwa and Beveridge 1998; Li et al. 1998). This predatory interaction indicates that within biofilms, where bacteria compete for growth with other bacteria in the surrounding microflora, OMVs might play a fitness role, providing an increased survival benefit (Kadurugamuwa and Beveridge 1997). This hypothesis is supported by the finding that P. aeruginosa OMVs play an important role in the formation of biofilms (Beveridge et al. 1997; Murphy et al. 2014). Cif, the cystic fibrosis (CF) transmembrane conductance regulator (CFTR) inhibitory factor, is associated with P. aeruginosa PA14 OMVs (MacEachran et al. 2007). The zinc-dependent leucine aminopeptidase PaAP, an enzyme involved in bacterial association with host cells, is also found in OMVs from two P. aeruginosa clinical strains, PAO1 and CF2 (Bauman and Kuehn 2006). The major outer membrane proteins (OMPs) that are associated with OMVs have been identified by mass spectrometry to be OprE, OprF, OprG, OprH, OprI, PcoB, and PagL (Bauman and Kuehn 2006; Choi et al. 2017a; Tashiro et al. 2010). Previously, it was thought that OMVs contain only outer membrane and periplasmic proteins; however, proteomic analysis of P. aeruginosa revealed the possible presence of cytoplasmic proteins in naturally released OMVs (Choi et al. 2017a). P. aeruginosa primarily occupies the mucus layer of the lung epithelium in cystic fibrosis patients. Released OMVs can deliver virulence factors into the cytoplasm of host cells, resulting in modified innate immune responses. A recent study investigated whether the antibiotic tobramycin, which is commonly used to treat CF patient lung infections caused by P. aeruginosa, affects the abundance of virulence factors in OMVs. The study demonstrated that in CF patients, tobramycin may improve lung function by decreasing the abundance of several key virulence factors in OMVs, which restores chloride ion secretion necessary for bacterial clearance from the lungs (Koeppen et al. 2019). It is thus likely that OMVs carrying virulence factors are important contributors to the in vivo survival and adaptability of P. aeruginosa in CF lung infection.
7.1.4 Acinetobacter baumannii
Acinetobacter baumannii is an opportunistic pathogen responsible for a wide range of nosocomial infections. A. baumannii secretes OMVs that contain phospholipases and exhibit both hemolytic and leukocytic activities against target host cells (Jha et al. 2017). In a lung infection mouse model, surface proteins of A. baumannii OMVs can induce pro-inflammatory immune responses (Jun et al. 2013). Release of the elongation factor Tu (EF-Tu) from bacterial cells associated with OMVs from A. baumannii may contribute to fibronectin-mediated binding on the host cell (Dallo et al. 2012). Vaccinating a sepsis mouse model with OMVs purified from A. baumannii triggers high levels of IgM, IgG1, and IgG2c immunoglobulins, while levels of the pro-inflammatory cytokines IL-1β and IL-6 remain low (McConnell et al. 2011). OMV vaccination protects mice against challenge with the A. baumannii ATCC 19606 strain (McConnell et al. 2011). Variation in OMV production is associated with a unique feature of A. baumanii, the reversible switching between formation of opaque and translucent colonies. In experiments assessing immune response in macrophages, OMVs from the A. baumannii opaque colony form appear to be more immunogenic than those from the translucent colony form (Ahmad et al. 2019). The rise in extensive antibiotic resistance to A. baumannii highlights the potential need for a vaccine against this organism (Li et al. 2006; Lei et al. 2019). Therefore, OMVs from A. baumannii represent a promising vaccine candidate due to its immunogenic properties.
7.1.5 Porphyromonas gingivalis
Porphyromonas gingivalis is a major pathogenic cause of adult periodontitis. The Gram-negative anaerobic bacterium P. gingivalis releases OMVs that contribute to pathogenesis due to their high proteolytic and hemagglutinating activities, as well as their ability to promote inter- and intra-bacterial species adherence (Olsen and Amano 2015). Multiple studies have shown that OMV-associated toxins and proteolytic enzymes have a major contribution to periodontal diseases (Bourgeau and Mayrand 1990; Duchesne et al. 1995; Ellen and Grove 1989; Kamaguchi et al. 2003; Patrick et al. 1996; Singh et al. 1989; Smalley et al. 1991). The specific OMV-associated virulence factor(s) involved in OMV-mediated pathogenesis of P. gingivalis are not well known despite the clinical importance of P. gingivalis. It is known that P. gingivalis produces gingipain proteinases that are preferentially packed into OMVs (Haurat et al. 2011; Veith et al. 2014). OMV-associated FimA, hemagglutinin A, and heat-stress protein (HtrA) are involved in the attachment of P. gingivalis to host cells and subsequent invasion (Belanger et al. 2012; Zhang et al. 2011). Also, major P. gingivalis outer membrane proteins are associated with OMVs, which are used to efficiently invade host cells (Ho et al. 2015; Mantri et al. 2015; Veith et al. 2014). Thus, it has been suggested that OMVs may be involved in the development of atherosclerosis and represent a “Trojan horse” strategy to cause an effect without employing intact bacterial cells (Xie 2015).
7.2 OMV-Mediated Virulence Factor Delivery by Other Gram-Negative Bacteria
OMVs from several different Gram-negative bacterial species effectively transport multifunctional cargo over long distances. For example, OMVs from Aggregatibacter actinomycetemcomitans carry proteins that function in antibiotic targeting, nutrient acquisition, and immune evasion, representing both offensive and defensive activities (Kieselbach et al. 2015). This phenomenon was described for the first time in Bordetella pertussis, whose OMVs carry a virulence complex that includes an adhesin (Imagawa et al. 1979). A causal organism of acute respiratory tract infection, B. pertussis harbors a wide range of virulence factors, including pertussis toxin, filamentous hemagglutinin, adenylate cyclase hemolysin, and tracheal cytotoxin, which are secreted in association with OMVs (Hozbor et al. 1999). OMVs carrying adenylate cyclase toxin can induce murine macrophage and CHO-K1 cell death independent of the toxin’s receptors (Donato et al. 2012). Immunization with B. pertussis OMVs may represent an effective next-generation pertussis vaccine strategy as evidenced by its ability to protect against bacterial colonization by eliciting antibody and Th1/Th17 type immune responses (Raeven et al. 2016).
One of the major virulence factors of Campylobacter jejuni and A. actinomycetemcommitans, cytolethal distending toxin (CDT), is secreted primarily from bacterial cells in association with OMVs, suggesting evolutionary conservation of this mode of CDT delivery (Berlanda Scorza et al. 2008; Lindmark et al. 2009; Rompikuntal et al. 2012). OMVs from C. jejuni carry three proteases, HtrA, Cj0511, and Cj1365c, and these OMV-associated proteases can cleave occludin and E-cadherin of T84 colon carcinoma cells (Elmi et al. 2016). In humans, C. jejuni can cause gastroenteritis, while in avian hosts, colonization is asymptomatic. The body temperature difference between human (37 °C) and avian (42 °C) hosts suggests that growth of C. jejuni at 37 °C potentially cues expression of bacterial virulence factors. Proteome analyses comparing OMVs from C. jejuni grown at 37 °C and at 42 °C revealed more virulence-related proteins associated with OMVs isolated from the bacteria grown at 37 °C (Taheri et al. 2018). The presence of bile in the growth medium also influences the selective packing of virulence factors in C. jejuni OMVs (Taheri et al. 2018), suggesting that the protein cargo of OMVs may also be regulated by the host environment.
The release and intracellular uptake of Bacteroides fragilis OMVs can activate caspase-11-dependent cell death and IL-1 responses to LPS (Vanaja et al. 2016). OMVs from B. fragilis carry polysaccharide A capsular antigen (PSA). PSA induces TLR2-mediated signaling in dendritic cells, which results in regulatory T cell maturation by production of the immunoregulatory cytokine IL-10 (Shen et al. 2012), thus implicating OMVs as an important mediator in establishing mutualism.
Bacteria employ various secretion systems to deliver virulence factors to target cells. The field of bacterial OMVs has become an exciting research area that is poised to improve our understanding of bacterial pathogenesis and provide alternative strategies to control infectious disease. Further investigation into the mechanisms and roles of OMV secretion systems may uncover novel targets and strategies for developing new antimicrobial therapies.
