Abstract
In the past years, artificial cellular models for origins-of-life research and synthetic biology have been extensively studied. At this aim, solute-filled lipid vesicles (liposomes) are widely used. Several evidences have been collected about the capture of water-soluble chemicals, the mechanism of vesicle self-reproduction, and the course of (bio)chemical reactions in the vesicle lumen. Among the several fascinating questions which emerged from these studies, here we focus on a peculiar feature, namely, the fact that a spontaneous heterogeneity of vesicle structure often emerges. In other words, vesicle populations created in the laboratory by classical batch methods include very ‘diverse’ vesicles with respect to size, morphology, and – importantly – solute content. The consequences of this between-vesicle diversity are shortly discussed.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Lipid Vesicles in Origin-of-Life Research and in Synthetic Biology
Lipid vesicles (liposomes) are versatile microscopic structures originated from the self-assembly of lipids. They are hollow aqueous micro-compartments, often spherical, whose boundary is a semi-permeable membrane composed by two juxtaposed lipid layers (the so-called ‘bilayer’), where lipids are arranged tail-to-tail. Lipid bilayers form spontaneously as soon as lipids are dissolved in an aqueous solution, and normally they bend and self-seal to form a closed spherical compartment (typical diameters from ca. 30 nm to ca. 100 \(\upmu \)m). Water-soluble compounds, if present in the aqueous phase, become passively encapsulated in the vesicle lumen, whereas hydrophobic and amphiphilic chemicals will self-localize in the membrane (Fig. 1).
Giant lipid vesicles are often used for constructing cell-like systems. The picture shows calcein-filled vesicles whose membranes have been stained by Trypan Blue. Reproduced from [38] according to the CC-BY license. (Color figure online)
The vesicle formation is spontaneous. This is due to entropic factors, and it represents a very remarkable exergonic mechanism for achieving highly ordered structures (the lipid bilayer, and thus the vesicle) at the expenses of the water molecules of the solvent, whose entropy increases. Moreover, the bilayer closure to form a vesicle further dissymmetrizes the system by generating a distinction between the intra-vesicle chemical composition and the external one.
Due to these features, lipid vesicles are used as simplified models of biological cells in two apparently diverse research areas, namely origins of life and synthetic biology. In origins of life investigations, lipid vesicles are taken as primitive cell models. The ‘constructive’ approach [6, 17] represents the unique way to study primitive cells – so to explore their structure, behavior, stability, reactivity, capacity of encapsulating other substances, and so on. On the other hand, the same approach can be adopted in synthetic biology to build simplified cellular models aiming at the engineering and construction of cell-like structures in controlled and programmed way. These ‘minimal synthetic cells’ can help understanding of how molecular biosystems work or can accomplish specific biotechnological tasks. As mentioned, these two research directions, which might appear almost at the antipodes, share as their common ground the theory and the practice of constructing cellular models in the laboratory [37].
2 Inquiring into the Cell Cycle of Primitive Cells
Experimental research on vesicles as primitive cell models was started in the 1990s essentially by the group of Pier Luigi Luisi at the ETH Zürich. Prior models were indeed based on coacervates [27] or microspheres [10] (see [13] for an historical review), but the discovery of liposomes by Alec Bangham [2] and especially the several studies on fatty acid vesicles [11, 15] strongly promoted the shift toward vesicles as biomimetic compartments.
Two elements of innovation were introduced by the Swiss group. Firstly, the target was the assembly of enzyme-filled vesicles (analogous to the Oparin enzyme-filled coacervates). Second, the vesicles were not static, but thanks to the incorporation of new membrane building blocks, they could grow and divide in order to simulate a protocellular proliferation. In other words, from the very beginning it was clear that primitive cell models have to incorporate two of the main features of living cells: an internal reaction network and the grow of the ‘shell’, and ideally perform a sort of primitive cell cycle, as shown in Fig. 2.
The hypothetical cycle of a primitive cell. 0th step: formation of primitive cell by lipid self-assembly and solute capture; 1st step: primitive cell growth at the expenses of external compounds, which should enter into the primitive cell (e.g., by diffusion, by the help of a primitive translocator), and being processed by the internal metabolic network so to produce primitive cell own components, including the membrane, so to allow a growth; 2nd step: division of the grown mother cell into two or more daughter cell, a process that implies solute partition; 3rd step: each one of the daughter cell re-start the cycle, giving rise to primitive cell proliferation. The transformations here depicted independent from the chemical nature of primitive cell components, but obey to an autopoietic mechanism whereby the internalized reaction network produces all its components and the boundary molecules as well. Note that an autopoietic mechanism is a prerequisite for a cell cycle, but does not imply it (an autopoietic cell can stay in a stationary homeostatic state without self-reproduction, but still displaying – as internal activity – a continuous production and degradation of its components, membrane ones included). For a discussion on autopoiesis, and reported experiments on artificial autopoietic self-reproduction and autopoietic homeostasis, see, respectively [21, 41, 43].
