Abstract
Allosteric ligands modulate binding and function of muscarinic receptors in a different way than orthosteric ligands. Unlike orthosteric ligands their effects are limited by a cooperativity factor. This imparts them unique properties, including cooperativity-based selectivity, functional selectivity and restoring of physiological-like space and time pattern of signaling under pathological conditions. Therefore, allosteric modulators of muscarinic receptor are intensively studied as possible therapeutics of pathological conditions including Alzheimer’s disease and schizophrenia. Research of allosteric modulation has pioneered the way for a whole class A of G-protein coupled receptors and has had an impact beyond its own field. We review principles of allosteric modulations and their implications for proper design of binding as well as functional experiments and for proper data analysis. We demonstrate immense complexity of allosteric modulation of functional responses. Such complexity is reflected in the inability to determine individual microscopic constants in allosterically modulated systems. Therefore, the effects of a given allosteric modulator can be characterized by only two macroscopic parameters, namely a change in the agonist potency and efficacy. We also discuss distinct properties of allosteric interactions that are specific to muscarinic receptors.
Access provided by CONRICYT – Journals CONACYT. Download protocol PDF
Similar content being viewed by others
Key words
1 Historic Overview
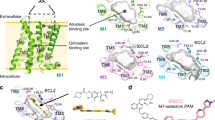
The concept of allosterism was formally introduced into the field of enzymology by Monod et al. [1, 2] in their description of a generalized model of oligomeric enzymes that contained a number of “stereo-specifically different, non-overlapping receptor sites.” The substrate was said to bind at the “primary” or “active” site. On the other hand, an “allosteric effector” (from the Greek word “allo” meaning “other”) was defined as a molecule that binds to a site other than the primary binding site. In pharmacology, the term orthosteric ligand denotes a compound that binds to the same binding site as endogenous ligand (neurotransmitter or hormone), while a ligand that binds to other sites on the receptor is termed allosteric. Allosteric ligands influence (modulate) binding and effects of orthosteric ligands in a different way than orthosteric ligands. In the pioneering work by Clark and Mitchelson, gallamine was found to shift the concentration-response curves of acetylcholine in inhibiting the heart atrium to the right but the magnitude of the shifts was smaller than expected for conventional competitive receptor antagonists [3]. They proposed that gallamine interacts with a secondary allosteric site on the receptor. This notion was later confirmed in radioligand binding studies [4]. Subsequently, a wide variety of allosteric modulators of muscarinic receptors was identified including toxiferous alkaloids [5], the L-calcium channel blocker verapamil [6], the potassium channel inhibitor 4-aminopyridine [7], inhibitors of acetylcholinesterase [8–10], strychnos and vinca alkaloids [11], and antibiotics like staurosporine [12]. Numerous site-directed mutagenesis studies located allosteric binding sites for most allosteric modulators to the extracellular domain of the receptor, namely between the second and the third extracellular loops [13–20]. For further details see review by Jakubík and El-Fakahany [21]. This has been recently confirmed by crystallographic studies [22]. Allosteric modulation of muscarinic receptors has been intensively studied for decades for its perspective role in therapy of many pathological conditions including Alzheimer’s disease and schizophrenia [23, 24].
2 Principles of Allosteric Modulation
2.1 Allosteric Modulation of Ligand Binding
By definition, allosteric ligands bind to a site on the receptor that is spatially distinct from that of endogenous ligands of the receptor. Consequently, binding of an allosteric ligand (A) and an orthosteric ligand (L) to the receptor (R) is not mutually exclusive, i.e., both ligands may bind to the receptor to form a ternary complex LRA (Scheme 1). Binding of allosteric modulators induces a change in the conformation of the receptor that results in changes in binding and/or effects of the orthosteric ligand. The law of microscopic reversibility of thermodynamics dictates that binding of orthosteric ligand L affects binding of allosteric ligand A in the same way in which the allosteric ligand A affects binding of the orthosteric ligand L. A situation when L and A mutually strengthen each other’s binding is called positive cooperativity, i.e., formation of the ternary complex LRA leads to increase in the affinity of both ligands, that is a decrease in the equilibrium dissociation constants of orthosteric ligand (K D) and allosteric ligand (K A) (thus α < 1) (Scheme 1, Fig. 1 upper graph, green curve). The opposite situation when L and A mutually weaken their binding is called negative cooperativity. Under negative cooperativity formation of the ternary complex LRA leads to a mutual decrease in affinity, that is an increase in equilibrium dissociation constants of orthosteric ligand (K D) and allosteric ligand (K A) (thus α > 1) (Scheme 1, Fig. 1 upper graph, red curve). In other words, binding of L to R is stronger or weaker in the presence of A in case of positive or negative cooperativity, respectively. In rare situations the two ligands form a ternary complex with the receptor without mutually changing their affinities is called neutral cooperativity (α = 1) (Fig. 1 upper graph, blue curve). In this particular case the affinity of L for R or AR is the same. This knowledge is very important in design of binding experiments.
Scheme of allosteric interaction. An orthosteric ligand L binds to the receptor R with equilibrium dissociation constant K D and an allosteric modulator A binds to the receptor R with equilibrium dissociation constant K A . The orthosteric ligand L and the allosteric modulator A can bind concurrently to the receptor R to form a ternary complex LRA. Binding of one ligand to the receptor changes the equilibrium dissociation constant of the other ligand by factor of cooperativity α
Allosteric modulation of binding. Theoretical curves of tracer equilibrium saturation binding (upper graph) and tracer dissociation (lower graph) in the absence (black) or in the presence of positive (green), negative (red) or neutral (blue) allosteric modulator. Tracer binding (in binary LR and ternary LRA complexes) is expressed as a fraction of total receptor number RT (upper graph) or fraction of complexes at start of dissociation (lower graph). A positive allosteric modulator causes an increase in tracer affinity (decrease in tracer equilibrium dissociation constant K D) (upper graph) and slow down of tracer dissociation (lower graph). A negative allosteric modulator causes a decrease in tracer affinity (increase in tracer equilibrium dissociation constant K D) (upper graph) and acceleration of tracer dissociation (lower graph)
In general ligand association is a fast process that closely parallels ligand diffusion to the receptor. Thus allosteric effects on ligand binding are usually manifested in changes in ligand dissociation; by slowing or accelerating dissociation in case of positive and negative cooperativity, respectively (Fig. 1 lower graph). Because changes in kinetics of ligand binding do not strictly follow changes in ligand affinity, binding of an allosteric agent with neutral cooperativity may be detected by changes in the rate of dissociation of an orthosteric ligand. As changes in ligand kinetics are not possible without formation of the ternary complex they become a hallmark of allosteric interaction and the most straightforward way to identify it.
It should be noted that effects of allosteric modulator on both equilibrium and kinetic binding and on functional effects of orthosteric ligands is limited by a cooperativity factor α. For example, in Scheme 1, with increasing concentrations of the allosteric modulator A the equilibrium dissociation constant of the orthosteric ligand L changes from its original value K D until it reaches a value α × K D. Further increase in the concentration of A does not bring further change in K D. That is in contrast to competition of two orthosteric ligands for the same site where changes in binding of one orthosteric ligand are proportional to the concentration of the other orthosteric ligand without a maximal limit (see Chapter 3, Fig. 5). The level of maximum effect of allosteric modulator (also known as “ceiling effect”) would confer safety under conditions of overdosage. This represents an advantage of development of allosteric receptor modulators for therapeutics purposes.
A conformational change of the receptor induced by an allosteric ligand has different effects on binding of structurally different orthosteric ligands as well as structurally different receptor subtypes. Thus, the factor of cooperativity α depends on combination of all three constituents of allosteric interaction: the receptor and the orthosteric and allosteric ligands. For example eburnamonine decreases affinity of the agonist arecoline at the muscarinic M2 receptor, but has no effect on the affinity of the agonist arecaidine propargyl ester. Interestingly, it increases the affinity of the agonist pilocarpine. In contrast, eburnamonine decreases the affinity of all three agonists at M1 and M3 muscarinic receptors [11].
