Abstract
The natural ability of Trichoderma species to secrete a wide array of enzymes, capable of targeting and hydrolysing complex plant biomass and fungal pathogens, finds diverse applications in biotechnology and agricultural, pharmaceutical and other industrial sectors. Secretion of lignocellulose-degrading enzymes in particular by Trichoderma species makes it one of the most explored fungi which has gained worldwide attention of researchers for biofuel production from agricultural biomass. In particular, Trichoderma reesei brought a paradigm shift for industrially relevant enzymes. Enzymes produced by Trichoderma species include cellulase complex which targets cellulose, xylanase which targets xylan and other non-glycosyl hydrolases. Mining genome and in silico analysis of reference strain T. reesei QM6a genomes and transcriptomes for carbohydrate-active enzymes (CAZyme) have led to the identification of several candidate genes. Besides this, laccase (phenol oxidase) and lytic mono-oxygenase system of Trichoderma have been explored for industrial applications. Here in this chapter, attempt has been made to discuss the role of extracellular carbohydrate-active enzymes of fungal origin with special emphasis on Trichoderma for their role in biofuel prpduction from non-edible biomass.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
12.1 Introduction
Constant depletion of fossil fuels, increasing world population and concerns for environments, in particular the impact of climate change on our ecosystem, demand futuristic sustainable technologies. India is presently ranked third in oil consumption. Moreover, growing population size, growth in automobile and other industrial sectors in India, led to increase in energy consumption. The need for environmental friendly and renewable energy resources such as biofuels produced from agricultural-based biomass can decrease our dependence on fossil fuels (Borin et al. 2017). Therefore, efforts for developing alternate energy resources are on high priority. As per the records of US Department of Energy, United States and Brazil contributed to approximately 80% (24,570 million gallons) of the global ethanol production (http://www.afdc.energy.gov) (Borin et al. 2017). The bioprospection of agricultural biomass in particular from non-edible sources can be a better alternative and sustainable approach with minimal environmental concerns in the future (Gaurav et al. 2017). Agricultural biomass which is often a major source for environmental pollution can be of vital importance for biofuel production as an alternate energy resource (Ning et al. 2016; Wan et al. 2001; Chirino-Valle et al. 2016). Limitations of biomass from grain-producing crops demand alternative second-generation biofuels from non-edible agricultural crops (Ayrinhac et al. 2011) and other biomass sources. These carbohydrates from different non-edible biomass can be explored for biofuel using a combination of enzymes (Gaurav et al. 2017). Biofuels are categorized into three generations on the basis of raw material. Initially for first generation biofuel, crops plants were explored followed by second generation agricultural by-products and marine resources such as seaweeds and cyanobacteria (Demirbas 2008; Kang et al. 2014; Gaurav et al. 2017).
Biofuels from second-generation agricultural wastes offer several benefits. Biofuels from renewable resources can be exploited as promising and almost carbon-neutral fuel enhancers of octane in unleaded gasoline for cleaner combustion which can reduce environmental pollution. Plant biomass containing lignocellulose in terrestrial ecosystems is one of the most potential raw materials due to its availability, price and high sugar content (Barros-Rios et al. 2016; Zhao et al. 2016). The basic constituents of lignocellulose include cellulose, hemicellulose and lignin (Sindhu et al. 2016) which are interconnected through covalent and non-covalent bonds (Gaurav et al. 2017; Zhang et al. 2017). Cellulose which is a major part of plant biomass has been widely recognized and explored for developing sustainable processes and can help in mitigating the impact of climate change, occurs through consumption of fossil fuels (Gupta and Verma 2015; Zhang et al. 2017). Conversion of lignocellulose-based plant biomass is a major bottleneck in developing sustainable processes for alternate energy resources and other value-addition products (Kuhad et al. 2011; Villares et al. 2017). The breakdown of recalcitrance lignocellulose and chitin containing biomass using chemical pretreatment often results in toxic side effects to the ecosystem (Margeot et al. 2009; Wang et al. 2017) (Fig. 12.1).
The conversion of plant biomass into value added products can be achieved through the breakdown of recalcitrant plant biomass via pretreatment and enzymatic hydrolysis (Zhang et al. 2017). Efficient utilization of the lignin, hemicellulose and cellulose can decrease the cost of biofuel production up to 25% (Zhao and Xia 2009, 2010; Zhao et al. 2018a).
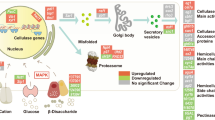
Enzymes in the CAZy database are categorized into four classes: glycoside hydrolases (GHs), polysaccharide lyases, glycosyl transferases and carbohydrate esterases. The glycosyl hydrolases (GHs) have potential to break the non-edible biomass into oligo- or monomers (Ferreira Filho et al. 2017). Additionally, a family of auxiliary enzymes known as lytic polysaccharide mono-oxygenases (LPMOs) which is a major component of saprophytic fungi like Trichoderma and Aspergillus catalyses copper-dependent oxidation of C-H bonds in complex polysaccharides (Obeng et al. 2017; Borin et al. 2017; Monclaro and Filho 2017; Cologna et al. 2018) (Fig. 12.2).
Overview of carbohydrate-active enyzmes as per cazy database http://www.cazy.org/
In general, the stains of Trichoderma are used as biocontrol agents due to their diverse attributes (Sharma and Shanmugam 2012; Sharma et al. 2013, 2016a, b, 2017a, b, 2018a, b). Besides this, enzymes from filamentous fungi such as T. reesei are paradigms for industrial application in paper, textile, pulp, food and biofuel processing industries (Kumar et al. 2008; Singhania et al. 2010; Seiboth et al. 2011; Marx et al. 2013; Tiwari et al. 2013) (Table 12.1).
Laccases or phenol oxidases and lytic mono-oxygenases can enhance the activity of lignocellulases and thus lower the enzyme required to break down alkali-pretreated agricultural biomass containing lignocellulose (Ladeira Azar et al. 2018). For examples, xylanase of A. niger and T. reesei are found to be inhibited by the presence of phenol at 1.5 mg and 0.3 mg per mg of protein, respectively. On the other hand, laccases of C. cubensis and Penicillium pinophilum are reported active at a concentration of 35 mg of phenol per mg of protein (Ladeira Azar et al. 2018). The glcyosyl hydrolase family plays a vital role in the breakdown of complex plant biomass, whereas the role of the auxiliary activity (AA) family has been discussed in recent studies (Levasseur et al. 2013). Among different CAZymes, β-1,4/(1,3)/(1,6)-type glycosyl hydrolase family breaks down complex plant polysaccharides to oligomers or monomers (Vu and Marletta 2016). Lytic mono-oxygenases belonging to AA9 (formerly GH-61), AA10 (formerly CBM-33) and AA11 enzymes are capable of targeting recalcitrant non-edible carbohydrates such as chitin, cellulose, starch and other polysaccharides containing β-linkages between glucose and substituted glucose units (Ravalason et al. 2012; Vu et al. 2014; Gong et al. 2015; Ning et al. 2016). The genomes of A. niger and T. reesei share about the same (2.5%) proportion of CAZymes in comparison to total predicted genes; still, the transcriptomic response of A. niger is found to be diverse and revealed upregulation of 190 CAZymes which belong to 62 different families, whereas for T. reesei, 105 CAZymes belonging to 51 families were upregulated (Borin et al. 2017).
The recent developments in genomic, transcriptomic, metabolomic or proteomic technologies have led to the identification of several CAZymes and other genes of Trichoderma which are active during agricultural biomass degradation. Keeping in view the importance of CAZymes in plant biomass degradation for various applications, attempt has been in present chapter to provide an overview of different lytic enzymes of Trichoderma strains in white biotechnology for biofuel production.
12.2 Biocatalysis of Plant Biomass Using Lignocellulases
Lignocellulose from plant biomass is the major raw material for biofuels, foods and other livestock feeds (Kumar et al. 2008). Studies on fungal lignocelluloses-mediated lysis have revealed several pathways for lignin metabolism (Mansur et al. 2003). The lignocellulose is a promising biomass pretreatment alternative, and fungal lignocellulases are one of the potential enzymes in debasing lignin of plants (dos Santos et al. 2007; Dias et al. 2007; Plácido and Capareda 2015;Martinez et al. 2009). Moreover, the lignocellulases are also explored for the removal of toxic compounds as well as supplementing the pre-existing technologies of sugar hydrolysates after conventional pretreatment (Plácido and Capareda 2015; Bilal et al. 2018). Higher white fungi are known to produce a plethora of lytic enzymes. The lignin-degrading enzyme complex in white fungi is mainly consists of lignin peroxidase, manganese peroxidase and laccase along with other enzymes which include peroxidase, aryl alcohol oxidase, glyoxal oxidase and oxalate. The broad specificity of substrates also makes them vital enzymes which are capable of breaking a wide range of xenobiotics and pollutants having structural similarities to lignin (Hofrichter 2002; Bilal et al. 2018). A combination of co-culture techniques is found to enhance production of these enzymes. For example, 2.6-fold enhancement in laccase activity compared to C. comatus monoculture with higher delignification of up to 66.5% and conversion of 82% of polysaccharides into fermentable sugars was recorded (Ma and Ruan 2015).