The two chemical systems (the ‘core’ and the ‘shell’) should act harmonically so to produce – inside the compartment – all constituents of the primitive cells. Only in this way the daughter cells will be able to re-start the life cycle.
To date, a vesicle system capable of performing a primitive cell cycle as that one depicted in Fig. 2 has not been realized yet. It is worth mentioning that such a system would be autopoietic [23], as it would be able to autonomously produce all its internal components from internal reactions. However several experimental studies have inquired into the several physico-chemical processes underlying that complex pattern.
Commenting on such results lies outside the scope of this paper. The interested readers can find a detailed account in a recently published review [35]. For the present discussion it is enough to mention that great efforts have been made (and are still made) to study internalized reactions either by employing allegedly primitive compounds (ribozymes, short peptides), either ‘modern’ molecules (protein, ribosomes, DNA). Ideally, an internal reaction network, irrespective from its material constitution, in order to be autopoietic should be able to synthesize all components of the network, and the boundary compounds as well. For the latter goal, lipid synthesis inside vesicles can be achieved [8, 19, 30] but not to the point of observing growth and division. Vice versa, if excess boundary-forming compounds (or their precursors) are added from outside, vesicle self-reproduction has been demonstrated [41, 44]. Moreover, it is of great interest the physical understanding of the 0th-step of the cell cycle in Fig. 2, namely, the ‘booting’ self-assembly process that leads to a solute-containing vesicle from separated components.
3 The Need of a Population Perspective: Vesicle ‘Diversity’
Overlaid to the dynamics illustrated in Fig. 2, which refers to the molecular and supra-molecular processes of chemical self-assembly, entrapment, transformation, growth-and-division, there is a central physical aspect that was neglected until a few years ago. This is the inescapable between-vesicle variability that emerges from the microscopic nature of these assemblies. The interplay between their small volume and the stochastic events at the molecular scale originate this additional level of description, that we refer as vesicle ‘diversity’ [39].
Although it was self-evident that a population of vesicles – intended as cell models – is clearly composed by different vesicles, because of different size, different content, different membrane composition, different number of lamellae, different metabolic efficiency, and so on, an adequate level of understanding of this diversity was not available.
Referring again to Fig. 2, vesicle diversity is generated and associated at each step of the cycle and its booting. In particular, at the 0th step, vesicle self-assembly and solute encapsulation generate probably the largest differences, but also the cycle steps such as the growth (due to diverse reaction rates of the network reactions), and the division (when compartmentalized solutes of the grown mother vesicle are distributed among daughter cells).
Each one of the above-mentioned events might generate large between-vesicle structural diversity, which might translate into functional diversity, and thus generating competition between vesicles (between protocells) - a typical ‘biological’ behavior. Moreover, the existence and the observation of diversity and of ‘extremes’ in vesicle populations can be highly informative about the mechanisms of life origins. This diversity is actually a faithful representation of primordial protocellular systems, where populations of lipid compartments spontaneously emerged from the multi-molecular milieu created by a contingent combination of local physico-chemical factors.
On the other hand, it should be noted that when vesicles are employed as synthetic cells (in the biotechnological or synthetic biology arenas), between-vesicle diversity should be kept at the lowest possible level, so to have homogeneous populations where all individuals behave the same manner.
4 The Numbers of Vesicle Diversity
Among the various sources of between-vesicle diversity, the number and the type of solutes encapsulated in their lumen provides the most striking effect of diverse functionality in these cell-like systems.
All steps of the primitive cell cycle (Fig. 2) influence (and are influenced by) the composition of the internal vesicle solution as specified in Table 1.
In the past few years we have explored the diversity of vesicle population with respect to the capture of solutes (0th step: bilayer self-assembly and vesicle formation) and partition of solutes among daughter vesicles (3rd step: vesicle division).
