Abstract
Microbial forensics is a field that has attracted tremendous interest off late. Bioterrorism, biowarfare, and weaponization of the microbiome have terrified the world. While microbial forensics deals with the study of microbes for legal purposes, its applicability in the other subfields of health and science is irrefutable. It has its application not only to bio-crime investigation but also for clinical and toxicology purposes. There are several risks associated with the handling and analysis of microbes. Hence, it is essential to follow the established guidelines to prevent contamination, cross-contamination, and accidental infections with limited or widespread drastic effects. These guidelines mainly ensure the safety, collection, preservation of microbial forensic samples. This chapter covers a broad overview of epidemiology and microbial forensics, the critical elements of microbial forensics, the sample collection methods and guidelines, the various detection methods (molecular), and the result interpretation.
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
Keywords
- Microbial forensics
- Microbial evidences
- Processing and detection methods
- DNA-based methods
- Massive parallel sequencing
- STR
- VNTR
- PCR
Introduction
Microbiology refers to “the study of microorganisms, i.e., the organisms that exist as single cells or cell clusters and must be viewed individually with the aid of a microscope” (Nema 2018). These diverse communities of microbes are manipulated by humans and are used as biological warfare agents. With the increase in tools and technologies to manipulate these organisms, their use or abuse has also increased. Thus, the need to investigate such acts has also increased; herein, forensic microbiology or microbial forensics plays a vital role.
Microbial forensics is the scientific discipline that analyzes evidence related to bioterrorism and bio-crimes, hoax, or inadvertent microorganism/toxin release for attribution purposes (Budowle et al. 2005b). Microbial forensics involves the characterization of microbial evidence with the help of microbiological methods to determine the source and assist in identifying incidences of bioterrorism, bio-attack, bio-crime, outbreaks and transmission of pathogens, or accidental release of a biological agent or a toxin (Budowle et al. 2003, 2005a; Oliveira and Amorim 2018; Schmedes and Budowle 2019; Smith 2019).
Microbial forensics has its application not only to bio-crime investigation but also for clinical and toxicology purposes. The human skin is believed to have a unique microbiome that may be individualistic to a person. Therefore, microbial forensics also encompasses identifying a person from their leftover microbial traces on the materials they were in contact with. In addition to this, it also has its application in determining the cause of death, helps to investigate drowning cases, toxicological cases, estimation of postmortem interval, etc. (Oliveira and Amorim 2018).
Bioterrorism
Bioterrorism is emerging as a global threat and one of the deadliest means used to cause mass disasters. This affects not only the economic condition but also the workforce of the country. The usage of bioweapon was prevalent for thousands of years. For example, the Romans were known to contaminate the water resource by decaying animal carcasses to harm their enemies. During the war of Kaffa, Tatar soldiers threw diseased bodies on the city’s walls to spread plague among enemies. Even in many wars such as WWI, WWII, French and Indian war, biological weapons were reportedly employed (Schmedes and Budowle 2019). There are other known incidences of potential bio-attacks. One such example is the bubonic plague attack in the middle of the fourteenth century in the Siege of Caffa (Budowle et al. 2005b; Barras and Greub 2014).
Another classic example is the use of plague as a biological weapon by the Japanese during the Sino-Japanese war in the 1930s–1940s (Budowle et al. 2005b; Barras and Greub 2014; The College of Physicians of Philadelphia 2018). Salmonella was used to contaminate the salad bars in the Dalles (Török et al. 1997; Schmedes and Budowle 2019). In 1996, Dallas, TX, hospital technician intentionally infected the muffins with Shigella and placed them in the eating area. Due to this, 12 people were infected, and 4 were hospitalized (Kolavic et al. 1997; Schmedes and Budowle 2019). In 1993, anthrax produced by bacterium B. anthracis became the most potential bioweapon disseminated in Tokyo by the Aum Shinrikyo Japanese cult (Schmedes and Budowle 2019).
The intentional use of pathogens and microorganisms to create biowarfare, the threat of terrorism, and detection of West Nile virus in New York City in 1999 was the major concern of the United States. In 2001, a 63-year-old employee of American Media in Boca Raton, Florida, suffered from fever, deprived sleep, emesis, and confusion. Later he was diagnosed with anthrax. The detection was confirmed by Laboratory Response Network, Department of Health, Florida. Meanwhile, at the commencement of the twentieth century, most cases of inhalation of anthrax in the United States were due to occupational exposure to infected animal skins or products. Thus, an urgent need was felt to improve the capabilities to collect, examine, and investigate the scene of incidence involving potential bio-crime acts (Morse and Budowle 2006).
Letters laced with anthrax (Bacillus anthracis spores) were sent through United States Postal Service along the eastern seaboard, to two senators, to news anchor of NBC News, and the New York Post, each containing B. anthracis spores. These spores infected 22 persons causing 5 deaths and disruptions (Barras and Greub 2014; Schmedes and Budowle 2019). As a result, the use of the US postal system for spreading endospores of B. anthracis became a national threat, and the FBI began the investigation.
Over the decades, the number of epidemic outbreaks has been increasing, such as H1N1 virus-swine-origin influenza A 2009, (Smith et al. 2009); Mycobacterium tuberculosis (Gardy et al. 2011); Vibrio cholerae (Hendriksen et al. 2011); MERS coronavirus, 2012, Saudi Arabia (Assiri et al. 2013); H7N9 virus, 2013, China (Kageyama et al. 2013); Escherichia coli O104:H4, 2011, Europe (Grad et al. 2012); Ebola virus 2014, Sierra Leone (Cenciarelli et al. 2015); Zika virus 2015, Brazil (Faria et al. 2016; Oliveira and Amorim 2018); Nipah virus, 2018, Kerala India; Ebola, 2018–2020, the Democratic Republic of the Congo and Uganda; Coronavirus disease 2019 COVID-19, worldwide 2019 to present; and Black fungus, 2021, India (Wikipedia Contributors 2021). The devastating effect of such epidemics , pandemics , and global bio-attacks led to the official launching of microbial forensics (Budowle et al. 2005b; Barras and Greub 2014).
Individualistic Microflora for Person Identification
The massive collection of microbes that reside on or inside a human body is termed as human microbiome. Personal identification is primordially made using DNA and fingerprints with a great degree of accuracy and validity. Nevertheless, some attempts have been made to individualize person based on the unique microbiome each human is believed to have (Fierer et al. 2010; Tridico et al. 2014; Meadow et al. 2014; Lax et al. 2015; Schmedes et al. 2017; Oliveira and Amorim 2018; Robinson et al. 2020). Several different microbial colonies reside in or on different parts of our body. To identify persons, skin microbiome or hair microbiome are potentially used. Oral microflora also has great significance for identification purposes (Robinson et al. 2020). Each individual has a unique skin microbiome composition, which can be collected from the items they came in contact with. Many studies are conducted on establishing the use of skin microbiome for individualization purposes (Fierer et al. 2010; Schmedes et al. 2017). As stated by Sir Edmond Locard in principle of exchange, “when two objects or entities or surfaces come into contact, there is always a mutual exchange of traces.”