7.3 Delivery of Bacterial Nucleic Acids by OMVs
7.3.1 DNA
In 1989, Neisseria gonorrhoeae was shown to release RNA and DNA in association with OMVs (Dorward and Garon 1989). It was suggested that the RNA and linear DNA are associated with the exterior of the vesicles because nuclease treatment eliminated them from OMV preparations. However, circular DNA inside the OMVs, and thus resistant to nuclease treatment, was capable of transforming recipient cells. Hence, it was suggested that OMVs can act as a mechanism by which cells can exchange genetic information (Dorward et al. 1989). Since the first description in 1989, an increasing number of reports have described the OMV-associated release of plasmid DNA and/or chromosomal DNA (Biller et al. 2014; Lee et al. 2007; Perez-Cruz et al. 2015; Renelli et al. 2004; Yaron et al. 2000). DNA purified from E. coli O157:H7 OMVs contain the virulence genes stx1, stx2, eae, and uidA (Kolling and Matthews 1999). Among bacteria, OMVs can contribute to antibiotic resistance spread by two different mechanisms (Ciofu et al. 2000; Mashburn-Warren and Whiteley 2006). Spread of antibiotic resistance may occur by direct transfer of a resistance protein (e.g., β-lactamase) to neighboring cells or by lateral transfer of resistance genes following fusion of the OMV with the recipient cell membrane (Fulsundar et al. 2014; Renelli et al. 2004; Rumbo et al. 2011; Yaron et al. 2000). In the case of P. aeruginosa, it has been suggested that OMVs can also be formed after cell lysis, when membrane fragments and cytosolic contents including DNA are released from spontaneously lysed bacteria (Turnbull et al. 2016). DNA associated with OMVs contributes to establishing bacterial biofilms to facilitate bacterial host colonization (Liao et al. 2014). OMVs can deliver DNA into eukaryotic cells, suggesting a role for bacteria–host cell interactions and demonstrating potential for OMV-based DNA vaccines (Bitto et al. 2017). Interestingly, integration of bacterial DNA has been detected in the host genome, suggesting transfer of bacterial genetic material into human somatic cells (Riley et al. 2013). It remains to be determined, however, if OMV-associated DNA integrates into the host genome. Further studies are needed to investigate whether OMVs are capable of delivering DNA into the host nucleus as well as whether OMV-delivered DNA can integrate into the host genome or modulate the innate immune response via DNA sensors (Hornung 2014). Earlier studies indicated that among similar bacterial species, i.e., N. gonorrheae, it may be possible for genetic material to be transferred by OMVs (Dorward and Garon 1989). Moreover, OMVs can transfer carbapenem-resistance genes to surrounding A. baumannii bacterial isolates (Rumbo et al. 2011). Research on Gram-positive bacterial membrane vesicles (MVs) has shown that Clostridium perfringens releases MV-containing DNA and protein components that can be internalized by macrophages and induce dramatic pro-inflammatory cytokines both in vitro and in vivo (Jiang et al. 2014). In addition to providing a potential mechanism for exchange of genetic material between prokaryotes and eukaryotes, these studies reveal a new perspective on the immunogenic properties of bacterial OMV and MV-based DNA vaccines.
7.3.2 RNA
Bacterial membrane vesicles share similarities with exomes, cell vesicles secreted by most mammalian cell types. Bacterial membrane vesicles and exosomes both carry payloads of proteins, lipids, and genetic material enclosed in membrane-bound spherical structures of similar size ranges. Both bacterial membrane vesicles and exosomes can deliver functional molecules to distant extracellular compartments and tissues. Exosomes are involved in the horizontal transfer of genetic material, such as mRNAs and miRNAs, from the donor cells to recipient cells (Valadi et al. 2007; Zhong et al. 2011; Zomer et al. 2010). Investigation of whether bacterial RNA associates with OMVs by us and other researchers revealed that RNA is indeed encapsulated inside the OMVs in the form of RNase-resistant secondary structures and/or is associated with proteins in RNase-stable complexes (Blenkiron et al. 2016; Choi et al. 2017a, b; Resch et al. 2016; Sjostrom et al. 2015). Emerging evidence indicates that OMVs contain short RNAs (sRNAs) that are differentially packaged and have the potential to target the function and/or stability of host mRNA. Interestingly, via a regulatory OMV-associated sRNA, a new mechanism of pathogen–host interaction attenuates the innate immune response in human airway epithelial cells as well as in mouse lung. A specific bacterial sRNA (sRNA52320) is transferred from P. aeruginosa OMVs to host cells, where in human airway epithelial cells it attenuates OMV-stimulated IL-8 secretion, and in the lungs of a mouse model it attenuates keratinocyte-derived cytokine secretion and neutrophil recruitment (Koeppen et al. 2016).
Different classes of RNA are present in OMV-associated fractions of Salmonella enterica serovar Typhimurium and are exported. These include rRNAs, mRNAs, tRNAs, and other ncRNAs (Malabirade et al. 2018). However, RNA associated with OMVs is clearly different when the bacteria are grown under host-mimic cultural conditions in comparison with ordinary laboratory culture media. At least a fraction of the extracellular RNA associated with OMVs is present as full-length transcripts, indicating that OMVs can protect RNA and that this RNA might be functionally active (Malabirade et al. 2018). Export of full-length transcripts via OMVs opens the possibility of numerous functional implications for bacteria–bacteria and bacteria–host communication.
7.3.2.1 How Does RNA Associate with OMVs?
Several possibilities for RNA association with OMVs have been suggested (Blenkiron et al. 2016). First, extracellular RNA released by general bacterial cell lysis may be tightly reassociated to the OMV surface after secretion from bacterial cells. Second, RNA incorporation into OMVs could occur through an active and selective mechanism. Third, RNA association with OMVs may merely represent nonspecific envelopment of RNA in the cytoplasm within vesicle blebs. Finally, the phenomenon may be due to RNA riding as passengers on OMV-bound proteins, as bacterial mRNAs are frequently found at the sites of their future protein products (Nevo-Dinur et al. 2012). There are indeed many mRNAs that encode many membrane proteins present in OMVs. These include mRNAs for ompA, lpp, and tonB in OMVs from uropathogenic E. coli and mRNAs for ompU, ompA, and tolC in OMVs from V. cholerae (Blenkiron et al. 2016; Sjostrom et al. 2015). The ability of OMVs to deliver their associated RNA cargo into host cells poses the interesting question of whether these RNAs can function as novel signaling molecules in bacteria–host interactions.
7.4 Bacterial Lipid Release in Association with OMVs
OMVs contain bacterial phospholipids such as phosphatidylglycerol, phosphatidylethanolamine, and cardiolipin. Moreover, the phospholipid composition of OMVs generally resembles that of the outer membrane (OM) from which they are derived (Horstman and Kuehn 2000). Phospholipid and fatty acid compositions between OMVs and the cellular OM of E. coli do not differ significantly (Hoekstra et al. 1976). However, the phospholipid head groups and acyl chains compositions between the bacterial OM and OMVs from P. aeruginosa are quite different. Therefore, the OMV membrane is considered rigid compared to the cellular OM of P. aeruginosa. Thus, it has been suggested that the OMV blebbing mechanism may not be conserved among Gram-negative bacteria (Tashiro et al. 2011). OMVs contain, in addition to phospholipids, abundant LPS, which normally comprise the majority of the OM outer leaflet. P. aeruginosa can express both a common antigen (A-band) and serotype-specific antigen (B-band) in the O-antigen portion of LPS. Thus, these OMVs are highly enriched in B-band LPS, in contrast to the lipid composition of the OM (Kadurugamuwa and Beveridge 1995). Based on these differences, B-band LPS has been proposed to sort into OMVs, similar to the sorting of LPS and proteins seen in Porphorymonas gingivalis (Haurat et al. 2011).
7.5 Small Molecule Delivery Via OMVs
In addition to lipids, proteins, and nucleic acids, small molecules associate with OMVs as well. Pseudomonas putida strains that are resistant to toluene produce more OMVs upon exposure to toluene and release toluene-enriched OMVs as a detoxification system (Kobayashi et al. 2000). In P. aeruginosa, PQS (Pseudomonas quinolone signal), a quorum-sensing molecule, associates with OMVs (Mashburn and Whiteley 2005). Because PQS is more hydrophobic than the quorum-sensing signal acylhomoserine-lactone of P. aeruginosa, it is concentrated in the OMV membrane where it can interact specifically with LPS. It appears that such interactions contribute to physically stimulate the formation of vesicles into which the PQS is subsequently packaged (Mashburn and Whiteley 2005).
Gram-negative bacteria can employ OMVs to deliver peptidoglycan to cytosolic nucleotide-binding oligomerization domain-containing protein 1 (NOD1) in host cells. OMVs (containing peptidoglycan) purified from P. aeruginosa, H. pylori, and Neisseria gonorrheae can upregulate NF-κB and NOD1-dependent responses in vitro (Irving et al. 2014; Kaparakis et al. 2010). Moreover, when administered to mice intragastrically, H. pylori OMVs trigger NOD1-dependent but TLR-independent innate and adaptive immune responses (Kaparakis et al. 2010). In mammalian cells, V. cholerae OMVs induce NOD1- and NOD2-mediated immune responses. Quorum-sensing machinery attenuates the inflammatory potential of OMVs, playing an important role in regulating this process during infection (Bielig et al. 2011a, b). In human embryonic kidney cells, A. actinomycetemcomitans OMVs strongly induce NOD1- and NOD2-dependent NF-κB activation. Moreover, in myeloid THP1 cells, NOD1, the primary sensor of peptidoglycan delivered by MVs, contributes to the overall inflammatory responses induced by the vesicles (Thay et al. 2014).
OMVs from P. aeruginosa, Shigella flexneri, and Myxococcus xanthus contain molecules with bacteriolytic properties (Evans et al. 2012; Kadurugamuwa and Beveridge 1995, 1997, 1999). Thus, OMVs are critical to intra- and inter-species communication, although in bacterial cell–cell interactions occurring via OMVs, the selectivity of the interaction between MVs and bacterial cells is not fully understood. Recently, employing OMVs isolated from the Enterobacterium Buttiauxella agrestis, OMVs selectively interacted with target bacterial cells (Tashiro et al. 2017). These results offer a new avenue by which particular bacterial species can be controlled using bacterial OMVs in microbial communities.
7.6 Gram-Positive Bacteria Membrane Vesicles (MVs)
MVs are also released by Gram-positive bacteria such as Staphylococcus aureus (Lee et al. 2009), Enterococcus faecium (Wagner et al. 2018), Streptococcus pneumoniae (Codemo et al. 2018), Streptococcus pyogenes (Resch et al. 2016), Mycobacterium ulcerans (Marsollier et al. 2007), Bacillus anthracis (Rivera et al. 2010), Listeria monocytogenes (Vdovikova et al. 2017), and Lactobacillus (Dean et al. 2019) (see Chap. 3). These MVs are released both in vivo and in vitro as spherical, bilayered structures with a diameter of approximately 20–150 nm (Gurung et al. 2011; Rivera et al. 2010; Vdovikova et al. 2017).
