Abstract
Mutations in the dystrophin gene cause Duchenne muscular dystrophy, a hereditary condition that leads to degeneration of both skeletal and cardiac muscle. All DMD patients inevitably develop heart disease (cardiomyopathy), and 30 % of them succumb to congestive heart failure. The mechanism of DMD heart failure is still elusive, partly due to the hard-to-access heart cells from patients. Because no effective treatment exists for DMD heart disease, patients with DMD represent an unmet medical need. To address this need, we modeled heart disease in DMD using a regenerative medicine technology called cellular reprogramming. Reprogramming involves converting an adult somatic cell into a pluripotent stem cell, called induced pluripotent stem cell (iPSC). IPSCs are further transformed or differentiated into heart cells for study. Reprogramming technology provides vast amounts of heart cells, offering an unprecedented opportunity to study genetic heart diseases like DMD heart failure. Establishing this regenerative medicine platform will not only facilitate exploration of disease etiology, but also provide unlimited personalized material to screen therapeutic compounds for individual patients.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Muscular dystrophy
- Duchenne
- Cardiomyopathy
- Dilated
- X-Linked
- Induced pluripotent stem cells
- Myocytes
- Cardiac
- Dystrophin
- Drug evaluation
- Preclinical
6.1 Introduction
Mutations in the dystrophin gene cause dystrophinopathy, a hereditary disorder with variable allelic clinical presentations including Duchenne muscular dystrophy (DMD), Becker muscular dystrophy (BMD), and X-linked dilated cardiomyopathy (XLDC). Though the disease is well known for the skeletal muscle involvement, most patients develop cardiomyopathy and eventually succumb to congestive heart failure. With ventilatory support preventing respiratory-related mortality, the greater cardiac workload associated with longer life expectancy is believed to increase the incidence of heart failure. Currently there are no effective therapies to contain the decline of heart function in these patients. Thus, it is urgent to devise new strategies to prevent, halt, or reverse the cardiomyopathy. To address this unmet medical need, we modeled dystrophin-deficient cardiomyopathy using cellular reprogramming technology, which involves converting adult somatic cells into pluripotent stem cells, termed induced pluripotent stem cells (iPSCs). Upon further induction, iPSCs can give rise to vast number of heart cells that can be used to study disease etiology and screen therapeutic compounds.
6.2 Dystrophin-Deficient Cardiomyopathy
6.2.1 Background
Dystrophinopathy refers to a group of genetic disorders, encompassing DMD, its milder variant BMD, and XLDC [1]. Though varied in clinical presentation, these diseases share common gene defects in dystrophin resulting in varying levels of dystrophin deficiency. Cardiac symptoms are invariably associated with all dystrophinopathy patients. Rare mutations cause localized dystrophin protein defects restricted to the heart, making the heart the only affected organ in XLDC. More commonly, mutations cause devastating skeletal muscle weakness for muscular dystrophy patients that overshadows any underlying cardiac abnormality. 98 % of DMD patients develop cardiac abnormalities, while congestive heart failure (CHF) and sudden cardiac death account for 10–20 % of the mortality in DMD patients. In contrast to their relatively mild skeletal muscle involvement, many BMD patients develop evident cardiac symptoms likely due to the cardiac workload imposed by the longer life span and vigorous physical activities. Cardiac complications are estimated to account for up to 50 % of the mortality in patients with DMD [1]. The mortality associated with cardiac failure is expected to rise even further, due to improved respiratory management that decreases fatal respiratory failure and extends the patients’ life span.
Other reports have highlighted the linkage of dystrophin with several forms of acquired cardiomyopathy [2–4], suggesting that dystrophin protein remodeling may represent a common pathway underlying contractile dysfunction in failing hearts [5]. Thus, restoring normal function of the dystrophin-associated glycoprotein complex (DGC) could serve as a potential therapeutic target for heart failure patients.
6.2.1.1 Dystrophin Gene and the Mutations
The gene encoding dystrophin locates to the X chromosome, spanning 79 exons and covering 2.4 Mbp [6]. While the shortest isoform, DP71, is ubiquitously expressed in multiple tissues, the full-length transcript variant Dp427m is mainly expressed in muscle tissue, including the heart [7]. By forming a dystrophin-associated glycoprotein complex (DGC) together with the sarcolemma, dystrophin mainly functions as the hub to connect the intracellular actin filament with the extracellular matrix (ECM), providing mechanical support to reinforce the sarcolemma. On the other hand, dystrophin also serves as a scaffold protein to organize molecules in proper position for function, such as membrane receptors and signaling proteins like neuronal nitric oxide synthase (nNOS) [8]. Hence, dystrophin plays a critical role in both mechanical membrane support and in proper function of certain cell signaling pathways.
Numerous genetic mutations have been identified across the whole length of the dystrophin gene, but mainly enriched within two “hotspots.” The most common region, 3’ end hotspot lies within exon 45–55 with genomic breakpoints at intron 44, while the 5’ end hotspot covers exons 2–19 with breakpoints in intron 2 and 7 [7]. Exon deletions and duplications are the most common forms of mutations. The severity of symptoms heavily depends on the maintenance of the open reading frame, rather than the size of mutated genomic regions. The frame shift hypothesis suggests that mutations maintaining the original open reading frame lead to the production of a truncated but partially functional protein, which usually leads to a milder clinical presentation in BMD. On the other hand, mutations shifting the reading frame completely cease protein production [7] with prominent disease manifestations.
6.2.1.2 Clinical Symptoms and Management
Cardiomyopathy in dystrophic patients is largely underdiagnosed and poorly managed [9], partly due to the fact that symptoms dynamically progress over time. To monitor disease progression, electrocardiogram, echocardiogram, and magnetic resonance imaging (MRI) can be used to determine the appropriate time and course of intervention. Cardiac manifestations associated with dystrophin deficiency include rhythmic disturbance, organ structural alteration, and hemodynamic abnormalities. A typical disease course includes three distinct but continuous stages [10]. The preclinical stage usually presents with an abnormal electrocardiogram, demonstrating a variety of findings such as sinus tachycardia, premature contractions, and conduction delays [1, 11]. As the disease progresses, imaging finds evidence of cardiac hypertrophy such as increase of ventricular septal thickness and left ventricular free wall/septum ratio in the hypertrophic stage. In the advanced dilated cardiomyopathy stage, echocardiogram usually reveals ventricular dilation coupled with hemodynamic disturbances that eventually progresses to congestive heart failure.
Available therapies are limited and palliative. Conventional anti-heart failure regimens, including ACE inhibitors (ACEI), angiotensin II receptor blockers (ARBs), beta-adrenergic receptor blockers, and aldosterone antagonists, are typically prescribed in an attempt to delay heart function decline [12–14]. Corticosteroids also demonstrated benefit in several reports, although most of these studies are retrospective observations with limited sample size. More recently a cohort of 86 patients was retrospectively analyzed, and the investigators concluded on top of ACEI therapy that the use of steroids was associated with a 76 % decrease of mortality, largely driven by the reduction of heart failure-associated death. However, the corticosteroid-treated group received ACEI treatment 3 years earlier, which may be the alternative explanation for the observed effect [15]. Other therapeutic modalities, such as pacemaker [16–18], ventricular assist device (VAD) [19–22], and cardiac resynchronization therapy (CRT) [23–25], may be beneficial in decreasing fatal arrhythmias and temporarily boost heart function.
With existing regimens, the majority of patients still face inevitable cardiac failure. This cruel reality makes it imperative to pursue new strategies. However, this attempt is largely hindered by the lack of a reliable disease model. Existing animal models such as the mdx mouse fail to precisely reproduce human pathophysiology. For example, pharmacotherapy proven effective in mdx mice failed to demonstrate the equivalent efficacy in DMD patients and even worsened heart performance [26]. On the other hand, utilization of primary human cardiomyocytes is limited by risky isolation procedures and poor proliferation capacity of cells captured from human biopsy material. Therefore, a disease model system that closely mimics human symptoms and is capable of predicting in vivo efficacy is invaluable.
6.2.2 Pathogenesis
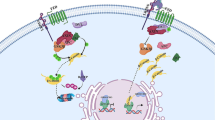
A plethora of evidence suggests that the outer cell membrane of the skeletal or cardiac muscle cell, the sarcolemma, is abnormally susceptible to mechanical stress in the face of dystrophin deficiency. This “vulnerable membrane” is characterized by the decrease of membrane stability when subjected to mechanical stretch during contraction, predisposing muscle cells to rupture. On the other hand, a spectrum of abnormal phenotypes across multiple physiological domains have also been linked to the absence of dystrophin, including but not limited to disturbance of calcium homeostasis, mitochondria dysfunction, and aberrant nNOS-cGMP signaling (Fig. 6.1). The pleiotropic effects of dystrophin deficiency are likely due to the multifaceted function of dystrophin and to some degree complicate the clear delineation of the pathogenic process. Consequently, it is still elusive how dystrophin deficiency leads to abnormal phenotypes. Earlier work suggests that those aforementioned phenotypes are not merely concomitants of dystrophin deficiency, but actively contributing to disease progression. Corrections of these abnormalities, such as amplifying cGMP signaling or correction of calcium mishandling, are accompanied by ameliorated tissue pathology and improved muscle function. Because successful disease modeling in vitro is determined by faithful reproduction of in vivo disease characteristics, confirmation of the presence of the disease features in dystrophin-deficient cardiac cells renders credibility to further exploration.
Mechanistic scheme of the dystrophin-deficient cardiomyopathy. The absence of dystrophin causes membrane instability, triggering the abnormal calcium influx through various calcium channels. Improper accumulation of extracellular calcium together with hypersensitive ryanodine receptor leads to calcium mishandling. Coupled with dysregulated nNOS-cGMP pathway and heightened oxidative stress, the disease network triggers cell death through calcium-dependent protein degradation and mitochondria-mediated apoptosis
6.2.2.1 Impairment of Membrane Barrier Function
Dystrophin is a cell membrane anchoring protein. Based on its sarcolemma localization, others postulated that the absence of dystrophin causes membrane barrier dysfunction. This idea is supported by the findings of “leaky sarcolemma.” For instance, measuring serum levels of intracellular proteins, such as creatine kinase (CK), has been widely employed to assess the degree of muscle damage in muscular dystrophy. Experimentally introduced membrane impermeable substances such as albumin [27–31] and Evans Blue dye were found to accumulate within damaged dystrophin-deficient myofibers. Although these early studies confirmed the presence of an altered membrane barrier function, they failed to precisely define the biophysical nature of the membrane lesion. Normally, skeletal and cardiac muscle endures mechanical strain during contraction. In dystrophin-deficient DMD muscle, discontinuation of the normal sarcolemmal membrane structure in non-necrotic muscle fibers was observed by transmission electron microscopy. This observation was coupled with pathological intracellular changes and hyper-contracture of the surrounding myofibers leading to the speculation that physical breakage of the membrane, or “micro-ruptures,” induced by mechanical strain during muscle contraction is the underlying membrane defect in dystrophin deficiency [32]. This notion is further supported by the observation that a synthetic polymer, poloxamer 188, seals membrane ruptures and reversed the dystrophic phenotype [33, 34]. In contrast to this prevailing view, Allen et al. argued that in dystrophin-deficient muscle, slow kinetics of trans-sarcolemmal calcium ingress following injury could not be accounted for by abrupt physical breakage. Instead, the investigators proposed that pathological activation of preexisting membrane channels are responsible for heightened membrane permeability [35]. Various calcium-permeable channels have been scrutinized for this purpose and will be discussed in the following section.