Similarly, when an individual touches any surface, they transfer their unique microbiome to that surface. This can be principally utilized in criminal identification. When a suspect touches any surface, it leaves traces there. Surfaces such as mobile phone screens, tablet screens, laptop screens, keyboards, mouse, floor, doorknobs, etc., act as an excellent source for harnessing the microflora of the person using it. Similarly, suspect’s unique microflora can be extracted from floors, victims clothing, or anything probable of being touched by him. Appropriate examination of the crime scene could assist in getting unique skin microbiome of the suspect (Meadow et al. 2014; Lax et al. 2015; Nema 2018; Oliveira and Amorim 2018; Robinson et al. 2020). Additionally, this can assist in finding geological locations of suspects based on the identification of microbes that are found explicitly in particular areas; however, this study is still in its stage of infancy (Oliveira and Amorim 2018; Robinson et al. 2020).
When individualizing a person using skin microbiome, one should be aware and cautious of the fact that whole human skin is not supposed to have a uniform microflora. The human body is heterogeneous and consists of different substances in or on the body. Likewise, the skin has various compositional changes moving from hair to toes every part having slightly different microflora. Thus, when using skin microflora for identification, the place where it is generated should be looked upon. Hair is a shred of persistent evidence found on several crime scenes. Some approaches also made to use scalp microflora (also found on hair) for individualization (Tridico et al. 2014; Robinson et al. 2020).
Postmortem Examination
PMI estimation: Determination of postmortem interval (PMI) is an integral part of postmortem examination. PMI estimation is predominantly done using sequential changes occurring in cadavers or entomological evidence. Many approaches are now directed to estimate PMI using microbial techniques (Metcalf et al. 2013; Hauther et al. 2015; Javan et al. 2016; Nema 2018; Oliveira and Amorim 2018; Zhang et al. 2019; Metcalf 2019). It is well known that microbes carry out decomposition, which is one of the postmortem changes in human cadavers. Decomposition occurs in several sequential steps, affecting the colonization of microbes on or in the body. This successional colonization of microbes can be used for the determination of PMI. This can be done using an ecological succession of microbes of skin or in internal organs of the body defined as thanatomicrobiome. Studies have been conducted to estimate PMI based on changing microbial community on the skin (Metcalf et al. 2013). Meanwhile, other approaches are based on the thanatomicrobiome, which harnesses the microbial community of internal organs and natural orifices, including gastrointestinal tracks, intestines, stomach, eyes, vagina, ears, lungs, etc. (Hauther et al. 2015; Javan et al. 2016; Metcalf 2019).
Estimation of PMI using microbiology has great advantages and can also be potentially applied to calculate submersion interval by studying successional colonization of marine microbes with great accuracy. Further, based on statistical regression models, there are also approaches to estimate PMI (Zhang et al. 2019). There are external factors affecting sequential colonization of microbes on the corpse, which includes that till what time after death PMI can be accurately determined, the weather conditions (sunlight, wind, humidity, etc.) the body is present in, whether the body is covered or open or is buried in soil (Metcalf 2019). These factors also need consideration and also standardization through further research.
Other postmortem examination: It includes determination of cause and manner of death. The cause and manner of death can also be potentially determined using microbial community. There are five manners of death, i.e., natural, accidental, suicidal, homicidal, and undetermined. Some studies show a predominance of a specific community of microbes for a particular manner of death, also some studies have shown that the abundance of specific taxa in different internal organs is also affected by the manner of death, which could be positively used to infer the manner of death (Oliveira and Amorim 2018; Robinson et al. 2020). However, the gender, age, and sex can also interfere in the interpretation of results. Thus, this cannot be used as a sole method of determination of cause and manner of death. It would require proper validation to be admissible in the court of law.
There are numerous causes of death. Death can occur due to prolonged infection, drowning, poisoning, etc. Microbial evidence cannot provide the definitive cause of death but has excellent potential in determining the cause of death. In death cases due to drowning, examination of diatoms is the gold standard in forensic investigations (Díaz-Palma et al. 2009; Oliveira and Amorim 2018). Some microbial communities can be utilized to investigate these cases (Oliveira and Amorim 2018). In this order, some studies establish the use of microflora in toxicological cases and hospital-acquired infections (Castle et al. 2017; Oliveira and Amorim 2018). In addition to determining the cause and manner of death, microbiome can also assist in determining the sex of the cadaver (Bell et al. 2018). Some approaches are being made to differentiate the postmortem changes in the body based on sex to be employed potentially for sex determination (Bell et al. 2018).
Detection of Body Fluids
Biological fluids such as saliva, semen, synovial fluid, blood, vaginal fluid, menstrual blood, urine, etc., are common evidence that forensic investigators encounter at crime scenes. Several preliminary and confirmatory tests achieve identification of these fluids. However, the microbial diversity also has its part in the identification of different body fluids (Zou et al. 2016; Hanssen et al. 2018; Oliveira and Amorim 2018; Robinson et al. 2020). Different body fluids have characteristic microbial profiles where some microbial communities are found in abundance and others in traces, thus can be used as bioindicators confirming the presence of a particular body fluid (Hanssen et al. 2018; Oliveira and Amorim 2018; Robinson et al. 2020). For instance, a study conducted on the Han Chinese population for detection of a particular microbial profile of saliva, vaginal fluid, and feces suggested that the presence of Lactobacillus crispatus, Lactobacillus gasseri, Bacteroides uniformis, Bacteroides thetaiotaomicron can be used as identification microbes for vaginal fluid and feces (Zou et al. 2016). Similarly, many approaches are being carried out and many more are needed to establish several biomarkers specific to a particular body fluid that can be positively used for fluid identification .
Microflora of Soil and Water for Forensic Use
Soil and water microflora can ultimately be used to identify geological locations as the microbial profile in soil and water changes over a few meters. Soil has its unique microflora, which can act as bioindicators to confirm two soil samples originating from the same site. Microflora of soil traces found on a crime scene, compared with the suspected soil site, can confirm the location (Oliveira and Amorim 2018; Robinson et al. 2020). In the same way, water microflora can also assist in determining geographical locations. Water microflora has its prime role in investigating drowning cases where diatoms are primarily employed for the purpose. The species of diatoms present in water and the composition of diatoms present in the lungs and internal organs of drowned bodies can confirm whether the person was dead or alive at the time of the drowning. Also, if some different diatom species or different microflora are seen in the body’s internal organs, it generates the line of doubt that the person was initially killed in some different environment. Other microbes can also be used as biomarker while investigating drowning cases (Levkov et al. 2017). Besides, there are studies conducted to examine the microflora of freshwater and seawater (Kakizaki et al. 2009). This application also needs some advancements and validation so that it can be potentially used to prove or disprove legal questions. Apart from diatoms, some other microbial markers should also be potentially included in such investigation .