The two cases are somehow similar because the solute capture and solute partition can be modeled by the same maths. In particular, we defined a simple model based on a very reasonable ‘null hypothesis’ \(H_0\), which states that when lipids self-assemble into a vesicle, it is expected that the average number \(\lambda \) of solutes captured in the vesicle lumen is proportional to the vesicle volume. This is equivalent to say that each vesicle ‘samples’ the aqueous solution where it forms, and capture the solutes dissolved in it, with a probability \(p = v/V_{\text {tot}}\), where v is the vesicle volume and \(V_{\text {tot}}\) is the total aqueous volume. Because there are \(N_{\text {tot}}\) molecules in \(V_{\text {tot}}\) (thus \(C_{\text {bulk}} = N_{\text {tot}} / V_{\text {tot}}\), then \(\lambda = p \times N_{\text {tot}}\) and \(c_{\text {vesicle}} = C_{\text {bulk}}\).
It follows that \(\lambda \) can be easily calculated as follows:
where \(N_{\text {A}}\) is the Avogadro’s number. For example, if \(C_{\text {bulk}} = 5\,\upmu \)M, \(\lambda \) is 12.6 and \(\sim 2 \times 10^5\), respectively, for vesicles with diameter of 200 nm and of 5 \(\upmu \)m.
Owing to stochastic fluctuations of the local (microscopic) \(C_{\text {bulk}}\) near the nascent vesicles, it is expected that the actual number n of entrapped solute molecules differs from \(\lambda \) (and \(c_{\text {vesicle}}\), the actual intra-vesicle solute concentration, which is proportional to n, will differ from \(C_{\text {bulk}}\)). As order of magnitude, it is expected that \(\varDelta \lambda /\lambda \) goes as \(\lambda ^{-{1/2}}\) (as obtained by the fluctuations theory). Given a certain \(C_{\text {bulk}}\), high variance should be observed for small vesicles, and small variance for large vesicles. Referring to the above-mentioned cases, \(\varDelta \lambda /\lambda \) should be \(\sim \) 30% and 0.2%, respectively.
As mentioned, according to \(H_0\), the entrapment process is equivalent to random sampling the solution by the nascent vesicle. Due to stochastic fluctuation of the solute concentration in the solution volume, a vesicle will capture n solutes when instead \(\lambda \) are expected on average. It follows that the number of entrapped solute in the vesicle population is distributed according to the Poisson distribution (Eq. 3):
where \(\lambda \) is the mean value, and \(\sqrt{\lambda }\) is the standard deviation. When \(\lambda \) is high, i.e., >10–15 (which in turn means large \(C_{\text {bulk}}\) and/or large V), the Poisson distribution becomes similar to the Normal (Gaussian) distribution.
However, the same mathematics can be applied to the question of solute partition during the division of a grown mother vesicle (2nd step in Fig. 2). If the distribution of solutes in the mother vesicle is spatially homogeneous, the expected number of solutes that will be found in the daughter cells will be proportional to their volume. Again, the \(\lambda \) value can be estimated by Eq. 2 and the distribution by Eq. 3.
The discussion becomes more complicated when solutes of different types (different chemical species) are considered. The simplest hypothesis is that the entrapment of a species does not depend on the presence of the others, so that the co-entrapment probability is simply given by the product of each entrapment probability (of each individual solute). In mathematical terms (for just two solutes A and B):
In general, for k different species, it is the product of k Poissonian terms with averages \(\lambda _{k} = N_{\text {A}} \cdot C_{k,\text {bulk}} \cdot v\) [16, 18, 31].
5 Experimental Evidences
There are not many studies specifically dealing with the question of between-vesicle diversity associated with the vesicle formation, and rare are those dealing with the steps of primitive cell cycle.
In a recent review [1], we have commented quite in detail the available experimental results. Only few authors have studied the solute occupancy distribution inside conventional and giant vesicles in the simple case of one-solute system. At this aim, non-averaging techniques are required, such as microscopy or flow cytometry. On the contrary, numerous reports are based on multiple-solute systems, but the studies do not address the measurement of the solutes concentration, rather focus on the outcome of complex reactions inside vesicles, such as the transcription-translation (TX-TL). TX-TL reactions lead to the production of a protein – a keystone step for this synthetic biology branch.
Anyway, most investigations, but not all, indicate that the between-vesicle diversity is often high, beyond the statistical expectations.
Let us summarize here a series of results, obtained by us, dealing with 0th and 2nd step of the primitive cell cycle (Fig. 2).