2.2 Anomalies in Allosteric Modulation of Functional Response
Allosteric modulators may affect receptor activation by mechanisms additional to effects on binding of an orthosteric agonist to the receptor. Thus for a given pair of allosteric modulator and orthosteric agonist, positive cooperativity in binding does not necessarily translate into an increase in agonist potency in functional assays and may even lead to “allosteric quenching of agonist efficacy” [25, 26]. Moreover, an allosteric modulator that exerts negative binding cooperativity with an agonist at equilibrium may potentiate agonist-induced activation of the receptor due to acceleration of agonist binding [27].
Another feature of allosteric modulators of muscarinic receptors is their ability to activate the receptors in the absence of agonists. For example, partial stimulation of accumulation of inositol phosphates and inhibition of accumulation of cAMP in response to strychnine-like allosteric modulators has been reported [28]. A whole new class of potent allosteric agonists has been reported recently [29–31].
3 Promises of Allosteric Targeting of Muscarinic Receptors
3.1 Selectivity by Targeting Less Conserved Domains
Muscarinic receptor subtypes share high structural homology in the transmembrane domains where the orthosteric binding site is located. On the other hand, domains out of membrane are less conserved. Targeting allosteric domains allows achieving binding selectivity to an extent that is not possible by orthosteric ligands. Allosteric modulators exhibit a wide range of selectivity for different muscarinic receptor subtypes. For example, while prototypical allosteric modulators like alcuronium and gallamine display selectivity towards M2 receptors [32], strychnine is M3 selective [11] and WIN compounds are M4 selective [33].
3.2 Conservation of Space and Time Pattern of Signaling
Theoretically, a positive allosteric modulator of acetylcholine that has no efficacy on its own would only induce an action when the endogenous acetylcholine is released. Consequently, its action would be restricted in space and time to those regions of the body where signaling is actually taking place. Thus, space and time pattern of signaling could be restored under pathological conditions of diminished acetylcholine release, e.g., degeneration of cholinergic neurons in certain brain areas in Alzheimer’s disease.
3.3 Absolute Selectivity
Theoretically, absolute selectivity is achieved by having an allosteric agent with the intended positive or negative cooperativity in combination with the orthosteric ligand at one subtype of the receptor and neutral cooperativity at the rest of the subtypes. Selectivity may be derived from binding and/or activation cooperativity. As proof of concept such cooperativity-based binding selectivity for thiochrome at the M4 muscarinic receptor has been reported [34].
4 Analyzing Allosteric Modulation of Ligand Binding
The methodology of radioligand binding at muscarinic receptors is described in detail in a previous chapter. Here we only describe the setup and data analysis of experiments with allosteric ligands. The major complication of radioligand binding experiments with allosteric modulators is the lack of suitable allosteric radioligands. So far only a few radiolabeled allosteric ligands are available. These include tritiated ABA-type like compounds (Fig. 2) dimethyl-W84 [35], derivatives of α-truxillic acid anatruxonium and truxillonium [36], and iodinated proteins like the muscarinic toxins MTX2 [37], MTX1 [38, 39], and MTX7 [40]. Tritiated ABA-type compounds have relatively low affinity and extremely high nonspecific binding in comparison to orthosteric antagonists that make their use as tracers difficult. Fluorescent labeling and detection of binding by FRET seems to be the way to reduce nonspecific binding [41, 42]. Muscarinic toxins display very slow kinetics that lead to kinetic artifacts [37, 40]. None of muscarinic radiolabeled ligands are available commercially. Thus almost all binding studies of muscarinic allosteric ligands are conducted indirectly and their binding parameters are inferred from changes in binding of orthosteric tracers.
Radiolabeled allosteric modulators. ABA-type compounds that have been experimentally tritiated. Dimethyl-W84 (6-[dimethyl-[3-(4-methyl-1,3-dioxoisoindol-2-yl)propyl]azaniumyl]hexyl-dimethyl-[3-(4-methyl-1,3-dioxoisoindol-2-yl)propyl]azanium), anatruxonium (1,1′-[(2,4-diphenylcyclobutane-1,3-diyl)bis(carbonyloxypropane-3,1-diyl)]bis(1-ethylpiperidinium)) and truxillonium (bis[4-(1-methylpiperidin-1-ium-1-yl)butyl]2,4-diphenylcyclobutane-1,3-dicarboxylate)
4.1 Allosteric Modulation of Tracer in Saturation Binding Experiments
Equilibrium dissociation constant of the allosteric ligand K A and factor of cooperativity α can be determined from a series of experiments of tracer saturation binding. Equilibrium dissociation constants in Scheme 1 are defined as follows:
Total number of receptors is sum of free receptors and binary and ternary complexes:
Fraction of receptors occupied by tracer L:
Multiplying the numerator and denominator of the fraction on the right side of Eq. (3) by 1/[L][R] gives:
Substituting Eq. (4) by Eq. (1a):
Substituting Eq. (5) by Eqs. (1b) and (1c):
After simplification:
where K ′D is the apparent equilibrium dissociation constant of the tracer in the presence of allosteric modulator A that is given as:
For saturating concentrations of the allosteric modulator ([A] ≫ K A) Eq. (7b) can be reduced to:
As can be seen, an allosteric modulator affects the tracer equilibrium dissociation constant (Eq. 7b) without a change in binding capacity (Eq. 7a). Maximum change in the equilibrium dissociation constant is given by the cooperativity factor α (Eq. 8). In case of positive cooperativity the tracer’s equilibrium dissociation constant decreases with increasing the concentration of the allosteric modulator (Eq. 7b; Fig. 3, upper graph). Such effects are unique to allosteric interaction and thus positive allosteric modulators are easily identified. In case of negative cooperativity the tracer’s equilibrium dissociation constant increases with increasing the concentration of an allosteric modulator (Eq. 7b; Fig. 3, lower graph). At first glance this is similar to competition of orthosteric ligand with the tracer for the same binding site (Fig. 4). However, as stated above, at negative cooperativity the decrease in tracer affinity has its limit given by the cooperativity factor α, while the effects of a classical competitive interaction are directly proportional to the competitor’s concentration without a limit. While plotting of tracer equilibrium dissociation constant against concentration of a competitor gives a straight line (with slope equal to 1 and constant equal to 1) (Fig. 5, circles) plotting tracer equilibrium constant against the concentration of a negative allosteric modulator gives a hyperbole with asymptote equal to the cooperativity factor α (Fig. 5, squares). It can be seen that deviations from competitive behavior are more obvious at high concentrations of a negative allosteric modulator. Likewise, plotting tracer equilibrium dissociation constant against the concentration of a positive allosteric modulator gives an inverse hyperbole with asymptote equal to the cooperativity factor α (Fig. 5, triangles).
Allosteric modulation of tracer saturation binding. Effects of positive (α = 0.1) (upper graph) and negative (α = 10) (lower graph) allosteric modulators on tracer saturation binding. Abscissa, the concentration of tracer L is expressed as a logarithm of the ratio to its equilibrium dissociation constant K D. Ordinate, tracer binding is expressed as a fraction of total receptor number RT. Legend, the concentration of allosteric modulator A is expressed as a ratio to its equilibrium dissociation constant K A. A positive allosteric modulator concentration dependently decreases tracer K D, while a negative allosteric modulator concentration dependently increases tracer K D
Competition with tracer saturation binding. Effects of a competitor on tracer saturation binding. Abscissa, concentration of tracer L is expressed as the logarithm of the ratio to its equilibrium dissociation constant K D. Ordinate, tracer binding is expressed as a fraction of total receptor number RT. Legend, concentration of competitor X is expressed as ratio to its equilibrium dissociation constant K X
Shifts in equilibrium dissociation constants. From Figs. 3 and 4 shifts in equilibrium dissociation constant caused by negative allosteric modulator (α = 10) (squares), positive allosteric modulator (α = 0.1) (triangles) and competitive ligand (circles) are expressed as the logarithm of the ratio of tracer apparent equilibrium dissociation constant K ′D in the presence of the second ligand to tracer equilibrium dissociation constant K D in its absence. Abscissa, concentration of the second ligand is expressed as logarithm of the ratio of second ligand concentration to its equilibrium dissociation constant
When equilibrium dissociation constant of the allosteric ligand K A and factor of cooperativity α are determined from a series of experiments of tracer saturation binding (like in Fig. 3) Eq. (7a) is fitted to data and apparent equilibrium dissociation constant of the tracer K ′D is determined for each concentration of the allosteric modulator A. Then the obtained K ′D values are plotted against the concentration of A (like in Fig. 5) and Eq. (7b) is fitted to data to determine the equilibrium dissociation constant of allosteric modulator K A and the cooperativity factor α.