6.2.2.2 Dysregulated Calcium Handling
Calcium is a critical ion with diversified biological functions in both physiological and pathological processes of muscle. As a result, muscle has evolved highly regulated machinery to keep calcium flow in check. Thus, malfunction of critical proteins in this system may jeopardize the delicate balance with a detrimental effect. Increase of intracellular calcium concentration triggers muscle contraction and attenuates cellular compliance [36]. Excessive activation of calcium signaling networks could lead to cell death [11], partly through the mitochondrial death pathway [5, 37]. It is also worth noting that skeletal muscle differs from cardiac muscle in calcium-handling processes. Opening of the membrane-bound l-type calcium channel in skeletal myocytes physically interacts and activates the ryanodine receptor (RyR), the gatekeeper of sarcolemma reticulum (SR) calcium storage. While in cardiomyocytes, it is the local elevation of subsarcolemmal calcium concentration, following calcium influx through the dihydropyridine receptor (DHPR), that activates the SR RyR. This difference in calcium handling could potentially contribute to different pathophysiologies and account for organ-specific phenotypes such as heart rhythm disturbances.
-
1.
[Ca2+]i handling in dystrophin-deficient cardiomyocytes
Aged mdx cardiomyocytes demonstrate elevated resting [Ca2+]i [38] and attenuated SR calcium storage [39] (Fig. 6.2), not present in young mdx mice. When young mdx cells were stressed by mechanical stretch, they responded with a profound calcium transient that largely surpassed the WT controls [34, 39, 40] (Fig. 6.2). Both adult and young mdx cardiomyocytes demonstrated prolonged calcium reuptake [38, 41, 42]. On the other hand, overexpression of sarco-/endoplasmic reticulum Ca2+-ATPase 2 (SERCA2), the pump responsible for sequestering calcium in SR during repolarization, normalized intracellular calcium load and corrected the abnormal EKG [43].
-
2.
Extracellular calcium entry
It is generally accepted that abnormal extracellular calcium entry through a disrupted membrane triggers downstream disease networks. In support of this idea, removal of extracellular calcium completely abrogates pathological [Ca2+]i accumulation [40]. However, the biophysical identity of the entry route is still obscure. Though membrane tears resulting from mechanical injury has long been suspected to be the culprit instigator, recently Whitehead et al. argued that the delayed [Ca2+]i uprise following mechanical stretch contradicts membrane micro-rupture to be the leading path of calcium entry [44]. Alternatively, they suggested a group of calcium-permeable cation channels. The so-called stretch-activated channels (SACs) were first described by Franco and Lansman to be abnormally active in mdx mice myotubes [45]. Squire et al. later confirmed that the occurrence and opening probability of this channel were greater in mdx myofibers [46]. Blocking this channel by gadolinium, streptomycin, and GsMT-4 significantly ameliorated intracellular calcium overload induced by mechanical stretch [38, 47, 48]. Unfortunately, though the functionality of SAC has been confirmed by electrophysiological exams, little is known about its protein identity. Some suggested involvement of a transient receptor potential channel (TRPC). For example, a recent study revealed in mdx heart a twofold increase of TRPC vanilloid channels type 2 (TRPV2) expression and a mislocalization from the cytoplasm to the sarcolemma. Knocking down TRPV2 by siRNA or antibody blocking TRPV2 channel abrogated mechanically induced intracellular calcium accumulation [49].
Other membrane-localized Ca2+ channels have also been investigated in the context of dystrophin deficiency. The sarcolemma l-type calcium channel, DHPR, is the main calcium channel that triggers excitation-contraction coupling and directly interacts with dystrophin in the t-tubule system [50] [51]. DHPR channel inactivation was delayed in cardiomyocytes from neonatal [50] and adult mdx mice [52, 53]. Moreover, on mdx cardiomyocytes, the l-type calcium channel mediates an enhanced calcium influx that contributes to prolonged action potential duration [52]. These findings inspired studies to test the effect of calcium inhibitors in DMD patients. However, neither animal experiments nor clinical trials have demonstrated significant benefit [54, 55].
-
3.
Intracellular calcium release through hyperactive RyR
Trans-sarcolemmal calcium influx is augmented in DMD cardiomyocytes. However, this change alone is insufficient to account for the cyclic abnormal calcium oscillations observed in DMD cardiomyocytes, as shown in Fig. 6.2 [36, 40]. The self-sustaining nature of the calcium oscillation suggests the involvement of intracellular calcium storage of the SR. Several groups have reported that the function of RyR, the molecule gating SR calcium release, is altered in DMD cardiomyocytes [36, 56]. A “leaky” RyR loses control over intracellular SR calcium release, contributing to calcium dysregulation. Ullrich et al. reported that the sensitivity of RyR is increased in mdx cardiomyocytes, responding to low concentration of extracellular calcium that could not activate wild-type RyR [37]. Prosser et al. further demonstrated that sensitized RyR responds to stimuli by a profound SR calcium liberation, which potentially accounts for DMD cardiac rhythm disturbances [36].
6.2.3 Excessive ROS Production
Cumulative evidence suggests a key role of elevated oxidative stress in the degeneration of dystrophin-deficient skeletal muscle [44, 57, 58]. Exposing mdx cardiomyocytes to hypoosmotic or stretch stress triggers higher-than-normal reactive oxygen species (ROS) production [36, 40], likely through upregulating key enzymes in ROS production such as NAPDH oxidase 2 (NOX2) [36], NOX4, and Lysyl oxidase (LOX) [59]. Antagonizing ROS by supplementing the ROS scavenger, N-acetylcysteine (NAC), attenuates calcium-handling abnormalities, preserves heart function, and mitigates inflammation and fibrosis [60].
The role of oxidative stress and its downstream molecular consequences has been well documented in other forms of cardiomyopathy [61]. Though direct evidence is lacking, it is reasonable to argue that a similar pathway may also be activated in dystrophin-deficient cardiomyocytes. Excessive oxidation of functional proteins inflicts cell damage and deteriorates heart function. Those susceptible targets include membrane channels, contractile proteins [62], signaling proteins [63], and EC coupling components [64]. For example, Prosser et al. revealed that the ROS generated via NOX2 oxidize RyR, which is the molecular basis of its hypersensitivity [36].
6.2.4 Mitochondria Dysfunction
Functional imaging studies through positron emission tomography (PET) identified regional abnormalities in the DMD heart, characterized by energy substrate shift from fatty acid to glucose [65–67]. Lately, alterations of metabolic substrate were experimentally confirmed in young mdx heart, prior to overt cardiomyopathy, coupled with increased oxygen consumption, glycolysis, and ATP production rate [68]. The same group also discovered that the mitochondria permeability transition pore (mPTP), an important mediator of mitochondrial apoptosis pathway, is more susceptible to open when challenged by stressors [5], suggesting that mitochondria are actively involved in disease progression and may represent an intervention target.
6.2.5 Abnormal nNOS-cGMP Signaling
In skeletal muscle, the neuronal nitric oxide synthase (nNOS) is anchored to the c-terminus of dystrophin to control nitric oxide (NO) production. While active NO modulates target proteins by binding to a thiol residue via s-nitrosylation, NO can also function through activating soluble guanylyl cyclase (sGC) to produce cGMP as the secondary messenger. In DMD patients, the absence of dystrophin causes nNOS mislocalization, which greatly dampens NOS function by up to 80 %. The benefits of nNOS overexpression in skeletal muscle include anti-inflammation, anti-fibrosis, and additional protection of myofibers against mechanical stress [69]. nNOS-mediated NO production also exerts a protective effect in the heart [8]. Differing from its sarcolemmal localization in skeletal myofiber, nNOS mainly associates with SR [70] and mitochondria [71] in the heart, without direct interaction with dystrophin. Surprisingly, Bia et al. discovered in dystrophin/utrophin double knockout mice an 80 % decrease of cardiac nNOS activity. Encouraged by their previous success in skeletal muscle [69], overexpressing nNOS in mdx heart demonstrated salutary effect to decrease inflammation and fibrosis [72]. Another group overexpressed the sGC and administration of the PDE5 inhibitor sildenafil with the goal to activate the sGC pathway. Remarkably, both these strategies improved mitochondrial function, attenuated stress-induced mPTP opening, and exerted sarcolemma protection against workload-induced damage [73, 74]. Sildenafil has also been shown capable to reverse cardiomyopathy in aged mdx mice, boosting depressed cardiac function to comparable levels to age-matched controls [75]. Taken together, evidence suggests that cGMP-mediated effects account for benefits associated with modulating the nNOS pathway. Commercially available PDE5 inhibitors may hold promise in treating heart disease in DMD.
6.3 Regenerative Cellular Technologies
Advances in genetics, epigenetics, and stem cell biology have empowered scientists with new tools to control cell fate. A variety of human tissues, including cardiomyocytes, neurons, hepatocytes, and endothelial cells can now be produced from stem cells in large quantities. This new technology paves the way for a myriad of downstream applications, such as studying disease mechanisms, screening therapeutic compounds, and enabling cellular replacement therapy.