Epidemiology and Microbial Forensics
Epidemiology is defined as the manifestation, topographies, and causes of disease among populations. Epidemiologic methods to investigate the infectious outbreak by examining the possible evidence of intentional and criminal behavior as contributing factors are termed forensic epidemiology (Goodman et al. 2003; Morse et al. 2019). Similar principles of epidemiology were being utilized to investigate the bio-crime, which involved the bioagent in creating a threat (Flowers et al. 2002).
Microbial forensic epidemiological investigation deals with the legal system that involves examining a crime scene, sustaining the chain of custody, validating the methods, and understanding of results and evolution of new methods to investigate bio-crimes. The investigator must use techniques that match the standard legal system, such as Daubert Standard, which will positively defend the cross-examination (Morse et al. 2019). There are important factors that need to be considered while investigating the outburst of contagious diseases, such as:
-
Occurrence of the outbreak
-
Identifying the population and community at risk
-
Mode of transmission and medium used for transmission
-
Illustrating the agent(s) responsible for the dissemination of infectious disease
An epidemiologic investigation would try to recognize the contributing agent and its source of disease outbreaks. Various molecular techniques are used to examine, identify, and characterize pathogens involved in deadly bio-crimes. However, in microbial forensics, the investigation is used for legal purposes (Morse et al. 2019). Numerous indicators of outbursts can be identified in the epidemiological investigation of infectious diseases. The below-listed factors are the potential clue to indicate the signs of an epidemic (Morse et al. 2019).
-
Disease caused by unknown agents and without explanation of epidemic.
-
The presence of uncommon strain and its antibiotic resistance pattern.
-
Higher rate of illness and death due to uncommon disease and patients showing failure in responding to the treatment.
-
The distribution of uncommon diseases affected by the season and geographical conditions like influenza spreading in the Northern Hemisphere in the summer season.
-
Transmission of disease through different mediums such as air, food, water, and aerosol, etc.
-
The presence of one or more strains of the disease in one patient and its reason remains unknown.
-
The transmission of the disease affecting a significant heterogeneous population.
-
The unfamiliar pattern of morbidity among animals is caused by the unexplained agent responsible for causing the same effect in humans.
-
The unexplained and uncommon illness and death occurring in humans by the agent responsible for causing illness and deaths in animals.
-
Origin of agents of illness from the source having the same genotype.
-
Dissemination of uncommon illness to the noninfectious area either, domestic and foreign.
-
A large number of senseless deaths and diseases.
The two essential aspects of microbial forensics are determining the reason for the release of pathogens, whether caused intentionally or due to some negligence and studying the application of protocols for monitoring the pathogens to distinguish between the unprompted and destructive spread of pathogens microorganisms.
The bio-agents are classified based on the hazards on humans. This classification was given by the National Institute of Health (NIH) in 2019 (Department of Health and Human Services, National Institutes of Health 2019). In 2013, they classified these microbes into three categories, but in 2019, these were divided into four main categories below in Table 1. In addition, the Centre for Disease Control and Prevention (CDC) classifies these biological agents into three categories: Category A, B, and C, given in Table 2 (Centres for Disease Control and Prevention 2018).
The Elements
Microbial forensics adopt genomic, microbiological, and epidemiological methods to characterize and determine biowarfare weapons and ascertain the intentional or unintentional release of destructive pathogens and toxins (Morse et al. 2019). Microbial forensics is an emerging field dedicated to the depiction, examination, and elucidation of evidence found during the act of biological terrorism, biowarfare, and involuntary release of endospores and microorganisms for the attribution purpose (Murch 2003; Morse and Budowle 2006).
The significant elements include:
-
(i)
Detection and identification : This is the first and the foremost step that incorporates detection and identification of the attack and the causing microbe behind it. For this, there should be proper collection and preservation of samples for analysis. Furthermore, for accurate analysis, powerful tools and techniques are needed to improve the sensitivity, accuracy, and specificity of results. These techniques can be categorized into three broad groups that are: (a) molecular techniques, (b) analytical techniques, and (c) physical analysis techniques (Budowle et al. 2005b; Varun et al. 2012).
-
(ii)
Information and database: The availability of information and databases, of course, increases the accuracy of results. Thus, the available database should be enhanced and expanded such that they contain the bioagent genomic sequence data, the whole genome of the agent used in the past, etc. This can be achieved by establishing the record systems at the national and international levels (Budowle et al. 2005b). In addition, interagency sharing of information and database must be encouraged.
-
(iii)
Development of strain repository : It is imperative to house these pathogens and near neighboring microorganisms in a strain repository. This information may prove essential in determining and identifying these near neighbors’ broad and narrow classes using different robust and sophisticated techniques (Budowle et al. 2005b). However, the security of such a repository would remain an active concern as any potential lapse may have devastating outcomes.
-
(iv)
Need for validation : This emphasizes the need for proper validation of new and existing techniques used in microbial forensics. All the methods used to analyze microbial samples must be accepted and validated (Budowle et al. 2005b; Varun et al. 2012). Furthermore, they must be robust enough to provide accurate results even in mutations and changes in the known strain.
-
(v)
Quality assurance guidelines : Safety and quality assurance must be practiced using the appropriate guidelines and norms. These guidelines are important to be followed by microbial forensic laboratories to guarantee reliable results and maintain safety during and after the analysis (Budowle et al. 2005b; Varun et al. 2012).
Sample Collection
Given the exchange principle, every contact leaves a trace; an investigator needs to collect evidence adequately; otherwise, it will lose its evidential value. Therefore, to avoid human error, the National Institute of Justice issued specific guidelines to minimize the chance of error in collecting microbial evidence. The guidelines mainly ensure the safety, collection, and preservation of microbial forensic samples. The main postulates of guidelines are mentioned below:
-
Valuation of the actual situation at a crime scene.
-
Planning related to sample collection, which includes assessing safety protocols for personnel, acquiescence with all guidelines and legal requirements, and discussing the prioritization of samples. It also mentioned selecting personnel and equipment utilized to collect and preserve samples with a proper time frame.
-
Documentation of place, area, subjects, whether human and animal. The possible and source need to find out and maintain the proper chain of custody of possession of evidence.