7.6.1 Staphylococcal Species
7.6.1.1 Staphylococcus aureus
S. aureus is responsible for a wide spectrum of human infections that range from superficial cutaneous infections to life-threatening bacteremia (Lowy 1998). MVs from S. aureus have been isolated and analyzed by mass spectrometry. Proteins identified include the IgG-binding protein, ferritin, ferrichrome-binding lipoprotein precursor, ABC transporter extracellular binding protein, β-lactamase, and membrane protein OxaA (Gurung et al. 2011). S. aureus MVs deliver protein A to host cells by interacting with host cell plasma membranes through a cholesterol-rich microdomain in the membrane (Gurung et al. 2011; Rivera et al. 2010). S. aureus α-toxin (α-hemolysin), a 33-kDa pore-forming protein, is also associated with MVs. S. aureus α-toxin can lyse a wide range of human cells and induce apoptosis in T cells (Berube and Bubeck Wardenburg 2013). S. aureus MVs containing α-toxin are cytotoxic to HeLa cells and induce erythrocyte lysis (Thay et al. 2013). S. aureus MVs have also been reported to contain δ-hemolysin (Hld), γ-hemolysin, leukocidin D, exfoliative toxin C, and exfoliative toxin A, identified by proteomic analysis (Jeon et al. 2016). Comparative proteomics identified a total of 131 and 617 proteins in MVs from S. aureus grown in Luria-Bertani and brain-heart infusion broths, respectively, suggesting that culture media components can influence MV protein composition (Askarian et al. 2018). A study of the roles of MVs in bacteria–host interactions led to the suggestion that during systemic infection, S. aureus MVs can influence bacteria–host interactions and that they provide protective immunity in murine infection models (Askarian et al. 2018).
7.6.1.2 Staphylococcus haemolyticus
S. haemolyticus is a skin commensal microorganism. S. haemolyticus nosocomial isolates are the most antibiotic-resistant members of the coagulase-negative Staphylococci. However, little is known about S. haemolyticus virulence factors. Potential virulence proteins associated with MVs have been compared to the S. haemolyticus total secretome. This comparison revealed that the cargo carried by MVs is enriched in proteins involved in adhesion, acquisition of iron, and antimicrobial resistance (Cavanagh et al. 2018).
7.6.2 Streptococcal Species
7.6.2.1 Streptococcus pyogenes
Comprehensive studies have been performed on MVs produced by the Gram-positive human pathogen S. pyogenes, the etiological agent of necrotizing fasciitis and streptococcal toxic shock syndrome. These studies have provided an explanation for the MV-associated secretion of S. pyogenes macromolecules, including RNAs, lipids, and proteins, as well as described a two-component system that modulates S. pyogenes MV production (Resch et al. 2016; Biagini et al. 2015).
7.6.2.2 Streptococcus pneumoniae
S. pneumoniae, a major Gram-positive respiratory pathogen, produces MVs that may serve as a vehicle for many bacterial proteins. Pneumolysin, a cytosolic pore-forming toxin, is significantly enriched in MVs (Codemo et al. 2018). Pneumococcal MVs are internalized into A549 lung epithelial cells and human monocyte-derived dendritic cells, where they trigger pro-inflammatory cytokine responses independent of pneumolysin content. It has been suggested that S. pneumoniae MVs act in an immunomodulatory manner by enabling transfer of vesicle-associated proteins and other macromolecules into host cells. In addition, MVs bind tightly to serum complement system components, sequestering complement factor C3 in human serum and decreasing pneumococcal opsonophagocytosis (Codemo et al. 2018).
7.6.3 Mycobacterial Species
The etiologic agent of Buruli ulcers, the mycobacterium Mycobacterium ulcerans is slow-growing and infects the skin and subcutaneous tissues (George et al. 1999). Mycolactone, a poliketide-derived macrolide, is the only virulence factor known to be responsible for Buruli ulcers (George et al. 1999). MVs from M. ulcerans are cytotoxic to mouse macrophages because the vesicles contain mycolactone (Marsollier et al. 2007). MVs from M. bovis BCG and M. tuberculosis H37Rv are enriched in proteins associated with bacterial virulence, revealed by proteomic analysis. These proteins include a remarkable abundance of putative Toll-like receptor 2 (TLR2) ligands, such as 19 kDa Mycobacterium lipoproteins LpqH, LprA, and LprG. Interaction of MVs from either M. bovis BCG or M. tuberculosis H37Rv with murine macrophages induces TLR2-dependent cytokine and chemokine release. This evidence demonstrated that mycobacterial vesicles serve as a delivery mechanism for immunologically active molecules that contribute to the virulence of mycobacteria (Prados-Rosales et al. 2011). It was recently reported that the protein VirR (encoded by the gene rv0431) in M. tuberculosis (Mtb) regulates the amount of Mtb-derived MVs containing TLR2 ligands such as the lipoproteins LpqH and SodC, suggesting that VirR plays a role in immunomodulating properties of Mtb via MVs (Lee et al. 2013; Rath et al. 2013).
7.6.4 Enterococcus faecium
E. faecium is a commensal organism that is inherently resistant to several antimicrobial agents and can become a bacteremia-causing pathogen. Like other Gram-positive bacteria, E. faecium strains produce MVs (Gao et al. 2018). E. faecium MV-associated proteins include virulence factors, such as biofilm-promoting proteins, extracellular matrix-binding proteins, and antimicrobial resistance-related proteins, suggesting that E. faecium may utilize MVs to release proteins promoting virulence, pathogenicity, and antimicrobial resistance (Wagner et al. 2018).
7.6.5 Bacillus anthracis
B. anthracis, a spore-producing bacillus, causes anthrax in a range of vertebrates. B. anthracis releases vesicles that contain components of the anthrax toxins, the protective antigen (PA), lethal factor (LF), and edema toxin (ET), as well as anthrolysin (Rivera et al. 2010). Immunizing mice with B. anthracis MVs protects them against subsequent challenge with B. anthracis (Marsollier et al. 2007).
7.6.6 Listeria monocytogenes
L. monocytogenes, a Gram-positive pathogen, causes listeriosis, an illness transmitted through the consumption of contaminated food. Similar to other Gram-positive bacteria, MVs are released by L. monocytogenes in a process that is regulated by the general stress transcription factor σB (Lee et al. 2013). Internalin B (InIB), which is responsible for L. monocytogenes entry into target cells, and listeriolysin O (LLO), a pore-forming toxin, were identified in MVs from L. monocytogenes (Lee et al. 2013). In a detailed study, Vdovikova et al. demonstrated that L. monocytogenes produces MVs both in vitro and in vivo (Vdovikova et al. 2017). The pore-forming hemolysin LLO is a major virulence factor that is tightly associated with MVs in an oxidized, inactive form. Autophagy induced by pure LLO, by other bacterial pore-forming toxins or by Torin1-stimulated macroautophagy is effectively abrogated by MVs. Thus, it has been suggested that L. monocytogenes may survive inside host cells by controlling LLO activity and avoiding destruction from the autophagy system via intracellular release of MVs (Vdovikova et al. 2017).
7.6.7 Lactobacillus
MVs from three different Lactobacillus species have been characterized for their physiochemical properties and protein compositions. A recent study identified more than 80 protein components from Lactobacillus-derived MVs, including bacteriocin, which is enriched in MVs, suggesting that the vesicles serve as vehicles for delivery of the antimicrobial molecule (Dean et al. 2019). Collectively, these studies highlight the role of MVs in the pathogenesis of Gram-positive bacterial infections.
7.7 Entry and Trafficking of OMVs and MVs into Host Cells
Bacterial OMVs and MVs can enter host cells using various pathways, including clathrin- or caveolin-mediated pathways, or through fusion with plasma membranes (Bielaszewska et al. 2017; Mulcahy et al. 2014; Olofsson et al. 2014). Despite extensive research to understand the mechanisms that regulate cellular uptake of OMVs, little is known about the cellular uptake of MVs. Interestingly, Gram-negative (V. cholerae) and Gram-positive (L. monocytogenes) bacterial vesicles were efficiently internalized into the intracellular compartments of epithelial cells, which accumulated primarily in the lysosomal compartment of host epithelial cells (Fig. 7.1a and b). Importantly, a recent study showed that E. coli O157 (EHEC) OMVs are quickly internalized into intracellular compartments, where they deliver a cocktail of bacterial factors to different host cell compartments (Bielaszewska et al. 2017).
Lysosomal accumulation of bacterial OMVs and MVs. (a) Cellular uptake of PKH2-labeled OMVs (Red) isolated from V. cholerae or MVs from L. monocytogenes. Arrow head indicates their vesicular uptake into HCT8 cells. Nucleus is counter-stained with Hoechst 33342. Scale bars = 10 μm. (b) Co-localization of OMVs or MVs (red) with lysosomal marker, Lysotracker (green). Arrow head indicates co-localized spots, seen as yellow in the cytoplasm of HCT8 cells. Scale bars = 10 μm
Endocytosis allows small molecules to traverse a cells membrane bilayer (Doherty and McMahon 2009). Host cells internalize OMVs and MVs from several bacteria mainly via various endocytic pathways. As described in recent reviews, endocytosis involves cell membrane invagination, and occurs through several different pathways that depend on the composition and cargo of the OMVs to be internalized (Bitto and Kaparakis-Liaskos 2017; Kaparakis-Liaskos and Ferrero 2015; Pathirana and Kaparakis-Liaskos 2016). Three primary cellular mechanisms regulating the cellular uptake of OMVs and MVs are: (i) clathrin-mediated endocytosis, (ii) cholesterol-enriched microdomains, also known as caveolae or lipid rafts, and (iii) F-actin-coated vacuoles, also known as macropinocytosis and phagocytosis (Table 7.1). These pathways produce endosomal compartments that allow cargo transfer to various subcellular sites in the host cell cytoplasm (Doherty and McMahon 2009). Furthermore, the size of OMVs has been recently shown to play an important role in the preferred mode of entry into host cells (Turner et al. 2018).