6.3.1 The Germination of Nuclear Reprogramming
Mammalian development is accompanied by diversification of progeny cell identity, realized by a gradual loss of cell plasticity. It was once believed that development is an irreversible process. Unidirectionality was ensured by a gradual loss of genetic material [76]. Consequently, cell fate was considered to be static and inter-lineage switch of cellular identity impossible. In the middle twentieth century, this dogma of a “one-way street” was challenged. John Gurdon discovered that differentiated tadpole muscle and intestinal nuclei could generate mature fertile adults after being transplanted into enucleated Xenopus eggs. This discovery unequivocally proved that the genome is relatively stable across the life span of individual organisms. The development process is not coupled with attrition of genetic material. When provided with the correct cue, an adult nucleus is capable to initiate and maintain normal development in a similar fashion as zygotes [77]. Miller and Ruddle had also shown that differentiated thymus cells could be reset to a primitive stage equivalent to pluripotent embryonic stem cells by fusing with embryonic carcinoma cells [78]. These findings broadened the “one-way street” to “two-way traffic” but were still constrained within a longitudinal developmental path. Later, Helen Blau utilized a similar cell fusion technique, termed heterokaryon, to successfully convert human non-muscle cell nuclei to acquire a skeletal muscle gene expression profile. This was the first demonstration of an inter-lineage switch [79]. Together, these findings revealed the surprising plasticity of nuclei and foreshadowed an era of nuclear reprogramming to manipulate cellular fate by altering gene expression.
6.3.2 Nuclear Reprogramming 2.0
6.3.2.1 Induced Pluripotency
A lesson learned from the somatic nuclear transfer and heterokaryon experiments is that certain molecules within the recipient’s cytoplasm redirect a terminally differentiated cell type to acquire a different identity. Consequently, identifying these limited molecules and overexpressing them in donor cells might simplify the process. The cosmologically vast number of molecules contained in the cytoplasm made this a formidable task. After decades of exploration, a seminal breakthrough occurred in 2006. Yamanaka and Takahashi identified ectopic expression of four transcription factors, Oct-4, Sox-2, c-Myc, and Klf4 [80], that converted both mouse [80] and human [81] fibroblasts to pluripotent stem cells. Another report around the same time demonstrated that nanog and Lin28 could replace c-Myc and Klf4 [82]. Converted somatic cells, termed, induced pluripotent stem cells (iPSCs), possess indefinite self-renewal capacity and the potential to generate virtually any cell types within the body. To date, although small differences have been reported between iPSC and hESC [83–87], these two behave largely the same. Not only do iPSC and ES cells share similar morphology and gene expression profiles, more importantly, iPSC cells acquire bona fide pluripotency, forming teratoma tumors after engrafting into immune-compromised animals. Mouse iPSCs could even pass the most stringent tetraploid complementation assay to generate progenies solely composed of cells differentiated from iPSCs [88].
Recent technique evolution has overcome some major hurdles for clinical translation of iPSC technology. The poor reprogramming efficiency and slow kinetics once the bottlenecks of iPSC derivation were overcome by optimizing the reprogramming factor combination and starting cell population [89–90]. Lately, a remarkable 100 % efficiency was achieved by simultaneously knocking down the MDB3 gene [91] together with reprogramming factor delivery. On the other hand, the concern of insertional mutagenesis, elicited by random viral integration, is addressed by “footprint-free” reprogramming technology. Non-integration delivery vectors, such as sendai virus, episome, or mRNA, can achieve efficient reprogramming without any genome perturbation.
6.3.2.2 Inter-Lineage Cell Fate Conversion
Derivation of iPSCs by transient, ectopic expression of transgenes demarcates the new era of nuclear reprogramming. Compared to previous SCNT experiments, the major improvement of this new version of nuclear reprogramming is that cell fate conversion is achieved through manipulating a small set of predefined factors. This breakthrough considerably lowered the technical barrier and encouraged further exploration. Following experiments demonstrated that other cell fates could also be rerouted via ectopic expression of a handful transgenes (Table 6.1). Inter-lineage cell fate conversion bypasses the pluripotent stem cell stage. This process, termed “trans-differentiation,” usually occurred with a donor cell developmentally related to the target cells, although trans-germ layer conversions have also been documented. The majority of these trans-differentiation transgenes are transcriptional factors, chromatin modifiers, and miRNAs critical to normal development of certain lineages. Surprisingly only a handful of factors are required to achieve these transformations, which may emphasize that cell identities are maintained through certain “master regulators,” such as MyoD for skeletal muscle [110]. Several studies have demonstrated that similar cell fate conversion could be achieved in vivo, such as cardiomyocytes [111], pancreatic beta cells [112, 113], and neurons [114, 115], showcasing the versatility of the reprogramming technology.
6.3.3 Guided Differentiation
An alternative strategy to achieve in vitro cell fate control is to guide pluripotent stem cells to differentiate into desired cell types. Insights gained from embryonic development have enabled scientists to harness the physiological cues to hijack the intrinsic developmental program. The key is to recapitulate those early events governing lineage commitment and germ layers’ specification, by temporally modulating critical signaling pathways [116]. A broad spectrum of cell types including cardiomyocytes, hepatocytes, pancreatic beta cells, endothelium, and dopaminergic neurons have been generated in this fashion. The advantages of this method are twofold. First, large quantity, highly purified target cells can be manufactured through an optimized induction protocol. For example, cardiomyocyte production can be over 90 % in purity with 3 output cardiomyocytes for every input stem cell. This scale is compatible with an industrial setting to produce clinically relevant cell numbers, usually in the billions [117]. Secondly, starting from a pluripotent stem cell offers a unique opportunity for genetic engineering to generate homogenous cell line with desired genetic modifications. The following section will focus on cardiac induction. The specification of other germ layers is well reviewed by Murry et al. [116].
It was noticed that beating cardiomyocytes could be generated from embryoid bodies (EB), stem cell aggregates mimicking early developing embryo structure. The efficiency of spontaneous cardiogenesis is low (less than 10 %) with large batch-to-batch variation [118, 119]. Not until insights gained from development were applied have investigators started to consistently generate adequate cardiomyocytes for downstream studies. In the embryo, cardiac progenitors are derived from brachyury T- and Emos-positive mesoderm cells, which further give rise to KDR- and PDGFRα-positive cardiac mesoderm cells and gradually turn on the cardiac master regulator MESP1. Further specification of cardiac mesoderm generates cardiac progenitors, characterized by the expression of a panel of cardiac-specific transcriptional factors such as Nkx2.5, GATA4, Tbx5, and Isl1. Temporally and spatially orchestrated Nodal, BMP4, and Wnt3 signaling ensure this organized sequential progression. Artificially supplementing these ligands in culture medium, mimicking the physiological strength and timing, can also differentiate pluripotent stem cells into cardiomyocytes. With some technical variation, this scheme consistently generates cardiomyocytes in around 2 weeks, with an efficiency ranging from 30 to 90 % [120]. It is important to note that the Wnt signaling has a unique biphasic role in cardiac development. Lately the Palecek group had shown that by fine-tuning Wnt pathway alone by two molecules, GSK-3β inhibitor CHIRO-99021 and Wnt inhibitor IWP-4 or IWR-2, cardiomyocytes can be generated from multiple cell lines with an efficiency over 90 % [121].
Various strategies have demonstrated that in vitro cardiac induction follows the same gene expression pattern that would occur during embryonic development. Downregulation of pluripotent genes is accompanied by upregulation of mesoderm markers Brachyury T and MESP1, which peak around day 2. Cardiac progenitor markers KDR, ISL1, and PDGFR-α peak at day 5, followed by a steady increase of mature myocyte markers such as Nkx2.5, myosin heavy chain (MHC), and troponin [122–125]. Differentiated cells manifest sarcomere striation, typical action potentials and associated ion channels [126], potent gap junctions for synchronized contraction, as well as positive humeral regulation response following isoproterenol stimulation [119, 127, 128]. Klug et al. demonstrated that stem cell-derived cardiomyocytes express dystrophin [129]. While this evidence suggests that these are bona fide cardiomyocytes, stem cell-derived cardiomyocytes largely mimic a developmentally immature cardiac phenotype. Transcriptional profiling suggested that the cardiac gene expression pattern was similar to 20-gestational-week fetal cardiomyocytes [130]. Electrophysiological assays indicated that stem cell-derived cardiomyocytes possess fetal-type ion channels [122], leading to a fetal-like negative force frequency relationship, blunted post-rest potentiation [131], higher resting potential, and slow action potential upstroke [130, 132]. Ultrastructure analysis revealed that the characteristic components, such as contraction machinery, sarcomeric reticulum, transverse tubules (t-tubule), and mitochondria, were all present, but their abundance, distribution, and organization failed to reach the level of adult cardiomyocytes [133].
6.3.4 From Urine to Dystrophin-Deficient Cardiomyocytes
Our group demonstrated that urine harbors a unique cell population capable to adhere to plastic and undergo extensive proliferation. These urine-derived cells manifest spindle-shape morphology and express classic MSC surface markers including CD44, CD73, CD90, CD105, and CD146 [134, 135]. Functional assays reveal that urine-derived cells possess progenitor properties and give rise to several somatic cell types [136]. When transduced with lentiviral vector containing classic OKSM reprogramming factors, these cells demonstrate faster reprogramming kinetics compared to mesenchymal stem cells, generating iPSC colonies within 2 weeks [134]. Urine-derived iPSC colonies, from both normal individual and DMD patient urine, can give rise to embryonic tumors containing characteristic tissue structures of all three germ layers, demonstrating bona fide pluripotency.
Exposing the urine-derived iPSC cells to a combination of growth factors, including Activin A, bone morphogenetic protein 4, and dickkopf-1, in a strict sequence and defined duration [137] forms beating cardiomyocytes with an efficiency from 40 to 90 %. These differentiated cardiomyocytes show typical sarcomere structure and are positive for sarcomeric α-actinin, cardiac myosin heavy chain, as well as membrane-localized connexin43. They also exhibit functional cardiac properties, spontaneous action potentials characteristic of nodal, ventricular, and atrial subtypes. On the other hand, dystrophin expression was absent from DMD cardiomyocytes, recapitulating the essential aspect of the DMD disease [134] (Figs. 6.3 and 6.4).