5.1 Spontaneous Formation of Solute-Filled Vesicles
Among the four traditional methods for vesicle preparation, the thin lipid film hydration (‘natural swelling’) is certainly the aptest for simulating the spontaneous formation of primitive cell-like structures. It consists in the deposition of a thin lipid film over a surface (e.g., in a glass test tube) followed by a lipid swelling in the presence of a solution of interest.Footnote 1
It is probably evident to most liposomists that vesicles formed by the thin lipid film hydration method are diverse in terms of morphology, size, lamellarity and – especially – solute content. No detailed discussions have been reported on the reasons for the latter heterogeneity (the width of the solute occupancy distribution) and on the expected-versus-observed variance.
We studied the encapsulation of ferritin, ribosomes and ribo-peptidic complexes (reviewed in [32]) in conventional phospholipid vesicles prepared by the film method. These are convenient macromolecular solutes – especially ferritin – because they can be visualized by cryo-transmission electronmicroscopy, and directly counted. Thus, the intravesicle solute occupancy distribution can be determined by analyzing a large number of images. Our expectation was to find the classical bell-shaped distribution (Poisson or Gaussian), but we surprisingly found a power-law distribution (Fig. 3) and interesting ‘super-filled’ vesicles whose formation cannot be easily explained by stochastic fluctuations.
Encapsulation of ferritin inside conventional lipid vesicles. (A-B-C) Typical super-filled vesicles which contain a higher-than-expected number of ferritin molecules. Such special vesicles are present in minor amount (ca. 0.1%). In particular, the intravesicle ferritin concentration is 8.5\(\times \) (A), 3.8\(\times \) (B), 11.8\(\times \) (C) the expected bulk concentration (\(C_{\text {bulk}}\), cf. Eq. 2). In the bottom panel (D), the expected (empty symbols) and the measured (filled symbols) solute occupancy distribution is reported. Note the logarithmic scale on the y-axis. The experimental data, when plotted on a double logarithmic plot, give a straight line (a power-law distribution). Reproduced from [22] with the permission of Wiley.
According to these observation, that have been confirmed qualitatively (but not quantitatively) when fluorescent proteins were encapsulated inside small (1–2 \(\upmu \)m) giant vesicles [5, 39], the actual solute occupancy distribution might vary significantly from expectations, and in particular the formation of solute-filled vesicles, although rare, can shed light on the self-organization drive to the onset of early cells.
When two or more solutes are considered, Eq. 4 extended to the \(k = 80\) different solutes of TX-TL reaction has shown limitations, because we recorded again solute rich vesicles even if statistically their occurrence should have been extremely rare. The solute-rich vesicles were those capable of synthesizing a protein. This was proved by running experiments in those conditions that give small \(\lambda _{k}\) values in two different ways, namely, using small vesicles (small v) [31], or using small \(C_{k,\text {bulk}}\) [36]. When conventional vesicles, with diameter of ca. 200 nm have been used, it results that although most vesicles cannot synthesize proteins, some still can, despite the very unfavorable co-entrapment probability (of the order of 10\(^{-26}\)). It has been shown that a power-law model can successfully explain the observation of TX-TL reactions in small GVs [25].
An attempt to explain the physics of solute entrapment and the reported power law [22] has been proposed, based on the Cox’s theory of renewal point processes [28], but a definitive clarification is still missing.
5.2 Division of a Grown Mother Vesicle into Two or More Daughter Vesicles
In order to model the spontaneous division of primitive cells we devised a mechanical approach whereby solute-filled phospholipid vesicles were fragmented in many small daughter by a mechanical non-spontaneous process called extrusion [9]. Although this strategy clearly represents a crude approximation of the spontaneous vesicle self-reproduction [41, 44], it includes the core mechanism of interest for our discussion, i.e., the stochastic partition of encapsulated solutes in the daughter vesicles. In the past, extrusion was applied to primitive cell models, providing information about average solute retention [14, 34].
We investigated how a population of giant vesicles (average diameter 5–8 \(\upmu \)m) behave when vesicles are extruded to give small (0.8 \(\upmu \)m) daughter vesicles. The study was carried out by firstly encapsulating fluorescent solutes of different size inside giant vesicles, then by extruding giant vesicles and measuring the internal solute concentration of the daughter vesicles (by confocal microscopy). According to \(H_0\) the solute concentration in the mother vesicles and in the daughter ones should be the same, although the spread around the mean value can differ due to stochastic fluctuations.