4.2 Displacement Binding Experiments
Determination of the equilibrium dissociation constant of the allosteric ligand K A and factor of cooperativity α from a series of experiments of tracer saturation binding is laborious and expensive. Binding parameters of an allosteric ligand can be determined in a simpler way by measuring the effects of increasing concentrations of an allosteric modulator on binding of a single concentration of the tracer. Binding of tracer L at fixed concentration in the presence of various concentrations of allosteric modulator A is described by Eq. (7) (Fig. 6). It is more convenient to express the data as a fraction of the tracer binding in the presence of the allosteric modulator than its binding in the absence of the allosteric modulator (Fig. 7) that is given:
Effects of an allosteric modulator on binding of the tracer at a fixed concentration. Effects of various concentrations of a positive (α = 0.1) (upper graph) and negative (α = 10) (lower graph) allosteric modulators at indicated on the abscissa on binding of the tracer at fixed concentration indicated in legend. Abscissa, concentration of allosteric modulator is expressed as logarithm of ratio to its equilibrium dissociation constant K A. Ordinate, the tracer binding is expressed as a fraction of total receptor number RT. Legend, concentration of the tracer L is expressed as ratio to its equilibrium dissociation constant K D. A positive allosteric modulator concentration dependently increases tracer binding, while a negative allosteric modulator concentration dependently decreases tracer binding. Changes in tracer binding are more obvious at lower concentrations of the tracer
Effects of an allosteric modulator on binding of the tracer at a fixed concentration. Effects of a positive (α = 0.1) (upper graph) and negative (α = 10) (lower graph) allosteric modulators at various concentrations indicated on the abscissa on binding of the tracer at a fixed concentration indicated in the legend. Abscissa, concentration of allosteric modulator is expressed as logarithm of the ratio to its equilibrium dissociation constant K A. Ordinate, tracer binding is expressed as a fraction of its binding in the absence of allosteric modulator. Legend, concentration of the tracer L is expressed as a ratio to its equilibrium dissociation constant K D. A positive allosteric modulator concentration dependently increases tracer binding while a negative allosteric modulator concentration dependently decreases tracer binding. Changes in tracer binding are greater at lower concentrations of the tracer
After substitution of Eq. (9) with Eq. (7b):
When the equilibrium dissociation constant of allosteric modulator K A and the cooperativity factor α are determined by measuring binding at a single concentration of the tracer and various concentrations of allosteric modulator Eq. (10) is fitted to data expressed as ratio of tracer binding in the presence of A to the tracer binding in the absence of A (like in Fig. 7). Equilibrium dissociation of the tracer K D has to be determined in separate measurements. Precise concentration of the tracer L used in the assay should be determined by counting total radioactivity added to the sample and division by specific radioactivity of the tracer and sample volume. It can be seen from equations describing allosteric binding that the equilibrium dissociation constant of an allosteric modulator K A and cooperativity factor α are interdependent parameters. Overestimation of α leads to underestimation of K A and vice versa. Thus a wide range of concentrations of A has to be used. For proper determination of α a saturating concentration of A has to be used to reach a curve plateau. This plateau defines α according to Eq. (9) where for a saturating concentration of A the apparent equilibrium dissociation constant of tracer K ′D becomes αK D. The value of αK A has to be determined properly for accurate determination of K A. The value of αK A corresponds to the inflection point of the binding curve. The latter is best determined by measuring binding at several concentrations close to the inflection point (αK A).
Positive cooperativity is easily spotted as an increase in tracer binding to free receptors (Figs. 6 and 7, upper graphs). The fraction of free receptors decreases with increasing tracer concentration. For positive allosteric modulators with strong cooperativity the fraction of free receptors may be limiting. If a saturating concentration of the tracer is used it is difficult to reliably determine the cooperativity factor α because subtle differences in the level of plateau represent huge differences in α. So in binding experiments with strong positive allosteric modulators low concentrations of the tracer (below its K D) are desired to prevent full receptor occupancy at saturating concentrations of the allosteric modulator. However too low concentration of the tracer gives low and unreliable control binding in the absence of allosteric modulator. Thus a compromise between the size of allosteric change and the quality of control binding has to be achieved. In the model case of positive cooperativity with α = 0.1 a tracer concentration equal to its K D gives sufficient (>80 %) increase in tracer binding and sufficient proportion (almost 20 %) of the receptors remain free at saturating concentrations of the allosteric ligand (Fig. 7, upper graph).
Negative binding cooperativity can be distinguished from competitive binding by incomplete inhibition of tracer binding, resulting in a plateau in the displacement curve (Figs. 6 and 7, lower graphs). The higher the concentration of tracer, the higher the level of the plateau (less complete inhibition of tracer binding) (Fig. 7). Thus, detection of strong negative cooperativity requires the use of a tracer concentration several times higher than its dissociation constant K D to get incomplete inhibition of tracer binding. However, the ratio of specific to nonspecific binding decreases with increasing tracer concentration, indicating that extremely high concentrations of the tracer should be avoided. It must be noted that allosteric modulators with very high cooperativity factors (α > 100) would cause almost complete inhibition of tracer binding, making them indistinguishable from competitive binding. A more experimental setup in this special case is to construct saturation curves for the tracer in the absence and in the presence of increasing concentrations of the allosteric modulator. The apparent equilibrium dissociation constant of the tracer K ′D is determined for each concentration of the allosteric modulator A (like in Fig. 3, lower graph) and plotted against concentration of A (like in Fig. 5, squares) and Eq. (7b) is fitted to data.
Ligands with weak (either positive or negative) cooperativity induce small changes in tracer binding. As can be seen in Eq. (9) the lower concentration of L the greater the change in fractional binding. Thus low concentrations of the tracer are desired to magnify changes induced by weak allosteric modulators. It can be demonstrated using Eq. (10) that even for ligands with very weak cooperativity (0.9 < α < 1.1) lowering tracer concentration below 0.1 × K D does not bring further increase in allosteric effects. Maximum attainable increase in tracer binding by positive allosteric modulators with α = 0.9 is 10 % of the control binding and, analogically, maximum attainable decrease in tracer binding is 10 % by very weak negative allosteric modulators with α = 1.1. Such small changes may be problematic to detect and are usually considered as neutral cooperativity.
Protocol A: Determination of K A and α in equilibrium experiments (96-well plate setup)
-
1.
Determine the equilibrium dissociation constant K D of [3H]NMS in saturation binding experiments in a buffer of your choice at 25 °C (see previous Chapter 3).
-
2.
Add membranes, about 20 fmol of receptors per well.
-
3.
Add [3H]NMS to a final concentration around 0.5 × K D (when positive cooperativity is expected) or 2 × K D (when negative cooperativity is expected) for 60 min at 25 °C in a final incubation volume of 0.4 ml.
-
4.
Add tested allosteric modulators to final concentrations ranging from 10 nM to 100 μM (9 concentrations at 0.5 log concentration steps). Make samples of control binding in the absence of allosteric modulator, samples of nonspecific binding in the presence of 1 μM atropine alone and in the presence of 100 μM allosteric modulator (to verify that the allosteric modulator does not change nonspecific binding).
-
5.
Seal the plate and incubate for 20 h at 25 °C (see justification below for the long incubation time).
-
6.
Filter samples though GF/C filters or a filtration plate. Wash with ice-cold deionized water for 6 s.
-
7.
Determine the exact tracer concentration used in the experiment by counting added radioactivity divided by specific radioactivity and incubation volume (0.4 ml).
-
8.
Fit Eq. (10) to specific binding expressed as a fraction of control. Use K D of [3H]NMS determined in saturation binding experiment and exact radioligand concentration L from step 7.