6.4 Dystrophin-Deficient Cardiomyocytes Generated via Cellular Reprogramming Are a Novel Biological Reagent for the Study of DMD Cardiomyopathy
The list of iPSC-based disease models is steadily growing. Impressively, different diseased cells manifest phenotypes analogous to symptoms in patients in culture dishes. A partial list includes models for amyotrophic lateral sclerosis, spinal muscular atrophy, familial dysautonomia [138], Rett syndrome [139], schizophrenia [140], Parkinson disease [141–144], Timothy syndrome [145], and long QT syndrome [146]. The explosion of the iPSC application is largely due to the fact that cellular reprogramming is a powerful tool that strongly resonates in the era of personalized medicine, which enables identifying and optimizing solutions for specific mutations. DMD cardiomyopathy is well suited to iPSC-based “disease-in-a-dish” platform. First, there is currently no effective treatment for DMD cardiomyopathy. Secondly, the molecular mechanism of the cardiomyopathy is not well understood. Though the disease root cause is clear, a variety of subdomains, such as membrane stability, calcium handling, and mitochondria abnormalities, have been suggested in the pathogenic process. The complexity of the underlying pathogenic network makes identifying therapeutic targets a daunting task. Thirdly, the most widely used DMD animal model, the mdx mouse, does not accurately reproduce the cardiac pathology observed in human patients [147]. Thus it is imperative to study this disease in a human context to render clinical relevance. Furthermore, the risk associated with cardiac biopsy makes acquiring samples from DMD patients nearly impossible. These limiting factors impose significant constraints in developing therapies for human patients. As a result, human cardiomyocytes differentiated from patient iPSCs represent an attractive option to overcome these limitations.
It is technically favorable to model Duchenne cardiomyopathy using the iPSC approach mainly because DMD is a classic Mendelian monogenic disease with complete penetrance in male patients. A large body of literature suggests that the disease phenotype is cell-autonomous, indicating that disease traits can be readily observed independent of the diseased body environment. Compared to polygenic genetic disorder or complex disorders involving strong environmental attributes, the technical complexity associated with modeling dystrophin-deficient cardiomyopathy ex vivo is considerably lower. Moreover, the existence of a highly efficient cardiac induction regimen bolsters this strategy by lowing the technical hurdle [121, 137].
6.4.1 Personalized Diagnosis: Exploring Disease Etiology
Presently, disease diagnosis has evolved from clinical and functional to molecular diagnosis using biomarkers [148, 149] and genome sequencing [150, 151]. However, genotype-phenotype relationships are not always strongly correlated, and sometimes it is difficult to elucidate a causal relationship. Toward this end, iPSC technology represents a unique opportunity to converge information from multiple levels and directly link them to a change in cellular function. For example, in iPSC-differentiated disease target cells, the mutated gene can be sequenced, biochemical reactions can be measured, and abnormal metabolites can be quantified. Readouts from these measurements can be further linked to cellular responses in a quantifiable fashion. In addition, cell culture systems allow intricate experimental interventions at virtually any level, from genetic to environmental, greatly aiding in determining the disease root cause. Lastly, because clinically relevant cells are differentiated stepwise from stem cells in a fashion that recapitulates the embryonic development, disease initiation and early progression, hard to assess by other modalities, can now be closely monitored. It is possible to exploit these models to uncover early disease markers before overt symptoms, increasing the opportunity for early intervention or even prevention. For instance, Kim et al. identified in patient iPSC-derived cardiomyocytes an abnormal peroxisome proliferator-activated receptor gamma (PPAR-γ) activation that underlies the pathogenesis of arrhythmogenic right ventricular dysplasia (ARVC), a previously unrecognized mechanism [152]. iPSC-based platforms also simplifies the diagnosis procedure since tests only need to be conducted on single diseased lineage in isolation, removed from potential secondary effects of residing within a sick animal.
6.4.2 Personalized Therapy: In Search for the Ideal Treatment
Because of its individualized nature, iPSC technology can also serve as an invaluable tool to identify new therapies. First, it can be employed as a platform to predict the potency and toxicity of existing therapies [153, 154]. It is not uncommon that a spectrum of drugs is available for a single disease, yet not every drug is equally effective for all patients. The potency and toxicity, which largely depend on individual’s unique genetic background, are hard to predict based on existing modalities. On the other hand, iPSC cells have the potential to be the stage for such forecasts, since the patients’ own cells are tested. Such tests are not necessarily limited to chemical compounds. As a cellular assay, other therapeutics, such as gene therapy vectors, can also be assayed as a quality control measure to predict its clinical efficacy. For those disorders without effective treatment, like DMD cardiomyopathy, de novo screening against an extensive compound library can potentially identify new treatments with no risk to the patient [155].
The differentiated cells themselves can also serve as therapeutics. One route to cure a genetic disorder may involve replacing the diseased cells with genetically corrected counterparts. The emergence of novel genetic engineering tools, such as CRISPR and TALEN enzymes, has greatly improved the feasibility of complex genome modification of human stem cells [144, 156–159]. Barrier of immunogenicity could be overcome by autologous transplantation of the progeny differentiated from iPSC. This strategy has been applied to improve skeletal muscle function in dystrophin-deficient mdx mice [160]. In terms of the heart, though previously it has been shown that genetic heart disease benefited from embryonic stem cell transplantation [161], the delivery method still imposes the biggest challenge for inherited disorders including DMD.
6.4.3 Dystrophin-Deficient Cardiomyocytes Generated via Cellular Reprogramming Recapitulate Certain Aspects of the Disease Phenotype
Though studies have shown that disease target cells differentiated from patient’s iPS cells faithfully reproduce disease-associated abnormalities, it is still questionable whether the cells created in laboratory, at a developmentally naïve stage, will be able to fully demonstrate a phenotype of late-onset pathology observed in DMD. Therefore the first task is to demonstrate progeny differentiated from iPSC cells that retain certain disease hallmarks, which will then lend credibility to novel insights gained on an iPSC platform. For example, ventricular cells differentiated from a long QT syndrome patient demonstrated the characteristic prolongation of the QT interval as well as an abnormal potassium channel [146, 154, 162].
In our hands, to model DMD cardiomyopathy, cardiomyocytes were assessed both molecularly and phenotypically for well-established DMD features. For example, the absence of dystrophin protein is the root cause of DMD. Confirmed both by immunostaining (Fig. 6.4), cardiomyocytes differentiated from DMD iPS cells were negative for dystrophin expression, recapitulating an essential aspect of the disease. It is well established that the absence of dystrophin causes a spectrum of physiologies deviating from normal control. As discussed earlier, membrane fragility is suspected to be a direct consequence of the absence of dystrophin and initiates downstream pathological cascades. One way to assess the membrane barrier function is measuring the release of intracellular content during a hypotonic stress challenge. In vitro, an increase of intracellular hydrostatic pressure passively stretches the cell’s sarcolemma. DMD cells are abnormally susceptible to mechanical stress, and their fragile membrane predisposes cells to rupture, leading to the liberation of intracellular contents. As an example of a hypotonic stress assay, red blood cells are subjected to a series of graded hypotonic solutions, and intracorpuscular hemoglobin release is assayed to diagnose hereditary spherocytosis [163]. We employed a similar strategy to mechanically stretch the cardiac sarcolemma by incubating normal and DMD cells in hypotonic solutions ranging from normal tonicity to 1/8 tonicity. The cardiac-specific injury marker CK-MB, a widely used clinical marker, was measured after 30 min. Dystrophin-deficient cardiomyocytes demonstrated an abnormally high release CK-MB profile. Measured as CK-MB concentration, increased levels were demonstrated even at relatively normal tonicity and became marked greater than normal cells as tonicity decreased (Fig. 6.4). This evidence supports the notion that dystrophin deficiency leads to sarcolemma fragility, which predisposes cardiomyocytes to mechanical stretch damage.
Calcium mishandling is another characteristic feature of DMD. Loaded with the calcium indicator Fluo-4, iPSC-derived DMD cardiomyocytes were paced by external field stimulation. Compared to dystrophin replete (normal) control cells, DMD cells demonstrated a prolonged calcium decay time T50, consistent with previous reports in the dystrophin-deficient mouse [38, 41, 42].
Mitochondria dysfunction observed in DMD patients was recently recognized as an important mediator of disease progression. Two prominent features, a lowered opening threshold for mitochondrial permeability transition pore (mPTP) and altered bioenergetics, have been linked to the disease in the mdx mouse. To evaluate mPTP pore opening, cardiomyocytes were loaded with mitochondria potential (Δψm) indicator tetramethylrhodamine ethyl ester (TMRE) and then exposed to focused laser to induce oxidative stress-mediated mPTP opening. The opening of mPTP leads to the decrease of Δψm, reflected by lowering of TMRE fluorescence. Dystrophin-deficient cardiomyocytes manifested a 50 % shorter mPTP time compared to normal cells. In other experiments, an extracellular biochemical analyzer (Seahorse) was employed to examine the bioenergetics of cardiomyocytes. Oxygen consumption rate (OCR) was measured as the parameter of metabolism activity. Surprisingly, DMD cells demonstrated augmented basal and maximal oxygen consumption, different from the report on mdx skeletal muscle but in agreement with a study using a Langendorff perfused mdx adult heart, in which the mdx heart exhibited elevated level of glycolysis, carbohydrate utilization, oxygen consumption, and overall ATP production [5].
6.5 Summary
Duchenne muscular dystrophy causes degeneration of both skeletal and cardiac muscle. All DMD patients inevitably develop heart disease, but the mechanism of DMD heart failure is still elusive, partly due to the hard-to-access heart cells from patients. We modeled heart disease in DMD using cellular reprogramming by converting patient urine cells into induced pluripotent stem cells or iPSCs. IPSCs were further transformed into heart cells for study. This regenerative medicine technology provided an unprecedented opportunity to study DMD heart disease from several physiological perspectives including membrane fragility, calcium handling, and energy metabolism. This regenerative medicine platform will facilitate exploration of DMD disease etiology and provide unlimited patient biologic material to screen new therapeutic compounds.
References
Hermans MC, Pinto YM, Merkies IS, de Die-Smulders CE, Crijns HJ, Faber CG. Hereditary muscular dystrophies and the heart. Neuromuscul Disord. 2010;20:479–92.
Armstrong SC, Latham CA, Shivell CL, Ganote CE. Ischemic loss of sarcolemmal dystrophin and spectrin: correlation with myocardial injury. J Mol Cell Cardiol. 2001;33:1165–79.
Kido M, Otani H, Kyoi S, Sumida T, Fujiwara H, Okada T, Imamura H. Ischemic preconditioning-mediated restoration of membrane dystrophin during reperfusion correlates with protection against contraction-induced myocardial injury. Am J Physiol Heart Circ Physiol. 2004;287:H81–90.
Lee GH, Badorff C, Knowlton KU. Dissociation of sarcoglycans and the dystrophin carboxyl terminus from the sarcolemma in enteroviral cardiomyopathy. Circ Res. 2000;87:489–95.
Burelle Y, Khairallah M, Ascah A, Allen BG, Deschepper CF, Petrof BJ, Des Rosiers C. Alterations in mitochondrial function as a harbinger of cardiomyopathy: lessons from the dystrophic heart. J Mol Cell Cardiol. 2010;48:310–21.
Hoffman EP, Brown Jr RH, Kunkel LM. Dystrophin: the protein product of the Duchenne muscular dystrophy locus. Cell. 1987;51:919–28.
Muntoni F, Torelli S, Ferlini A. Dystrophin and mutations: one gene, several proteins, multiple phenotypes. Lancet Neurol. 2003;2:731–40.
Percival JM, Adamo CM, Beavo JA, Froehner SC. Evaluation of the therapeutic utility of phosphodiesterase 5A inhibition in the mdx mouse model of duchenne muscular dystrophy. Handb Exp Pharmacol. 2011;323–344.
Spurney C, Shimizu R, Morgenroth LP, Kolski H, Gordish-Dressman H, Clemens PR, Investigators C. Cooperative International Neuromuscular Research Group Duchenne Natural History Study demonstrates insufficient diagnosis and treatment of cardiomyopathy in Duchenne muscular dystrophy. Muscle Nerve. 2014;50:250–6.
Nigro G, Comi LI, Politano L, Bain RJ. The incidence and evolution of cardiomyopathy in Duchenne muscular dystrophy. Int J Cardiol. 1990;26:271–7.
Townsend D, Yasuda S, Metzger J. Cardiomyopathy of Duchenne muscular dystrophy: pathogenesis and prospect of membrane sealants as a new therapeutic approach. Expert Rev Cardiovasc Ther. 2007;5:99–109.
Ishikawa Y, Bach JR, Minami R. Cardioprotection for Duchenne's muscular dystrophy. Am Heart J. 1999;137:895–902.
Matsumura T, Tamura T, Kuru S, Kikuchi Y, Kawai M. Carvedilol can prevent cardiac events in Duchenne muscular dystrophy. Intern Med. 2010;49:1357–63.
Rhodes J, Margossian R, Darras BT, Colan SD, Jenkins KJ, Geva T, Powell AJ. Safety and efficacy of carvedilol therapy for patients with dilated cardiomyopathy secondary to muscular dystrophy. Pediatr Cardiol. 2008;29:343–51.
Schram G, Fournier A, Leduc H, Dahdah N, Therien J, Vanasse M, Khairy P. All-cause mortality and cardiovascular outcomes with prophylactic steroid therapy in Duchenne muscular dystrophy. J Am Coll Cardiol. 2013;61:948–54.
Fayssoil A, Orlikowski D, Nardi O, Annane D. Complete atrioventricular block in Duchenne muscular dystrophy. Europace. 2008;10:1351–2.
Fayssoil A, Orlikowski D, Nardi O, Annane D. Pacemaker implantation for sinus node dysfunction in a young patient with Duchenne muscular dystrophy. Congest Heart Fail. 2010;16:127–8.
Takano N, Honke K, Hasui M, Ohno I, Takemura H. A case of pacemaker implantation for complete atrioventricular block associated with Duchenne muscular dystrophy. No To Hattatsu. 1997;29:476–80.
Amodeo A, Adorisio R. Left ventricular assist device in Duchenne cardiomyopathy: can we change the natural history of cardiac disease? Int J Cardiol. 2012;161, e43.
Davies JE, Winokur TS, Aaron MF, Benza RL, Foley BA, Holman WL. Cardiomyopathy in a carrier of Duchenne's muscular dystrophy. J Heart Lung Transplant. 2001;20:781–4.
Smith MC, Arabia FA, Tsau PH, Smith RG, Bose RK, Woolley DS, Rhenman BE, Sethi GK, Copeland JG. CardioWest total artificial heart in a moribund adolescent with left ventricular thrombi. Ann Thorac Surg. 2005;80:1490–2.
Webb ST, Patil V, Vuylsteke A. Anaesthesia for non-cardiac surgery in patient with Becker's muscular dystrophy supported with a left ventricular assist device. Eur J Anaesthesiol. 2007;24:640–2.
Andrikopoulos G, Kourouklis S, Trika C, Tzeis S, Rassias I, Papademetriou C, Katsivas A, Theodorakis G. Cardiac resynchronization therapy in Becker muscular dystrophy. Hellenic J Cardiol. 2013;54:227–9.
Fayssoil A, Abasse S. Cardiac resynchronization therapy in Becker muscular dystrophy: for which patients? Hellenic J Cardiol. 2010;51:377–8.
Stollberger C, Finsterer J. Left ventricular synchronization by biventricular pacing in Becker muscular dystrophy as assessed by tissue Doppler imaging. Heart Lung. 2005;34:317–20.
Leung DG, Herzka DA, Thompson WR, He B, Bibat G, Tennekoon G, Russell SD, Schuleri KH, Lardo AC, Kass DA, Thompson RE, Judge DP, Wagner KR. Sildenafil does not improve cardiomyopathy in Duchenne/Becker muscular dystrophy. Ann Neurol. 2014;76(4):541–9.
Childers MK, Okamura CS, Bogan DJ, Bogan JR, Petroski GF, McDonald K, Kornegay JN. Eccentric contraction injury in dystrophic canine muscle. Arch Phys Med Rehabil. 2002;83:1572–8.
Childers MK, Okamura CS, Bogan DJ, Bogan JR, Sullivan MJ, Kornegay JN. Myofiber injury and regeneration in a canine homologue of Duchenne muscular dystrophy. Am J Phys Med Rehabil. 2001;80:175–81.
Childers MK, Staley JT, Kornegay JN, McDonald KS. Skinned single fibers from normal and dystrophin-deficient dogs incur comparable stretch-induced force deficits. Muscle Nerve. 2005;31:768–71.
McNeil PL, Khakee R. Disruptions of muscle fiber plasma membranes. Role in exercise-induced damage. Am J Pathol. 1992;140:1097–109.
Petrof BJ, Shrager JB, Stedman HH, Kelly AM, Sweeney HL. Dystrophin protects the sarcolemma from stresses developed during muscle contraction. Proc Natl Acad Sci U S A. 1993;90:3710–4.
Mokri B, Engel AG. Duchenne dystrophy: electron microscopic findings pointing to a basic or early abnormality in the plasma membrane of the muscle fiber. Neurology. 1975;25:1111–20.
Townsend D, Turner I, Yasuda S, Martindale J, Davis J, Shillingford M, Kornegay JN, Metzger JM. Chronic administration of membrane sealant prevents severe cardiac injury and ventricular dilatation in dystrophic dogs. J Clin Invest. 2010;120:1140–50.
Yasuda S, Townsend D, Michele DE, Favre EG, Day SM, Metzger JM. Dystrophic heart failure blocked by membrane sealant poloxamer. Nature. 2005;436:1025–9.
Allen DG, Whitehead NP. Duchenne muscular dystrophy--what causes the increased membrane permeability in skeletal muscle? Int J Biochem Cell Biol. 2011;43:290–4.
Prosser BL, Ward CW, Lederer WJ. X-ROS signaling: rapid mechano-chemo transduction in heart. Science. 2011;333:1440–5.
Ullrich ND, Fanchaouy M, Gusev K, Shirokova N, Niggli E. Hypersensitivity of excitation-contraction coupling in dystrophic cardiomyocytes. Am J Physiol Heart Circ Physiol. 2009;297:H1992–2003.
Williams IA, Allen DG. Intracellular calcium handling in ventricular myocytes from mdx mice. Am J Physiol Heart Circ Physiol. 2007;292:H846–55.
Kyrychenko S, Polakova E, Kang C, Pocsai K, Ullrich ND, Niggli E, Shirokova N. Hierarchical accumulation of RyR post-translational modifications drives disease progression in dystrophic cardiomyopathy. Cardiovasc Res. 2013;97:666–75.
Jung C, Martins AS, Niggli E, Shirokova N. Dystrophic cardiomyopathy: amplification of cellular damage by Ca2+ signalling and reactive oxygen species-generating pathways. Cardiovasc Res. 2008;77:766–73.
Cheng YJ, Lang D, Caruthers SD, Efimov IR, Chen J, Wickline SA. Focal but reversible diastolic sheet dysfunction reflects regional calcium mishandling in dystrophic mdx mouse hearts. Am J Physiol Heart Circ Physiol. 2012;303:H559–68.
Koenig X, Dysek S, Kimbacher S, Mike AK, Cervenka R, Lukacs P, Nagl K, Dang XB, Todt H, Bittner RE, Hilber K. Voltage-gated ion channel dysfunction precedes cardiomyopathy development in the dystrophic heart. PLoS ONE. 2011;6, e20300.
Shin JH, Bostick B, Yue Y, Hajjar R, Duan D. SERCA2a gene transfer improves electrocardiographic performance in aged mdx mice. J Transl Med. 2011;9:132.
Whitehead NP, Yeung EW, Allen DG. Muscle damage in mdx (dystrophic) mice: role of calcium and reactive oxygen species. Clin Exp Pharmacol Physiol. 2006;33:657–62.
Franco Jr A, Lansman JB. Calcium entry through stretch-inactivated ion channels in mdx myotubes. Nature. 1990;344:670–3.
Squire S, Raymackers JM, Vandebrouck C, Potter A, Tinsley J, Fisher R, Gillis JM, Davies KE. Prevention of pathology in mdx mice by expression of utrophin: analysis using an inducible transgenic expression system. Hum Mol Genet. 2002;11:3333–44.
Fanchaouy M, Polakova E, Jung C, Ogrodnik J, Shirokova N, Niggli E. Pathways of abnormal stress-induced Ca2+ influx into dystrophic mdx cardiomyocytes. Cell Calcium. 2009;46:114–21.
Teichmann MD, Wegner FV, Fink RH, Chamberlain JS, Launikonis BS, Martinac B, Friedrich O. Inhibitory control over Ca(2+) sparks via mechanosensitive channels is disrupted in dystrophin deficient muscle but restored by mini-dystrophin expression. PLoS ONE. 2008;3, e3644.
Lorin C, Vögeli I, Niggli E. Dystrophic cardiomyopathy - role of TRPV2 channels in stretch-induced cell damage. Cardiovasc Res. 2015;106(1):153–62.
Sadeghi A, Doyle AD, Johnson BD. Regulation of the cardiac L-type Ca2+ channel by the actin-binding proteins alpha-actinin and dystrophin. Am J Physiol Cell Physiol. 2002;282:C1502–11.
Friedrich O, von Wegner F, Chamberlain JS, Fink RH, Rohrbach P. L-type Ca2+ channel function is linked to dystrophin expression in mammalian muscle. PLoS ONE. 2008;3, e1762.
Koenig X, Rubi L, Obermair GJ, Cervenka R, Dang XB, Lukacs P, Kummer S, Bittner RE, Kubista H, Todt H, Hilber K. Enhanced currents through L-type calcium channels in cardiomyocytes disturb the electrophysiology of the dystrophic heart. Am J Physiol Heart Circ Physiol. 2014;306:H564–73.
Woolf PJ, Lu S, Cornford-Nairn R, Watson M, Xiao XH, Holroyd SM, Brown L, Hoey AJ. Alterations in dihydropyridine receptors in dystrophin-deficient cardiac muscle. Am J Physiol Heart Circ Physiol. 2006;290:H2439–45.
Cohn RD, Durbeej M, Moore SA, Coral-Vazquez R, Prouty S, Campbell KP. Prevention of cardiomyopathy in mouse models lacking the smooth muscle sarcoglycan-sarcospan complex. J Clin Invest. 2001;107:R1–7.
Phillips MF, Quinlivan R. Calcium antagonists for Duchenne muscular dystrophy. Cochrane Database Syst Rev. 2008, CD004571.
Fauconnier J, Thireau J, Reiken S, Cassan C, Richard S, Matecki S, Marks AR, Lacampagne A. Leaky RyR2 trigger ventricular arrhythmias in Duchenne muscular dystrophy. Proc Natl Acad Sci U S A. 2010;107:1559–64.
Menazza S, Blaauw B, Tiepolo T, Toniolo L, Braghetta P, Spolaore B, Reggiani C, Di Lisa F, Bonaldo P, Canton M. Oxidative stress by monoamine oxidases is causally involved in myofiber damage in muscular dystrophy. Hum Mol Genet. 2010;19:4207–15.
Rando TA. Oxidative stress and the pathogenesis of muscular dystrophies. Am J Phys Med Rehabil. 2002;81:S175–86.
Spurney CF, Knoblach S, Pistilli EE, Nagaraju K, Martin GR, Hoffman EP. Dystrophin-deficient cardiomyopathy in mouse: expression of Nox4 and Lox are associated with fibrosis and altered functional parameters in the heart. Neuromuscul Disord. 2008;18:371–81.
Williams IA, Allen DG. The role of reactive oxygen species in the hearts of dystrophin-deficient mdx mice. Am J Physiol Heart Circ Physiol. 2007;293:H1969–77.
Nediani C, Raimondi L, Borchi E, Cerbai E. Nitric oxide/reactive oxygen species generation and nitroso/redox imbalance in heart failure: from molecular mechanisms to therapeutic implications. Antioxid Redox Signal. 2011;14:289–331.
Gao WD, Liu Y, Marban E. Selective effects of oxygen free radicals on excitation-contraction coupling in ventricular muscle. Implications for the mechanism of stunned myocardium. Circulation. 1996;94:2597–604.
Aikawa R, Komuro I, Yamazaki T, Zou Y, Kudoh S, Tanaka M, Shiojima I, Hiroi Y, Yazaki Y. Oxidative stress activates extracellular signal-regulated kinases through Src and Ras in cultured cardiac myocytes of neonatal rats. J Clin Invest. 1997;100:1813–21.
Zima AV, Blatter LA. Redox regulation of cardiac calcium channels and transporters. Cardiovasc Res. 2006;71:310–21.
Momose M, Iguchi N, Imamura K, Usui H, Ueda T, Miyamoto K, Inaba S. Depressed myocardial fatty acid metabolism in patients with muscular dystrophy. Neuromuscul Disord. 2001;11:464–9.
Perloff JK, Henze E, Schelbert HR. Alterations in regional myocardial metabolism, perfusion, and wall motion in Duchenne muscular dystrophy studied by radionuclide imaging. Circulation. 1984;69:33–42.
Quinlivan RM, Lewis P, Marsden P, Dundas R, Robb SA, Baker E, Maisey M. Cardiac function, metabolism and perfusion in Duchenne and Becker muscular dystrophy. Neuromuscul Disord. 1996;6:237–46.
Khairallah M, Labarthe F, Bouchard B, Danialou G, Petrof BJ, Des RC. Profiling substrate fluxes in the isolated working mouse heart using 13C-labeled substrates: focusing on the origin and fate of pyruvate and citrate carbons. Am J Physiol Heart Circ Physiol. 2004;286:H1461–70.
Wehling M, Spencer MJ, Tidball JG. A nitric oxide synthase transgene ameliorates muscular dystrophy in mdx mice. J Cell Biol. 2001;155:123–31.
Xu KY, Huso DL, Dawson TM, Bredt DS, Becker LC. Nitric oxide synthase in cardiac sarcoplasmic reticulum. Proc Natl Acad Sci U S A. 1999;96:657–62.
Elfering SL, Sarkela TM, Giulivi C. Biochemistry of mitochondrial nitric-oxide synthase. J Biol Chem. 2002;277:38079–86.
Wehling-Henricks M, Jordan MC, Roos KP, Deng B, Tidball JG. Cardiomyopathy in dystrophin-deficient hearts is prevented by expression of a neuronal nitric oxide synthase transgene in the myocardium. Hum Mol Genet. 2005;14:1921–33.
Ascah A, Khairallah M, Daussin F, Bourcier-Lucas C, Godin R, Allen BG, Petrof BJ, Des Rosiers C, Burelle Y. Stress-induced opening of the permeability transition pore in the dystrophin-deficient heart is attenuated by acute treatment with sildenafil. Am J Physiol Heart Circ Physiol. 2011;300:H144–53.
Khairallah M, Khairallah RJ, Young ME, Allen BG, Gillis MA, Danialou G, Deschepper CF, Petrof BJ, Des RC. Sildenafil and cardiomyocyte-specific cGMP signaling prevent cardiomyopathic changes associated with dystrophin deficiency. Proc Natl Acad Sci U S A. 2008;105:7028–33.
Adamo CM, Dai DF, Percival JM, Minami E, Willis MS, Patrucco E, Froehner SC, Beavo JA. Sildenafil reverses cardiac dysfunction in the mdx mouse model of Duchenne muscular dystrophy. Proc Natl Acad Sci U S A. 2010;107:19079–83.
Briggs R, King TJ. Nuclear transplantation studies on the early gastrula (Rana pipiens). I. Nuclei of presumptive endoderm. Dev Biol. 1960;2:252–70.
Gurdon JB, Elsdale TR, Fischberg M. Sexually mature individuals of Xenopus laevis from the transplantation of single somatic nuclei. Nature. 1958;182:64–5.
Miller RA, Ruddle FH. Pluripotent teratocarcinoma-thymus somatic cell hybrids. Cell. 1976;9:45–55.
Blau HM, Chiu CP, Webster C. Cytoplasmic activation of human nuclear genes in stable heterocaryons. Cell. 1983;32:1171–80.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–76.
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–72.
Yu J, Vodyanik MA, Smuga-Otto K, Antosiewicz-Bourget J, Frane JL, Tian S, Nie J, Jonsdottir GA, Ruotti V, Stewart R, Slukvin II, Thomson JA. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917–20.
Kim K, Doi A, Wen B, Ng K, Zhao R, Cahan P, Kim J, Aryee MJ, Ji H, Ehrlich LI, Yabuuchi A, Takeuchi A, Cunniff KC, Hongguang H, McKinney-Freeman S, Naveiras O, Yoon TJ, Irizarry RA, Jung N, Seita J, Hanna J, Murakami P, Jaenisch R, Weissleder R, Orkin SH, Weissman IL, Feinberg AP, Daley GQ. Epigenetic memory in induced pluripotent stem cells. Nature. 2010;467:285–90.
Kim K, Zhao R, Doi A, Ng K, Unternaehrer J, Cahan P, Hongguang H, Loh YH, Aryee MJ, Lensch MW, Li H, Collins JJ, Feinberg AP, Daley GQ. Donor cell type can influence the epigenome and differentiation potential of human induced pluripotent stem cells. Nat Biotechnol. 2011;29(12):1117–9.
Marchetto MC, Yeo GW, Kainohana O, Marsala M, Gage FH, Muotri AR. Transcriptional signature and memory retention of human-induced pluripotent stem cells. PLoS ONE. 2009;4, e7076.
Ohi Y, Qin H, Hong C, Blouin L, Polo JM, Guo T, Qi Z, Downey SL, Manos PD, Rossi DJ, Yu J, Hebrok M, Hochedlinger K, Costello JF, Song JS, Ramalho-Santos M. Incomplete DNA methylation underlies a transcriptional memory of somatic cells in human iPS cells. Nat Cell Biol. 2011;13:541–9.
Polo JM, Liu S, Figueroa ME, Kulalert W, Eminli S, Tan KY, Apostolou E, Stadtfeld M, Li Y, Shioda T, Natesan S, Wagers AJ, Melnick A, Evans T, Hochedlinger K. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nat Biotechnol. 2010;28:848–55.
Zhao XY, Li W, Lv Z, Liu L, Tong M, Hai T, Hao J, Guo CL, Ma QW, Wang L, Zeng F, Zhou Q. iPS cells produce viable mice through tetraploid complementation. Nature. 2009;461:86–90.
Kim JB, Greber B, Arauzo-Bravo MJ, Meyer J, Park KI, Zaehres H, Scholer HR. Direct reprogramming of human neural stem cells by OCT4. Nature. 2009;461:649–53.
Liu H, Ye Z, Kim Y, Sharkis S, Jang YY. Generation of endoderm-derived human induced pluripotent stem cells from primary hepatocytes. Hepatology. 2010;51:1810–9.
Mukamel Z, Hagai T, Gilad S, Amann-Zalcenstein D, Tanay A, Amit I, Novershtern N, Hanna JH. Deterministic direct reprogramming of somatic cells to pluripotency. Nature. 2013;502:65–70.
Yang R, Zheng Y, Li L, Liu S, Burrows M, Wei Z, Nace A, Herlyn M, Cui R, Guo W, Cotsarelis G, Xu X. Direct conversion of mouse and human fibroblasts to functional melanocytes by defined factors. Nat Commun. 2014;5:5807.
Lemper M, Leuckx G, Heremans Y, German MS, Heimberg H, Bouwens L, Baeyens L. Reprogramming of human pancreatic exocrine cells to beta-like cells. Cell Death Differ. 2014;22(7):1117–30.
Pulecio J, Nivet E, Sancho-Martinez I, Vitaloni M, Guenechea G, Xia Y, Kurian L, Dubova I, Bueren J, Laricchia-Robbio L, Izpisua Belmonte JC. Conversion of human fibroblasts into monocyte-like progenitor cells. Stem Cells. 2014;32:2923–38.