Figure 4, taken from [9], summarizes the experimental results. In particular, the plots compare the experimental solute occupancy distribution with the theoretical one, obtained by stochastic partition of the solutes initially present in giant vesicles. In this manner not only the average but also the variance of the expected and measured distributions is compared.
Comparison between the solute occupancy distribution as obtained experimentally (grey area) and by numerical modeling (red lines). The distributions refer to extruded filled with (a) pyranine; (b) calcein; (c) fluorescein-marked bovine serum albumin; (d) fluorescein-marked dextran. Reproduced from [9] with permission from The Royal Society of Chemistry. (Color figure online)
Results show that the solute partition pattern depends on the solute size. Small solutes such as pyranine (0.52 kDa) and calcein (0.62 kDa) are largely lost during the extrusion of giant vesicles (−48% and −76%, respectively).Footnote 2 In contrary, large solutes such as bovine serum albumin (69 kDa) and dextran (150 kDa) – both marked by fluorescein) – experience a lower lost (−11% and −30%, respectively).
Data have been explained on the basis of the different diffusion coefficient (D) of small and large solutes. The high D of small solutes implies a greater movement in the time interval of \(\sim \)1 ms when the daughter vesicles seal their membrane (after being pinched off from the mother vesicle). In most cases then, the solutes will diffuse away from the volume captured by the daughter vesicle. Vice versa, large solutes will mainly stay and be captured due to their low D.
6 Relevance, Future Challenges, and the Role of Bioinformatics
In this work we have emphasized the role of vesicle diversity as derived from spontaneous processes such as vesicle formation, growth, division. We have shown that when this aspect is specifically investigated, interesting observations are collected – but most of them are currently without a clear explanation.
What are the very generative mechanisms and what is the role of vesicle diversity when primitive cell models or synthetic cells are built in the laboratory?
While undergoing and future research tools need to be sharpened in order to solve the first question, it is already evident that vesicle diversity is a conceptually important factor to consider. Focusing on this topic represents already a cultural shift that can lead to interesting developments. For example, the emergence of primitive cells, often discussed from an individualist viewpoint should be instead discussed from a population perspective, and the factors that amplify or propagate any between-individual variations should be taken into account.
The new vision based on protocell ensembles makes the story more interesting. The duality between competition and cooperation, two central ideas in biology, makes sense only when structural and functional diversities are explicitly considered. In the prebiotic context, such a diversity is strongly linked to structure and precedes the onset of genotype variations. The structural difference between protocells is more radical as it originates from the mechanisms of vesicle formation (by self-assembly or by division of a parent vesicle). Therefore, protocell diversity has realistically impacted on early pre-biological evolution.
Protocell assemblies models have been already put forward [4, 12, 33] but more research is needed. Vesicles could be included in a gel matrix [29] so to create large tissue-like models.
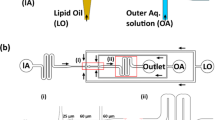
Synthetic biology applications, on the other hand, require highly homogeneous samples. Although most of published work generally rely on batch vesicle preparation methods (and thus display high between-vesicle variability), modern trends suggest that the microfluidic technologies [7, 40, 42] will revolution the experimental approaches to synthetic cells. We expect indeed that synthetic cell construction by microfluidics will overcome soon the traditional batch methods [35].
The evidences here summarized strongly call for integrate theoretical, experimental, and stochastic modeling approaches [3, 9, 20, 24, 25].
Bioinformatics can be an important tool for understanding the behavior of vesicle populations, and realize a qualitative jump in the direction of a systemic and multi-scale view. Bioinformatics can be introduced at three different levels.
-
1.
In the theory of minimal living systems, the conceptualization of the so-called “organizational closure”, which characterize autopoietic systems, has been central. Now that synthetic approaches have become realistic, organizational closure can be designed by bioinformatics. What is the minimal complexity of a reaction network capable of self-sustainment? How to understand whether the production of certain component(s) can be omitted from the autopoietic system and instead be ‘outsourced’ to the environment? What are the network topology features that define autonomy?
-
2.
As modeling support for the experimental approaches, numerical simulations can be very useful in two important aspects that determine the course of micro-compartmentalized reactions, namely extrinsic and intrinsic stochastic effects. The phenomena discussed in Sect. 5 are based on extrinsic stochastic effects. Reactions inside microcompartments, however, are subjected to relevant intrinsic stochastic effects, due to the small number of molecules confined in the vesicle lumen or in the membrane. This fact implies that the reactions occurring in primitive cell models and synthetic cells are best simulated by stochastic kinetics (i.e., master equations, Monte-Carlo methods, etc.), rather than with ordinary differential equations.