-
Mention the proper method and equipment that will utilize to collect the microbial forensic evidence.
-
Method of proper preservation of samples (Smith 2019).
The techniques for collecting microbial forensic evidence involves strategic planning, logistic support, and statistic data in collecting microbes that will not produce a toxic effect on other microorganisms. Some principles and guidelines need to be followed to properly handle, collect, and preserve microbial forensic evidence. Tools used to collect the microbial evidence should be validated first and should not react with the sample of interest (Schutzer et al. 2011). The preservation procedure is also based upon the same principle that it will not affect the targeted sample. After the preservation process, samples are packaged and sent for analysis to the laboratory (Schutzer et al. 2011). This is the primary step of the analysis as the quality of the result may vary if the sample is not collected and preserved well. In addition, contamination and cross-contamination should be avoided. The sampling process includes collection and preservation, but several other steps need to be followed given in Fig. 1 (National Research Council 2014; Smith 2020).
Sampling can be achieved by one of the two strategies mentioned below (National Research Council 2014; Smith 2020). Choosing the sampling strategy varies depending on the location, type, and the extent of symptoms in clinical and agricultural settings (Budowle et al. 2006)
-
(i)
Targeted sampling : This is the sampling procedure where a sample is collected from a targeted area. In this process, the sample is collected from an area that is believed to be contaminated based on past knowledge. It is also known as judgmental sampling (Budowle et al. 2006; Sego et al. 2007; Smith 2020).
-
(ii)
Random sampling : This is the type of sampling in which the sample is collected from random areas without any prior knowledge (Budowle et al. 2006; Sego et al. 2007; Smith 2020).
Actual sample collection is the process of taking the sample for analysis. Sample collection can be carried out using three approaches that are:
-
(i)
Collecting the whole item (is also termed as a bulk collection): In this, the whole item is collected and transported to the laboratory for analysis. This approach reduces the extra time required for the collection, but this can only be used if the object or the evidence can be easily removed from the scene (Budowle et al. 2006; Smith 2020).
-
(ii)
Collecting a portion of an item: This applies to immovable objects that cannot be transported to the laboratory. This includes methods such as vacuuming using high-efficiency particulate air vacuums, filtration, etc.
-
(iii)
Swabbing or wiping surfaces: This approach is best suited for trace pieces of evidence. It can be achieved using relevant sample collection devices. For this purpose, dry swabs, premoistened swabs or wipes can be employed (Budowle et al. 2006; Smith 2020).
There are three main issues with these collection methods: (a) many of these collection methods and devices are not rigorously validated; (b) some of the methods are validated, but some security restrictions hamper sharing of this validation data to authorities who need them; (c) the collector must be well acquainted with the analyte or target signatures that are to be analyzed and accordingly the method should be chosen.
Once the evidence is collected, they are packed appropriately and labelled along with the tags indicating the biohazard materials. The preservation of these samples can be achieved using preservative or transport media such as buffered tryptose broth, buffered glycerine, phosphate-buffered sucrose, etc. (Budowle et al. 2006). These media should be chosen based on the type of pathogen to not interfere with the analysis. In some cases, postmortem sampling is also required. Table 3 shows the viscera sample and quantity to be collected related to some common pathogens (Fernández-Rodríguez et al. 2019).
Detection Methods
Microbial forensic evidence encompasses various samples such as food, water, air, swab, soil, animal food, tissue, and clinical samples like blood, urine, stool, tissue, sputum and saliva, etc. The microbial samples are analyzed using various methods such as culture, microscopy, immunoassays, mass spectrometry, real-time PCR, microarray, genetic typing, whole-genome sequencing, and targeted sequencing. Meanwhile, the culturing method is considered the gold standard, but sometimes there is a delay in growth, and it may compromise the safety of an individual. The culture method also suffers problems when dealing with novel and uncharacterized microorganisms (Schmedes and Budowle 2019). Microbial analysis can be carried out using three methodologies: (a) molecular methods, (b) analytical methods, and (c) physical analysis (Budowle et al. 2005b; Varun et al. 2012).
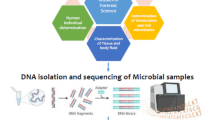
Molecular techniques can be either protein-based, such as microarray assays, immunoassays, etc., or DNA-based techniques (Fig. 2) such as PCR, SNP, etc. (Budowle et al. 2005a; National Research Council 2014; Nema 2018; Blondeau et al. 2019; Kieser and Budowle 2020). Analytical techniques include the use of various instrumental techniques such as matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF), gas chromatography-mass spectroscopy (GC-MS), and liquid chromatography-mass spectroscopy (LC-MS) (National Research Council 2014; Nema 2018; Blondeau et al. 2019). The physical analysis includes microscopic analysis of materials such as soil, water, etc. (National Research Council 2014; Nema 2018; Schmedes and Budowle 2019).
Polymerase chain reaction (PCR) and real-time PCR: PCR-based techniques are the easiest to perform and require a minimal sample quantity. This includes in vitro amplification of DNA carried out in a specified instrument (Budowle et al. 2005a). The instrument performs programable cycling at controlled temperatures to generate millions of copies of targeted DNA that can be detected using different techniques such as hybridization or electrophoresis (Budowle et al. 2005a). These assays can screen many samples with a specific target DNA sequence (Budowle et al. 2005a). Real-time PCR (RT-PCR) is employed for those microbes having RNA as genetic material; this is because PCR can only be proceeded on DNA, and therefore in RT-PCR, reverse transcription is carried out to generate DNA from RNA to procced with PCR. With real-time PCR, amplification and detection of a specific variant of a microbe can be done simultaneously. This is generally done using fluorescent chemistries (Budowle et al. 2005a). Pyrosequencing PCR can also be helpful in many ways. This process is based on detecting luminescence by releasing pyrophosphate on nucleotide addition into the strand (Kieser and Budowle 2020).