4.3 Allosteric Modulation of Tracer Binding Kinetics
Usually association of substrates or ligands with enzymes or receptors is a fast process, being controlled by diffusion. Under such conditions a change in the affinity of the orthosteric radioligand by an allosteric modulator is manifested mainly as a change in the dissociation rate of the tracer. Allosteric modulation of the radioligand rate of dissociation in an ideal model system with constant association rate by positive (α = 0.1) and negative (α = 10) allosteric modulators is shown in Fig. 8. A positive allosteric modulator concentration dependently slows down tracer dissociation. The slowdown is limited by the factor of cooperativity α, in this case 10-times at maximum. Inversely, a negative allosteric modulator concentration dependently speeds up tracer dissociation. Again, the effect is limited by the factor of cooperativity α, reaching a maximum of a tenfold change in this particular case.
Allosteric modulation of tracer binding kinetics. Effects of a positive (α = 0.1) (upper graph) and negative (α = 10) (lower graph) allosteric modulator on binding kinetics of the tracer. Abscissa, time is expressed as folds of dissociation halftime of tracer-receptor complex LR in the absence of allosteric modulator A. Ordinate, the tracer binding is expressed as a ratio of tracer L binding to its binding at the start of dissociation. Legend, concentration of allosteric modulator A is expressed as a ratio to its equilibrium dissociation constant K A. A positive allosteric modulator concentration dependently slows down tracer dissociation while a negative allosteric modulator concentration dependently speeds up tracer dissociation
However, muscarinic receptors are far from ideal. Almost all muscarinic allosteric ligands, regardless of being positive or negative modulators, slow down both association and dissociation of orthosteric ligands. The orthosteric binding site of the receptor is located deep within transmembrane helices so association is relatively slow and may be accelerated by allosteric modulators [27]. The binding site for allosteric ligands is located between the second and the third extracellular loops, being close to the path that an orthosteric ligand takes during association and dissociation. Thus, bound allosteric ligands usually represent a physical obstacle (steric hindrance) for orthosteric ligands on their way to and away from the muscarinic receptor [43]. It is noteworthy that effects on the kinetics of binding of an orthosteric ligand allows for identification of agents with neutral cooperativity, since these agents do not change affinity of the tracer in equilibrium experiments. Furthermore, effects on the kinetics of tracer binding distinguish allosteric modulators with very strong negative cooperativity from competitive agents.
Mechanistically, orthosteric ligand L may bind only to free receptor R and is not able to bind to complex of receptor and allosteric ligand RA. In Scheme 1 reaction L + RA to LRA is not possible (Scheme 2).
The orthosteric ligand L and allosteric modulator A can bind concurrently to receptor R and form a ternary complex. The orthosteric ligand L has to bind first to R followed by binding of A to form the ternary complex LRA. If the receptor is already occupied by an allosteric ligand the orthosteric ligand has to “wait” until the allosteric ligand dissociates. Dissociation of the orthosteric ligand from the ternary complex LRA must take place in the reverse order, i.e., the allosteric ligand has to dissociate first to make way for dissociation of the orthosteric ligand. Thus, this sequential arrangement of binding has profound effects on binding kinetics of an orthosteric ligand as demonstrated in the example of slowing down of binding kinetics of [3H]NMS at M2 muscarinic receptors by alcuronium. With increasing concentrations of the allosteric modulator alcuronium the proportion of free receptors decreases and thus association of the orthosteric tracer [3H]NMS decelerates, even though alcuronium is a positive allosteric modulator (Fig. 9, upper graph). Concurrently, increasing the concentration of alcuronium is associated with exerting stronger steric hindrance of [3H]NMS dissociation (Fig. 9, lower graph). Retardation of the on and off biding kinetics of orthosteric ligands by allosteric ligands is proportional to the concentration of the allosteric ligand and is unlimited. Thus extremely long incubation times (hours or even days) are required to reach equilibrium of an orthosteric radioligand at high concentrations (over 100-times of equilibrium dissociation constant) of an allosteric ligand [11].
Slowdown of [3H]NMS binding kinetics by alcuronium. Time courses of association (upper graph) and dissociation (lower graph) of 100 pM [3H]NMS at M2 muscarinic receptors in the absence (black curves) or in the presence of alcuronium at the concentrations indicated in the legend. Upper graph: [3H]NMS binding at the time points indicated on the abscissa is expressed as pmol per mg of protein. Alcuronium concentration dependently increases equilibrium binding and slows down the rate of association of [3H]NMS. Lower graph: binding at the time point indicated on the abscissa is expressed as percent of [3H]NMS binding at the start of dissociation. Alcuronium concentration dependently slows down the rate of dissociation
Non-equilibrium binding leads to kinetic artifacts as shown in Fig. 10 where equilibrium binding is not reached at 10 μM of alcuronium and higher, causing the binding curve to appear bell-shaped (black curve) instead of being sigmoidal (red curve). In such experiment equilibrium may be reached faster by preincubation of receptors with the orthosteric tracer (e.g., 1 h with [3H]NMS) to allow it to bind without slowing-down by the allosteric modulator. Adding a positive allosteric modulator to the preformed [3H]NMS-receptor complex will result in an increase in bound radioactivity. Thus binding lower than control binding (like in Fig. 10) cannot be observed. The situation is more complicated in the case of negative allosteric modulators, where prolonged time may be necessary to allow the tracer to dissociate from the receptor and for binding the allosteric agent to the receptor to reach equilibrium. Lack of equilibrium after the addition of a negative allosteric modulator would lead to underestimation of the factor of cooperativity.
Artifacts of non-equilibrium binding. Binding of 100 pM [3H]NMS to M2 receptors after 3 h of incubation at 25 °C in the presence of alcuronium at the concentrations indicated on the asbcissa is expressed as percent of [3H]NMS binding in the absence of alcuronium (circles). Equilibrium is not reached in the presence of alcuronium at a concentration of 10 μM and higher. The red curve represents binding under equilibrium (K D = 250 pM, α = 0.32)
Extremely long incubation times needed to reach equilibrium may be avoided by inferring allosteric modulator binding from changes in tracer kinetics. Because of arrangement of allosteric and orthosteric sites on muscarinic receptors dissociation of the orthosteric ligand from ternary complex is impossible. Thus observed rate of dissociation limits to zero with increase in concentration of allosteric modulator. Observed dissociation rate is inversely proportional to receptor occupancy by allosteric modulator. Receptor occupancy is given by saturation binding isotherm with apparent equilibrium dissociation constant of allosteric modulator K ′A . Observed dissociation rate k ′Off is thus given by Eq. (11)
where k 0 is dissociation rate constant of the tracer in the absence of allosteric modulator. At saturating concentrations of A dissociation of tracer occurs only from ternary complexes and thus K ′A becomes αK A. For [A] ≫ αK A Eq. (10) simplifies to:
Moreover, at high concentrations of A dissociation of tracer is monophasic and thus observed dissociation rate constant k ′Off can be determined in a single time-point measurement and the equilibrium dissociation constant of allosteric modulator at the ternary complex (αK A) determined without prolonged incubation needed to reach equilibrium (Protocol B). An example of determination of apparent equilibrium dissociation constant of the allosteric modulator methoctramine at M2 muscarinic receptors from changes in [3H]NMS dissociation is shown in Fig. 11. The disadvantage of this approach is that only αK A can be determined (not α and K A separately). However, this approach is sufficient for screening purposes (e.g., in structure–function relationship studies where similar α and K A values are expected for similar compounds). Moreover, in specific conditions (like in the case of methoctramine that binds with high affinity to the orthosteric site and with low affinity to the allosteric site) kinetic experiments are the only way for assessing the apparent equilibrium dissociation constant [42].
Protocol B: Determination of apparent equilibrium dissociation constant in dissociation experiments (96-well plate setup)
-
1.
Determine the equilibrium dissociation constant K D of [3H]NMS in a buffer of your choice at 25 °C in saturation binding experiment and the dissociation rate constant k off of [3H]NMS in dissociation experiments (see previous Chapter 3).
-
2.
Add membranes, about 10 fmol of receptors per well.
-
3.
Add [3H]NMS to a final concentration 3 × K D for 60 min at 25 °C in a final incubation volume of 0.2 ml.
-
4.
Initiate dissociation by the addition of 0.2 ml of atropine in a final concentration of 1 μM either alone or in combination with the tested allosteric modulator in concentrations ranging from 10 μM to 1 mM (5 concentrations at 0.5 log concentration steps). Make samples of control binding (no addition/dilution) and samples of nonspecific binding in the presence of 1 μM (added prior step 3). Sums to 32 samples, one third of 96-well plate, when performed in quadruplicates.
-
5.
Incubate samples for three times of the half-life of ligand dissociation (about 10 min for M2, 45 min for M1, M3, and M4, and 3 h for M5).
-
6.
End dissociation by filtering samples through GF/C filters or filtration plates. Wash with ice-cold deionized water for 6 s.
-
7.
Calculate the observed dissociation rates k ′Off from decrease in specific binding as negative natural logarithm of fractional binding divided by dissociation time.
-
8.
Plot calculated k ′Off values against the concentrations of the tested allosteric modulator and fit Eq. (12) to data.
4.4 Three Ligand System
Many compounds of interest like orthosteric agonists are not candidates as useful tracers because of their low affinity. As explained above, neither are allosteric modulators suitable for radiolabeling. For investigation of allosteric interaction of non-labeled orthosteric ligands and non-labeled allosteric modulators a procedure employing three ligands (orthosteric tracer L, non-labeled orthosteric ligand B, and non-labeled allosteric modulator A) has been devised as depicted in Scheme 3 [11].
Orthosteric tracer L binds to the receptor R with equilibrium dissociation constant K D , orthosteric non-labeled ligand binds to the receptor R with equilibrium dissociation constant K B and allosteric modulator binds to the receptor R with equilibrium dissociation constant K A . The binding of orthosteric ligands L and B is mutually exclusive but the allosteric modulator A can bind concurrently to the receptor R occupied by either of the orthosteric ligands and form a ternary complex LRA or BRA. Alpha and β are factors of binding cooperativity of A with L and A with B, respectively.
In this procedure allosterically induced changes in the affinity for non-labeled orthosteric ligands are reflected in changes in the binding of an orthosteric tracer. The following relations apply besides those described in Eq. (1) in the three ligand system:
And Eq. (2) becomes to:
Analogously to derivation in Eqs. (3)–(7a, b) apparent dissociation constant of the orthosteric tracer L in the presence of the allosteric modulator A and the orthosteric ligand B is derived as:
And the ratio of tracer binding in the presence of A and B to the absence of A and B becomes:
Experimental setup is similar to measurement at a fixed concentration of the tracer and various concentrations of the allosteric modulator. In this three ligand system two curves are measured. One in the absence of the non-labeled orthosteric ligand and one in the presence of a fixed concentration of the orthosteric ligand (Fig. 12). In control curve (in the absence of B) equilibrium dissociation constant of allosteric modulator K A and factor of cooperativity α are determined by fitting Eq. (10) to data. These parameters are then used for fitting Eq. (16) to binding data measured in the presence of B. Equilibrium dissociation constants of the tracer K D and of the orthosteric ligand K B have to be determined in separate measurements. The precise concentration of the tracer L used in the assay should be determined by counting total radioactivity added to the sample and division by specific radioactivity of the tracer and sample volume. Inhibition of tracer binding by B in the absence of A has to correspond to inhibition calculated using equilibrium dissociation constants K D and K B and concentrations of L and B (see Chapter 3, Eq. (13)).
A low concentration of the tracer has to be used when studying allosteric modulators that exert positive cooperativity with the tracer (Fig. 12 upper graph) to get a clear increase in tracer binding by allosteric modulators. Vice versa, a high concentration of the tracer has to be used in case of allosteric modulators with negative cooperativity (Fig. 12 lower graph) to get incomplete inhibition of tracer binding by the allosteric modulator. The concentration of B should be chosen based on the factor of cooperativity β. Low concentrations of tracer (in relation to K B) are suitable in case of positive cooperativity between A and B but leave little room for quantification of negative cooperativity and vice versa. Moreover the concentration of L (in relation to K D) also affects the range of appropriate concentrations of B. Higher concentration of B is required when a higher concentrations of L (in relation to K D) is used to reach the same inhibition of tracer binding. In case of negative cooperativity between A and B with increase in concentration of A binding of B becomes weaker and thus inhibition of the tracer binding smaller. As a result the curves of inhibition of tracer binding in the absence and in the presence of B move closer with increasing the concentration of A (Fig. 12, red and green curves). On the other hand, positive cooperativity between A and B leads to strengthening of binding of B and therefore stronger inhibition of tracer binding. As a result the curves of tracer binding in the absence and the presence of B show more diversion with increasing the concentration of A (Fig. 12, magenta and yellow curves).
There are several parameter combinations under which it is very hard or impossible to determine the factor of cooperativity β. One is the case of studying a combination of an allosteric modulator with strong cooperativity with another with weak cooperativity (negative or positive). This is because on the one hand a low concentration of the tracer is a prerequisite for measurements of weak (either positive or negative) cooperativity and on the other hand a high concentration of the tracer is required for measurements of strong cooperativity systems. In such situation a series of tracer saturation binding has to be performed to determine the apparent equilibrium dissociation constant of the tracer K ′D in the presence of various concentrations of A and one concentration of B. Then K ′D has to be plotted against concentration of A and Eq. (15) fitted to data (Fig. 11).
Dependence of the dissociation rate of [3H]NMS from M2 receptors on the concentration of methoctramine. Observed rate dissociation constants (k ′Off ) are plotted against the concentrations of methoctramine. Fitting Eq. (12) to data yields K ′A around 2.8 μM (\( p{K}_{\mathrm{A}}^{\prime }=5.55\pm 0.5 \); means ± SEM, n = 4). Data are means ± SEM of 3–4 independent experiments performed in quadruplicates
Effects of an allosteric modulator on binding of a tracer at a fixed concentration in the presence of a non-labeled orthosteric ligand. Effects of a positive (upper graph) and negative (lower graph) allosteric modulator A at various concentrations indicated on the abscissa on binding of the tracer L at a fixed concentration indicated in the legend, in the absence (black curves) or in the presence of orthosteric ligand B at a fixed concentration indicated in the legend. Abscissa, concentration of allosteric modulator is expressed as logarithm of ratio to its equilibrium dissociation constant K A. Ordinate, the tracer binding is expressed as a fraction of tracer binding in the absence of allosteric modulator. Legend, factors of cooperativity β of A and B binding. Orthosteric ligand B decreases tracer binding (color curves). In case of negative cooperativity between A and B (red and green curves) A concentration dependently weakens the binding of B that results in smaller inhibition of tracer binding (curves are getting closer to control curve). In case of positive cooperativity between A and B (magenta and yellow curves) A concentration dependently strengthens the binding of B that results in greater inhibition of tracer binding (curves are getting apart from control curve)
Bitopic ligands that bind both to the orthosteric and allosteric sites at muscarinic receptors have been proposed [44] and subsequently identified [42, 45, 46]. Binding of a bitopic ligand B to the allosteric binding site prevents binding of the allosteric ligand A and vice versa (Scheme 4).
The orthosteric ligand L and allosteric modulator A can bind concurrently to the receptor R and form a ternary complex LRA. Binding of the bitopic ligand B is mutually exclusive both with binding of L and A. The ternary complex cannot be formed upon binding of B. Equation (16) does not fit the data as fractional binding of the tracer in the presence of A and B and the absence of A and B is:
5 Analyzing Allosteric Modulation of Functional Responses
5.1 Effects of Allosteric Modulators on Functional Response Under Equilibrium
Muscarinic receptors are spontaneously active that is manifested by activation of second messenger pathways in the absence of agonists [47–49]. Such receptor spontaneous activity implicates that in the absence of the agonist there is a balance between two forms of the receptor (active RA and inactive RI) with non-zero RA number. The thermodynamically complete description of interaction between a agonist and an allosteric modulator on functional receptor is thus described by the cubic ternary complex model (Scheme 5).
The receptor exists in an inactive conformation RI and an active conformation RA. The ratio of RA to RI is given by the activation constant K ACT. The agonist L and allosteric modulator A bind to inactive receptor RI with equilibrium dissociation constants K D and K A, respectively. Effects of an allosteric modulator on the functional response of the receptor to the agonist are complex. Besides allosteric modulation of agonist binding (factor of cooperativity α) an allosteric modulator affects spontaneous activation of the receptor (factor cooperativity γ) and agonist-induced receptor activation (factor of cooperativity δ). If effects of the allosteric modulator on spontaneous activation of the receptor are positive then the allosteric modulator activates receptors even in the absence of agonists. Such allosteric modulators have been identified [28–31]. The overall effect of an allosteric modulator on the formation of the ternary complex with the active receptor LRAA is given by multiplication of these three factors of cooperativity (α, γ, δ). If the resultant of multiplication is greater than 1 then the effect of the allosteric modulator on agonist potency is negative even if L and A have positive binding cooperativity.
G-proteins bind to the receptor in both activation states (spontaneously active and agonist bound). Effects of agonists on G-protein binding to receptor are described by next cubic ternary complex model (Scheme 6).
G-protein G binds to the receptor in its inactive state RI with equilibrium dissociation constant K G. There is mutual allosteric modulation of G-protein binding and receptor activation (factor of cooperativity ε). Agonist L allosterically modulates binding of G to RI (factor of cooperativity η), receptor activation (factor of cooperativity β) and G-protein binding induced by receptor activity (factor cooperativity ζ). The aggregate effect of agonist on G-protein binding is given by multiplication of these three factors of cooperativity (ζ, η, β). It is obvious that an increase in concentration of G (overexpression of G) leads to activation of receptor. Thus the aggregate effect of agonist is dependent on the receptor to G-protein ratio and is, generally speaking, system dependent.
Besides modulation of receptor activation and agonist binding described in Scheme 5 an allosteric agent can also allosterically modulate G-protein binding to the receptor and receptor agonist complexes (Scheme 7).
G-proteins can bind both to receptors in inactive (Scheme 7, left cube) and active (Scheme 7, right cube) conformation. As described in Scheme 6 G-protein binds to free receptor in the inactive conformation with equilibrium dissociation constant K G and to the active conformation with εK G. In addition to modulation of agonist binding (α), receptor spontaneous activation (γ) and agonist-induced receptor activation (δ) an allosteric modulator affects G-protein binding to RI (κ) and RA (πκ) and effects of agonist on G-protein binding (λ). Similar to an agonist, effects of an allosteric modulator depend on the receptor to G-protein ratio. Moreover, effects of an allosteric modulator depend on the direction (activation or inhibition) and magnitude of agonist effects. The overall effect of an allosteric modulator is given by multiplication of all factors of cooperativity involved in transition from state in the absence of A (Scheme 6) to state in the presence of A (Scheme 7) (all factors of cooperativity except ε, β, η and ζ).
Conversions between complexes with bound G-protein have to be added for the scheme describing interactions between receptor, G-protein, agonist and allosteric modulator in order for the model to be thermodynamically complete (Scheme 8).
All eight receptor-G-protein complexes are interchangeable in a step-by-step manner with equilibrium dissociation constants resulting from Scheme 5 through 7. Moreover, G-protein activation is initiated by release of GDP from the G-protein as a result of negative cooperativity between agonist and GDP [50]. There are four receptor-G-protein complexes with bound allosteric modulator in the interaction scheme. Allosteric modulators may affect GDP affinity (and thus activation of G-protein) differently at these four complexes.
Effects of allosteric modulators on functional response depend on the nature of the agonist and system. The ratio of receptor to G proteins affects system basal activity. Systems with high R to G ratio have low basal activity and high receptor reserve. As a result agonists have high efficacy and potency. On the other hand, systems with low R to G ratio have high basal activity and no receptor reserve. As a result agonists have both low efficacy and potency. Effects of both positive and negative allosteric modulators on agonist potency (shift in apparent K G in the presence versus in the absence of L) (α, γ, δ) are weaker at systems with high R to G ratio due to high receptor reserve. High basal activity of the system decreases agonist efficacy (shift in ratio of active species in the presence versus in the absence of L). At systems with high basal activity effects of positive allosteric modulator on efficacy are weaker. On the other hand, effects of negative allosteric modulators on agonist efficacy may be stronger (if δ < πκλ).
Full agonists, due to strong positive cooperativity β, act as agonists at all systems (regardless of R to G ratio). Allosteric modulators may act as agonists (activate receptors in the absence of an orthosteric agonist) [28] or inverse agonists depending on the factor of cooperativity γ, activation constant K ACT and receptor to G-protein ratio [51]. Allosteric modulators that have weak positive cooperativity γ act as partial agonists in a system with high R to G ratio and as inverse agonists at a low R to G ratio. As evident, effects of allosteric modulators on functional responses to an agonist are very complex. It is technically unfeasible to experimentally isolate and determine individual constants and factors of cooperativity. From a practical point of view only the overall effect of an allosteric modulator on the potency and efficacy of a given agonist in a given system could be determined.
5.2 Effects of Allosteric Modulators on the Kinetics of Functional Responses
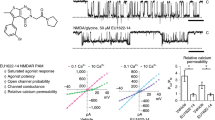
Although a change in agonist potency induced by an allosteric modulator usually follows change in agonist affinity [52, 53] it has been reported that allosteric modulators may have different effects on agonist binding and agonist-mediated functional responses [25]. Moreover, effects of allosteric modulators on functional responses may also differ over time. This is exemplified by the dichotomous effects of the allosteric modulator rapacuronium on acetylcholine equilibrium binding on the one hand and on the kinetics of acetylcholine binding on the other hand [27]. For example, although rapacuronium exerts negative cooperativity with binding of acetylcholine to all muscarinic receptor subtypes at equilibrium it accelerates the rate of acetylcholine binding at odd-numbered subtypes. At low concentrations it transiently increases the potency and efficacy of functional responses to acetylcholine at odd-numbered subtypes (Fig. 13). The time between acetylcholine release and termination of its action by acetylcholinesterase is in the range of a fraction of a second. Therefore, effects of allosteric modulators in the early non-equilibrium stage of receptor signaling are physiologically more relevant than effects on acetylcholine equilibrium binding that does not occur in vivo. Thus fast functional assays that much better simulate physiological conditions are more suitable for screening of potential allosteric modulators of neurotransmission than long-lasting equilibrium binding experiments.
Effects of rapacuronium on the kinetics of [35S]GTPγS binding. Membranes were preincubated for 60 min in the presence (open symbols) or in the absence (closed symbols) of 1 μM rapacuronium. Then [35S]GTPγS was added simultaneously with buffer (circles) or 10 μM acetylcholine (squares). Incubations were terminated at the times indicated on the abscissa. The increase of specific [35S]GTPγS binding is expressed as fmol per μg of protein (top) and as fold increase of specific binding under basal conditions (bottom). Data are means ± SE of values from three independent experiments performed in quadruplicates
The advantage of [35S]GTPγS binding as a measure of receptor functional response (Fig. 13) is that it can be easily scaled up for high-throughput screening. Its disadvantage is that an agonist must be present during incubation lasting minutes that is still far from physiological conditions. Moreover, the resulting signal is the sum of functional response over the whole time-course of incubation. Methods that allow real-time measurement of functional responses and transient application of agonist are more appropriate. Such methods include microfluorometry of intracellular calcium release [54] (see Protocol C) or measurement of conformation changes of the receptor by fluorescence resonance energy transfer between two dyes attached to one receptor (see Chapter 8).
Protocol C: Measurement of allosteric modulation of a functional response by microfluorometry of intracellular calcium
-
1.
Seed CHO cells stably expressing muscarinic receptors on 24 mm-diameter microscope cover glass in 35 mm-diameter Petri dish and cultivate them until about 80 % confluency.
-
2.
Optional: If CHO cells express M2 or M4 receptors transfect cells with G15/16 G-protein to couple these subtypes to phospholipase C [55].
-
3.
Prepare DMSO solutions of 2 mM Fura-2AM and 20 % Pluronic F-68 and mix them 1:1.
-
4.
Wash cells with warm Krebs-HEPES buffer (KHB; final concentrations in mM: NaCl 138; KCl 4; CaCl2 1.3; MgCl2 1; NaH2PO4 1.2; Hepes 20; glucose 10; Probenecid 1; pH adjusted to 7.4).
-
5.
Load cells with Fura-2 by incubating them in 0.5 ml KHB and 10 μl of solution from step 4 for 1 h at 37 °C.
-
6.
Remove KHB, wash cells with fresh KHB.
-
7.
Assemble the cover glass in superfusion chamber and place under a fluorescence microscope. Record Fura-2 emission (>470 nm) at 380 and 340 nm emissions twice a second.
-
8.
Expose cells to increasing concentrations of an agonist (e.g., carbachol or acetylcholine 10 nM to 10 μM) for 5 s. Allow cells to rest for 3–5 min between stimuli.
-
9.
Analyze the ratio of Fura-2 emissions at 380 and 340 nm excitation to determine half-efficient concentration (EC50) and maximum stimulation.
-
10.
Expose cells to the tested allosteric modulator (1 μM to 1 mM) for 5 s to check for possible agonist/inverse agonist effects of the allosteric modulator by itself. Allow cells to rest for 3–5 min between stimuli.
-
11.
To determine the effects of an allosteric modulator on agonist potency expose cells for 5 s to agonist at a concentration around its EC50 (for positive cooperativity slightly below and for negative cooperativity slightly above EC50) alone and then in mixture with increasing concentrations of the tested allosteric modulator. Allow cells to rest for 3–5 min between stimuli.
-
12.
To determine effects of allosteric modulator on maximum response to agonist expose cells for 5 s to agonist in saturating concentration alone and then in mixture with increasing concentrations of the tested allosteric modulator. Allow cells to rest for 3–5 min between stimuli.
6 Conclusions
Allosteric modulation of muscarinic receptors is an interesting phenomenon with great potential for drug discovery and pharmaceutical application. However, detailed studies and understanding are limited to simple systems due to complexity of allosteric interactions. Another limitation is that allosteric modulators of muscarinic receptors generally have affinity that is too low to make them suitable radiolabeled tracers. This necessitates complex experimental arrangements to quantify binding parameters of these agents.
References
Monod J, Changeux JP, Jacob F (1963) Allosteric proteins and cellular control systems. J Mol Biol 6:306–329
Monod J, Wyman J, Changeux JP (1965) On the nature of allosteric transitions: a plausible model. J Mol Biol 12:88–118
Clark AL, Mitchelson F (1976) The inhibitory effect of gallamine on muscarinic receptors. Br J Pharmacol 58:323–331
Stockton JM, Birdsall NJ, Burgen AS, Hulme EC (1983) Modification of the binding properties of muscarinic receptors by gallamine. Mol Pharmacol 23:551–557
Dalton DW, Tyers MB (1982) A comparison of the muscarinic antagonist actions of pancuronium and alcuronium. J Auton Pharmacol 2:261–266
Waelbroeck M, Robberecht P, De Neef P, Christophe J (1984) Effects of verapamil on the binding properties of rat heart muscarinic receptors: evidence for an allosteric site. Biochem Biophys Res Commun 121:340–345
Lai WS, Ramkumar V, El-Fakahany EE (1985) Possible allosteric interaction of 4-aminopyridine with rat brain muscarinic acetylcholine receptors. J Neurochem 44:1936–1942
Kloog Y, Sokolovsky M (1985) Allosteric interactions between muscarinic agonist binding sites and effector sites demonstrated by the use of bisquaternary pyridinium oximes. Life Sci 36:2127–2136
Nedoma J, Tucek S, Danilov AF, Shelkovnikov SA (1986) Stabilization of antagonist binding to cardiac muscarinic acetylcholine receptors by gallamine and other neuromuscular blocking drugs. J Pharmacol Exp Ther 236:219–223
Flynn DD, Mash DC (1989) Multiple in vitro interactions with and differential in vivo regulation of muscarinic receptor subtypes by tetrahydroaminoacridine. J Pharmacol Exp Ther 250:573–581
Jakubík J, Bačáková L, El-Fakahany EE, Tuček S (1997) Positive cooperativity of acetylcholine and other agonists with allosteric ligands on muscarinic acetylcholine receptors. Mol Pharmacol 52:172–179
Lazareno S, Popham A, Birdsall NJ (2000) Allosteric interactions of staurosporine and other indolocarbazoles with N-[methyl-(3)H]scopolamine and acetylcholine at muscarinic receptor subtypes: identification of a second allosteric site. Mol Pharmacol 58:194–207
Leppik RA, Miller RC, Eck M, Paquet JL (1994) Role of acidic amino acids in the allosteric modulation by gallamine of antagonist binding at the m2 muscarinic acetylcholine receptor. Mol Pharmacol 45:983–990
Matsui H, Lazareno S, Birdsall NJ (1995) Probing of the location of the allosteric site on m1 muscarinic receptors by site-directed mutagenesis. Mol Pharmacol 47:88–98
Krejčí A, Tuček S (2001) Changes of cooperativity between N-methylscopolamine and allosteric modulators alcuronium and gallamine induced by mutations of external loops of muscarinic M3 receptors. Mol Pharmacol 60:761–767
Voigtländer U, Jöhren K, Mohr M, Raasch A, Tränkle C, Buller S, Ellis J, Höltje H, Mohr K (2003) Allosteric site on muscarinic acetylcholine receptors: identification of two amino acids in the muscarinic M2 receptor that account entirely for the M2/M5 subtype selectivities of some structurally diverse allosteric ligands in N-methylscopolamine-occupied receptors. Mol Pharmacol 64:21–31
Jakubík J, Krejčí A, Doležal V (2005) Asparagine, valine, and threonine in the third extracellular loop of muscarinic receptor have essential roles in the positive cooperativity of strychnine-like allosteric modulators. J Pharmacol Exp Ther 313:688–696
Huang X, Prilla S, Mohr K, Ellis J (2005) Critical amino acid residues of the common allosteric site on the M2 muscarinic acetylcholine receptor: more similarities than differences between the structurally divergent agents gallamine and bis(ammonio)alkane-type hexamethylene-bis-[dimethyl-(3-phthalimidopropyl)ammonium]dibromide. Mol Pharmacol 68:769–778
Tränkle C, Dittmann A, Schulz U, Weyand O, Buller S, Jöhren K, Heller E, Birdsall NJM, Holzgrabe U, Ellis J, Höltje HD, Mohr K (2005) Atypical muscarinic allosteric modulation: cooperativity between modulators and their atypical binding topology in muscarinic M2 and M2/M5 chimeric receptors. Mol Pharmacol 68:1597–1610
Prilla S, Schrobang J, Ellis J, Höltje H, Mohr K (2006) Allosteric interactions with muscarinic acetylcholine receptors: complex role of the conserved tryptophan M2 422Trp in a critical cluster of amino acids for baseline affinity, subtype selectivity, and cooperativity. Mol Pharmacol 70:181–193
Jakubík J, El-Fakahany EE (2010) Allosteric modulation of muscarinic acetylcholine receptors. Pharmaceuticals 9:2838–2860
Kruse AC, Ring AM, Manglik A, Hu J, Hu K, Eitel K, Hübner H, Pardon E, Valant C, Sexton PM, Christopoulos A, Felder CC, Gmeiner P, Steyaert J, Weis WI, Garcia KC, Wess J, Kobilka BK (2013) Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504:101–106
Fisher A (2012) Cholinergic modulation of amyloid precursor protein processing with emphasis on M1 muscarinic receptor: perspectives and challenges in treatment of Alzheimer’s disease. J Neurochem 120(Suppl 1):22–33
Jones CK, Byun N, Bubser M (2012) Muscarinic and nicotinic acetylcholine receptor agonists and allosteric modulators for the treatment of schizophrenia. Neuropsychopharmacology 37:16–42
Zahn K, Eckstein N, Tränkle C, Sadée W, Mohr K (2002) Allosteric modulation of muscarinic receptor signaling: alcuronium-induced conversion of pilocarpine from an agonist into an antagonist. J Pharmacol Exp Ther 301:720–728
Jäger D, Schmalenbach C, Prilla S, Schrobang J, Kebig A, Sennwitz M, Heller E, Tränkle C, Holzgrabe U, Höltje H, Mohr K (2007) Allosteric small molecules unveil a role of an extracellular E2/transmembrane helix 7 junction for G protein-coupled receptor activation. J Biol Chem 282:34968–34976
Jakubík J, Randaková A, El-Fakahany EE, Doležal V (2009) Divergence of allosteric effects of rapacuronium on binding and function of muscarinic receptors. BMC Pharmacol 9:15
Jakubík J, Bačáková L, Lisá V, El-Fakahany EE, Tuček S (1996) Activation of muscarinic acetylcholine receptors via their allosteric binding sites. Proc Natl Acad Sci U S A 93:8705–8709
Lebois EP, Bridges TM, Lewis LM, Dawson ES, Kane AS, Xiang Z, Jadhav SB, Yin H, Kennedy JP, Meiler J, Niswender CM, Jones CK, Conn PJ, Weaver CD, Lindsley CW (2010) Discovery and characterization of novel subtype-selective allosteric agonists for the investigation of M1 receptor function in the central nervous system. ACS Chem Neurosci 1:104–121
Lebois EP, Digby GJ, Sheffler DJ, Melancon BJ, Tarr JC, Cho HP, Miller NR, Morrison R, Bridges TM, Xiang Z, Daniels JS, Wood MR, Conn PJ, Lindsley CW (2011) Development of a highly selective, orally bioavailable and CNS penetrant M1 agonist derived from the MLPCN probe ML071. Bioorg Med Chem Lett 21:6451–6455
Digby GJ, Noetzel MJ, Bubser M, Utley TJ, Walker AG, Byun NE, Lebois EP, Xiang Z, Sheffler DJ, Cho HP, Davis AA, Nemirovsky NE, Mennenga SE, Camp BW, Bimonte-Nelson HA, Bode J, Italiano K, Morrison R, Daniels JS, Niswender CM, Olive MF, Lindsley CW, Jones CK, Conn PJ (2012) Novel allosteric agonists of M1 muscarinic acetylcholine receptors induce brain region-specific responses that correspond with behavioral effects in animal models. J Neurosci 32:8532–8544
Jakubík J, Bačáková L, El-Fakahany EE, Tuček S (1995) Subtype selectivity of the positive allosteric action of alcuronium at cloned M1-M5 muscarinic acetylcholine receptors. J Pharmacol Exp Ther 274:1077–1083
Lazareno S, Popham A, Birdsall NJM (2002) Analogs of WIN 62,577 define a second allosteric site on muscarinic receptors. Mol Pharmacol 62:1492–1505
Lazareno S, Doležal V, Popham A, Birdsall NJM (2004) Thiochrome enhances acetylcholine affinity at muscarinic M4 receptors: receptor subtype selectivity via cooperativity rather than affinity. Mol Pharmacol 65:257–266
Tränkle C, Mies-Klomfass E, Cid MH, Holzgrabe U, Mohr K (1998) Identification of a [3H]Ligand for the common allosteric site of muscarinic acetylcholine M2 receptors. Mol Pharmacol 54:139–145
Lysíková M, Fuksová K, Elbert T, Jakubík J, Tuček S (1999) Subtype-selective inhibition of [methyl-3H]-N-methylscopolamine binding to muscarinic receptors by alpha-truxillic acid esters. Br J Pharmacol 127:1240–1246
Jerusalinsky D, Cerveñasky C, Peña C, Raskovsky S, Dajas F (1992) Two polypeptides from Dendroaspis angusticeps venom selectively inhibit the binding of central muscarinic cholinergic receptor ligands. Neurochem Int 20:237–246
Jolkkonen M, Adem A, Hellman U, Wernstedt C, Karlsson E (1995) A snake toxin against muscarinic acetylcholine receptors: amino acid sequence, subtype specificity and effect on guinea-pig ileum. Toxicon 33:399–410
Waelbroeck M, De Neef P, Domenach V, Vandermeers-Piret MC, Vandermeers A (1996) Binding of the labelled muscarinic toxin 125I-MT1 to rat brain muscarinic M1 receptors. Eur J Pharmacol 305:187–192
Fruchart-Gaillard C, Mourier G, Marquer C, Ménez A, Servent D (2006) Identification of various allosteric interaction sites on M1 muscarinic receptor using 125I-Met35-oxidized muscarinic toxin 7. Mol Pharmacol 69:1641–1651
Ilien B, Franchet C, Bernard P, Morisset S, Weill CO, Bourguignon J, Hibert M, Galzi J (2003) Fluorescence resonance energy transfer to probe human M1 muscarinic receptor structure and drug binding properties. J Neurochem 85:768–778
Jakubík J, Zimčík P, Randáková A, Fuksová K, El-Fakahany EE, Doležal V (2014) Molecular mechanisms of methoctramine binding and selectivity at muscarinic acetylcholine receptors. Mol Pharmacol 86:180–192
Proška J, Tuček S (1994) Mechanisms of steric and cooperative actions of alcuronium on cardiac muscarinic acetylcholine receptors. Mol Pharmacol 45:709–717
Melchiorre C, Minarini A, Angeli P, Giardinà D, Gulini U, Quaglia W (1989) Polymethylene tetraamines as muscarinic receptor probes. Trends Pharmacol Sci. Suppl: 55–59
Tahtaoui C, Parrot I, Klotz P, Guillier F, Galzi J, Hibert M, Ilien B (2004) Fluorescent pirenzepine derivatives as potential bitopic ligands of the human M1 muscarinic receptor. J Med Chem 47:4300–4315
Daval SB, Valant C, Bonnet D, Kellenberger E, Hibert M, Galzi J, Ilien B (2012) Fluorescent derivatives of AC-42 to probe bitopic orthosteric/allosteric binding mechanisms on muscarinic M1 receptors. J Med Chem 55:2125–2143
Burstein ES, Spalding TA, Braüner-Osborne H, Brann MR (1995) Constitutive activation of muscarinic receptors by the G-protein Gq. FEBS Lett 363:261–263
Jakubík J, Bačáková L, El-Fakahany EE, Tuček S (1995) Constitutive activity of the M1-M4 subtypes of muscarinic receptors in transfected CHO cells and of muscarinic receptors in the heart cells revealed by negative antagonists. FEBS Lett 377:275–279
Burstein ES, Spalding TA, Brann MR (1997) Pharmacology of muscarinic receptor subtypes constitutively activated by G proteins. Mol Pharmacol 51:312–319
Jakubík J, Janíčková H, El-Fakahany EE, Doležal V (2011) Negative cooperativity in binding of muscarinic receptor agonists and GDP as a measure of agonist efficacy. Br J Pharmacol 162:1029–1044
Jakubík J, Haga T, Tuček S (1998) Effects of an agonist, allosteric modulator, and antagonist on guanosine-gamma-[35S]thiotriphosphate binding to liposomes with varying muscarinic receptor/Go protein stoichiometry. Mol Pharmacol 54:899–906
Birdsall NJ, Farries T, Gharagozloo P, Kobayashi S, Lazareno S, Sugimoto M (1999) Subtype-selective positive cooperative interactions between brucine analogs and acetylcholine at muscarinic receptors: functional studies. Mol Pharmacol 55:778–786
Lazareno S, Birdsall B, Fukazawa T, Gharagozloo P, Hashimoto T, Kuwano H, Popham A, Sugimoto M, Birdsall NJ (1999) Allosteric effects of four stereoisomers of a fused indole ring system with 3H-N-methylscopolamine and acetylcholine at M1-M4 muscarinic receptors. Life Sci 64:519–526
Santrůčková E, Doležal V, El-Fakahany EE, Jakubík J (2014) Long-term activation upon brief exposure to xanomeline is unique to M1 and M4 subtypes of muscarinic acetylcholine receptors. PLoS One 9, e88910
Milligan G, Marshall F, Rees S (1996) G16 as a universal G protein adapter: implications for agonist screening strategies. Trends Pharmacol Sci 17:235–237
Acknowledgments
This research was supported by Academy of Sciences of the Czech Republic support RVO: 67985823 and Grant Agency of the Czech Republic grant P304/12/G069.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Jakubik, J., El-Fakahany, E.E. (2016). Allosteric Modulation of Muscarinic Receptors. In: Myslivecek, J., Jakubik, J. (eds) Muscarinic Receptor: From Structure to Animal Models. Neuromethods, vol 107. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-2858-3_6
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2858-3_6
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-2857-6
Online ISBN: 978-1-4939-2858-3
eBook Packages: Springer Protocols