OMVs from several microorganisms, H. pylori, A. actinomycetemcomitans, enterohemorrhagic E. coli (EHEC), enteroaggregative E. coli (EAEC), and Brucella abortus, use clathrin-mediated endocytosis as their major mode of entry into host cells (Bielaszewska et al. 2013; Canas et al. 2016; O’Donoghue and Krachler 2016; Olofsson et al. 2014; Pollak et al. 2012; Thay et al. 2014). In addition to clathrin-mediated endocytosis, it has been proposed that dynamin also plays an important role in host cell-mediated uptake of OMVs (Bielaszewska et al. 2017; Kunsmann et al. 2015).
Membrane microdomains, called lipid rafts, are dynamic and abundant in several types of lipids, including cholesterol and sphingolipids, and proteins such as caveolin. The importance of membrane cholesterol for delivery of OMV cargo to the intracellular compartment of host cells has been reported by several investigators. OMVs from V. cholerae (Mondal et al. 2016), V. vulnificus (Kim et al. 2010), A. actinomycetemcomitans (Rompikuntal et al. 2012), ETEC (Johnson et al. 2009), H. influenzae (Sharpe et al. 2011), P. gingivalis (Furuta et al. 2009), Moraxella catharralis (Schaar et al. 2011), H. pylori (Kaparakis et al. 2010; Olofsson et al. 2014), and C. jejuni (Elmi et al. 2012) rely on lipid rafts to mediate internalization by host cells via endocytosis. Bacteria shed OMVs and MVs of different sizes, with the larger OMVs being engulfed by the host cell through ruffled cell membrane protrusions driven by actin polymerization (Kaparakis-Liaskos and Ferrero 2015). Inhibiting actin polymerization using cytochalasin D or wiskostatin decreases entry of P. aeruginosa OMVs into the epithelial cells lining the airway tract (Bomberger et al. 2009).
The different pathways involved in entry of vesicles into host cells have been elucidated through experiments employing a large repertoire of inhibitors/binders specific for different components of each pathway, e.g., dynasore for dynamin, chlorpromazine for clathrin, filipin III, wortmannin, nystatin for lipid rafts, and cytochalasin D for pinocytosis (Amano et al. 2010; Canas et al. 2016; O'Donoghue and Krachler 2016; Rompikuntal et al. 2012). Vesicles make ideal delivery vehicles due to their ability to enter eukaryotic host cells and transfer their cargo to intracellular compartments. In addition, both Gram-positive and Gram-negative bacteria benefit from using vesicles for pathogenesis, intracellular communication and regulating host immunity (Bitto and Kaparakis-Liaskos 2017). Thus, discovering strategies to block vesicle entry into host cells may inhibit membrane vesicle-mediated pathogenesis of bacterial infections.
Membrane fusion is a mechanism by which Gram-negative and Gram-positive bacterial vesicles are internalized into host cells. Membrane fusion enables vesicles to deliver multiple virulence factors directly and simultaneously into the host cell cytoplasm in a coordinated fashion. This phenomenon was first reported by Bomberger et al. (2009) who demonstrated delivery of β-lactamase, alkaline phosphatase, hemolytic phospholipase C, and Cif by P. aeruginosa OMVs into human airway epithelial cells. Membrane fusion between P. aeruginosa vesicles and epithelial cells appear to occur preferentially at lipid raft domains on target host cells. Concomitantly, using filipin III, which sequesters cholesterol and disrupts lipid rafts, the membrane fusion events can be eliminated (Bomberger et al. 2009). A. actinomycetemcomitans OMVs deliver cytolethal distending toxin (CDT) in its biologically active form, and other proteins, including OmpA, into HeLa cells and human gingival fibroblasts, respectively. The OMV-mediated delivery of these proteins occurs in a cholesterol-dependent manner (Rompikuntal et al. 2012). Membrane fusion as a mechanism to deliver virulence factors into host cells has also been observed with MVs from Gram-positive bacteria, i.e., delivery of α-toxin (Hla) by S. aureus MVs into HeLa cells that occurs in a cholesterol-dependent manner and triggers death of the host cell (Thay et al. 2013). Despite the utility of filipin III in studying the dependence on lipid rafts for fusion of bacterial membrane vesicles with host cell vesicles, there is a limitation in its use. Filipin III affects a major component of eukaryotic cell plasma membranes, thus its inhibition of membrane fusion may also extend to processes beyond lipid rafts. In a study of the interaction of L. pneumophila OMVs with model membranes, the membrane material of the MV became incorporated into liposomes composed of different eukaryotic phospholipids, revealing that MVs have an inherent tendency to fuse with eukaryotic membranes (Jager et al. 2015).
7.8 Conclusions
Taken together, the present literature provides ample evidence that OMVs and MVs are capable of employing multiple routes to enter mammalian host cells. Due to their small size, adhesive properties, immunomodulating activity, and ability to carry and deliver specific effectors into mammalian cells, membrane vesicles of bacterial pathogens are well-suited to contribute significantly in the host interaction. Bacterial membrane vesicles allow the extracellular dispersal of particular proteins, as part of complexes of proteins, as well as lipids that can function synergistically to activate different pathways, either toxic or protective, in the host. Further investigations to dissect mechanisms of vesicle adhesion and entry, vesicle trafficking, and vesicle-associated contents will provide a critical foundation for future exploitation of OMVs and MVs for medical use. To date, innovative approaches based on engineered bacterial membrane vesicles have shown great clinical potential, and progress is being made to gain further insight and “know how” in using bacterial membrane vesicle-based technologies to enhance global human health.
References
Ahmad I, Karah N, Nadeem A, Wai SN, Uhlin BE (2019) Analysis of colony phase variation switch in Acinetobacter baumannii clinical isolates. PLoS One 14:e0210082
Aldick T, Bielaszewska M, Uhlin BE, Humpf HU, Wai SN, Karch H (2009) Vesicular stabilization and activity augmentation of enterohaemorrhagic Escherichia coli haemolysin. Mol Microbiol 71:1496–1508
Alvarez CS, Badia J, Bosch M, Gimenez R, Baldoma L (2016) Outer membrane vesicles and soluble factors released by probiotic Escherichia coli Nissle 1917 and commensal ECOR63 enhance barrier function by regulating expression of tight junction proteins in intestinal epithelial cells. Front Microbiol 7:1981
Amano A, Takeuchi H, Furuta N (2010) Outer membrane vesicles function as offensive weapons in host-parasite interactions. Microbes Infect 12:791–798
Askarian F, Lapek JD Jr, Dongre M, Tsai CM, Kumaraswamy M, Kousha A, Valderrama JA, Ludviksen JA, Cavanagh JP, Uchiyama S, Mollnes TE, Gonzalez DJ, Wai SN, Nizet V, Johannessen M (2018) Staphylococcus aureus membrane-derived vesicles promote bacterial virulence and confer protective immunity in murine infection models. Front Microbiol 9:262
Balsalobre C, Silvan JM, Berglund S, Mizunoe Y, Uhlin BE, Wai SN (2006) Release of the type I secreted alpha-haemolysin via outer membrane vesicles from Escherichia coli. Mol Microbiol 59:99–112
Bauman SJ, Kuehn MJ (2006) Purification of outer membrane vesicles from Pseudomonas aeruginosa and their activation of an IL-8 response. Microbes Infect 8:2400–2408
Bauman SJ, Kuehn MJ (2009) Pseudomonas aeruginosa vesicles associate with and are internalized by human lung epithelial cells. BMC Microbiol 9:26
Behrouzi A, Vaziri F, Riazi Rad F, Amanzadeh A, Fateh A, Moshiri A, Khatami S, Siadat SD (2018) Comparative study of pathogenic and non-pathogenic Escherichia coli outer membrane vesicles and prediction of host-interactions with TLR signaling pathways. BMC Res Notes 11:539
Belanger M, Kozarov E, Song H, Whitlock J, Progulske-Fox A (2012) Both the unique and repeat regions of the Porphyromonas gingivalis hemagglutin A are involved in adhesion and invasion of host cells. Anaerobe 18:128–134
Berlanda Scorza F, Doro F, Rodriguez-Ortega MJ, Stella M, Liberatori S, Taddei AR, Serino L, Gomes Moriel D, Nesta B, Fontana MR, Spagnuolo A, Pizza M, Norais N, Grandi G (2008) Proteomics characterization of outer membrane vesicles from the extraintestinal pathogenic Escherichia coli DeltatolR IHE3034 mutant. Mol Cell Proteomics 7:473–485
Berleman J, Auer M (2013) The role of bacterial outer membrane vesicles for intra- and interspecies delivery. Environ Microbiol 15:347–354
Berube BJ, Bubeck Wardenburg J (2013) Staphylococcus aureus alpha-toxin: nearly a century of intrigue. Toxins (Basel) 5:1140–1166
Beveridge TJ, Makin SA, Kadurugamuwa JL, Li Z (1997) Interactions between biofilms and the environment. FEMS Microbiol Rev 20(3–4):291–303
Biagini M, Garibaldi M, Aprea S, Pezzicoli A, Doro F, Becherelli M, Taddei AR, Tani C, Tavarini S, Mora M, Teti G, D’Oro U, Nuti S, Soriani M, Margarit I, Rappuoli R, Grandi G, Norais N (2015) The human pathogen Streptococcus pyogenes releases lipoproteins as lipoprotein-rich membrane vesicles. Mol Cell Proteomics 14:2138–2149
Bielaszewska M, Ruter C, Kunsmann L, Greune L, Bauwens A, Zhang W, Kuczius T, Kim KS, Mellmann A, Schmidt MA, Karch H (2013) Enterohemorrhagic Escherichia coli hemolysin employs outer membrane vesicles to target mitochondria and cause endothelial and epithelial apoptosis. PLoS Pathog 9:e1003797
Bielaszewska M, Ruter C, Bauwens A, Greune L, Jarosch KA, Steil D, Zhang W, He X, Lloubes R, Fruth A, Kim KS, Schmidt MA, Dobrindt U, Mellmann A, Karch H (2017) Host cell interactions of outer membrane vesicle-associated virulence factors of enterohemorrhagic Escherichia coli O157: intracellular delivery, trafficking and mechanisms of cell injury. PLoS Pathog 13:e1006159
Bielaszewska M, Marejkova M, Bauwens A, Kunsmann-Prokscha L, Mellmann A, Karch H (2018) Enterohemorrhagic Escherichia coli O157 outer membrane vesicles induce interleukin 8 production in human intestinal epithelial cells by signaling via toll-like receptors TLR4 and TLR5 and activation of the nuclear factor NF-kappaB. Int J Med Microbiol 308:882–889
Bielig H, Dongre M, Zurek B, Wai SN, Kufer TA (2011a) A role for quorum sensing in regulating innate immune responses mediated by Vibrio cholerae outer membrane vesicles (OMVs). Gut Microbes 2:274–279
Bielig H, Rompikuntal PK, Dongre M, Zurek B, Lindmark B, Ramstedt M, Wai SN, Kufer TA (2011b) NOD-like receptor activation by outer membrane vesicles from Vibrio cholerae non-O1 non-O139 strains is modulated by the quorum-sensing regulator HapR. Infect Immun 79:1418–1427
Biller SJ, Schubotz F, Roggensack SE, Thompson AW, Summons RE, Chisholm SW (2014) Bacterial vesicles in marine ecosystems. Science 343:183–186
Bitto NJ, Kaparakis-Liaskos M (2017) The therapeutic benefit of bacterial membrane vesicles. Int J Mol Sci 18:E1287
Bitto NJ, Chapman R, Pidot S, Costin A, Lo C, Choi J, D’Cruze T, Reynolds EC, Dashper SG, Turnbull L, Whitchurch CB, Stinear TP, Stacey KJ, Ferrero RL (2017) Bacterial membrane vesicles transport their DNA cargo into host cells. Sci Rep 7:7072
Blenkiron C, Simonov D, Muthukaruppan A, Tsai P, Dauros P, Green S, Hong J, Print CG, Swift S, Phillips AR (2016) Uropathogenic Escherichia coli releases extracellular vesicles that are associated with RNA. PLoS One 11:e0160440
Bomberger JM, Maceachran DP, Coutermarsh BA, Ye S, O'Toole GA, Stanton BA (2009) Long-distance delivery of bacterial virulence factors by Pseudomonas aeruginosa outer membrane vesicles. PLoS Pathog 5:e1000382
Bourgeau G, Mayrand D (1990) Aggregation of Actinomyces strains by extracellular vesicles produced by Bacteroides gingivalis. Can J Microbiol 36:362–365
Brown EA, Hardwidge PR (2007) Biochemical characterization of the enterotoxigenic Escherichia coli LeoA protein. Microbiology 153:3776–3784
Canas MA, Gimenez R, Fabrega MJ, Toloza L, Baldoma L, Badia J (2016) Outer membrane vesicles from the probiotic Escherichia coli Nissle 1917 and the commensal ECOR12 enter intestinal epithelial cells via clathrin-dependent endocytosis and elicit differential effects on DNA damage. PLoS One 11:e0160374
Canas MA, Fabrega MJ, Gimenez R, Badia J, Baldoma L (2018) Outer membrane vesicles from probiotic and commensal Escherichia coli activate NOD1-mediated immune responses in intestinal epithelial cells. Front Microbiol 9:498
Cavanagh JP, Pain M, Askarian F, Bruun JA, Urbarova I, Wai SN, Schmidt F, Johannessen M (2018) Comparative exoproteome profiling of an invasive and a commensal Staphylococcus haemolyticus isolate. J Proteomics 195:33–40
Chatterjee D, Chaudhuri K (2011) Association of cholera toxin with Vibrio cholerae outer membrane vesicles which are internalized by human intestinal epithelial cells. FEBS Lett 585:1357–1362
Choi JW, Kim SC, Hong SH, Lee HJ (2017a) Secretable Small RNAs via outer membrane vesicles in periodontal pathogens. J Dent Res 96:458–466
Choi JW, Um JH, Cho JH, Lee HJ (2017b) Tiny RNAs and their voyage via extracellular vesicles: secretion of bacterial small RNA and eukaryotic microRNA. Exp Biol Med (Maywood) 242:1475–1481
Ciofu O, Beveridge TJ, Kadurugamuwa J, Walther-Rasmussen J, Hoiby N (2000) Chromosomal beta-lactamase is packaged into membrane vesicles and secreted from Pseudomonas aeruginosa. J Antimicrob Chemother 45:9–13
Codemo M, Muschiol S, Iovino F, Nannapaneni P, Plant L, Wai SN, Henriques-Normark B (2018) Immunomodulatory effects of pneumococcal extracellular vesicles on cellular and humoral host defenses. MBio 9:e00559–e00518
Dallo SF, Zhang B, Denno J, Hong S, Tsai A, Haskins W, Ye JY, Weitao T (2012) Association of Acinetobacter baumannii EF-Tu with cell surface, outer membrane vesicles, and fibronectin. Sci World J 2012:128705
Dean SN, Leary DH, Sullivan CJ, Oh E, Walper SA (2019) Isolation and characterization of lactobacillus-derived membrane vesicles. Sci Rep 9:877
Doherty GJ, McMahon HT (2009) Mechanisms of endocytosis. Annu Rev Biochem 78:857–902
Donato GM, Goldsmith CS, Paddock CD, Eby JC, Gray MC, Hewlett EL (2012) Delivery of Bordetella pertussis adenylate cyclase toxin to target cells via outer membrane vesicles. FEBS Lett 586:459–465
Dorward DW, Garon CF (1989) DNA-binding proteins in cells and membrane blebs of Neisseria gonorrhoeae. J Bacteriol 171:4196–4201
Dorward DW, Garon CF, Judd RC (1989) Export and intercellular transfer of DNA via membrane blebs of Neisseria gonorrhoeae. J Bacteriol 171:2499–2505
Duchesne P, Grenier D, Mayrand D (1995) Demonstration of adherence properties of Porphyromonas gingivalis outer membrane vesicles using a new microassay. Oral Microbiol Immunol 10:76–80
Ellen RP, Grove DA (1989) Bacteroides gingivalis vesicles bind to and aggregate Actinomyces viscosus. Infect Immun 57:1618–1620
Ellis TN, Leiman SA, Kuehn MJ (2010) Naturally produced outer membrane vesicles from Pseudomonas aeruginosa elicit a potent innate immune response via combined sensing of both lipopolysaccharide and protein components. Infect Immun 78:3822–3831
Elluri S, Enow C, Vdovikova S, Rompikuntal PK, Dongre M, Carlsson S, Pal A, Uhlin BE, Wai SN (2014) Outer membrane vesicles mediate transport of biologically active Vibrio cholerae cytolysin (VCC) from V. cholerae strains. PLoS One 9:e106731
Elmi A, Watson E, Sandu P, Gundogdu O, Mills DC, Inglis NF, Manson E, Imrie L, Bajaj-Elliott M, Wren BW, Smith DG, Dorrell N (2012) Campylobacter jejuni outer membrane vesicles play an important role in bacterial interactions with human intestinal epithelial cells. Infect Immun 80:4089–4098
Elmi A, Nasher F, Jagatia H, Gundogdu O, Bajaj-Elliott M, Wren B, Dorrell N (2016) Campylobacter jejuni outer membrane vesicle-associated proteolytic activity promotes bacterial invasion by mediating cleavage of intestinal epithelial cell E-cadherin and occludin. Cell Microbiol 18:561–572
Evans AG, Davey HM, Cookson A, Currinn H, Cooke-Fox G, Stanczyk PJ, Whitworth DE (2012) Predatory activity of Myxococcus xanthus outer-membrane vesicles and properties of their hydrolase cargo. Microbiology 158:2742–2752
Fabrega MJ, Rodriguez-Nogales A, Garrido-Mesa J, Algieri F, Badia J, Gimenez R, Galvez J, Baldoma L (2017) Intestinal anti-inflammatory effects of outer membrane vesicles from Escherichia coli Nissle 1917 in DSS-experimental colitis in mice. Front Microbiol 8:1274
Fulsundar S, Harms K, Flaten GE, Johnsen PJ, Chopade BA, Nielsen KM (2014) Gene transfer potential of outer membrane vesicles of Acinetobacter baylyi and effects of stress on vesiculation. Appl Environ Microbiol 80:3469–3483
Furuta N, Tsuda K, Omori H, Yoshimori T, Yoshimura F, Amano A (2009) Porphyromonas gingivalis outer membrane vesicles enter human epithelial cells via an endocytic pathway and are sorted to lysosomal compartments. Infect Immun 77:4187–4196
Galka F, Wai SN, Kusch H, Engelmann S, Hecker M, Schmeck B, Hippenstiel S, Uhlin BE, Steinert M (2008) Proteomic characterization of the whole secretome of Legionella pneumophila and functional analysis of outer membrane vesicles. Infect Immun 76(5):1825–1836
Gankema H, Wensink J, Guinee PA, Jansen WH, Witholt B (1980) Some characteristics of the outer membrane material released by growing enterotoxigenic Escherichia coli. Infect Immun 29:704–713
Gao W, Howden BP, Stinear TP (2018) Evolution of virulence in Enterococcus faecium, a hospital-adapted opportunistic pathogen. Curr Opin Microbiol 41:76–82
George KM, Chatterjee D, Gunawardana G, Welty D, Hayman J, Lee R, Small PL (1999) Mycolactone: a polyketide toxin from Mycobacterium ulcerans required for virulence. Science 283:854–857
Ghosh A, Saha DR, Hoque KM, Asakuna M, Yamasaki S, Koley H, Das SS, Chakrabarti MK, Pal A (2006) Enterotoxigenicity of mature 45-kilodalton and processed 35-kilodalton forms of hemagglutinin protease purified from a cholera toxin gene-negative Vibrio cholerae non-O1, non-O139 strain. Infect Immun 74:2937–2946
Green ER, Mecsas J (2016) Bacterial secretion systems: an overview. Microbiol Spectr 4
Gurung M, Moon DC, Choi CW, Lee JH, Bae YC, Kim J, Lee YC, Seol SY, Cho DT, Kim SI, Lee JC (2011) Staphylococcus aureus produces membrane-derived vesicles that induce host cell death. PLoS One 6:e27958
Hase CC, Finkelstein RA (1991) Cloning and nucleotide sequence of the Vibrio cholerae hemagglutinin/protease (HA/protease) gene and construction of an HA/protease-negative strain. J Bacteriol 173:3311–3317
Haurat MF, Aduse-Opoku J, Rangarajan M, Dorobantu L, Gray MR, Curtis MA, Feldman MF (2011) Selective sorting of cargo proteins into bacterial membrane vesicles. J Biol Chem 286:1269–1276
Ho MH, Chen CH, Goodwin JS, Wang BY, Xie H (2015) Functional advantages of Porphyromonas gingivalis vesicles. PLoS One 10:e0123448
Hoekstra D, van der Laan JW, de Leij L, Witholt B (1976) Release of outer membrane fragments from normally growing Escherichia coli. Biochim Biophys Acta 455:889–899
Hornung V (2014) SnapShot: nucleic acid immune sensors, part 1. Immunity 41:868, 868 e1
Horstman AL, Kuehn MJ (2000) Enterotoxigenic Escherichia coli secretes active heat-labile enterotoxin via outer membrane vesicles. J Biol Chem 275:12489–12496
Horstman AL, Kuehn MJ (2002) Bacterial surface association of heat-labile enterotoxin through lipopolysaccharide after secretion via the general secretory pathway. J Biol Chem 277:32538–32545
Hozbor D, Rodriguez ME, Fernandez J, Lagares A, Guiso N, Yantorno O (1999) Release of outer membrane vesicles from Bordetella pertussis. Curr Microbiol 38:273–278
Imagawa T, Sonoda S, Kanoh M, Utsumi S (1979) Polymorphonuclear leukocyte-inhibitory factor of Bordetella pertussis. II. Localization in the outer membrane. Biken J 22:1–10
Irving AT, Mimuro H, Kufer TA, Lo C, Wheeler R, Turner LJ, Thomas BJ, Malosse C, Gantier MP, Casillas LN, Votta BJ, Bertin J, Boneca IG, Sasakawa C, Philpott DJ, Ferrero RL, Kaparakis-Liaskos M (2014) The immune receptor NOD1 and kinase RIP2 interact with bacterial peptidoglycan on early endosomes to promote autophagy and inflammatory signaling. Cell Host Microbe 15:623–635
Jager J, Keese S, Roessle M, Steinert M, Schromm AB (2015) Fusion of Legionella pneumophila outer membrane vesicles with eukaryotic membrane systems is a mechanism to deliver pathogen factors to host cell membranes. Cell Microbiol 17:607–620
Jeon H, Oh MH, Jun SH, Kim SI, Choi CW, Kwon HI, Na SH, Kim YJ, Nicholas A, Selasi GN, Lee JC (2016) Variation among Staphylococcus aureus membrane vesicle proteomes affects cytotoxicity of host cells. Microb Pathog 93:185–193
Jha C, Ghosh S, Gautam V, Malhotra P, Ray P (2017) In vitro study of virulence potential of Acinetobacter baumannii outer membrane vesicles. Microb Pathog 111:218–224
Jiang Y, Kong Q, Roland KL, Curtiss R 3rd (2014) Membrane vesicles of Clostridium perfringens type a strains induce innate and adaptive immunity. Int J Med Microbiol 304:431–443
Johnson AM, Kaushik RS, Francis DH, Fleckenstein JM, Hardwidge PR (2009) Heat-labile enterotoxin promotes Escherichia coli adherence to intestinal epithelial cells. J Bacteriol 191:178–186
Jun SH, Lee JH, Kim BR, Kim SI, Park TI, Lee JC, Lee YC (2013) Acinetobacter baumannii outer membrane vesicles elicit a potent innate immune response via membrane proteins. PLoS One 8:e71751
Kadurugamuwa JL, Beveridge TJ (1995) Virulence factors are released from Pseudomonas aeruginosa in association with membrane vesicles during normal growth and exposure to gentamicin: a novel mechanism of enzyme secretion. J Bacteriol 177:3998–4008
Kadurugamuwa JL, Beveridge TJ (1997) Natural release of virulence factors in membrane vesicles by Pseudomonas aeruginosa and the effect of aminoglycoside antibiotics on their release. J Antimicrob Chemother 40:615–621
Kadurugamuwa JL, Beveridge TJ (1998) Delivery of the non-membrane-permeative antibiotic gentamicin into mammalian cells by using Shigella flexneri membrane vesicles. Antimicrob Agents Chemother 42:1476–1483
Kadurugamuwa JL, Beveridge TJ (1999) Membrane vesicles derived from Pseudomonas aeruginosa and Shigella flexneri can be integrated into the surfaces of other gram-negative bacteria. Microbiology 145(Pt 8):2051–2060
Kamaguchi A, Nakayama K, Ichiyama S, Nakamura R, Watanabe T, Ohta M, Baba H, Ohyama T (2003) Effect of Porphyromonas gingivalis vesicles on coaggregation of Staphylococcus aureus to oral microorganisms. Curr Microbiol 47:485–491
Kaparakis M, Turnbull L, Carneiro L, Firth S, Coleman HA, Parkington HC, Le Bourhis L, Karrar A, Viala J, Mak J, Hutton ML, Davies JK, Crack PJ, Hertzog PJ, Philpott DJ, Girardin SE, Whitchurch CB, Ferrero RL (2010) Bacterial membrane vesicles deliver peptidoglycan to NOD1 in epithelial cells. Cell Microbiol 12:372–385
Kaparakis-Liaskos M, Ferrero RL (2015) Immune modulation by bacterial outer membrane vesicles. Nat Rev Immunol 15:375–387
Kaper JB, Morris JG Jr, Levine MM (1995) Cholera. Clin Microbiol Rev 8:48–86
Kesty NC, Mason KM, Reedy M, Miller SE, Kuehn MJ (2004) Enterotoxigenic Escherichia coli vesicles target toxin delivery into mammalian cells. EMBO J 23:4538–4549
Kieselbach T, Zijnge V, Granstrom E, Oscarsson J (2015) Proteomics of Aggregatibacter actinomycetemcomitans outer membrane vesicles. PLoS One 10:e0138591
Kim YR, Kim BU, Kim SY, Kim CM, Na HS, Koh JT, Choy HE, Rhee JH, Lee SE (2010) Outer membrane vesicles of Vibrio vulnificus deliver cytolysin-hemolysin VvhA into epithelial cells to induce cytotoxicity. Biochem Biophys Res Commun 399:607–612
Kim OY, Park HT, Dinh NTH, Choi SJ, Lee J, Kim JH, Lee SW, Gho YS (2017) Bacterial outer membrane vesicles suppress tumor by interferon-gamma-mediated antitumor response. Nat Commun 8:626
Kobayashi H, Uematsu K, Hirayama H, Horikoshi K (2000) Novel toluene elimination system in a toluene-tolerant microorganism. J Bacteriol 182:6451–6455
Koeppen K, Hampton TH, Jarek M, Scharfe M, Gerber SA, Mielcarz DW, Demers EG, Dolben EL, Hammond JH, Hogan DA, Stanton BA (2016) A novel mechanism of host-pathogen interaction through sRNA in bacterial outer membrane vesicles. PLoS Pathog 12:e1005672
Koeppen K, Barnaby R, Jackson AA, Gerber SA, Hogan DA, Stanton BA (2019) Tobramycin reduces key virulence determinants in the proteome of Pseudomonas aeruginosa outer membrane vesicles. PLoS One 14:e0211290
Kolling GL, Matthews KR (1999) Export of virulence genes and Shiga toxin by membrane vesicles of Escherichia coli O157:H7. Appl Environ Microbiol 65:1843–1848
Kondo K, Takade A, Amako K (1993) Release of the outer membrane vesicles from Vibrio cholerae and Vibrio parahaemolyticus. Microbiol Immunol 37:149–152
Kouokam JC, Wai SN, Fallman M, Dobrindt U, Hacker J, Uhlin BE (2006) Active cytotoxic necrotizing factor 1 associated with outer membrane vesicles from uropathogenic Escherichia coli. Infect Immun 74:2022–2030
Kunsmann L, Ruter C, Bauwens A, Greune L, Gluder M, Kemper B, Fruth A, Wai SN, He X, Lloubes R, Schmidt MA, Dobrindt U, Mellmann A, Karch H, Bielaszewska M (2015) Virulence from vesicles: novel mechanisms of host cell injury by Escherichia coli O104:H4 outbreak strain. Sci Rep 5:13252
Lee EY, Bang JY, Park GW, Choi DS, Kang JS, Kim HJ, Park KS, Lee JO, Kim YK, Kwon KH, Kim KP, Gho YS (2007) Global proteomic profiling of native outer membrane vesicles derived from Escherichia coli. Proteomics 7:3143–3153
Lee EY, Choi DY, Kim DK, Kim JW, Park JO, Kim S, Kim SH, Desiderio DM, Kim YK, Kim KP, Gho YS (2009) Gram-positive bacteria produce membrane vesicles: proteomics-based characterization of Staphylococcus aureus-derived membrane vesicles. Proteomics 9:5425–5436
Lee JH, Choi CW, Lee T, Kim SI, Lee JC, Shin JH (2013) Transcription factor sigmaB plays an important role in the production of extracellular membrane-derived vesicles in Listeria monocytogenes. PLoS One 8:e73196
Lee J, Yoon YJ, Kim JH, Dinh NTH, Go G, Tae S, Park KS, Park HT, Lee C, Roh TY, Di Vizio D, Gho YS (2018) Outer membrane vesicles derived from Escherichia coli regulate neutrophil migration by induction of endothelial IL-8. Front Microbiol 9:2268
Lei L, Yang F, Zou J, Jing H, Zhang J, Xu W, Zou Q, Zhang J (2019) Wang X (2019) DNA vaccine encoding OmpA and Pal from Acinetobacter baumannii efficiently protects mice against pulmonary infection. Mol Biol Rep 46(5):5397–5408
Li Z, Clarke AJ, Beveridge TJ (1998) Gram-negative bacteria produce membrane vesicles which are capable of killing other bacteria. J Bacteriol 180:5478–5483
Li J, Nation RL, Turnidge JD, Milne RW, Coulthard K, Rayner CR, Pater-son DL (2006) Colistin: the re-emerging antibiotic for multidrug-resistant Gram-negative bacterial infections. Lancet Infect Dis 6:589–601
Liao S, Klein MI, Heim KP, Fan Y, Bitoun JP, Ahn SJ, Burne RA, Koo H, Brady LJ, Wen ZT (2014) Streptococcus mutans extracellular DNA is upregulated during growth in biofilms, actively released via membrane vesicles, and influenced by components of the protein secretion machinery. J Bacteriol 196:2355–2366
Lindmark B, Rompikuntal PK, Vaitkevicius K, Song T, Mizunoe Y, Uhlin BE, Guerry P, Wai SN (2009) Outer membrane vesicle-mediated release of cytolethal distending toxin (CDT) from Campylobacter jejuni. BMC Microbiol 9:220
Lowy FD (1998) Staphylococcus aureus infections. N Engl J Med 339:520–532
MacEachran DP, Ye S, Bomberger JM, Hogan DA, Swiatecka-Urban A, Stanton BA, O'Toole GA (2007) The Pseudomonas aeruginosa secreted protein PA2934 decreases apical membrane expression of the cystic fibrosis transmembrane conductance regulator. Infect Immun 75:3902–3912
Malabirade A, Habier J, Heintz-Buschart A, May P, Godet J, Halder R, Etheridge A, Galas D, Wilmes P, Fritz JV (2018) The RNA complement of outer membrane vesicles from Salmonella enterica Serovar Typhimurium under distinct culture conditions. Front Microbiol 9:2015
Mantri CK, Chen CH, Dong X, Goodwin JS, Pratap S, Paromov V, Xie H (2015) Fimbriae-mediated outer membrane vesicle production and invasion of Porphyromonas gingivalis. Microbiology 4:53–65
Marsollier L, Brodin P, Jackson M, Kordulakova J, Tafelmeyer P, Carbonnelle E, Aubry J, Milon G, Legras P, Andre JP, Leroy C, Cottin J, Guillou ML, Reysset G, Cole ST (2007) Impact of Mycobacterium ulcerans biofilm on transmissibility to ecological niches and Buruli ulcer pathogenesis. PLoS Pathog 3:e62
Mashburn LM, Whiteley M (2005) Membrane vesicles traffic signals and facilitate group activities in a prokaryote. Nature 437:422–425
Mashburn-Warren LM, Whiteley M (2006) Special delivery: vesicle trafficking in prokaryotes. Mol Microbiol 61:839–846
McConnell MJ, Rumbo C, Bou G, Pachon J (2011) Outer membrane vesicles as an acellular vaccine against Acinetobacter baumannii. Vaccine 29:5705–5710
Mondal A, Tapader R, Chatterjee NS, Ghosh A, Sinha R, Koley H, Saha DR, Chakrabarti MK, Wai SN, Pal A (2016) Cytotoxic and inflammatory responses induced by outer membrane vesicle-associated biologically active proteases from Vibrio cholerae. Infect Immun 84:1478–1490
Mulcahy LA, Pink RC, Carter DR (2014) Routes and mechanisms of extracellular vesicle uptake. J Extracell Vesicles 3. https://doi.org/10.3402/jev.v3.24641.eCollection
Murphy K, Park AJ, Hao Y, Brewer D, Lam JS, Khursigara CM (2014) Influence of O polysaccharides on biofilm development and outer membrane vesicle biogenesis in Pseudomonas aeruginosa PAO1. J Bacteriol 196:1306–1317
Nakao R, Myint SL, Wai SN, Uhlin BE (2018) Enhanced biofilm formation and membrane vesicle release by Escherichia coli expressing a commonly occurring plasmid gene, kil. Front Microbiol 9:2605
Nevo-Dinur K, Govindarajan S, Amster-Choder O (2012) Subcellular localization of RNA and proteins in prokaryotes. Trends Genet 28:314–322
O’Donoghue EJ, Krachler AM (2016) Mechanisms of outer membrane vesicle entry into host cells. Cell Microbiol 18:1508–1517
Olier M, Marcq I, Salvador-Cartier C, Secher T, Dobrindt U, Boury M, Bacquie V, Penary M, Gaultier E, Nougayrede JP, Fioramonti J, Oswald E (2012) Genotoxicity of Escherichia coli Nissle 1917 strain cannot be dissociated from its probiotic activity. Gut Microbes 3:501–509
Olivier V, Haines GK 3rd, Tan Y, Satchell KJ (2007) Hemolysin and the multifunctional autoprocessing RTX toxin are virulence factors during intestinal infection of mice with Vibrio cholerae El Tor O1 strains. Infect Immun 75:5035–5042
Olofsson A, Nygard Skalman L, Obi I, Lundmark R, Arnqvist A (2014) Uptake of Helicobacter pylori vesicles is facilitated by clathrin-dependent and clathrin-independent endocytic pathways. MBio 5:e00979–e00914
Olsen I, Amano A (2015) Outer membrane vesicles - offensive weapons or good Samaritans? J Oral Microbiol 7:27468
Pathirana RD, Kaparakis-Liaskos M (2016) Bacterial membrane vesicles: biogenesis, immune regulation and pathogenesis. Cell Microbiol 18:1518–1524
Patrick S, McKenna JP, O'Hagan S, Dermott E (1996) A comparison of the haemagglutinating and enzymic activities of Bacteroides fragilis whole cells and outer membrane vesicles. Microb Pathog 20:191–202
Perez-Cruz C, Delgado L, Lopez-Iglesias C, Mercade E (2015) Outer-inner membrane vesicles naturally secreted by gram-negative pathogenic bacteria. PLoS One 10:e0116896
Pollak CN, Delpino MV, Fossati CA, Baldi PC (2012) Outer membrane vesicles from Brucella abortus promote bacterial internalization by human monocytes and modulate their innate immune response. PLoS One 7:e50214
Prados-Rosales R, Baena A, Martinez LR, Luque-Garcia J, Kalscheuer R, Veeraraghavan U, Camara C, Nosanchuk JD, Besra GS, Chen B, Jimenez J, Glatman-Freedman A, Jacobs WR Jr, Porcelli SA, Casadevall A (2011) Mycobacteria release active membrane vesicles that modulate immune responses in a TLR2-dependent manner in mice. J Clin Invest 121:1471–1483
Raeven RH, Brummelman J, Pennings JL, van der Maas L, Tilstra W, Helm K, van Riet E, Jiskoot W, van Els CA, Han WG, Kersten GF, Metz B (2016) Bordetella pertussis outer membrane vesicle vaccine confers equal efficacy in mice with milder inflammatory responses compared to a whole-cell vaccine. Sci Rep 6:38240
Rasti ES, Schappert ML, Brown AC (2018) Association of Vibrio cholerae 569B outer membrane vesicles with host cells occurs in a GM1-independent manner. Cell Microbiol 20:e12828
Rath P, Huang C, Wang T, Wang T, Li H, Prados-Rosales R, Elemento O, Casadevall A, Nathan CF (2013) Genetic regulation of vesiculogenesis and immunomodulation in Mycobacterium tuberculosis. Proc Natl Acad Sci U S A 110:E4790–E4797
Renelli M, Matias V, Lo RY, Beveridge TJ (2004) DNA-containing membrane vesicles of Pseudomonas aeruginosa PAO1 and their genetic transformation potential. Microbiology 150:2161–2169
Resch U, Tsatsaronis JA, Le Rhun A, Stubiger G, Rohde M, Kasvandik S, Holzmeister S, Tinnefeld P, Wai SN, Charpentier E (2016) A two-component regulatory system impacts extracellular membrane-derived vesicle production in group A Streptococcus. MBio 7:e00207–e00216
Riley DR, Sieber KB, Robinson KM, White JR, Ganesan A, Nourbakhsh S, Dunning Hotopp JC (2013) Bacteria-human somatic cell lateral gene transfer is enriched in cancer samples. PLoS Comput Biol 9:e1003107