PCR and RT-PCR show tremendous significance in forensic microbiology (Bauer et al. 1999; Power et al. 2010; Wang et al. 2013; Park et al. 2014; Rajalakshmi 2017; Aqeel and Omran 2018; Jung et al. 2018). There are numerous species for which PCR markers are readily available, i.e., Lactobacillus, Gardnerella vaginalis, Mycoplasma hominis, etc. (Rajalakshmi 2017). Some studies have positively used PCR markers for the identification of fluids such as blood, saliva, menstrual blood, other vaginal secretions, etc. (Bauer et al. 1999; Wang et al. 2013). Studies have been carried out on messenger RNA (mRNA) profiling using PCR and RTPCR. SPTB, PBGD, HBB, HBA, ALAS2, CD3G, ANK1, PBGD, SPTB, AQP9 can be potentially used as RT-PCR mRNA markers for identification of blood form dried as well as wet stains (Bauer et al. 1999; Wang et al. 2013). Some other attempts are also made using streptococcal bacteria present in saliva as a marker to identify expirated blood (Power et al. 2010). Similarly, matrix metalloproteinase (MMP) and protamine mRNA markers can be used to detect menstrual blood (Wang et al. 2013). One study used oral bacteria Streptococcus salivarius, Streptococcus sanguinis, and Neisseria subflava to identify saliva (Jung et al. 2018). Other fluids such as semen, vaginal fluid, saliva also have such identification mRNA biomarkers (Wang et al. 2013). PCR and RT-PCR markers are also available for the skin microbiome (Hanson et al. 2012; Aqeel and Omran 2018). These are some standard PCR and RT-PCR markers that can be potentially used in microbial forensics. Other than those mentioned here, there are several other PCR and RT-PCR markers available for different species as well.
Single nucleotide polymorphism (SNP) array: This technique detects the variation at a single DNA site. It is a multiplex analysis of SNP (Budowle et al. 2005a; Schmedes and Budowle 2019; Kieser and Budowle 2020). After the PCR product is achieved, it is subjected to SNP primer and fluorescently labelled terminator nucleotide. The polymerase reaction is ceased by terminator nucleotide. This product is then separated by capillary or slab electrophoresis (Budowle et al. 2005a). This microarray can be a highly efficient screening technique with species to strain-level detection (Schmedes and Budowle 2019).
SNPs (single nucleotide polymorphisms) and other genetic signatures, which can be highly efficient screening and characterization tools, have been used precisely for bacterial and viral detection and can attain species to strain-level identification. Whole-genome shotgun sequencing (WGSS) is based on the sequencing method, requiring any previous sequence information to be determined. It can analyze any number of genetic markers such as SNPs, insertion, duplication, deletion, rearrangement, genetically and engineered genomes. Previously the WGSS used Sanger sequencing but is required to use cloning vector, was time-consuming, had low output, and was more expensive (Sanger et al. 1977). PCR-based assays are one of the less expensive and easy methods to perform being utilized to analyze the strain. Although real-time PCR allows analysis of the microbial sample, this technique is limited to detecting few microbes variants (Schmedes and Budowle 2019).
SNP has the potentials of providing strain-specific information about the bacterial or any microbial genome being examined. A study conducted on Bacillus anthracis Ames strain for its detection by SNP concluded that among 88 other B. anthracis strains, they successfully found 6 different SNP characteristics to Ames strain, and 5 of them were even capable of differentiating Ames strain from its close isolates. Thus, this shows that SNP can be utilized to gain strain-level information about the microbial community (Van Ert et al. 2007). Similarly, another study on Mycobacterium tuberculosis using IS6110 SNP and some other analyses showed that SNP assays have also concluded its use for establishing ancestry in the microbial community (Faksri et al. 2011). While SNP assays having application in providing information about these disease-causing microbes, it also has potential in providing species or strain-specific information in 16s rRNA gene which is the most used gene for forensic analysis (Gu et al. 2017). One other such approach was also made to discover potential SNP of Cutibacterium acnes (Propionibacterium acnes) 16S rRNA, which is the most used microbial marker from the human skin microbiome to establish a contact of a person. It would assist in identifying different strains of C. acnes and could be incorporated for determining ownership. This resulted in discovering many such SNPs that can be utilized for the defined purpose (Yang et al. 2019). SNPs can be used as informative markers in Microbial forensic .
Variable number of tandem repeats (VNTR): Multilocus VNTR can be performed to analyze polymorphism at the minisatellite region of DNA (Budowle et al. 2005a; National Research Council 2014; Schmedes and Budowle 2019). This polymorphism is unique to a species and can be used to screen samples for a particular species or strain. This technique encompasses amplification of fragments of DNA that differ in size by the number of repeat units present within the sample (Budowle et al. 2005a). These are then separated using electrophoresis and are viewed under fluorescently labelled primer incorporated during PCR. This ensures the separation of pathogen DNA but has limitations in phylogenic isolation (Budowle et al. 2005a).
MLVA (multilocus variable number tandem repeat analysis) is another method to detect polymorphisms found in minisatellite regions found in bacterial genomes. It is found to affect the discrimination of strains of highly monomorphic species such as B. anthracis (Schmedes and Budowle 2019). MLVA also has a great species to strain identification capacity as SNP and thus can differentiate distantly related isolates and closely related isolates (Klevytska et al. 2001; Noller et al. 2003; Keim et al. 2008; Thierry et al. 2014). Varied approaches are being made to generate different VNTR markers that can be used to identify microbes. MLVA markers for microbes causing disease outbreaks such as Bacillus anthracis and for Yersinia pestis are studied. A study on the genome of B. anthracis using 31 VNTR loci showed that this technique, combined others, can potentially differentiate different strains of B. anthracis (Thierry et al. 2014).
Similarly, many such studies have shown positive discrimination of B. anthracis using multiple locus VNTR (Keim et al. 2008). Another study on Yersinia pestis using 42 VNTR loci analysis from one chromosomal and two plasmid pMT1 and pCD1 DNA sequences also suggested the potential use of MLVA for differentiation of closely and distantly related isolates (Klevytska et al. 2001). Another similar study on Escherichia coli O157:H7 suggested that MLVA is as potent a method as pulsed-field gel electrophoresis (PFGE) (Noller et al. 2003). The studies mentioned here shows that multilocus VNTR analysis can also be positively utilized for microbial analysis .
Massively parallel sequencing (MPS): Massively parallel sequencing (MPS) is another genetic tool available in the hand of a scientist. MPS is the technique used to analyze gigabase sequences of data in a short period. MPS is the technique in which millions of sequencing reactions can be carried out in a massively parallel way in a single run (National Research Council 2014; Kieser and Budowle 2020). This technique offers a complete characterization of the viral or bacterial genome. Much deeper genetic information can be achieved using this technique (Schmedes and Budowle 2019). It provides a high outturn, culture-independent method for whole genome sequencing (WGS). MPS has been a potent tool to detect and identify various disease outbreaks (Schmedes and Budowle 2019). Also, it provides an advantage that it has no requirement for enriched DNA for sequencing. MPS technique detects SNP and other genetic variants among the selected agents such as B. anthracis and Y. pestis. This technique can detect and differentiate among four different variants of both the microbial agent in a single sequencing run (Schmedes and Budowle 2019; Wetterstrand 2020).