Kim YJ, Lim H, Li Z, Oh Y, Kovlyagina I, Choi IY, Dong X, Lee G. Generation of multipotent induced neural crest by direct reprogramming of human postnatal fibroblasts with a single transcription factor. Cell Stem Cell. 2014;15:497–506.
Sandler VM, Lis R, Liu Y, Kedem A, James D, Elemento O, Butler JM, Scandura JM, Rafii S. Reprogramming human endothelial cells to haematopoietic cells requires vascular induction. Nature. 2014;511:312–8.
Huang P, Zhang L, Gao Y, He Z, Yao D, Wu Z, Cen J, Chen X, Liu C, Hu Y, Lai D, Hu Z, Chen L, Zhang Y, Cheng X, Ma X, Pan G, Wang X, Hui L. Direct reprogramming of human fibroblasts to functional and expandable hepatocytes. Cell Stem Cell. 2014;14:370–84.
Du Y, Wang J, Jia J, Song N, Xiang C, Xu J, Hou Z, Su X, Liu B, Jiang T, Zhao D, Sun Y, Shu J, Guo Q, Yin M, Sun D, Lu S, Shi Y, Deng H. Human hepatocytes with drug metabolic function induced from fibroblasts by lineage reprogramming. Cell Stem Cell. 2014;14:394–403.
Zhang K, Liu GH, Yi F, Montserrat N, Hishida T, Esteban CR, Izpisua Belmonte JC. Direct conversion of human fibroblasts into retinal pigment epithelium-like cells by defined factors. Protein Cell. 2014;5:48–58.
Liu ML, Zang T, Zou Y, Chang JC, Gibson JR, Huber KM, Zhang CL. Small molecules enable neurogenin 2 to efficiently convert human fibroblasts into cholinergic neurons. Nat Commun. 2013;4:2183.
Wada R, Muraoka N, Inagawa K, Yamakawa H, Miyamoto K, Sadahiro T, Umei T, Kaneda R, Suzuki T, Kamiya K, Tohyama S, Yuasa S, Kokaji K, Aeba R, Yozu R, Yamagishi H, Kitamura T, Fukuda K, Ieda M. Induction of human cardiomyocyte-like cells from fibroblasts by defined factors. Proc Natl Acad Sci U S A. 2013;110:12667–72.
Hendry CE, Vanslambrouck JM, Ineson J, Suhaimi N, Takasato M, Rae F, Little MH. Direct transcriptional reprogramming of adult cells to embryonic nephron progenitors. J Am Soc Nephrol. 2013;24:1424–34.
Nam YJ, Song K, Luo X, Daniel E, Lambeth K, West K, Hill JA, DiMaio JM, Baker LA, Bassel-Duby R, Olson EN. Reprogramming of human fibroblasts toward a cardiac fate. Proc Natl Acad Sci U S A. 2013;110:5588–93.
Islas JF, Liu Y, Weng KC, Robertson MJ, Zhang S, Prejusa A, Harger J, Tikhomirova D, Chopra M, Iyer D, Mercola M, Oshima RG, Willerson JT, Potaman VN, Schwartz RJ. Transcription factors ETS2 and MESP1 transdifferentiate human dermal fibroblasts into cardiac progenitors. Proc Natl Acad Sci U S A. 2012;109:13016–21.
Ring KL, Tong LM, Balestra ME, Javier R, Andrews-Zwilling Y, Li G, Walker D, Zhang WR, Kreitzer AC, Huang Y. Direct reprogramming of mouse and human fibroblasts into multipotent neural stem cells with a single factor. Cell Stem Cell. 2012;11:100–9.
Liu X, Li F, Stubblefield EA, Blanchard B, Richards TL, Larson GA, He Y, Huang Q, Tan AC, Zhang D, Benke TA, Sladek JR, Zahniser NR, Li CY. Direct reprogramming of human fibroblasts into dopaminergic neuron-like cells. Cell Res. 2012;22:321–32.
Son EY, Ichida JK, Wainger BJ, Toma JS, Rafuse VF, Woolf CJ, Eggan K. Conversion of mouse and human fibroblasts into functional spinal motor neurons. Cell Stem Cell. 2011;9:205–18.
Ambasudhan R, Talantova M, Coleman R, Yuan X, Zhu S, Lipton Stuart A, Ding S. Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell. 2011;9:113–8.
Yoo AS, Sun AX, Li L, Shcheglovitov A, Portmann T, Li Y, Lee-Messer C, Dolmetsch RE, Tsien RW, Crabtree GR. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature. 2011;476:228–31.
Tapscott SJ, Davis RL, Thayer MJ, Cheng PF, Weintraub H, Lassar AB. MyoD1: a nuclear phosphoprotein requiring a Myc homology region to convert fibroblasts to myoblasts. Science. 1988;242:405–11.
Qian L, Huang Y, Spencer CI, Foley A, Vedantham V, Liu L, Conway SJ, Fu J-D, Srivastava D. In vivo reprogramming of murine cardiac fibroblasts into induced cardiomyocytes. Nature. 2012;485:593–8.
Li W, Nakanishi M, Zumsteg A, Shear M, Wright C, Melton DA, Zhou Q. In vivo reprogramming of pancreatic acinar cells to three islet endocrine subtypes. Elife. 2014;3, e01846.
Zhou Q, Brown J, Kanarek A, Rajagopal J, Melton DA. In vivo reprogramming of adult pancreatic exocrine cells to beta-cells. Nature. 2008;455:627–32.
De la Rossa A, Bellone C, Golding B, Vitali I, Moss J, Toni N, Luscher C, Jabaudon D. In vivo reprogramming of circuit connectivity in postmitotic neocortical neurons. Nat Neurosci. 2013;16:193–200.
Guo Z, Zhang L, Wu Z, Chen Y, Wang F, Chen G. In vivo direct reprogramming of reactive glial cells into functional neurons after brain injury and in an Alzheimer's disease model. Cell Stem Cell. 2014;14:188–202.
Murry CE, Keller G. Differentiation of embryonic stem cells to clinically relevant populations: lessons from embryonic development. Cell. 2008;132:661–80.
Laflamme MA, Murry CE. Regenerating the heart. Nat Biotechnol. 2005;23:845–56.
Kehat I, Kenyagin-Karsenti D, Snir M, Segev H, Amit M, Gepstein A, Livne E, Binah O, Itskovitz-Eldor J, Gepstein L. Human embryonic stem cells can differentiate into myocytes with structural and functional properties of cardiomyocytes. J Clin Invest. 2001;108:407–14.
Zhang J, Wilson GF, Soerens AG, Koonce CH, Yu J, Palecek SP, Thomson JA, Kamp TJ. Functional cardiomyocytes derived from human induced pluripotent stem cells. Circ Res. 2009;104:e30–41.
Burridge Paul W, Keller G, Gold Joseph D, Wu Joseph C. Production of de novo cardiomyocytes: human pluripotent stem cell differentiation and direct reprogramming. Cell Stem Cell. 2012;10:16–28.
Lian X, Hsiao C, Wilson G, Zhu K, Hazeltine LB, Azarin SM, Raval KK, Zhang J, Kamp TJ, Palecek SP. Robust cardiomyocyte differentiation from human pluripotent stem cells via temporal modulation of canonical Wnt signaling. Proc Natl Acad Sci U S A. 2012;109(27):E1848–57.
Beqqali A, Kloots J, Ward-van Oostwaard D, Mummery C, Passier R. Genome-wide transcriptional profiling of human embryonic stem cells differentiating to cardiomyocytes. Stem Cells. 2006;24:1956–67.
Burridge PW, Thompson S, Millrod MA, Weinberg S, Yuan X, Peters A, Mahairaki V, Koliatsos VE, Tung L, Zambidis ET. A universal system for highly efficient cardiac differentiation of human induced pluripotent stem cells that eliminates interline variability. PLoS ONE. 2011;6, e18293.
Uosaki H, Fukushima H, Takeuchi A, Matsuoka S, Nakatsuji N, Yamanaka S, Yamashita JK. Efficient and scalable purification of cardiomyocytes from human embryonic and induced pluripotent stem cells by VCAM1 surface expression. PLoS ONE. 2011;6, e23657.
Yang L, Soonpaa MH, Adler ED, Roepke TK, Kattman SJ, Kennedy M, Henckaerts E, Bonham K, Abbott GW, Linden RM, Field LJ, Keller GM. Human cardiovascular progenitor cells develop from a KDR+ embryonic-stem-cell-derived population. Nature. 2008;453:524–8.
Ma J, Guo L, Fiene SJ, Anson BD, Thomson JA, Kamp TJ, Kolaja KL, Swanson BJ, January CT. High purity human-induced pluripotent stem cell-derived cardiomyocytes: electrophysiological properties of action potentials and ionic currents. Am J Physiol Heart Circ Physiol. 2011;301:H2006–17.
Liu J, Sun N, Bruce MA, Wu JC, Butte MJ. Atomic force mechanobiology of pluripotent stem cell-derived cardiomyocytes. PLoS ONE. 2012;7, e37559.
Pillekamp F, Haustein M, Khalil M, Emmelheinz M, Nazzal R, Adelmann R, Nguemo F, Rubenchyk O, Pfannkuche K, Matzkies M, Reppel M, Bloch W, Brockmeier K, Hescheler J. Contractile properties of early human embryonic stem cell-derived cardiomyocytes: beta-adrenergic stimulation induces positive chronotropy and lusitropy but not inotropy. Stem Cells Dev. 2012;21(12):2111–21.
Klug MG, Soonpaa MH, Koh GY, Field LJ. Genetically selected cardiomyocytes from differentiating embronic stem cells form stable intracardiac grafts. J Clin Invest. 1996;98:216–24.
Cao F, Wagner RA, Wilson KD, Xie X, Fu JD, Drukker M, Lee A, Li RA, Gambhir SS, Weissman IL, Robbins RC, Wu JC. Transcriptional and functional profiling of human embryonic stem cell-derived cardiomyocytes. PLoS ONE. 2008;3, e3474.
Binah O, Dolnikov K, Sadan O, Shilkrut M, Zeevi-Levin N, Amit M, Danon A, Itskovitz-Eldor J. Functional and developmental properties of human embryonic stem cells-derived cardiomyocytes. J Electrocardiol. 2007;40:S192–6.
He JQ, Ma Y, Lee Y, Thomson JA, Kamp TJ. Human embryonic stem cells develop into multiple types of cardiac myocytes: action potential characterization. Circ Res. 2003;93:32–9.
Satin J, Itzhaki I, Rapoport S, Schroder EA, Izu L, Arbel G, Beyar R, Balke CW, Schiller J, Gepstein L. Calcium handling in human embryonic stem cell-derived cardiomyocytes. Stem Cells. 2008;26:1961–72.
Guan X, Mack DL, Moreno CM, Strande JL, Mathieu J, Shi Y, Markert CD, Wang Z, Liu G, Lawlor MW, Moorefield EC, Jones TN, Fugate JA, Furth ME, Murry CE, Ruohola-Baker H, Zhang Y, Santana LF, Childers MK. Dystrophin-deficient cardiomyocytes derived from human urine: new biologic reagents for drug discovery. Stem Cell Res. 2014;12:467–80.
Zhang Y, McNeill E, Tian H, Soker S, Andersson KE, Yoo JJ, Atala A. Urine derived cells are a potential source for urological tissue reconstruction. J Urol. 2008;180:2226–33.
Bharadwaj S, Liu G, Shi Y, Wu R, Yang B, He T, Fan Y, Lu X, Zhou X, Liu H, Atala A, Rohozinski J, Zhang Y. Multipotential differentiation of human urine-derived stem cells: potential for therapeutic applications in urology. Stem Cells. 2013;31:1840–56.
Laflamme MA, Chen KY, Naumova AV, Muskheli V, Fugate JA, Dupras SK, Reinecke H, Xu C, Hassanipour M, Police S, O'Sullivan C, Collins L, Chen Y, Minami E, Gill EA, Ueno S, Yuan C, Gold J, Murry CE. Cardiomyocytes derived from human embryonic stem cells in pro-survival factors enhance function of infarcted rat hearts. Nat Biotechnol. 2007;25:1015–24.
Lee G, Papapetrou EP, Kim H, Chambers SM, Tomishima MJ, Fasano CA, Ganat YM, Menon J, Shimizu F, Viale A, Tabar V, Sadelain M, Studer L. Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature. 2009;461:402–6.
Marchetto MC, Carromeu C, Acab A, Yu D, Yeo GW, Mu Y, Chen G, Gage FH, Muotri AR. A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell. 2010;143:527–39.
Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N, Sangar S, Li Y, Mu Y, Chen G, Yu D, McCarthy S, Sebat J, Gage FH. Modelling schizophrenia using human induced pluripotent stem cells. Nature. 2011;473:221–5.
Cooper O, Seo H, Andrabi S, Guardia-Laguarta C, Graziotto J, Sundberg M, McLean JR, Carrillo-Reid L, Xie Z, Osborn T, Hargus G, Deleidi M, Lawson T, Bogetofte H, Perez-Torres E, Clark L, Moskowitz C, Mazzulli J, Chen L, Volpicelli-Daley L, Romero N, Jiang H, Uitti RJ, Huang Z, Opala G, Scarffe LA, Dawson VL, Klein C, Feng J, Ross OA, Trojanowski JQ, Lee VM-Y, Marder K, Surmeier DJ, Wszolek ZK, Przedborski S, Krainc D, Dawson TM, Isacson O. Pharmacological rescue of mitochondrial deficits in iPSC-derived neural cells from patients with familial Parkinson’s disease. Sci Transl Med. 2012;4, 141ra190.
Kaplitt MG, Feigin A, Tang C, Fitzsimons HL, Mattis P, Lawlor PA, Bland RJ, Young D, Strybing K, Eidelberg D, During MJ. Safety and tolerability of gene therapy with an adeno-associated virus (AAV) borne GAD gene for Parkinson's disease: an open label, phase I trial. Lancet. 2007;369:2097–105.
Ryan Scott D, Dolatabadi N, Chan Shing F, Zhang X, Akhtar Mohd W, Parker J, Soldner F, Sunico Carmen R, Nagar S, Talantova M, Lee B, Lopez K, Nutter A, Shan B, Molokanova E, Zhang Y, Han X, Nakamura T, Masliah E, Yates John R, Nakanishi N, Andreyev Aleksander Y, Okamoto S-i, Jaenisch R, Ambasudhan R, Lipton Stuart A. Isogenic human iPSC Parkinson s model shows nitrosative stress-induced dysfunction in MEF2-PGC1± transcription. Cell. 2013;155:1351–64.
Soldner F, Laganière J, Cheng Albert W, Hockemeyer D, Gao Q, Alagappan R, Khurana V, Golbe Lawrence I, Myers Richard H, Lindquist S, Zhang L, Guschin D, Fong Lauren K, Vu BJ, Meng X, Urnov Fyodor D, Rebar Edward J, Gregory Philip D, Zhang HS, Jaenisch R. Generation of isogenic pluripotent stem cells differing exclusively at two early onset Parkinson point mutations. Cell. 2011;146:318–31.
Yazawa M, Hsueh B, Jia X, Pasca AM, Bernstein JA, Hallmayer J, Dolmetsch RE. Using induced pluripotent stem cells to investigate cardiac phenotypes in Timothy syndrome. Nature. 2011;471:230–4.
Malan D, Friedrichs S, Fleischmann BK, Sasse P. Cardiomyocytes obtained from induced pluripotent stem cells with long-QT syndrome 3 recapitulate typical disease-specific features in vitro. Circ Res. 2011;109:841–7.
Sacco A, Mourkioti F, Tran R, Choi J, Llewellyn M, Kraft P, Shkreli M, Delp S, Pomerantz JH, Artandi SE, Blau HM. Short telomeres and stem cell exhaustion model Duchenne muscular dystrophy in mdx/mTR mice. Cell. 2010;143:1059–71.
Braunwald E. Biomarkers in heart failure. N Engl J Med. 2008;358:2148–59.
Coca SG, Yalavarthy R, Concato J, Parikh CR. Biomarkers for the diagnosis and risk stratification of acute kidney injury: a systematic review. Kidney Int. 2008;73:1008–16.
Berg JS, Khoury MJ, Evans JP. Deploying whole genome sequencing in clinical practice and public health: meeting the challenge one bin at a time. Genet Med. 2011;13:499–504.
Rizzo JM, Buck MJ. Key principles and clinical applications of “next-generation” DNA sequencing. Cancer Prev Res. 2012;5:887–900.
Kim C, Wong J, Wen J, Wang S, Wang C, Spiering S, Kan NG, Forcales S, Puri PL, Leone TC, Marine JE, Calkins H, Kelly DP, Judge DP, Chen HS. Studying arrhythmogenic right ventricular dysplasia with patient-specific iPSCs. Nature. 2013;494:105–10.
Lan F, Lee AS, Liang P, Sanchez-Freire V, Nguyen PK, Wang L, Han L, Yen M, Wang Y, Sun N, Abilez OJ, Hu S, Ebert AD, Navarrete EG, Simmons CS, Wheeler M, Pruitt B, Lewis R, Yamaguchi Y, Ashley EA, Bers DM, Robbins RC, Longaker MT, Wu JC. Abnormal calcium handling properties underlie familial hypertrophic cardiomyopathy pathology in patient-specific induced pluripotent stem cells. Cell Stem Cell. 2013;12:101–13.
Wang Y, Liang P, Lan F, Wu H, Lisowski L, Gu M, Hu S, Kay MA, Urnov FD, Shinnawi R, Gold JD, Gepstein L, Wu JC. Genome editing of isogenic human induced pluripotent stem cells recapitulates long QT phenotype for drug testing. J Am Coll Cardiol. 2014;64:451–9.
Yang YM, Gupta SK, Kim KJ, Powers BE, Cerqueira A, Wainger BJ, Ngo HD, Rosowski KA, Schein PA, Ackeifi CA, Arvanites AC, Davidow LS, Woolf CJ, Rubin LL. A small molecule screen in stem-cell-derived motor neurons identifies a kinase inhibitor as a candidate therapeutic for ALS. Cell Stem Cell. 2013;12:713–26.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339:819–23.
Gonzalez F, Zhu Z, Shi ZD, Lelli K, Verma N, Li QV, Huangfu D. An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells. Cell Stem Cell. 2014;15:215–26.
Li HL, Fujimoto N, Sasakawa N, Shirai S, Ohkame T, Sakuma T, Tanaka M, Amano N, Watanabe A, Sakurai H, Yamamoto T, Yamanaka S, Hotta A. Precise correction of the dystrophin gene in Duchenne muscular dystrophy patient induced pluripotent stem cells by TALEN and CRISPR-Cas9. Stem Cell Rep. 2014;4(1):143–54.
Veres A, Gosis BS, Ding Q, Collins R, Ragavendran A, Brand H, Erdin S, Cowan CA, Talkowski ME, Musunuru K. Low incidence of off-target mutations in individual CRISPR-Cas9 and TALEN targeted human stem cell clones detected by whole-genome sequencing. Cell Stem Cell. 2014;15:27–30.
Filareto A, Parker S, Darabi R, Borges L, Iacovino M, Schaaf T, Mayerhofer T, Chamberlain JS, Ervasti JM, McIvor RS, Kyba M, Perlingeiro RC. An ex vivo gene therapy approach to treat muscular dystrophy using inducible pluripotent stem cells. Nat Commun. 2013;4:1549.
Yamada S, Nelson TJ, Crespo-Diaz RJ, Perez-Terzic C, Liu XK, Miki T, Seino S, Behfar A, Terzic A. Embryonic stem cell therapy of heart failure in genetic cardiomyopathy. Stem Cells. 2008;26:2644–53.
Moretti A, Bellin M, Welling A, Jung CB, Lam JT, Bott-Flugel L, Dorn T, Goedel A, Hohnke C, Hofmann F, Seyfarth M, Sinnecker D, Schomig A, Laugwitz KL. Patient-specific induced pluripotent stem-cell models for long-QT syndrome. N Engl J Med. 2010;363:1397–409.
Young LE, Platzer RF, Ervin DM, Izzo MJ. Hereditary spherocytosis. II. Observations on the role of the spleen. Blood. 1951 Nov;6(11):1099–1113.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this chapter
Cite this chapter
Guan, X., Mack, D., Childers, M.K. (2016). Patient-Derived Induced Pluripotent Stem Cells Provide a Regenerative Medicine Platform for Duchenne Muscular Dystrophy Heart Failure. In: Childers, M. (eds) Regenerative Medicine for Degenerative Muscle Diseases. Stem Cell Biology and Regenerative Medicine. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3228-3_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3228-3_6
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3227-6
Online ISBN: 978-1-4939-3228-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)