12.2.1 Cellulases
Cellulase , a complex of three enzymes, leads to the complete breakdown of cellulose to glucose units which can be used as fermentable sugar for biofuel production. The cellulose is degraded initially through endoglucanase (EG) (1,4-β- D-glucan-4-glucano-hydrolases) (EC 3.2.1.74) by random action into oligomers which are then targeted by exoglucanase (EC 3.2.1.74 and EC3.2.1.91) into cellobiose and glucose units. The β-glucosidases belonging to EC 3.2.1.21 hydrolyse the cellodextrins, cellobiose into glucose units (Keshwani and Cheng 2009; Jeya et al. 2009). The cellobiohydrolases (CBHs, named as CBH1 and CBH2), β-glucosidases (BGLs) and endoglucanases (EGs) act in a coordinated and complementary fashion to hydrolyse cellulose (Cavaco-Paulo et al. 1997; Gusakov et al. 2007; Jørgensen et al. 2007; Ma et al. 2011). The cocktail of different cellulolytic enzymes play vital role in the hydrolysis of complex plant polysaccharides. For example, a mixture of CBH1, CBH2 and EG1 is found to responsible for up to 80% of cellulose breakdown (Rosgaard et al. 2007). T. reesei, an industrial strain, is known to secrete CBH1, CBH2, EG1, EG2, EG3 and EG5 which act in a synergistic manner to completely hydrolyse the lignocellulose (Fang and Xia 2013). CBH1 and CBH2 are reported as major components of cellulase complex and accounts for 50–60% and 10–15% of the secreted protein, respectively (Rosgaard et al. 2007). Compared to CBH1, the specificity of CBH2 is approximately twice for crystalline cellulose (Zhou et al. 2008), and optimum synergism is reported at a 2:1 molar ratio (Zhou et al. 2009).
The other components of cellulase complex in T. reesei such as endo-β-1,4-D-glucanases are reported from glycosyl hydrolase families GH5, GH7, GH12 and GH45, whereas cellobiohydrolases are reported from families GH6 and GH7. The GH7 family contains endo-β-1,4-D-glucanases of CEL7B, previously known as EGL1 and CBHs (CEL7A, named as CBH1). The family GH5 cellulases is mostly explored from fungi strains (Li and Walton 2017), and three candidates of this family have been reported from T. reesei. The enzymes of GH7 family are distributed commonly. The orthologues of CEL7A cellulases are prevalent in the secretome of fungi-degrading biomass. The members of GH6 family comprise cellulase which acts exclusively from the non-reducing end of cellulose chain. The synergistic action of CEL7A and CEL6A is considered to play a key role in biomass degradation. The members of GH12 are typically low molecular weight (25 kDa) and do not contain cellulose-binding domain (CBM1) and glycosylation site. Due to their small size, GH12 can diffuse deeper into cellulosic material, and hence preferred for their role in laundry industry. On the other hand, members of GH45 cellulases are in general small and have a wide substrate range compared to families GH5 and GH7. The members of GH45 enzymes share interesting structural similarities to plant expansins. Further intensive research efforts with genetic engineering strategies for single-enzyme cellulase components have increased the scope of T. reesei strain’s improvement (Pryor and Nahar 2015; Qian et al. 2016, 2017; Wang and Xia 2011; Zhang et al. 2010).
12.2.2 β-Glucosidase
A heterogeneous family containing exo-glycosyl hydrolases catalyses the cleavage of β-glycosidic bonds in disaccharide or glucose-substituted molecules (Bhatia et al. 2002; Chandra et al. 2013; Cheng et al. 2017; Leah et al. 1995; Zagrobelny et al. 2008). According to the classification of CAZy (http://www.cazy.org) (Henrissat 1991; Cantarel et al. 2009), β-glucosidases are classified into two families: 1 and 3 of glycosyl hydrolases (Jeng et al. 2011). These enzymes enhance the action of cellulose-degrading enzymes by releasing phenolic compounds and hence are an attractive choice for renewable bioenergy. β-glucosidases hydrolyse the oligosaccharides and cellobiose oligomeric units obtained after the endoglucanases and cellobiohydrolases activities into monomeric glucose (Chandra et al. 2013).
The β-glucosidases of T. reesei are categorized into GH1 and GH3. The members belonging to family GH1 are exclusively intracellular in nature, whereas GH3 β-glucosidases are predominantly extracellular (Guo et al. 2016). CEL3A previously categorized as BGL1 is responsible for majority of the β-glucosidase activity. The ‘exo/endo’ concept revealed that CEL7A is also able to act in endo-manner; therefore, it is not a true exocellulase (Stahlberg et al. 1993; Kurasin and Valjamae 2011). However, neither the EGs nor the CBHs from fungi can cause massive cellulose decomposition (Payne et al. 2015). The lytic polysaccharide mono-oxygenases which were identified previously as endoglucanases belonging to GH61 (Sharma et al. 2018b) are now known as auxiliary family and cleave β-glucan in an oxidative fashion. The members of the family GH61 are also reported for their weak endoglucanase activity. The genome of T. reesei (http://www.genome.jgipsf.org/Trire2/Trire2.home.html) is reported to contain at least 10, β-glucosidases-encoded genes which include cel1A, cel1B, cel3A, cel3B, cel3C, cel3D, cel3E, cel3F, cel3G and cel3H. The gene encoding cel3A (bgl1) was found to be major extracellular β-glucosidase, whereas cel1A (bgl2) (Saloheimo et al. 2002a, b) and cel1B (Zhou et al. 2012) were reported to be intracellular. Additionally, cel3B, cel3E, cel3F, cel3G and cel3H are assumed to be extracellular, and cel3C, cel3D and cel3H are depicted as intracellular (Guo et al. 2016). Different knockouts, amino acid substitution and mutation of the BglR transcription factor in the PC-3-7 strain have been used to reveal the function of β-glucosidases (Fowler and Brown 1992; Zhou et al. 2012; Nitta et al. 2012; Xu et al. 2014; de Porciuncula et al. 2013; Shida et al. 2015; Li et al. 2016).
12.2.3 Xylanases
With a backbone of β-(1-4)-linked xylose units, polysaccharide xylan are structurally diverse and complex polysaccharides and predominantly composed of hemicelluloses which are linked to cellulose microfibrils (Scheller and Ulvskov 2010). The side chains are connected through C2 and C3 positions of D-xylosyl units (Puls and Schuseil 1993), and these chains can be substituted with acetyl, 4-methyl-D-glucuronosyl or L-arabinosyl units (Wong et al. 1988; Dodd and Cann 2009). Endo-β-1,4-xylanases or β-1,4-D-xylan xylanohydrolases (EC 3.2.1.8) are one of the important lytic components which can target the glycoside bonds in xylan backbone internally (Biely 1985; Polizeli et al. 2005; Mangan et al. 2017). Members of xylanase family belong to glycoside hydrolase (GH) families 5–12, 16, 26, 30, 43, 44, 51 and 62. Enzymes classified in 16, 51 and 62 families contain two catalytic domains compared to 5–11 and 43 families which have a true catalytic domain with endo-1,4-β-xylanase activity. The 9, 12, 26, 30 and 44 families may possess residual or secondary xylanase activity.
In recent classifications based on hydrophobic cluster analysis of catalytic domains and amino acid sequence similarities, xylanases are classified as GH10 and 11 and have a retaining type of mechanism. The information on catalytic properties of families 5, 7, 8 and 43 are very limited. The members of GH families 5, 7, 8, 10, 11 and 43 are different in their structure, mode of action, physicochemical properties and substrate specificities (Collins et al. 2005). The members of GH 10 family include high-molecular-mass proteins with cellulose-binding and catalytic domains and are connected through linker peptides. The estimated pI is 8–9.5 with (α/β)8 fold TIM barrel structure. On the other side, the GH11 family with low molecular mass and pI are further divided into two, alkaline and acidic (Buchert et al. 1995; Juturu and Wu 2012). The GH11 members exclusively catalyse endo-β-1,4–mediated cleavage (EC 3.2.1.8) in xylan and hence are also known as true xylanases. The high catalytic efficiencies of these enzymes due to small size, vast temperature and pH optima provide them an edge for their exploitation in various biotechnological applications (Paes et al. 2012).
Xylanases of Trichoderma are one of the widely explored enzymes, and Rut C-30 strain of T. reesei is well explored for commercial applications of xylanase and cellulase production (Gerber et al. 1997). The xylanases produced by T. harzianum, T. lignorum, T. koningii, T. longibrachiatum, T. pseudokoningii and T. viride also have been investigated (Silveira et al. 1999; Chen et al. 2009). Xylanases from a psychrotrophic Trichoderma strain have been characterized (Zhou et al. 2011) and genes encoding them have been cloned from Trichoderma species and expressed in heterologous hosts such as E. coli (Min et al. 2002), S. cerevisiae (Ahmed et al. 2005) and P. pastoris (He et al. 2009). In the T. reesei genome, three xylanases belonging to the GH11 family have been identified, and two of these were reported in the early 1990s, whereas the third GH11 xylanase XYN5 was identified in a recent study (Martinez et al. 2008; Dos Santos Castro et al. 2014; Peciulyte et al. 2014; Saloheimo and Pakula 2012).
12.2.4 Lytic Polysaccharide Mono-oxygenases (LPMOs)
The recently discovered enzyme class, the LPMOs , stimulates the hydrolysis of plant biomass and enhances the efficacy of glycosyl hydrolases (Hu et al. 2014). Unlike cellulases which target glycosidic bonds by hydrolysis, LPMOs are copper dependent and catalyse the breakdown of polysaccharides through oxidation at C1 or C4 glucose units in the presence of external electron donors.
12.2.5 Laccases
Laccases are also known as phenol oxidases or benzenediol: oxygen oxidoreductase (EC 1.10.3.2) belongs to the multicopper oxidase (MCO) family and represents a group of metalloenzymes. These enzymes are used in various biotechnological applications. The search for strains producing such laccases has gained increased attention in recent times. In general, laccases are monomeric glycoproteins of 60–70 kDa in size, and carbohydrates approximately contribute to 30% of their molecular weight (Cázares-García et al. 2013). Laccases oxidize compounds containing a variety of phenolic, diamines and aromatic amines (Abd El Monssef et al. 2016). In lignocelluloses containing biomass, laccases play an important role in developing a clean biocatalytic process and improve cellulose recovery from feedstocks containing lignocellulose (Avanthi and Banerjee 2016). Additionally, the affinity of laccases for different aromatic compounds make them a promising and attractive tool for de-colouration and detoxification of different synthetic dyes and phenolic pollutants. These chemicals are often a source of water contamination and thus can cause problems to public health and our environment (Anbia and Ghaffari 2011). A combination of laccases and cellulases enhances delignification and thus increases the efficiency of developing enzymatic processes for biofuels and other value-added product generations such as coal solubilization (Chakroun et al. 2010). The extracellular laccase of T. virens is reported for their role in mycoparasitism against the sclerotia of plant pathogens such as Botrytis cinerea and Sclerotinia sclerotiorum (Catalano et al. 2011; Cázares-García et al. 2013).
Fungi of basidiomycetes and ascomycetes division are known to degrade lignin, xenobiotics, chemicals used for guaiacol synthesis and vanillin metabolites at industrial scales (Dekker et al. 2002; Halaburgi et al. 2011; Younes and Sayadi 2011). The wood-rotting fungi such as Trametes spp., Cerrena maxima, Lentinus tigrinus, Coriolopsis polyzona and Pleurotus eryngii are prominent laccase producers (Saloheimo and Niku-Paavola 1991; Morozova et al. 2007; Madhavi and Lele 2009). In general, fungal laccases are known to possess high redox -potential and broad substrate specificity compared to laccases of bacterial origin. The pH optima of fungal laccases is reported at acidic pH, whereas for bacterial laccases, like oxidases, it operates close to neutral-alkaline pH (Kolomytseva et al. 2017). Laccases can target phenolic constituents of lignin and have compatibility to work at industrial pH, in solvents and at especially high temperatures and therefore are potential source for wood delignification for bioethanol production (Shanmugam et al. 2018). The laccase-encoding genes from other fungi such as Pycnoporus sanguineus and Phlebia radiata have been cloned and expressed using the Pcbh1 promoter and the Tcbh1 terminator of T. reesei (Zhao et al. 2018b).
Among ascomycetes, Trichoderma species have been extensively explored for cellulase production (Tsao and Chiang 1983). Trichoderma strains with laccase activity are more efficient in breaking natural substrates than strains without these enzymes (Assavanig et al. 1992). The laccases from ascomycetes have characteristic features which are not present in basidiomycetes laccases. The presence of additional L1–L4 signature domains (Kumar et al. 2003) helps their differentiation from other multicopper oxidases. The laccase activity has been reported in strains of T. atroviride, T. reesei, T. viride, T. longibrachiatum and T. virens (Assavanig et al. 1992; Krastanov et al. 2007; Gochev and Krastanov 2007; Catalano et al. 2011; Cázares-García et al. 2013). In addition, the conidia of T. atroviride, T. viride and T. harzianum are also reported for laccase activity (Holker et al. 2002; Pokorny et al. 2005). Studies on purification and characterization of laccases of extracellular nature have been conducted in T. harzianum (Sadhasivam et al. 2009), T. atroviride (Chakroun et al. 2010) and T. reesei (Levasseur et al. 2010) strains. The infection in Pleurotus ostreatus cultures with T. viride spores is also reported to induce higher laccase activity (Divya et al. 2013).
12.3 Distribution and Identification of CAZy Genes in Trichoderma Genome
The comprehensive information on carbohydrate-active enzymes glycoside hydrolases, carbohydrate esterases, polysaccharide lyases and glycosyltransferases which contribute to the breakdown and modification of glycosidic bonds can be gained from CAZy database. A number of enzymes and active transcripts involved in plant biomass degradation have been identified using genomics, transcriptomics or proteomics approaches. The majority of these transcripts have been identified as glycosyl hydrolases and carbohydrate esterases (Fig. 12.3).
In industrial strain T. reesei, a limited number of carbohydrate active enzymes (CAZymes) have been characterized, whereas genome sequencing revealed presence of several candidates genes which may have been transferred horizontally from bacteria (Häkkinen et al. 2012). Phylogenetic analysis of different CAZy genes has identified around 201 glycoside hydrolase-encoding genes, 22 carbohydrate-encoding esterase genes and 5 polysaccharide lyase genes. Among glycosyl hydrolases, β-glucosidases of GH3, α-galactosidases of GH27 and chitinases of GH18 have been reported in abundance (Häkkinen et al. 2012). In the genome of T. reesei, 61 CAZy families were predicted which exclude family CE10. The complete list of CAZy families in T. reesei can be obtained from a study conducted by Häkkinen et al. (2012). A comparison overview of T. reesei CAZY enzymes with other fungi revealed that cluster containing AGLIII and other four candidate α-galactosidases are restricted to T. reesei. The cluster for β-glucuronidase genes of families GH79, GH18 and GH92 also revealed expansion in T. reesei, whereas families GH43 and Gh61 showed reduction (Häkkinen et al. 2012). The CBHI/CEL7A and CBHII/CEL6A acts in exo-fashion on cellobiohydrolases whereas five endo-acting cellulases such as EGII/CEL5A, EGI/ CEL7B, EGIII/CEL12A, EGV/CEL45A and EGIV/CEL61A are also reported form T. reesei strain (Penttilä et al. 1986; Saloheimo et al. 1994; Saloheimo et al. 1997; Okada et al. 1998). Additionally three putative endoglucanases (CEL74A, CEL61B and CEL5B) were reported (Foreman et al. 2003). In the genome of T. reesei, two β-glucosidases (BGLI/CEL3A and BGLII/ CEL1A) (Barnett et al. 1991; Fowler and Brown 1992; Takashima et al. 1999; Saloheimo et al. 2002a, b) and five β-glucosidases (CEL3B, CEL3D, CEL1B, CEL3C, CEL3E) also have been reported (Foreman et al. 2003). A protein named as swollenin (SWOI) involved in the biomass degradation by disrupting cellulose crystalline structure without the release of sugars has been also reported (Häkkinen et al. 2012). On the other hand, a number of other enzymes such as xylanases (XYNI, XYNII, XYNIII and XYNIV), mannanase (MANI) (Stalbrand et al. 1995), acetyl xylan esterase (Foreman et al. 2003; Margolles-Clark et al. 1996a), α-glucuronidase (GLRI) (Margolles-Clark et al. 1996a), arabinofuranosidases (ABFII and ABFIII) (Margolles-Clark et al. 1996b; Foreman et al. 2003; Herpoël-Gimbert et al. 2008), α-galactosidases (AGLI, AGLII and AGLIII) (Margolles-Clark et al. 1996b; Zeilinger et al. 1993) and β-xylosidase (BXLI) (Margolles-Clark et al. 1996c, d) has also been reported from T. reesei and other filamentoru fungi (Tenkanen et al. 1992; Torronen et al. 1992; Xu et al. 1998; Knob et al. 2010). These proteins are known to play a vital role in breaking xylan-derived oligosaccharides. Also, several novel candidate lignocellulose-degrading genes have been identified from T. reesei genome (Martinez et al. 2008).
Screening of T. harzianum isolate for CAZymes via RNA-Seq and bioinformatics approach revealed around 259 transcripts related to glycoside hydrolases, 101 transcripts for glycosyl transferases, 6 for polysaccharide lyases, 22 for carbohydrate esterases, 42 for auxiliary activities (AAs) and 46 for carbohydrate-binding proteins when cellulose was used as substrate. The highest number of genes has been reported from GH18, GH3, GH16, GH2 and GH5 families. For hemicellulases, 24 glycosyl hydrolases belonging to families GH10, GH11, GH26, GH43, GH54, GH62, GH67 and GH95 were identified. The maximum enzymes were reported from GH43 and GH95 families, whereas the lowest number was identified from GH67, GH62, GH54, GH26 and GH10 families (Ferreira Filho et al. 2017).
12.4 Strain Improvements
Strains of T. reesei have been the topic of investigation for its cellulases. Higher enzyme production cost is one of the key hurdles involved in commercial applications of these enzymes for biofuel production. Screening for high level of cellulase-producing strains is an efficient strategy to address this issue. Due to high porduction cost of enzymes, efforts are required to enhance the production, intrinsic activity and reinforcing the existing biomass degrading enzymes with auxiliary proteins (Wilson 2009; Horn et al. 2012; Peterson and Nevalainen 2012; Hu et al. 2015; Müller et al. 2015; Payne et al. 2015). A number of tools such as genetic engineering advance genetic transformation based on use of marker or marker-free selections, or RNA interference has been discussed by Bischof and Seiboth (2014). T. reesei Rut-C30 and T. reesei D-7 mutants developed by the use of basic chemicals such as ethyl methyl sulfonate (EMS), and other methods such as plasma irradiation are already used for high cellulase production. The filter paper activity and corn starch hydrolysate higher cellulase production in T. reesei strain D-7. Mutant-based study has been successful in obtaining potential cellulase-producing mutants (Zhang et al. 2017). In the last decades, efforts on strain improvement using traditional mutagenesis and screening methods have resulted in T. reesei strains RUT-C30 capable of producing up to 30 g/l of extracellular cellulases (Eveleigh and Montenecourt 1979; Eveleigh 1982) and even producing as high as 100 g/l of extracellular protein (Cherry and Fidantsef 2003). The commercial formulation for enhanced cellulase production such as Novozymes and Dupont are also obtained through mutations in T. reesei. In recent studies, the advancements of molecular tools in gene/genome engineering using specific insertion or deletion or mutation of nucleotides have been explored to meet the growing demands of different biomolecules including enzymes. The discovery of Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (cas) 9 genes (CRISPR/cas9) system has democratized the genome engineering in a flexible manner either at a single- or multi-loci-based genome-wide modification. The CRISPR/cas9 system nowadays has emerged as a powerful tool for strain improvement in filamentous fungi such as T. reesei (Liu et al. 2015; Donohoue et al. 2018).
12.5 Conclusion and Future Prospects
The exploration of microbe’s innate capacity to convert complex polysaccharides into biofuels with octane value is one of the predominant research areas presently. The filamentous fungi such as T. reesei have been widely extensively for cellulase and hemicellulase production. The genetic manipulation of T. reesei using mutagenesis has led to improved strains with higher cellulase production. Advancements in biotechnological tools have significantly contributed in developing alternate and efficient technologies. Enzymatic treatment offers advantage over chemical and physical methods being environmentally friendly. In several studies, either single or a combination of physical and chemical methods of mutations such as UV irradiation, ethyl methanesulfonate and N-Methyl-N′-nitro-N-nitrosoguanidine had been deployed in Trichoderma, Aspergillus and other fungi. The commercial formulation developed by the enzyme industry in companies such as Novozymes and Dupont was obtained through mutations for enhanced cellulase production in T. reesei.
Enzyme-mediated delignification has been used for enhancing enzyme production using rational, semi-rational and directed evolution-based molecular and protein engineering strategies. In rational approach, modification through the use of site direct mutagenesis for lignolytic enzymes such as laccases has been used successfully. Alternatively, a mixture of two filamentous fungi such as T.reesei and A. niger has been found better for cellulase production. Despite the challenge associated with the expression of active recombinant proteins in heterologous system, paucity of signal peptides and expression system, genetic engineering through the use of codon optimization and substitution with unnatural amino acids in recombinant proteins is emerging field and can provide us enzyme systems with better catalytic property and enhanced self-life. However, the concern for low or lack of production of potent hemicellulases and β-glucosidases in T. reesei secretome needs alternative potential strategies which could either replace or supplement T. reesei enzyme system. Therefore, efforts are required for exploring microbial enzymes for biofuel production from agricultural biomass.
References
Abd El Monssef RA, Hassan EA, Ramadan EM (2016) Production of laccase enzyme for their potential application to decolorize fungal pigments on aging paper and parchment. Ann Agric Sci 61(1):145–154
Ahmed S, Aslam N, Latif F, Rajoka MI, Jamil A (2005) Molecular cloning of cellulase genes from Trichoderma harzianum. Front Nat Prod Chem 1:73–75
Anbia M, Ghaffari A (2011) Removal of malachite green from dye wastewater using mesoporous carbon adsorbent. J Iran Chem Soc 8:67–76
Assavanig A, Amornkitticharoen B, Ekpaisal N, Meevootisom V, Flegel TW (1992) Isolation, characterization and function of laccase from Trichoderma. Appl Microbiol Biotechnol 38:198–202
Avanthi A, Banerjee R (2016) A strategic laccase mediated lignin degradation of lignocellulosic feedstocks for ethanol production. Ind Crops Prod 92:174–185
Ayrinhac C, Margeot A, Ferreira NL, Chaabane FB, Monot F, Ravot G et al (2011) Improved saccharification of wheat straw for biofuel production using an engineered secretome of Trichoderma reesei. Org Process Res Dev 15(1):275–278
Barnett CC, Berka RM, Fowler T (1991) Cloning and amplification of the gene encoding an extracellular β-glucosidase from Trichoderma reesei: evidence for improved rates of saccharification of cellulosic substrates. Nat Biotechnol 9(6):562–567
Barros-Rios J, Romani A, Peleteiro S, Garrote G, Ordas B (2016) Second-generation bioethanol of hydrothermally pretreated stover biomass from maize genotypes. Biomass Bioenergy 90:42–49
Bhatia R, Dogra RC, Sharma PK (2002) Construction of green fluorescent protein (GFP)-marked strains of Bradyrhizobium for ecological studies. J Appl Microbiol 93:835–839
Biely P (1985) Microbial xylanolytic systems. Trends Biotechnol 3:286–290
Bilal M, Iqbal HMN, Hu H, Wang W, Zhang X (2018) Metabolic engineering and enzyme-mediated processing: A biotechnological venture towards biofuel production – A review. Renew Sustain Energy Rev 82:436–447
Bischof R, Seiboth B (2014) Molecular tools for strain improvement of Trichoderma spp. In: Gupta VK, Schmoll M, Herrera-Estrella A, Upadhyay RS, Druzhinina I, Tuohy MG (eds) Biotechnology and biology of trichoderma. Elsevier, Amsterdam, pp 179–191
Borin GP, Sanchez CC, de Santana ES, Zanini GK, Dos Santos RAC, de Oliveira PA, de Souza AT, Dal'Mas RMMTS, Riaño-Pachón DM, Goldman GH, Oliveira JVC (2017) Comparative transcriptome analysis reveals different strategies for degradation of steam-exploded sugarcane bagasse by Aspergillus niger and Trichoderma reesei. BMC Genomics 18:501
Buchert JTM, Kantelinen A, Viikari L (1995) Application of xylanases in the pulp and paper industry. Bioresour Technol 50:65–72
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B (2009) The carbohydrate-active enzymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res 37:D233–D238
Catalano V, Vergara M, Hauzenberger JR, Seiboth B, Sarrocco S et al (2011) Use a non-homologous end-joining-deficient strain (delta-ku70) of the biocontrol fungus Trichoderma virens to investigate the function of the laccase gene lcc1 in sclerotia degradation. Curr Genet 57:13–23
Cavaco-Paulo A, Cortez J, Almeida L (1997) The effect of cellulase treatment in textile washing processes. J Soc Dyers Colour 113(7–8):218–222
Cázares-García SV, Vázquez-Garcidueñas MS, Vázquez-Marrufo G (2013) Structural and phylogenetic analysis of laccases from Trichoderma: A bioinformatic approach. PLoS One 8(1):e55295
Chakroun H, Mechichi T, Martinez MJ, Dhouib A, Sayadi S (2010) Purification and characterization of a novel laccase from the ascomycete Trichoderma atroviride: application on bioremediation of phenolic compounds. Process Biochem 45(4):507–513
Chandra M, Kalra A, Sangwan NS, Sangwan RS (2013) Biochemical and proteomic characterization of a novel extracellular β-glucosidase from Trichoderma citrinoviride. Mol Biotechnol 53(3):289–299
Chen LL, Zhang M, Zhang DH, Chen XL, Sun CY, Zhou BC et al (2009) Purification and enzymatic characterization of two β-endoxylanases from Trichoderma sp. K9301 and their actions in xylooligosaccharide production. Bioresour Technol 100:5230–5236
Cheng P, Liu B, Su Y, Hu Y, Hong Y, Yi X, Chen L, Su S, Chu JSC, Chen N, Xiong X (2017) Genomics insights into different cellobiose hydrolysis activities in two Trichoderma hamatum strains. Microb Cell Fact 16(1):1–16
Cherry JR, Fidantsef AL (2003) Directed evolution of industrial enzymes: an update. Curr Opin Biotechnol 14(4):438–443
Chirino-Valle I, Kandula D, Littlejohn C, Hill R, Walker M, Shields M, Wratten S (2016) Potential of the beneficial fungus Trichoderma to enhance ecosystem-service provision in the biofuel grass Miscanthus × giganteus in agriculture. Sci Rep 6:1–8
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Cologna NMD, Gómez-Mendoza DP, Zanoelo FF, Giannesi GC, Guimarães NCA, Moreira LRS, Filho EXF, Ricart CAO (2018) Exploring Trichoderma and Aspergillus secretomes: Proteomics approaches for the identification of enzymes of biotechnological interest. Enzym Microb Technol 109:1–10
de Porciuncula JO, Furukawa T, Shida Y, Mori K, Kuhara S, Morikawa Y, Ogasawara W (2013) Identification of major facilitator transporters involved in cellulase production during lactose culture of Trichoderma reesei PC-3-7. Biosci Biotechnol Biochem 77(5):1014–1022
dos Santos AB, Cervantes FJ, van Lier JB (2007) Review paper on current technologies for decolourisation of textile wastewaters: perspectives for anaerobic biotechnology. Bioresour Technol 98(12):2369–2385
Dahiya N, Tewari R, Hoondal GS (2006) Biotechnological aspects of chitinolytic enzymes: a review. Appl Microbiol Biotechnol 71:773–782
Dekker RFH, Barbosa AM, Sargent K (2002) The effect of lignin-related compounds on the growth and production of laccases by the ascomycete Botryosphaeria sp. Enzym Microb Technol 30:374–380
Demirbas A (2008) Biofuels sources, biofuel policy, biofuel economy and global biofuel projections. Energy Convers Manag 49:2106–2116
Dias AA, Sampaio A, Bezerra RM (2007) Environmental applications of fungal and plant systems: decolourisation of textile wastewater and related dyestuffs. In: Environmental bioremediation technologies. Springer, Berlin/Heidelberg, pp 445–463
Divya LM, Prasanth GK, Sadasivan C (2013) Isolation of a salt tolerant laccase secreting strain of Trichoderma sp. NFCCI-2745 and optimization of culture conditions and assessing its effectiveness in treating saline phenolic effluents. J Environ Sci (China) 25(12):2410–2416
Dodd D, Cann IKO (2009) Enzymatic deconstruction of xylan for biofuel production. Glob Change Biol Bioenergy 1:2–17
Donohoue PD, Barrangou R, May AP (2018) Advances in industrial biotechnology using CRISPR-Cas Systems. Trends Biotechnol 36(2):134–146
Dos Santos Castro L, Pedersoli WR, Antoniêto ACC, Steindorff AS, Silva-Rocha R, Martinez-Rossi NM et al (2014) Comparative metabolism of cellulose, sophorose and glucose in Trichoderma reesei using high-throughput genomic and proteomic analysis. Biotechnol Biofuels 7:41
Eveleigh DE, Montenecourt BS (1979) Increasing yields of extracellular enzymes. Adv Appl Microbiol 25:57–74
Eveleigh DE (1982) Reducing the cost of cellulase production-selection of the hypercellulolytic Trichoderma reesei RUT-C30 mutant. Rutgers University, New Brunswick
Fang H, Xia L (2013) High activity cellulase production by recombinant Trichoderma reesei ZU-02 with the enhanced cellobiohydrolase production. Bioresour Technol 144:693–697
Ferreira Filho JA, Horta MAC, Beloti LL, Dos Santos CA, de Souza AP (2017) Carbohydrate-active enzymes in Trichoderma harzianum: a bioinformatic analysis bioprospecting for key enzymes for the biofuels industry. BMC Genomics 18(1):779
Foreman PK, Brown D, Dankmeyer L, Dean R, Diener S, Dunn-Coleman NS, Goedegebuur F, Houfek TD, England GJ, Kelley AS, Meerman HJ, Mitchell T, Mitchinson C, Olivares HA, Teunissen PJM, Yao J, Ward M (2003) Transcriptional regulation of biomass-degrading enzymes in the filamentous fungus Trichoderma reesei. J Biol Chem 278(34):31988–31997
Fowler T, Brown RD (1992) The bgI1 gene encoding extracellular β-glucosidase from Trichoderma reesei is required for rapid induction of the cellulase complex. Mol Microbiol 6(21):3225–3235
Gao J et al. (2017) Biotechnology for biofuels production of the versatile cellulase for cellulose bioconversion and cellulase inducer synthesis by genetic improvement of Trichoderma Reesei. Biotechnol Biofuels 10:274
Gaurav N, Sivasankari S, Kiran GS, Ninawe A, Selvin J (2017) Utilization of bioresources for sustainable biofuels: a review. Renew Sust Energ Rev 73:205–214
Gerber PJ, Heitmann JA, Joyce TW (1997) Purification and characterization of xylanases from Trichoderma. Bioresour Technol 61(2):127–140
Gochev VK, Krastanov AI (2007) Isolation of laccase producing Trichoderma spp. Bulg J Agric Sci 13:171–176
Gong W, Zhang H, Liu S, Zhang L, Gao P, Chen G, Wang LS (2015) Comparative secretome analysis of Aspergillus niger, Trichoderma reesei, and Penicillium oxalicum during solid-state fermentation. Appl Biochem Biotechnol 177(6):1252–1271
Guo B, Sato N, Biely P, Amano Y, Nozaki K (2016) Comparison of catalytic proper- ties of multiple β-glucosidases of Trichoderma reesei. Appl Microbiol Biotechnol 100:4959–4968
Gupta A, Verma JP (2015) Sustainable bio-ethanol production from agro-residues: a review. Renew Sustain Energy Rev 41:550–567
Gusakov AV, Salanovich TN, Antonov AI, Ustinov BB, Okunev ON, Burlingame R, Emalfarb M, Baez M, Sinitsyn AP (2007) Design of highly efficient cel- lulase mixtures for enzymatic hydrolysis of cellulose. Biotechnol Bioeng 97(5):1028–1038
Häkkinen M, Arvas M, Oja M, Aro N, Penttilä M, Saloheimo M, Pakula TM (2012) Re-annotation of the CAZy genes of Trichoderma reesei and transcription in the presence of lignocellulosic substrates. Microb Cell Fact 11:1–26
Halaburgi VM, Sharma S, Sinha M, Singh TP, Karegoudar TB (2011) Purification and characterization of a thermostable laccase from the ascomycetes Cladosporium cladosporioides and its applications. Process Biochem 46:1146–1152
He J, Yu B, Zhang K, Ding X, Chen D (2009) Expression of endo-1,4-beta-xylanase from Trichoderma reesei in Pichia pastoris and functional characterization of the produced enzyme. BMC Biotechnol 9:56
Henrissat B (1991) A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 280:309–316
Herpoël-Gimbert I, Margeot A, Dolla A, Jan G, Mollé D, Lignon S, Mathis H, Sigoillot J, Monot F, Asther M (2008) Comparative secretome analyses of two Trichoderma reesei RUT-C30 and CL847 hypersecretory strains. Biotechnol Biofuels 1:18
Hofrichter M (2002) Review: lignin conversion by manganese peroxidase (MnP). Enzym Microb Technol 30(4):454–466
Holker H, Dohse J, Hofer M (2002) Extracellular laccases in ascomycetes Trichoderma atroviride and Trichoderma harzianum. Folia Microbiol 47:423–427
Horn SJ, Vaaje-Kolstad G, Westereng B, Eijsink VG (2012) Novel enzymes for the degradation of cellulose. Biotechnol Biofuels 5:45
Hu J, Chandra R, Arantes V, Gourlay K, van Dyk JS, Saddler JN (2015) The addition of accessory enzymes enhances the hydrolytic performance of cellulase enzymes at high solid loadings. Bioresour Technol 186:149–153
Hu JG, Arantes V, Pribowo A, Gourlay K, Saddler JN (2014) Substrate factors that infuence the synergistic interaction of AA9 and cellulases during the enzymatic hydrolysis of biomass. Energ Environ Sci 7:2308–2315
Jeng WY, Wang NC, Lin MH, Lin CT, Liaw YC, Chang WJ et al (2011) Structural and functional analysis of three β-glucosidases from bacterium Clostridium cellulovorans, fungus Trichoderma reesei and termite Neotermes koshunensis. J Struct Biol 173(1):46–56
Jeya M, Zhang WY, Kin IW, Lee JK (2009) Enhanced saccharification of alkali-treated rice straw by cellulase from Trametes hirsuta and statistical optimization of hydrolysis conditions by RSM. Bioresour Technol 100(21):5155–5161
Jørgensen H, Kristensen JB, Felby C (2007) Enzymatic conversion of lignocellulose into fermentable sugars: challenges and opportunities. Biofuels Bioprod Biorefin 1(2):119–134
Juturu V, Wu JC (2012) Microbial xylanases: engineering, production and industrial applications. Biotechnol Adv 30:1219–1227
Kang Q, Appels L, Tan T, Dewil R (2014) Bioethanol from lignocellulosic biomass: currentcurrent findings determine research priorities. Sci World J 2014:1–13
Keshwani DR, Cheng JJ (2009) Switchgrass for bioethanol and other value-added applications: a review. Bioresour Technol 100:1515–1523
Knob A, Terrasan C, Carmona E (2010) β-xylosidases from filamentous fungi: an overview. World J Microbiol Biotechnol 26:389–407
Kolomytseva M, Myasoedova N, Samoilova A, Podieiablonskaia E, Chernykh A, Classen T, Golovleva L (2017) Rapid identification of fungal laccases/oxidases with different pH-optimum. Process Biochem 62:174–183
Krastanov AI, Gochev VK, Girova TD (2007) Nutritive medium dependent biosynthesis of extracellular laccase from Trichoderma spp. Bulg J Agric Sci 13:349–355
Kuhad RC, Gupta R, Singh A (2011) Microbial cellulases and their industrial applications. Enzym Res 2:280696
Kumar S, Phale P, Durani S, Wangikar PP (2003) Combined sequence and structure analysis of the fungal laccase family. Biotechnol Bioeng 83:386–394
Kumar R, Singh S, Singh OV (2008) Bioconversion of lignocellulosic biomass: biochemical and molecular perspectives. J Ind Microbiol Biotechnol 35:377–391
Kurasin M, Valjamae P (2011) Processivity of cellobiohydrolases is limited by the substrate. J Biol Chem 286:169–178
Ladeira Ázar RIS, Morgan T, dos Santos ACF, de Aquino XE, Ladisch MR, Guimarães VM (2018) Deactivation and activation of lignocellulose degrading enzymes in the presence of laccase. Enzyme Microb Technol 109:25–30
Leah R, Kigel J, Svendsen I, Mundy J (1995) Biochemical and molecular characterization of a barley seed β-glucosidase. J Biol Chem 270:15789–15797
Levasseur A, Saloheimo M, Navarro D, Andberg M, Pontarotti P et al (2010) Exploring laccase-like multicopper oxidase genes from the ascomycete Trichoderma reesei: a functional, phylogenetic and evolutionary study. BMC Biochem 11:32
Levasseur A, Drula E, Lombard V, Coutinho PM, Henrissat B (2013) Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes. Biotechnol Biofuels 6(1):41
Li B, Walton JD (2017) Functional diversity for bio- mass deconstruction in family 5 subfamily 5 (GH5_5) of fungal endo-β-1,4-glucanases. Appl Microbiol Biotechnol 101:4093–4101
Li C, Lin F, Li Y, Wei W, Wang H, Qin L et al (2016) A β-glucosidase hyper-production Trichoderma reesei mutant reveals a potential role of cel3D in cellulase production. Microb Cell Fact 15(1):1–13
Liu R, Chen L, Jiang Y, Zhou Z, Zou G (2015) Efficient genome editing in filamentous fungus Trichoderma reesei using the CRISPR/Cas9 system. Cell Discov 1–11:15007
Ma K, Ruan Z (2015) Production of a lignocellulolytic enzyme system for simultaneous bio-delignification and saccharification of corn stover employing co-culture of fungi. Bioresour Technol 175:586–593
Ma L, Zhang J, Zou G, Wang C, Zhou Z (2011) Improvement of cellulase activity in Trichoderma reesei by heterologous expression of a beta- glucosidase gene from Penicillium decumbens. Enzyme Microb Technol 49(4):366–371
Madhavi V, Lele SS (2009) Laccase: properties and applications. Bioresources 4:1694–1717
Mangan D, Cornaggia C, Liadova A, McCormack N, Ivory R, McKie VA et al (2017) Novel substrates for the automated and manual assay of endo-1,4-β-xylanase. Carbohydr Res 445:14–22
Mansur M, Arias ME, Copa-Patiño JL, Flärdh M, González AE (2003) The white-rot fungus Pleurotus ostreatus secretes laccase isozymes with different substrate specificities. Mycologia 95(6):1013–1020
Margeot A, Hahn-Hagerdal B, Edlund M, Slade R, Monot F (2009) New improvements for lignocellulosic ethanol. Curr Opin Biotechnol 20(3):372–380
Margolles-Clark E, Saloheimo M, Siika-aho M, Penttilä M (1996a) The α-glucuronidase-encoding gene of Trichoderma reesei. Gene 172(1):171–172
Margolles-Clark E, Tenkanen M, Luonteri E, Penttilä M (1996b) Three α- galactosidase genes of Trichoderma reesei cloned by expression in yeast. Eur J Biochem 240(1):104–111
Margolles-Clark E, Tenkanen M, Nakari-Setala T, Penttila M (1996c) Cloning of genes encoding alpha-L-arabinofuranosidase and beta-xylosidase from Trichoderma reesei by expression in Saccharomyces cerevisiae. Appl Environ Microbiol 62(10):3840–3846
Margolles-Clark E, Tenkanen M, Soderlund H, Penttila M (1996d) Acetyl xylan esterase from Trichoderma reesei contains an active-site serine residue and a cellulose-binding domain. Eur J Biochem 237(3):553–560
Martinez D, Berka RM, Henrissat B, Saloheimo M, Arvas M, Baker SE, Chapman J, Chertkov O, Coutinho PM, Cullen D, Danchin EGJ, Grigoriev IV, Harris P, Jackson M, Kubicek CP, Han CS, Ho I, Larrondo LF, de Leon AL, Magnuson JK, Merino S, Misra M, Nelson B, Putnam N, Robbertse B, Salamov AA, Schmoll M, Terry A, Thayer N, Westerholm-Parvinen A, Schoch CL, Yao J, Barabote R, Nelson MA, Detter C, Bruce D, Kuske CR, Xie G, Richardson P, Rokhsar DS, Lucas SM, Rubin EM, Dunn-Coleman N, Ward M, Brettin TS (2008) Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol 26(5):553–560
Martinez AT, Ruiz-Dueñas FJ, Martínez MJ, del Río JC, Gutierrez A (2009) Enzymatic delignification of plant cell wall: from nature to mill. Curr Opin Biotechnol 20(3):348–357
Marx IJ, vanWyk N, Smit S, Jacobson D, Viljoen-Bloom M, Volschenk H (2013) Comparative secretome analysis of Trichoderma asperellum S4F8 and Trichoderma reesei Rut C30 during solid-state fermentation on sugarcane bagasse. Biotechnol Biofuels 6(1):1–13
Min SY, Kim BG, Lee C, Hur H-G, Ahn JH (2002) Purification, characterization and cDNA cloning of xylanase from fungus Trichoderma strain SY. J Microbiol Biotechnol 12:890–894
Monclaro AV, Filho EXF (2017) Fungal lytic polysaccharide monooxygenases from family AA9: Recent developments and application in lignocelullose breakdown. Int J Biol Macromol 102:771–778
Morozova OV, Shumakovich GP, Gorbacheva MA, Shleev SV, Yaropolov AI (2007) Blue laccases. Biochemistry (Mosc) 72:1136–1150
Müller H, Berg C, Landa BB, Auerbach A, Moissl-Eichinger C, Berg C (2015) Plant genotype-specific archaeal and bacterial endophytes but similar Bacillus antagonists colonize Mediterranean olive trees. Front Microbiol 6:138
Ning Z, Liu J, Yang J, Lin Y, Yi Y, Lei J, Li M, Yuan HL (2016) Comparative analysis of the secretomes of Schizophyllum commune and other wood-decay basidiomycetes during solid-state fermentation reveals its unique lignocellulose-degrading enzyme system. Biotechnol Biofuels 9(1):1–22
Nitta M, Furukawa T, Shida Y, Mori K, Kuhara S, Morikawa Y, Ogasawara W (2012) A new Zn(II)(2)Cys(6)-type transcription factor BglR regulates beta-glucosidase expression in Trichoderma reesei. Fungal Genet Biol 49(5):388–397
Obeng EM, Adam SNN, Budiman C. Ongkudon CM, Jose RMJ (2017) Lignocellulases: a review of emerging and developing enzymes, systems, and practices. Bioresour Bioprocess 4:41–22
Okada H, Tada K, Sekiya T, Yokoyama K, Takahashi A, Tohda H, Kumagai H, Morikawa Y (1998) Molecular characterization and heterologous expression of the gene encoding a low-molecular-mass endoglucanase from Trichoderma reesei QM9414. Appl Environ Microbiol 64(2):555–563
Paes G, Berrin J-G, Beaugrand J (2012) GH11 xylanases: Structure/function/properties relationships and applications. Biotechnol Adv 30(3):564–592
Payne CM, Knott BC, Mayes HB, Hansson H, Himmel ME, Sandgren M et al (2015) Fungal cellulases. Chem Rev 115:1308–1448
Peciulyte A, Anasontzis GE, Karlström K, Larsson PT, Olsson L (2014) Morphology and enzyme production of Trichoderma reesei rut C-30 are affected by the physical and structural characteristics of cellulosic substrates. Fungal Genet Biol 72:64–72
Penttilä M, Lehtovaara P, Nevalainen H, Bhikhabhai R, Knowles J (1986) Homology between cellulase genes of Trichoderma reesei: complete nucleotide sequence of the endoglucanase I gene. Gene 45(3):253–263
Peterson R, Nevalainen H (2012) Trichoderma reesei RUT-C30– thirty years of strain improvement. Microbiology 158(Pt 1):58–68
Plácido J, Capareda S (2015) Ligninolytic enzymes: a biotechnological alternative for bioethanol production. Bioresour Bioprocess 2(1):23
Pokorny R, Vargovic P, Holker U, Janssen M, Bend J et al (2005) Developmental changes in Trichoderma viride enzymes abundant in conidia and the light-induced conidiation signalling pathway. J Basic Microbiol 45:219–229
Polizeli MLT, Rizzati ACS, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591
Pryor SW, Nahar N (2015) β-glucosidase supplementation during biomass hydrolysis: how low can we go? Biomass Bioenergy 80:298–302
Puls J, Schuseil J (1993) Chemistry of hemicellulose: relationship between hemicellulose structure and enzymes required for hydrolysis. In: Coughlan MP, Hazlewood GP (eds) Hemicellulose and hemicellulases. Portland Press, London, pp 1–27
Qian Y, Zhong L, Hou Y, Qu Y, Zhong Y (2016) Characterization and strain improvement of a hypercellulytic variant, Trichoderma reesei SN1, by genetic engineering for optimized cellulase production in biomass conversion improvement. Front Microbiol 7:1349
Qian Y, Zhong L, Gao J, Sun N, Wang Y, Sun G et al (2017) Production of highly efficient cellulase mixtures by genetically exploiting the potentials of Trichoderma reesei endogenous cellulases for hydrolysis of corncob residues. Microb Cell Fact 16(1):1–16
Ramoni J, Marchetti-Deschmann M, Seidl-Seiboth V, Seiboth B (2017) Trichoderma reesei xylanase 5 is defective in the reference strain QM6a but functional alleles are present in other wild-type strains. Appl Microbiol Biotechnol 101(10):4139–4149
Ravalason H, Grisel S, Chevret D, Favel A, Berrin JG, Sigoillot JC, Herpoël-Gimbert I (2012) Fusarium verticillioides secretome as a source of auxiliary enzymes to enhance saccharification of wheat straw. Bioresour Technol 114(2):589–596
Rosgaard L, Pedersen S, Langston J, Akerhielm D, Cherry JR, Meyer AS (2007) Evaluation of minimal Trichoderma reesei cellulase mixtures on differently pretreated barley straw substrates. Biotechnol Prog 23(6):1270–1276
Sadhasivam S, Savitha S, Swwaminathan K (2009) Redox-mediated decolorization of recalcitrant textile dyes by Trichoderma harzianum WL1 laccase. World J Microbiol Biotechnol 25:1733–1741
Saloheimo M, Niku-Paavola ML (1991) Heterologous production of a ligninolytic enzyme: Expression of the Phlebia radiata laccase gene in Trichoderma reesei. Nat Biotechnol 9(10):987–990
Saloheimo A, Henrissat B, Hoffrén AM, Teleman O, Penttilä M (1994) A novel, small endoglucanase gene, egl5, from Trichoderma reesei isolated by expression in yeast. Mol Microbiol 13(2):219–228
Saloheimo M, Pakula TM (2012) The cargo and the transport system: secreted proteins and protein secretion in Trichoderma reesei (Hypocrea jecorina). Microbiology 158:46–57
Saloheimo M, Nakari-Setälä T, Tenkanen M, Penttilä M (1997) cDNA cloning of a Trichoderma reesei cellulase and demonstration of endoglucanase activity by expression in yeast. Eur J Biochem 249(2):584–591
Saloheimo M, Kuja-Panula J, Ylosmaki E, Ward M, Penttila M (2002a) Enzymatic properties and intracellular localization of the novel Trichoderma reesei beta-glucosidase BGLII (cel1A). Appl Microbiol Biotechnol 68(9):4546–4553
Saloheimo M, Paloheimo M, Hakola S, Pere J, Swanson B, Nyyssönen E, Bhatia A, Ward M, Penttilä M (2002b) Swollenin, a Trichoderma reesei protein with sequence similarity to the plant expansins, exhibits disruption activity on cellulosic materials. Eur J Biochem 269(17):4202–4211
Scheller HV, Ulvskov P (2010) Hemicelluloses. Annu Rev Plant Biol 61:263–289
Seiboth B, Ivanova C, Seidl-Seiboth V (2011) Trichoderma reesei: a fungal enzyme producer for cellulosic biofuels. In: dos Santos Bernardes MA (ed) Biofuel production recent developments and prospects. InTech, Rijeka, pp 309–340
Shanmugam S, Hari A, Ulaganathan P, Yang F, Krishnaswamy S, Wu YR (2018) Potential of biohydrogen generation using the delignified lignocellulosic biomass by a newly identified thermostable laccase from Trichoderma asperellum strain BPLMBT1. Int J Hydrogen Energy 43(7):3618–3628
Sharma V, Shanmugam V (2012) Purification and characterization of an extracellular 24 kDa chitobiosidase from the mycoparasitic fungus Trichoderma saturnisporum. J Basic Microbiol 52:324–331
Sharma V, Bhandari P, Singh B, Bhatacharya A, Shanmugam V (2013) Chitinase expression due to reduction in fusaric acid level in an antagonistic Trichoderma harzianum S17TH. Indian J Microbiol 53(2):214–220
Sharma V, Salwan R, Sharma PN (2016a) Differential response of extracellular proteases of Trichoderma harzianum against fungal phytopathogens. Curr Microbiol 73(3):419–425
Sharma V, Salwan R, Sharma PN, Kanwar SS (2016b) Molecular cloning and characterization of ech46 endochitinase from Trichoderma harzianum. Int J Biol Macromol 92:615–624
Sharma V, Salwan R, Sharma PN, Gulati A (2017a) Integrated translatome and proteome: approach for accurate portraying of widespread multifunctional aspects of Trichoderma. Front Microbiol 8:1602
Sharma V, Salwan R, Sharma PN (2017b) The comparative mechanistic aspects of Trichoderma and Probiotics: Scope for future research. Physiol Mol Plant Pathol 100:84–96
Sharma V, Salwan R, Sharma PN, Kanwar SS (2017c) Elucidation of biocontrol mechanisms of Trichoderma harzianum against different plant fungal pathogens: Universal yet host specific response. Int J Biol Macromol 95:72–79
Sharma V, Salwan R, Shanmugam V (2018a) Unraveling the multilevel aspects of least explored plant beneficial Trichoderma saturnisporum isolate GITX-Panog (C). Eur J Plant Pathol 152(1):169–183
Sharma V, Salwan R, Shanmugam V (2018b) Molecular characterization of β-endoglucanase from antagonistic Trichoderma saturnisporum isolate GITX-Panog (C) induced under mycoparasitic conditions. In Press Pest Biochem Physiol 149:73–80
Shida Y, Yamaguchi K, Nitta M, Nakamura A, Takahashi M, Kidokoro S, Mori K, Tashiro K, Kuhara S, Matsuzawa T et al (2015) The impact of a single- nucleotide mutation of bgl2 on cellulase induction in a Trichoderma reesei mutant. Biotechnol Biofuels 8:230
Silveira FQP, Sousa MV, Ricart CAO, Milagres AMF, Medeiros CL, Filho EXF (1999) A new xylanase from a Trichoderma harzianum strain. J Ind Microbiol Biotechnol 23:682–685
Sindhu R, Kuttiraja M, Prabisha TP, Binod P, Sukumaran RK, Pandey A (2016) Development of a combined pretreatment and hydrolysis strategy of rice straw for the production of bioethanol and biopolymer. Bioresour Technol 215:110–116
Singhania RR, Sukumaran RK, Patel AK, Larroche C, Pandey A (2010) Advancement and comparative profiles in the production technologies using solid-state and submerged fermentation for microbial cellulases. Enzym Microb Technol 46(7):541–549
Song B et al. (2018) Real-Time imaging reveals that lytic polysaccharide monooxygenase promotes cellulase activity by increasing cellulose accessibility. Biotechnol Biofuels 11(1):41
Stahlberg J, Johansson G, Pettersson G (1993) Trichoderma reesei has no true exo-cellulase: all intact and truncated cellulases produce new reducing end groups on cellulose. Biochim Biophys Acta 1157:107–113
Stalbrand H, Saloheimo A, Vehmaanpera J, Henrissat B, Penttila M (1995) Cloning and expression in Saccharomyces cerevisiae of a Trichoderma reesei beta- mannanase gene containing a cellulose binding domain. Appl Environ Microbiol 61(3):1090–1097
Takashima S, Nakamura A, Hidaka M, Masaki H, Uozumi T (1999) Molecular cloning and expression of the novel fungal β-glucosidase genes from Humicola grisea and Trichoderma reesei. J Biochem 125(4):728–736
Tenkanen M, Puls J, Poutanen K (1992) Two major xylanases of Trichoderma reesei. Enzyme Microb Technol 14(7):566–574
Tiwari P, Misra BN, Sangwan NS (2013) β-glucosidases from the fungus Trichoderma: An efficient cellulase machinery in biotechnological applications. Biomed Res Int 2013:203735
Torronen A, Mach RL, Messner R, Gonzalez R, Kalkkinen N, Harkki A, Kubicek CP (1992) The two major xylanases from Trichoderma reesei: characterization of both enzymes and genes. Nat Biotech 10(11):1461–1465
Trichoderma reesei genome database v2.0. http://genome.jgi-psf.org/Trire2/Trire2.home.html
Tsao GT, Chiang L-C (1983) Cellulose and hemicellulose technol-ogy. In: Smith JE, Berry DR, Kristiansen B (eds) The filamentous fungi, vol 4. Edward Arnold, London, pp 296–326
Villares A, Moreau C, Bennati-Granier C, Garajova S, Foucat L, Falourd X et al (2017) Lytic polysaccharide monooxygenases disrupt the cellulose fibers structure. Sci Rep 7:1–9
Vu VV, Marletta MA (2016) Starch-degrading polysaccharide monooxygenases. Cell Mol Life Sci 73(14):2809–2819
Vu VV, Beeson WT, Span EA, Farquhar ER, Marletta MA (2014) A family of starch-active polysaccharide monooxygenases. Proc Natl Acad Sci U S A 111(38):13822–13827
Wan C, Zhou Y, Li Y (2001) Liquid hot water and alkaline pretreatment of soybean straw for improving cellulose digestibility. Bioresour Technol 102(10):6254–6259
Wang B, Xia L (2011) High efficient expression of cellobiase gene from Aspergillus niger in the cells of Trichoderma reesei. Bioresour Technol 102(6):4568–4572
Wang Q, Chen L, Yu D, Lin H, Shen Q, Zhao Y (2017) Excellent waste biomass-degrading performance of Trichoderma asperellum T-1 during submerged fermentation. Sci Total Environ 609:1329–1339
Wilson DB (2009) Cellulases and biofuels. Curr Opin Biotechnol 20(3):295–299
Wong KKY, Tan LUL, Saddler JN (1988) Multiplicity of β-1,4-xylanase in microorganisms: Functions and applications. Microbiol Rev 52:305–317
Xu J, Takakuwa N, Nogawa M, Okada H, Okada H (1998) A third xylanase from Trichoderma reesei PC-3-7. Appl Microbiol Biotechnol 49(6):718–724
Xu J, Zhao G, Kou Y, Zhang W, Zhou Q, Chen G, Liu W (2014) Intracellular beta- glucosidases CEL1a and CEL1b are essential for cellulase induction on lactose in Trichoderma reesei. Eukaryot Cell 13(8):1001–1013
Younes SB, Sayadi S (2011) Purification and characterization of a novel trimeric and thermotolerant laccase produced from the ascomycete Scytalidium thermophilum strain. J Mol Catal B Enzym 73:35–42
Zagrobelny M, Bak S, Moller BL (2008) Cyanogenesis in plants and arthropods. Phytochemistry 69:1457–1468
Zeilinger S, Kristufek D, Arisan-Atac I, Hodits R, Kubicek CP (1993) Conditions of formation, purification, and characterization of an alpha-galactosidase of Trichoderma reesei RUT C-30. Appl Environ Microbiol 59(5):1347–1353
Zhang J, Zhong Y, Zhao X, Wang T (2010) Development of the cellulolytic fungus Trichoderma reesei strain with enhanced beta-glucosidase and filter paper activity using strong artificial cellobiohydrolase 1 promoter. Bioresour Technol 101(24):9815–9818
Zhang Q, Huang H, Han H, Qiu Z, Achal V (2017) Stimulatory effect of in-situ detoxification on bioethanol production by rice straw. Energy 135:32–39
Zhang XY, Zi LH, Ge XM, Li YH, Liu CG, Bai FW (2017) Development of Trichoderma reesei mutants by combined mutagenesis and induction of cellulase by low-cost corn starch hydrolysate. Process Biochem 54:96–101
Zhao J, Xia L (2009) Simultaneous saccharification and fermentation of alkaline- pretreated corn stover to ethanol using a recombinant yeast strain. Fuel Process Technol 90:1193–1197
Zhao J, Xia L (2010) Ethanol production from corn stover hemicellulosic hydrolysate using immobilized recombinant yeast cells. Biochem Eng J 49:28–32
Zhao LL, Ou XM, Chang SY (2016) Life-cycle greenhouse gas emission and energy use of bioethanol produced from corn stover in China: current perspectives and future prospectives. Energy 115:303–313
Zhao C, Zou Z, Li J, Jia H, Liesche J, Chen S, Fang H (2018a) Efficient bioethanol production from sodium hydroxide pretreated corn stover and rice straw in the context of on-site cellulase production. Renew Energy 118:14–24
Zhao J, Zeng S, Xia Y, Xia L (2018b) Expression of a thermotolerant laccase from Pycnoporus sanguineus in Trichoderma reesei and its application in the degradation of bisphenol A. J Biosci Bioeng 125(4):371–376
Zhou J, Wang Y, Chu J, Zhuang Y, Zhang S, Yin P (2008) Identification and purification of the main components of cellulases from a mutant strain of Trichoderma viride T 100-14. Bioresour Technol 99(15):6826–6833
Zhou J, Wang Y, Chu J, Luo L, Zhuang Y, Zhang S (2009) Optimization of cellulase mixture for efficient hydrolysis of steam-exploded corn stover by statistically designed experiments. Bioresour Technol 100(2):819–825
Zhou P, Zhu H, Yan Q, Katrolia P, Jiang Z (2011) Purification and properties of a psychrotrophic Trichoderma sp. xylanase and its gene sequence. Appl Biochem Biotechnol 164:944–956
Zhou Q, Xu J, Kou Y, Lv X, Zhang X, Zhao G, Zhang W, Chen G, Liu W (2012) Differential involvement of beta-glucosidases from Hypocrea jecorina in rapid induction of cellulase genes by cellulose and cellobiose. Eukaryot Cell 11(11):1371–1381
Acknowledgements
The authors are thankful to Chandigarh University for providing necessary infrastructure and SEED Division, Department of Science and Technology, GOI for providing financial benefits (SP/YO/125/2017).
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Sharma, V., Salwan, R. (2019). Extracellular Carbohydrate-Active Enzymes of Trichoderma and Their Role in the Bioconversion of Non-edible Biomass to Biofuel. In: Yadav, A., Singh, S., Mishra, S., Gupta, A. (eds) Recent Advancement in White Biotechnology Through Fungi. Fungal Biology. Springer, Cham. https://doi.org/10.1007/978-3-030-14846-1_12
Download citation
DOI: https://doi.org/10.1007/978-3-030-14846-1_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-14845-4
Online ISBN: 978-3-030-14846-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)