Metagenomics: Metagenomics involves the application of sequencing genetic material collected from an environmentally generated source such as water (Biers et al. 2009), soil (Mocali and Benedetti 2010), and human linked samples (Huttenhower et al. 2012; Schmedes and Budowle 2019). Metagenomic samples are applied in the determination of various forensic aspects such as to cause of death (Kakizaki et al. 2012), identification of human (Fierer et al. 2010), time since death (Hyde et al. 2013), characterization of biological fluids (Benschop et al. 2012), and pathogenic outburst investigation (Loman et al. 2013). In the forensic investigation, the microbial samples encountered are variable, mixed profile with other organisms, and low quantity samples. Forensic metagenomics is used to analyzed target microorganisms from complex matrices (Schmedes and Budowle 2019).
These were the commonly used DNA-based techniques that can assist in the investigation of microbial attacks. As a result, these techniques are widely used in microbial forensics.
Interpretation of Results
Interpretation is a crucial step that validates the findings and provides confidence to withstand trial and scrutiny. Forensic science is based upon the comparative study between the questioned and specimen/reference samples. Three types of interpretation are generally acceptable viz., inclusion, exclusion, and inconclusive result. Inclusion is the similarity between the compared samples and shows the exact antecedent beyond a reasonable doubt. Exclusion show dissimilarity between the compared samples and different origin beyond a reasonable doubt. An inconclusive result signifies the insufficient information is obtained to summarize interpretation and reach any specific conclusion. The result should be statistically strengthened by using various tools to validate and concrete scientists’ observations and should be able to endure strongly before the legal system. When dealing with microbial data, interpretation of results is critical as it requires high knowledge about sequencing and other techniques. Different techniques have their interpretation guidelines. For instance, in the case of analysis using commercially available RT-PCR kits, the presence of a particular microbe is or can be confirmed using fluorescence which confirms its presence. This is relatively easy and does not require much knowledge about sequencing data. However, when dealing with more sophisticated techniques such as SNP, VNTR, RFLP, MPS, metagenomics, etc., prior knowledge about sequencing phenomena and other basics of technique is needed to interpret results accurately. While interpreting results, the other thing that should be looked upon is reviewing sampling technique, sample condition, external factors affecting samples such as geographical location, temperature, humidity, etc., affecting microbial community colonizing at the site of colonization. These are how interpretation can be interfered by different factors needed to be kept in hand while interpreting results .
Conclusion
Microbial forensics is the emerging branch of forensic science. It plays a vital role in investigating bio-crimes. The increasing use of microorganisms as war agents has also increased the need to have a body that separately analyses such outbreaks. Microbial forensics deals with the collection, preservation, storage, transport, and analysis of microbial forensic evidence. Forensic analysis of such evidence incorporates many techniques, among which molecular techniques may be helpful for analysis. Different molecular techniques used in forensic microbiology are discussed in the above sections. Further standardization is needed to strengthen the system to make it more robust and enabling high throughput outcomes.
References
Aqeel L, Omran R (2018) Detection of staphylococcal species diversity within human skin microbiome using a simple PCR-SSCP technique. J Univ BABYLON Pure Appl Sci 26:1–6. https://doi.org/10.29196/jubpas.v26i8.1646
Assiri A, McGeer A, Perl TM et al (2013) Hospital outbreak of Middle East respiratory syndrome coronavirus. N Engl J Med 369:407–416. https://doi.org/10.1056/NEJMoa1306742
Barras V, Greub G (2014) History of biological warfare and bioterrorism. Clin Microbiol Infect 20:497–502. https://doi.org/10.1111/1469-0691.12706
Bauer M, Kraus A, Patzelt D (1999) Detection of epithelial cells in dried blood stains by reverse transcriptase-polymerase chain reaction. J Forensic Sci 44:1232–1236
Bell CR, Wilkinson JE, Robertson BK, Javan GT (2018) Sex-related differences in the thanatomicrobiome in postmortem heart samples using bacterial gene regions V1-2 and V4. Lett Appl Microbiol 67:144–153. https://doi.org/10.1111/lam.13005
Benschop CCG, Quaak FCA, Boon ME et al (2012) Vaginal microbial flora analysis by next generation sequencing and microarrays; can microbes indicate vaginal origin in a forensic context? Int J Legal Med 126:303–310. https://doi.org/10.1007/s00414-011-0660-8
Biers EJ, Sun S, Howard EC (2009) Prokaryotic genomes and diversity in surface ocean waters: interrogating the global ocean sampling metagenome. Appl Environ Microbiol 75:2221–2229. https://doi.org/10.1128/AEM.02118-08
Blondeau LD, Rubin JE, Deneer H et al (2019) Forensic, investigative and diagnostic microbiology: similar technologies but different priorities. Future Microbiol 14:553–558. https://doi.org/10.2217/fmb-2019-0088
Budowle B, Schutzer SE, Einseln A et al (2003) Public health. Building microbial forensics as a response to bioterrorism. Science 301:1852–1853. https://doi.org/10.1126/science.1090083
Budowle B, Johnson MD, Fraser CM et al (2005a) Genetic analysis and attribution of microbial forensics evidence. Crit Rev Microbiol 31:233–254. https://doi.org/10.1080/10408410500304082
Budowle B, Schutzer S, Breeze R (eds) (2005b) Microbial forensics, 1st edn. Academic Press
Budowle B, Schutzer SE, Burans JP et al (2006) Quality sample collection, handling, and preservation for an effective microbial forensics program. Appl Environ Microbiol 72:6431–6438. https://doi.org/10.1128/AEM.01165-06
Castle JW, Butzbach DM, Walker GS et al (2017) Microbial impacts in postmortem toxicology. In: Forensic microbiology. Wiley, Chichester, pp 212–244
Cenciarelli O, Pietropaoli S, Malizia A et al (2015) Ebola virus disease 2013–2014 outbreak in West Africa: an analysis of the epidemic spread and response. Int J Microbiol 2015:1–12. https://doi.org/10.1155/2015/769121