Rivera J, Cordero RJ, Nakouzi AS, Frases S, Nicola A, Casadevall A (2010) Bacillus anthracis produces membrane-derived vesicles containing biologically active toxins. Proc Natl Acad Sci U S A 107:19002–19007
Rompikuntal PK, Thay B, Khan MK, Alanko J, Penttinen AM, Asikainen S, Wai SN, Oscarsson J (2012) Perinuclear localization of internalized outer membrane vesicles carrying active cytolethal distending toxin from Aggregatibacter actinomycetemcomitans. Infect Immun 80:31–42
Rompikuntal PK, Vdovikova S, Duperthuy M, Johnson TL, Ahlund M, Lundmark R, Oscarsson J, Sandkvist M, Uhlin BE, Wai SN (2015) Outer membrane vesicle-mediated export of processed PrtV protease from Vibrio cholerae. PLoS One 10:e0134098
Rumbo C, Fernandez-Moreira E, Merino M, Poza M, Mendez JA, Soares NC, Mosquera A, Chaves F, Bou G (2011) Horizontal transfer of the OXA-24 carbapenemase gene via outer membrane vesicles: a new mechanism of dissemination of carbapenem resistance genes in Acinetobacter baumannii. Antimicrob Agents Chemother 55:3084–3090
Schaar V, de Vries SP, Perez Vidakovics ML, Bootsma HJ, Larsson L, Hermans PW, Bjartell A, Morgelin M, Riesbeck K (2011) Multicomponent Moraxella catarrhalis outer membrane vesicles induce an inflammatory response and are internalized by human epithelial cells. Cell Microbiol 13:432–449
Sharpe SW, Kuehn MJ, Mason KM (2011) Elicitation of epithelial cell-derived immune effectors by outer membrane vesicles of nontypeable Haemophilus influenzae. Infect Immun 79:4361–4369
Shen Y, Giardino Torchia ML, Lawson GW, Karp CL, Ashwell JD, Mazmanian SK (2012) Outer membrane vesicles of a human commensal mediate immune regulation and disease protection. Cell Host Microbe 12:509–520
Singh U, Grenier D, McBride BC (1989) Bacteroides gingivalis vesicles mediate attachment of streptococci to serum-coated hydroxyapatite. Oral Microbiol Immunol 4:199–203
Sjostrom AE, Sandblad L, Uhlin BE, Wai SN (2015) Membrane vesicle-mediated release of bacterial RNA. Sci Rep 5:15329
Smalley JW, Birss AJ, McKee AS, Marsh PD (1991) Haemin-restriction influences haemin-binding, haemagglutination and protease activity of cells and extracellular membrane vesicles of Porphyromonas gingivalis W50. FEMS Microbiol Lett 69:63–67
Svennerholm K, Park KS, Wikstrom J, Lasser C, Crescitelli R, Shelke GV, Jang SC, Suzuki S, Bandeira E, Olofsson CS, Lotvall J (2017) Escherichia coli outer membrane vesicles can contribute to sepsis induced cardiac dysfunction. Sci Rep 7:17434
Syngkon A, Elluri S, Koley H, Rompikuntal PK, Saha DR, Chakrabarti MK, Bhadra RK, Wai SN, Pal A (2010) Studies on a novel serine protease of a DeltahapADeltaprtV Vibrio cholerae O1 strain and its role in hemorrhagic response in the rabbit ileal loop model. PLoS One 5:e13122
Taheri N, Mahmud A, Sandblad L, Fallman M, Wai SN, Fahlgren A (2018) Campylobacter jejuni bile exposure influences outer membrane vesicles protein content and bacterial interaction with epithelial cells. Sci Rep 8:16996
Tashiro Y, Ichikawa S, Shimizu M, Toyofuku M, Takaya N, Nakajima-Kambe T, Uchiyama H, Nomura N (2010) Variation of physiochemical properties and cell association activity of membrane vesicles with growth phase in Pseudomonas aeruginosa. Appl Environ Microbiol 76:3732–3739
Tashiro Y, Inagaki A, Shimizu M, Ichikawa S, Takaya N, Nakajima-Kambe T, Uchiyama H, Nomura N (2011) Characterization of phospholipids in membrane vesicles derived from Pseudomonas aeruginosa. Biosci Biotechnol Biochem 75:605–607
Tashiro Y, Hasegawa Y, Shintani M, Takaki K, Ohkuma M, Kimbara K, Futamata H (2017) Interaction of bacterial membrane vesicles with specific species and their potential for delivery to target cells. Front Microbiol 8:571
Thay B, Wai SN, Oscarsson J (2013) Staphylococcus aureus alpha-toxin-dependent induction of host cell death by membrane-derived vesicles. PLoS One 8:e54661
Thay B, Damm A, Kufer TA, Wai SN, Oscarsson J (2014) Aggregatibacter actinomycetemcomitans outer membrane vesicles are internalized in human host cells and trigger NOD1- and NOD2-dependent NF-kappaB activation. Infect Immun 82:4034–4046
Turnbull L, Toyofuku M, Hynen AL, Kurosawa M, Pessi G, Petty NK, Osvath SR, Carcamo-Oyarce G, Gloag ES, Shimoni R, Omasits U, Ito S, Yap X, Monahan LG, Cavaliere R, Ahrens CH, Charles IG, Nomura N, Eberl L, Whitchurch CB (2016) Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nat Commun 7:11220
Turner L, Bitto NJ, Steer DL, Lo C, D'Costa K, Ramm G, Shambrook M, Hill AF, Ferrero RL, Kaparakis-Liaskos M (2018) Helicobacter pylori outer membrane vesicle size determines their mechanisms of host cell entry and protein content. Front Immunol 9:1466
Tyrer PC, Frizelle FA, Keenan JI (2014) Escherichia coli-derived outer membrane vesicles are genotoxic to human enterocyte-like cells. Infect Agent Cancer 9:2
Vaitkevicius K, Lindmark B, Ou G, Song T, Toma C, Iwanaga M, Zhu J, Andersson A, Hammarstrom ML, Tuck S, Wai SN (2006) A Vibrio cholerae protease needed for killing of Caenorhabditis elegans has a role in protection from natural predator grazing. Proc Natl Acad Sci U S A 103:9280–9285
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ, Lotvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9:654–659
Vanaja SK, Russo AJ, Behl B, Banerjee I, Yankova M, Deshmukh SD, Rathinam VAK (2016) Bacterial outer membrane vesicles mediate cytosolic localization of LPS and caspase-11 activation. Cell 165:1106–1119
Vdovikova S, Luhr M, Szalai P, Nygard Skalman L, Francis MK, Lundmark R, Engedal N, Johansson J, Wai SN (2017) A novel role of Listeria monocytogenes membrane vesicles in inhibition of autophagy and cell death. Front Cell Infect Microbiol 7:154
Vdovikova S, Gilfillan S, Wang S, Dongre M, Wai SN, Hurtado A (2018) Modulation of gene transcription and epigenetics of colon carcinoma cells by bacterial membrane vesicles. Sci Rep 8:7434
Veith PD, Chen YY, Gorasia DG, Chen D, Glew MD, O’Brien-Simpson NM, Cecil JD, Holden JA, Reynolds EC (2014) Porphyromonas gingivalis outer membrane vesicles exclusively contain outer membrane and periplasmic proteins and carry a cargo enriched with virulence factors. J Proteome Res 13:2420–2432
Vidakovics ML, Jendholm J, Morgelin M, Mansson A, Larsson C, Cardell LO, Riesbeck K (2010) B cell activation by outer membrane vesicles-a novel virulence mechanism. PLoS Pathog 6:e1000724
Wagner T, Joshi B, Janice J, Askarian F, Skalko-Basnet N, Hagestad OC, Mekhlif A, Wai SN, Hegstad K, Johannessen M (2018) Enterococcus faecium produces membrane vesicles containing virulence factors and antimicrobial resistance related proteins. J Proteome 187:28–38
Wai SN, Lindmark B, Soderblom T, Takade A, Westermark M, Oscarsson J, Jass J, Richter-Dahlfors A, Mizunoe Y, Uhlin BE (2003) Vesicle-mediated export and assembly of pore-forming oligomers of the enterobacterial ClyA cytotoxin. Cell 115:25–35
Xie H (2015) Biogenesis and function of Porphyromonas gingivalis outer membrane vesicles. Future Microbiol 10:1517–1527
Yaron S, Kolling GL, Simon L, Matthews KR (2000) Vesicle-mediated transfer of virulence genes from Escherichia coli O157:H7 to other enteric bacteria. Appl Environ Microbiol 66:4414–4420
Zhang W, Ju J, Rigney T, Tribble GD (2011) Fimbriae of Porphyromonas gingivalis are important for initial invasion of osteoblasts, but not for inhibition of their differentiation and mineralization. J Periodontol 82:909–916
Zhong H, Yang Y, Ma S, Xiu F, Cai Z, Zhao H, Du L (2011) Induction of a tumour-specific CTL response by exosomes isolated from heat-treated malignant ascites of gastric cancer patients. Int J Hyperth 27:604–611
Zomer A, Vendrig T, Hopmans ES, van Eijndhoven M, Middeldorp JM, Pegtel DM (2010) Exosomes: fit to deliver small RNA. Commun Integr Biol 3:447–450
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Nadeem, A., Oscarsson, J., Wai, S.N. (2020). Delivery of Virulence Factors by Bacterial Membrane Vesicles to Mammalian Host Cells. In: Kaparakis-Liaskos, M., Kufer, T. (eds) Bacterial Membrane Vesicles. Springer, Cham. https://doi.org/10.1007/978-3-030-36331-4_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-36331-4_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-36330-7
Online ISBN: 978-3-030-36331-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)