-
3.
In the emerging field of bio-chemical-based Information Technologies (bio-chem-ITs) and Molecular Communications [26], bioinformatics becomes functional to develop synthetic cells capable of manipulating chemical signals in terms of the communication theory, from relatively simple systems such as signal exchanges between synthetic cells, to coordinated or synchronized behavior of synthetic cell assemblies, up to more complex pattern as can be a rudimentary differentiation – which are experimental goals probably not so distant in the future.
7 Concluding Remarks
In this paper we have emphasized and discussed the important but often neglected topic of between-vesicle diversity, referred to the current experimental approaches to primitive cell models and synthetic cells. Experiments show that experimentally obtained vesicle populations display quite high variances (except when new microfluidics methods are employed). Accordingly, the definition of an ‘average behavior’ might become almost meaningless. But instead of being a bad news, these evidences offer an opportunity for better understanding the generative mechanism of protocells, the paths that lead to prebiotic evolution, and the early cell communities.
Finally, we remark that as laboratory-made artificial cells are very simple systems when compared to biological living cells, the application of rigorous bioinformatic models is not only helpful, but also very meaningful.
Notes
- 1.
Ideally, one would like to have a single lipid monolayer deposited over a large surface so that all parts of the film would experience the same conditions. This corresponds, in most cases, to work well below the \(\mu \)M lipid concentration range, with consequent vesicle losses and other impractical complications. Thus, in the most common experimental conditions the film is rarely so perfect and different regions of the film will experience different micro-environments. Actually, realistic laboratory conditions might affect the measured heterogeneity of vesicle formation paths.
- 2.
Note that this loss does not refer to the volume loss which follows from the vesicle size reduction, but it is an authentic concentration reduction due to the reduction of the average number per unit of volume.
References
Altamura, E., Carrara, P., D’Angelo, F., Mavelli, F., Stano, P.: Extrinsic stochastic factors (solute partition) in gene expression inside lipid vesicles and lipid-stabilized water-in-oil droplets. Synth. Biol. (OUP) p. (2018, in press)
Bangham, A.D., Horne, R.W.: Negative staining of phospholipids and their structural modification by surface-active agents as observed in the electron microscope. J. Mol. Biol. 8, 660–668 (1964)
Calviello, L., Stano, P., Mavelli, F., Luisi, P.L., Marangoni, R.: Quasi-cellular systems: stochastic simulation analysis at nanoscale range. BMC Bioinf. 14, S7 (2013). https://doi.org/10.1186/1471-2105-14-S7-S7
Carrara, P., Stano, P., Luisi, P.L.: Giant vesicles “colonies”: a model for primitive cell communities. Chembiochem Eur. J. Chem. Biol. 13(10), 1497–1502 (2012). https://doi.org/10.1002/cbic.201200133
D’Aguanno, E., Altamura, E., Mavelli, F., Fahr, A., Stano, P., Luisi, P.L.: Physical routes to primitive cells: an experimental model based on the spontaneous entrapment of enzymes inside micrometer-sized liposomes. Life (Basel, Switzerland) 5(1), 969–996 (2015). https://doi.org/10.3390/life5010969
Damiano, L., Canamero, L.: The frontier of synthetic knowledge: toward a constructivist science. World Futures 68(3), 171–177 (2012). https://doi.org/10.1080/02604027.2012.668409
Elani, Y.: Construction of membrane-bound artificial cells using microfluidics: a new frontier in bottom-up synthetic biology. Biochem. Soc. Trans. 44(3), 723–730 (2016). https://doi.org/10.1042/BST20160052
Exterkate, M., Caforio, A., Stuart, M.C.A., Driessen, A.J.M.: Growing membranes in vitro by continuous phospholipid biosynthesis from free fatty acids. ACS Synth. Biol. 7(1), 153–165 (2018). https://doi.org/10.1021/acssynbio.7b00265
Fanti, A., Gammuto, L., Mavelli, F., Stano, P., Marangoni, R.: Do protocells preferentially retain macromolecular solutes upon division/fragmentation? A study based on the extrusion of POPC giant vesicles. Integr. Biol. (Camb) 10(1), 6–17 (2018). https://doi.org/10.1039/c7ib00138j
Fox, S.V.: Self-assembly of the protocell from a self-ordered polymer. J. Sci. Ind. Res. 27, 267–274 (1968)
Gebicki, J.M., Hicks, M.: Ufasomes are stable particles surrounded by unsaturated fatty acid membranes. Nature 243(5404), 232–234 (1973)
Hadorn, M., Boenzli, E., Sørensen, K.T., De Lucrezia, D., Hanczyc, M.M., Yomo, T.: Defined DNA-mediated assemblies of gene-expressing giant unilamellar vesicles. Langmuir 29(49), 15309–15319 (2013). https://doi.org/10.1021/la402621r
Hanczyc, M.M.: The early history of protocells - the search for the recipe of life. In: Rasmussen, S., et al. (eds.) Protocells Bridging Nonliving and Living Matter. MIT Press, Cambridge (2009)
Hanczyc, M.M., Fujikawa, S.M., Szostak, J.W.: Experimental models of primitive cellular compartments: encapsulation, growth, and division. Science 302(5645), 618–622 (2003). https://doi.org/10.1126/science.1089904
Hargreaves, W.R., Deamer, D.W.: Liposomes from ionic, single-chain amphiphiles. Biochemistry 17(18), 3759–3768 (1978)
Hosoda, K., Sunami, T., Kazuta, Y., Matsuura, T., Suzuki, H., Yomo, T.: Quantitative study of the structure of multilamellar giant liposomes as a container of protein synthesis reaction. Langmuir 24(23), 13540–13548 (2008). https://doi.org/10.1021/la802432f
Kaneko, K.: Life: An Introduction to Complex Systems Biology. UCS. Springer, Heidelberg (2006). https://doi.org/10.1007/978-3-540-32667-0
Kita, H., et al.: Replication of genetic information with self-encoded replicase in liposomes. ChemBioChem. 9(15), 2403–2410 (2008). https://doi.org/10.1002/cbic.200800360
Kuruma, Y., Stano, P., Ueda, T., Luisi, P.L.: A synthetic biology approach to the construction of membrane proteins in semi-synthetic minimal cells. Biochim. Biophys. Acta 1788(2), 567–574 (2009). https://doi.org/10.1016/j.bbamem.2008.10.017
Lazzerini-Ospri, L., Stano, P., Luisi, P., Marangoni, R.: Characterization of the emergent properties of a synthetic quasi-cellular system. BMC Bioinform. 13(Suppl 4), S9 (2012). https://doi.org/10.1186/1471-2105-13-S4-S9
Luisi, P.L.: Autopoiesis: a review and a reappraisal. Naturwissenschaften 90(2), 49–59 (2003). https://doi.org/10.1007/s00114-002-0389-9
Luisi, P.L., Allegretti, M., Pereira de Souza, T., Steiniger, F., Fahr, A., Stano, P.: Spontaneous protein crowding in liposomes: a new vista for the origin of cellular metabolism. ChemBioChem 11(14), 1989–1992 (2010). https://doi.org/10.1002/cbic.201000381
Maturana, H.R., Varela, F.J.: Autopoiesis and Cognition: The Realization of the Living. D. Reidel Publishing Company, 1st edn. (1980)
Mavelli, F.: Stochastic simulations of minimal cells: the Ribocell model. BMC Bioinform. 13(Suppl 4), S10 (2012). https://doi.org/10.1186/1471-2105-13-S4-S10
Mavelli, F., Stano, P.: Experiments on and numerical modeling of the capture and concentration of transcription-translation machinery inside vesicles. Artif. Life 21(4), 445–463 (2015)
Nakano, T., Eckford, A.W., Haraguchi, T.: Molecular Communications. Cambridge University Press, Cambridge UK (2013)
Oparin, A.I.: The pathways of the primary development of metabolism and artificial modeling of this development in coacervate drops. In: The Origins of Prebiological Systems and of their Molecular Matrices, pp. 331–345. S. W. Fox, New York, Academic Press edn. (1965)
Paradisi, P., Allegrini, P., Chiarugi, D.: A renewal model for the emergence of anomalous solute crowding in liposomes. BMC Syst. Biol. 9(Suppl 3), S7 (2015). https://doi.org/10.1186/1752-0509-9-S3-S7
Rampioni, G., et al.: Synthetic cells produce a quorum sensing chemical signal perceived by Pseudomonas aeruginosa. Chem. Commun. 54, 2090–2093 (2018). https://doi.org/10.1039/C7CC09678J
Schmidli, P., Schurtenberger, P., Luisi, P.: Liposome-mediated enzymatic-synthesis of phosphatidylcholine as an approach to self-replicating liposomes. J. Am. Chem. Soc. 113(21), 8127–8130 (1991). https://doi.org/10.1021/ja00021a043
Pereira de Souza, T., Stano, P., Luisi, P.L.: The minimal size of liposome-based model cells brings about a remarkably enhanced entrapment and protein synthesis. Chembiochem 10(6), 1056–1063 (2009). https://doi.org/10.1002/cbic.200800810
de Souza, T.P., Fahr, A., Luisi, P.L., Stano, P.: Spontaneous encapsulation and concentration of biological macromolecules in liposomes: an intriguing phenomenon and its relevance in origins of life. J. Mol. Evol. 79(5–6), 179–192 (2014). https://doi.org/10.1007/s00239-014-9655-7
Souza, T.P.D., et al.: Vesicle aggregates as a model for primitive cellular assemblies. Phys. Chem. Chem. Phys. 19(30), 20082–20092 (2017). https://doi.org/10.1039/C7CP03751A
Stano, P., Wehrli, E., Luisi, P.L.: Insights into the self-reproduction of oleate vesicles. J. Phys. Condens. Matter 18(33), S2231 (2006). https://doi.org/10.1088/0953-8984/18/33/S37
Stano, P., Carrara, P., Kuruma, Y., de Souza, T.P., Luisi, P.L.: Compartmentalized reactions as a case of soft-matter biotechnology: synthesis of proteins and nucleic acids inside lipid vesicles. J. Mater. Chem. 21(47), 18887–18902 (2011). https://doi.org/10.1039/c1jm12298c
Stano, P., D’Aguanno, E., Bolz, J., Fahr, A., Luisi, P.L.: A remarkable self-organization process as the origin of primitive functional cells. Angew. Chemie-Int. Ed. 52(50), 13397–13400 (2013). https://doi.org/10.1002/anie.201306613
Stano, P., Luisi, P.: Theory and construction of semi-synthetic minimal cells. In: Nesbeth, D.N. (ed.) Synthetic Biology Handbook, pp. 209–258. CRC Press, London (2016)
Stano, P., Mavelli, F.: Protocells models in origin of life and synthetic biology. Life 5(4), 1700–1702 (2015). https://doi.org/10.3390/life5041700
Stano, P., et al.: Recent biophysical issues about the preparation of solute-filled lipid vesicles. Mech. Adv. Mater. Struct. 22(9), 748–759 (2015). https://doi.org/10.1080/15376494.2013.857743
van Swaay, D., deMello, A.: Microfluidic methods for forming liposomes. Lab. Chip. 13(5), 752–767 (2013). https://doi.org/10.1039/c2lc41121k
Walde, P., Wick, R., Fresta, M., Mangone, A., Luisi, P.: Autopoietic self-reproduction of fatty-acid vesicles. J. Am. Chem. Soc. 116(26), 11649–11654 (1994). https://doi.org/10.1039/c2lc41121k
Weiss, M., et al.: Sequential bottom-up assembly of mechanically stabilized synthetic cells by microfluidics. Nat. Mater. 17(1), 89–96 (2018). https://doi.org/10.1038/nmat5005
Zepik, H.H., Blochliger, E., Luisi, P.L.: A chemical model of homeostasis. Angew. Chem.-Int. Edit. 40(1), 199–202 (2001)
Zhu, T.F., Szostak, J.W.: Coupled growth and division of model protocell membranes. J. Am. Chem. Soc. 131(15), 5705–5713 (2009). https://doi.org/10.1021/ja900919c
Acknowledgments
Collaboration among the authors has been fostered by the European COST Action CM1304 Emergence and Evolution of Complex Chemical Systems.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Stano, P., Marangoni, R., Mavelli, F. (2019). Experimental Evidences Suggest High Between-Vesicle Diversity of Artificial Vesicle Populations: Results, Models and Implications. In: Bartoletti, M., et al. Computational Intelligence Methods for Bioinformatics and Biostatistics. CIBB 2017. Lecture Notes in Computer Science(), vol 10834. Springer, Cham. https://doi.org/10.1007/978-3-030-14160-8_17
Download citation
DOI: https://doi.org/10.1007/978-3-030-14160-8_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-14159-2
Online ISBN: 978-3-030-14160-8
eBook Packages: Computer ScienceComputer Science (R0)