Centres for Disease Control and Prevention (2018) Bioterrorism agents/diseases. In: Centers Dis. Control Prev. https://emergency.cdc.gov/agent/agentlist.asp. Accessed 10 Jun 2021
Department of Health and Human Services, National Institutes of Health (2019) NIH guidelines for research involving recombinant or synthetic nucleic acid molecules. NIH Guidel 142
Díaz-Palma PA, Alucema A, Hayashida G, Maidana NI (2009) Development and standardization of a microalgae test for determining deaths by drowning. Forensic Sci Int 184:37–41. https://doi.org/10.1016/j.forsciint.2008.11.015
Faksri K, Drobniewski F, Nikolayevskyy V et al (2011) Genetic diversity of the Mycobacterium tuberculosis Beijing family based on IS6110, SNP, LSP and VNTR profiles from Thailand. Infect Genet Evol 11:1142–1149. https://doi.org/10.1016/j.meegid.2011.04.007
Faria NR, Azevedo R d. S d. S, Kraemer MUG et al (2016) Zika virus in the Americas: early epidemiological and genetic findings. Science 352:345–349. https://doi.org/10.1126/science.aaf5036
Fernández-Rodríguez A, Burton JL, Andreoletti L et al (2019) Post-mortem microbiology in sudden death: sampling protocols proposed in different clinical settings. Clin Microbiol Infect 25:570–579. https://doi.org/10.1016/j.cmi.2018.08.009
Fierer N, Lauber CL, Zhou N et al (2010) Forensic identification using skin bacterial communities. Proc Natl Acad Sci U S A 107:6477–6481. https://doi.org/10.1073/pnas.1000162107
Flowers LK, Mothershead JL, Blackwell TH (2002) Bioterrorism preparedness. II: the community and emergency medical services systems. Emerg Med Clin North Am 20:457–476. https://doi.org/10.1016/s0733-8627(01)00009-8
Gardy JL, Johnston JC, Sui SJH et al (2011) Whole-genome sequencing and social-network analysis of a tuberculosis outbreak. N Engl J Med 364:730–739. https://doi.org/10.1056/NEJMoa1003176
Goodman RA, Munson JW, Dammers K et al (2003) Forensic epidemiology: law at the intersection of public health and criminal investigations. J Law Med Ethics 31:684–700. https://doi.org/10.1111/j.1748-720x.2003.tb00135.x
Grad YH, Lipsitch M, Feldgarden M et al (2012) Genomic epidemiology of the Escherichia coli O104:H4 outbreaks in Europe, 2011. Proc Natl Acad Sci 109:3065–3070. https://doi.org/10.1073/pnas.1121491109
Gu Y, Zha L, Yun L (2017) Potential usefulness of SNP in the 16S rRNA gene serving as informative microbial marker for forensic attribution. Forensic Sci Int Genet Suppl Ser 6:e451–e452. https://doi.org/10.1016/j.fsigss.2017.09.176
Hanson E, Haas C, Jucker R, Ballantyne J (2012) Specific and sensitive mRNA biomarkers for the identification of skin in “touch DNA” evidence. Forensic Sci Int Genet 6:548–558. https://doi.org/10.1016/j.fsigen.2012.01.004
Hanssen EN, Liland KH, Gill P, Snipen L (2018) Optimizing body fluid recognition from microbial taxonomic profiles. Forensic Sci Int Genet 37:13–20. https://doi.org/10.1016/j.fsigen.2018.07.012
Hauther KA, Cobaugh KL, Jantz LM et al (2015) Estimating time since death from Postmortem human gut microbial communities. J Forensic Sci 60:1234–1240. https://doi.org/10.1111/1556-4029.12828
Hendriksen RS, Price LB, Schupp JM et al (2011) Population genetics of vibrio cholerae from Nepal in 2010: evidence on the origin of the Haitian outbreak. MBio 2. https://doi.org/10.1128/mBio.00157-11
Huttenhower C, Gevers D, Knight R et al (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214. https://doi.org/10.1038/nature11234
Hyde ER, Haarmann DP, Lynne AM et al (2013) The living dead: bacterial community structure of a cadaver at the onset and end of the bloat stage of decomposition. PLoS One 8:e77733. https://doi.org/10.1371/journal.pone.0077733
Javan GT, Finley SJ, Can I et al (2016) Human Thanatomicrobiome succession and time since death. Sci Rep 6:29598. https://doi.org/10.1038/srep29598
Jung JY, Yoon HK, An S et al (2018) Rapid oral bacteria detection based on real-time PCR for the forensic identification of saliva. Sci Rep 8:10852. https://doi.org/10.1038/s41598-018-29264-2
Kageyama T, Fujisaki S, Takashita E et al (2013) Genetic analysis of novel avian a(H7N9) influenza viruses isolated from patients in China, February to April 2013. Euro Surveill 18:20453
Kakizaki E, Kozawa S, Sakai M, Yukawa N (2009) Bioluminescent bacteria have potential as a marker of drowning in seawater: two immersed cadavers retrieved near estuaries. Legal Med 11:91–96. https://doi.org/10.1016/j.legalmed.2008.10.004
Kakizaki E, Ogura Y, Kozawa S et al (2012) Detection of diverse aquatic microbes in blood and organs of drowning victims: first metagenomic approach using high-throughput 454-pyrosequencing. Forensic Sci Int 220:135–146. https://doi.org/10.1016/j.forsciint.2012.02.010
Keim P, Pearson T, Okinaka R (2008) Microbial forensics: DNA fingerprinting of bacillus anthracis (anthrax). Anal Chem 80:4791–4799. https://doi.org/10.1021/ac086131g
Kieser RE, Budowle B (2020) Select methods for microbial forensic nucleic acid analysis of trace and uncultivable specimens. In: Microbial forensics. Elsevier, pp 195–205
Klevytska AM, Price LB, Schupp JM et al (2001) Identification and characterization of variable-number tandem repeats in the Yersinia pestis genome. J Clin Microbiol 39:3179–3185. https://doi.org/10.1128/JCM.39.9.3179-3185.2001
Kolavic SA, Kimura A, Simons SL et al (1997) An outbreak of Shigella dysenteriae type 2 among laboratory workers due to intentional food contamination. JAMA 278:396–398
Lax S, Hampton-Marcell JT, Gibbons SM et al (2015) Forensic analysis of the microbiome of phones and shoes. Microbiome 3:21. https://doi.org/10.1186/s40168-015-0082-9
Levkov Z, Williams DM, Nikolovska D et al (2017) The use of diatoms in forensic science: advantages and limitations of the diatom test in cases of drowning. In: The archaeological and forensic applications of microfossils: a deeper understanding of human history. The Geological Society of London on behalf of The Micropalaeontological Society, pp 261–277
Loman NJ, Constantinidou C, Christner M et al (2013) A culture-independent sequence-based Metagenomics approach to the investigation of an outbreak of Shiga-toxigenic Escherichia coli O104:H4. JAMA 309:1502. https://doi.org/10.1001/jama.2013.3231
Meadow JF, Altrichter AE, Green JL (2014) Mobile phones carry the personal microbiome of their owners. Peer J 2:e447. https://doi.org/10.7717/peerj.447
Metcalf JL (2019) Estimating the postmortem interval using microbes: knowledge gaps and a path to technology adoption. Forensic Sci Int Genet 38:211–218. https://doi.org/10.1016/j.fsigen.2018.11.004
Metcalf JL, Wegener Parfrey L, Gonzalez A et al (2013) A microbial clock provides an accurate estimate of the postmortem interval in a mouse model system. Elife 2. https://doi.org/10.7554/eLife.01104
Mocali S, Benedetti A (2010) Exploring research frontiers in microbiology: the challenge of metagenomics in soil microbiology. Res Microbiol 161:497–505. https://doi.org/10.1016/j.resmic.2010.04.010
Morse SA, Budowle B (2006) Microbial forensics: application to bioterrorism preparedness and response. Infect Dis Clin N Am 20:455–473. https://doi.org/10.1016/j.idc.2006.03.004
Morse SA, Budowle B, Schutzer SE (2019) Microbial forensics: what next? In: Microbial forensics
Murch RS (2003) Microbial forensics: building a national capacity to investigate bioterrorism. Biosecur Bioterror Biodef Strateg Pract Sci 1:117–122. https://doi.org/10.1089/153871303766275781
National Research Council (2014) Science needs for microbial forensics: developing initial international research priorities. National Academies Press, Washington, DC
Nema V (2018) Microbial forensics: beyond a fascination. In: DNA fingerprinting: advancements and future endeavors. Springer Singapore, Singapore, pp 295–306
Noller AC, McEllistrem MC, Pacheco AGF et al (2003) Multilocus variable-number tandem repeat analysis distinguishes outbreak and sporadic Escherichia coli O157:H7 isolates. J Clin Microbiol 41:5389–5397. https://doi.org/10.1128/JCM.41.12.5389-5397.2003
Oliveira M, Amorim A (2018) Microbial forensics: new breakthroughs and future prospects. Appl Microbiol Biotechnol 102:10377–10391. https://doi.org/10.1007/s00253-018-9414-6
Park J-L, Park S-M, Kwon O-H et al (2014) Microarray screening and qRT-PCR evaluation of microRNA markers for forensic body fluid identification. Electrophoresis 35:3062–3068. https://doi.org/10.1002/elps.201400075
Power DA, Cordiner SJ, Kieser JA et al (2010) PCR-based detection of salivary bacteria as a marker of expirated blood. Sci Justice 50:59–63. https://doi.org/10.1016/j.scijus.2009.04.006
Rajalakshmi S (2017) Different types of PCR techniques and its applications. Int J Pharm Chem Biol Sci 7:285–292
Robinson JM, Pasternak Z, Mason CE, Elhaik E (2020) Forensic applications of microbiomics: a review. Front Microbiol 11:608101. https://doi.org/10.3389/fmicb.2020.608101
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A 74:5463–5467. https://doi.org/10.1073/pnas.74.12.5463
Schmedes S, Budowle B (2019) Microbial forensics☆. In: Reference module in biomedical sciences: encyclopedia of microbiology, 4th edn. Elsevier, pp 134–145
Schmedes SE, Woerner AE, Budowle B (2017) Forensic human identification using skin microbiomes. Appl Environ Microbiol 83. https://doi.org/10.1128/AEM.01672-17
Schutzer SE, Budowle B, Keim PS (2011) Microbial forensics: educating the workforce and the community. In: Microbial forensics. Elsevier, pp 667–680
Sego LH, Anderson KK, Matzke BD et al (2007) An environmental sampling model for combining judgment and randomly placed samples, Richland
Smith JAL (2019) Collection and preservation of microbial forensic samples. In: Microbial forensics
Smith JAL (2020) Collection and preservation of microbial forensic samples. In: Microbial forensics. Elsevier, pp 313–322
Smith GJD, Vijaykrishna D, Bahl J et al (2009) Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459:1122–1125. https://doi.org/10.1038/nature08182
The College of Physicians of Philadelphia (2018) Biological weapons, bioterrorism, and vaccines. Hist. Vaccines An Educ. Resour. by Coll. Physicians Philadelphia
Thierry S, Tourterel C, Le Flèche P et al (2014) Genotyping of French Bacillus anthracis strains based on 31-loci multi locus VNTR analysis: epidemiology, marker evaluation, and update of the internet genotype database. PLoS One 9:e95131. https://doi.org/10.1371/journal.pone.0095131
Török TJ, Tauxe RV, Wise RP et al (1997) A large community outbreak of salmonellosis caused by intentional contamination of restaurant salad bars. JAMA 278:389–395. https://doi.org/10.1001/jama.1997.03550050051033
Tridico SR, Murray DC, Addison J et al (2014) Metagenomic analyses of bacteria on human hairs: a qualitative assessment for applications in forensic science. Investig Genet 5:16. https://doi.org/10.1186/s13323-014-0016-5
Van Ert MN, Easterday WR, Simonson TS et al (2007) Strain-specific single-nucleotide polymorphism assays for the Bacillus anthracis Ames strain. J Clin Microbiol 45:47–53. https://doi.org/10.1128/JCM.01233-06
Varun CN, Kuruvilla Thomas S, Zevita F (2012) Microbial forensics- past, present and future. Int J Biol Med Res 3:1546–1549
Wang Z, Zhang S, Di Z et al (2013) Messenger RNA profiling for forensic body fluid identification: research and applications. Fa Yi Xue Za Zhi 29:368–374
Wetterstrand KA (2020) DNA sequencing costs: data from the NHGRI Genome Sequencing Program (GSP). http://www.genome.gov/sequencingcostsdata. Accessed 10 Jun 2021
Wikipedia Contributors (2021) List of epidemics. In: Wikipedia free encyclopedia. https://en.wikipedia.org/w/index.php?title=List_of_epidemics&oldid=1028613106. Accessed 15 Jun 2021
Yang J, Tsukimi T, Yoshikawa M et al (2019) Cutibacterium acnes (Propionibacterium acnes) 16S rRNA genotyping of microbial samples from possessions contributes to owner identification. mSystems 4. https://doi.org/10.1128/mSystems.00594-19
Zhang Y, Pechal JL, Schmidt CJ et al (2019) Machine learning performance in a microbial molecular autopsy context: a cross-sectional postmortem human population study. PLoS One 14:e0213829. https://doi.org/10.1371/journal.pone.0213829
Zou K-N, Ren L-J, Ping Y et al (2016) Identification of vaginal fluid, saliva, and feces using microbial signatures in a Han Chinese population. J Forensic Legal Med 43:126–131. https://doi.org/10.1016/j.jflm.2016.08.003
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Singapore Pte Ltd.
About this entry
Cite this entry
Kapoor, N., Sulke, P., Badiye, A. (2022). Molecular Techniques in Microbial Forensics. In: Dash, H.R., Shrivastava, P., Lorente, J.A. (eds) Handbook of DNA Profiling. Springer, Singapore. https://doi.org/10.1007/978-981-16-4318-7_44
Download citation
DOI: https://doi.org/10.1007/978-981-16-4318-7_44
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-4317-0
Online ISBN: 978-981-16-4318-7
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences