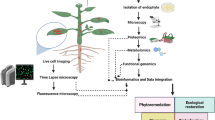
Abstract
Plants are home to a wide assemblage of nonpathogenic microbial community belonging to different phyla, bacteria, fungi, actinomycetes and viruses, the collective term for which is called endophyte. These endosymbiotic individuals exhibit endophytism principally by assisting in vigor and endurance to host plant and protect them from biotic (pathogenic infections) and abiotic stress (water, heat, nutrient, salinity, and herbivory). In return, these endosymbionts receive energy in the form of carbon from the host tissue. Colonization of endophyte in the internal tissues has been reported almost in every plant examined so far either in intercellular or intracellular mode. The form of relationships established with the host plant may be mutualistic, symbiotic, commensalistic, and trophobiotic. These are either rhizospheric or phyllospheric in origin. To establish such mutualistic relationships between plants and endophytes, certain chemical signals play important role in inducing production of the enhanced amount of secondary metabolites in host plant tissues. These novel metabolites act as a very good source of stress relievers to host and protect from grazing animals. The renewed interest in endophyte is due to the biotechnological relevance of these signal molecules as these have been used as a good source for production of biochemical compounds of industrial importance more specifically in agriculture and medicine. Additionally, their capacity to decontaminate soil bacteria and bring in soil fertility invites huge application in phytoremediation. However, the physiology, biochemistry, and genetics behind such complex interactions, exchange of chemical signals, and their production (the endophytism of plan-microbiome) are still half-understood. With the advent of new efficient analytical technology in molecular biology and genomics, the basic information on the existing diversity, phylogenetic lineage, evolution, and ecophysiological information about these endophytes has been tried to understand. However, the functional gene expression, posttranslational modifications, and protein turnover under various environmental circumstances are only revealed through transcriptome and proteomics analysis. Soon, high-throughput next-generation sequencing technology has remarkably changed the whole scenario of solving the intricate issues entangled with the complexity underlying endophytism. Sequencing of the whole genome of individuals following cultivable method (genomics), multiple host plants and their microbiome (comparative genomics), non-cultivable methods (metagenomics and metatranscriptomics), and microarray has been proved to be potential approaches to unravel the truth behind the plant-endophyte interactions. The present script deals with scopes, prospects, and outcomes of use of these “omics tools” to understand the deep insight into the mechanism of plant host infestation, biological reason behind the mutualism between host and endophytes, exchange of biochemical compounds, enhanced production of secondary metabolite, and host plant ecology.
Access provided by Autonomous University of Puebla. Download reference work entry PDF
Similar content being viewed by others
Keywords
1 Introduction
The compelling interest in endophytes has been for the secondary metabolites they are able to produce. These molecules of natural origin generally hold overabundance of beneficial properties useful as a source of potential drug [1, 2]. With time, molecules with desired bioactivities have been identified and isolated from a large number of endophytes from a list of medicinal plants which are yet to be explored for their large-scale commercial production. Accumulation of these secondary metabolites, nutrients, and hormones might have been produced in host plants associated with endophytes in response to the biotic and abiotic stress or due to some unknown reason during such mutualism exhibited by the endophytes [3, 4].
Therefore, understanding the whole science behind the establishment of endophytism is the prime effort to be taken so as to utilize the incredible potential of these high valued molecules produced by endophytes having potential applications in pharmaceutical, food, agriculture, and medical industry. So far, attempts are taken to establish their identity and diversity and to unravel the metabolite potential. But, there has been a paradigm shift among the scientific community toward understanding the physiology, biochemistry, and the genetics behind the plant-endophyte relationship of several ecological niches.
Endophytes are basically bacteria or fungi which reside as intercellular or intracellular in rhizospheric or phyllospheric tissues of the host plant under symbiosis or commensal type of association. Horizontally transmitted endophytes, the most ubiquitous fungal endophyte, inhabiting major plants studied for their potential production of bioactive molecules, have been subjected to unanswered questions on interactions of endophytes with their plant hosts, phytophagous insects, and other fungi. The present review highlights the possible role of modern omics-based methods in understanding the gray areas of endophytism and their potential exploration in different avenues of biotechnology.
With the advent of new efficient analytical technology in molecular biology and genomics, the basic information on the existing diversity, phylogenetic lineage, evolution, and ecophysiological information about these endophytes has been understood [5]. Although the genomic study provides the information on molecular machinery and functional expression is only revealed through transcriptome analysis under various environmental circumstances without any information on posttranslational modifications and protein turnover, etc. proteomics deals with the study of functional gene expression products. Alone, the transcriptomic or proteomic study is incomplete in interpretation in absence of genomic information. Moreover, supplementing the information generated from the metagenomic study with those of metatranscriptomics and metaproteomics may help to find detailed intricacies involved in the establishment of endophytism. All these techniques although self-sufficient are inter-reliant, and thus the information obtained from individual method or technique is the accompaniment to each other. Thus, the combinatorial approach of analyzing the data produced from various recent “omics” tools will help in resolving the enigma existing in the endophyte-host relationship. Genome sequencing options, metagenomics and metatranscriptomics, have increased the perspective of analyzing the microbial community. These meta-omics methodologies explore the community having the genes, transcripts, and proteins from millions of microbes and provide a scope to analyze their biochemical functions as well as systems-level microbial interactions. Functional assays involving whole community analysis in addition to metagenomics and metatranscriptomics offer new avenues to understand biogeochemical environments, complex ecosystems involving host organisms, their metabolism, and the possible interactions among them. These meta-omics studies characteristically aim to recognize a panel of microorganisms, genes, their variants, and metabolic pathways of the microbial community inhabiting an uncultivated sample. These abovementioned analytical methods complemented with advanced computational tools (systems biology science) are the key approaches to understanding significant biochemical and environmental interactions occurring in a community. Thus, we have described here the current skills, recent technological advances, and unresolved challenges involved in the functional analysis of microbial community.
2 Conventional Techniques Used in Endophyte Studies
2.1 Direct Observation Method
This is the most common, simple, and preliminary method of observing the endophyte harboring the living host plant tissue directly under light or electron microscope. This method reveals the limited morphological features of infested microorganism inside the intercellular and rarely intracellular tissue of the plant which is generally restricted to the hyphal structure or the shape of the bacteria. It excludes isolation of endophyte in vitro and further possible characterization. Since it cannot provide any information about taxonomically distinguishable features like spores/endospores/conidia or spore-producing structures, this method can hardly be used for understanding the phylogenetic identification and biodiversity analysis of the endophyte [6, 7].
2.2 Cultivation-Dependent Method
Typical methodology of isolation and cultivation of endophyte through in vitro culture-dependent techniques involves the following few steps (Fig. 1): (i) surface sterilization of host plant tissue infested with endophyte adopting different protocols [8], (ii) isolation of endophyte grown out of the incubated plant sample on suitable media, (iii) manipulation of cultivation and incubation parameters to promote sporulation, and finally (iv) identification through morphological, microscopic, and biochemical analysis [9,10,11,12,13]. This cultivation-dependent method has been followed across the globe since it is one of the rapid effective methods of isolation of endophytes from the plant tissue under changeable parameters during the whole process of sterilization, inoculation, and incubation in artificial culture media. Cultivation and characterization of endophyte isolates have been inevitable not only to understand the population structure and species diversity [13,14,15,16,17] but also to unravel the physiology behind its role in plant growth and protection through the production of secondary metabolic compounds [18,19,20,21].
Flow chart showing the cultivation-dependent analysis of endophyte, Source: [65]
Reports reveal that the enhanced recovery of endophyte from the host plant using the smaller size of tissue incubated [22] or whole leaves instead of leaf disk [13, 23]. However, retrieval and growth of the higher amount of endophyte without spores (sterile isolates) add problem in detailed characterization and identification as no taxonomic units have been assigned based on limited morphological features. This urges implementation of different means to promote sporulation or production of the fruiting body. Guo and his coworkers could enhance the rate of sporulation from 48% to 59.5% by incubating the palm leaf tissue onto media surface and again to 83.5% though longer incubation of isolates for 3 months onto pieces of petiole of the leaves [24]. It has been also observed that some of the endophyte species of a community might be suppressed by fast-growing isolates in vitro due to competition for nutrients in artificial media.
However cultivation-dependent method has been subjected to methodological shortcomings and technical biases. Characterization, more specifically sporulation of endophyte, gets affected by the techniques followed for sterilization, the conditions maintained for incubation, and the type of media used. The adaptability of the plant type, the tissue size, their number, and the endophyte community to the overall procedure of isolation also bring in limitations in revealing the facts and features about the harboring endophyte in the host tissue.
3 Omics Intervention in Endophyte Studies
3.1 Genomic Analysis by the Cultivation-Dependent Method
In absence of omics-based analytical methods, different isolates obtained from conventional in vitro cultivation procedures having similarities in morphological (color and texture of colonies) and growth characteristics had been named as different “morphotypes” and were designated as “Mycelia sterilia” (where the sporulation could not be obtained). But these morphotypes could not be accepted as units of the taxon to classify and establish the diversity existing within them and failed as the criteria to establish phylogenetic lineage [13, 16, 25, 26]. With the intervention of molecular techniques, the bottlenecks that generally cropped up through traditional protocols for identification and diversity analysis could, however, be overcome.
Molecular identification of sporulating and non-sporulating endophytes basing on DNA markers like ITS, 23S, and 18S for fungus and 16S for bacteria may be the suitable solution in detecting the diversity existing in the community. Using ITS marker, 19 non-sporulating morphotypes of L. chinensis could be identified and grouped into three genera Mycosphaerella, Xylaria, and Diaporthe [16]. Similarly, 221, 74, and 18 non-sporulating fungal endophytes were grouped into 37, 64, and 3 taxa, respectively [13, 25, 26]. González and Tello in Spain could assign taxonomic identifier at the level of genus and species for non-sporulating Vitis vinifera employing ITS sequences [27]. In this way, these DNA marker-based molecular analyses will not only help to assign a taxonomic place for the community present in phyllosphere and rhizosphere but also understand the species diversity existing within them. ITS analysis supported with the morphological information became the preferred practice specifically for understanding biodiversity among the isolates present in host tissues and their ecology [28, 29]. As uses of 18S and 28S genes are generally employed to find out the higher taxonomic level (order and suborder) for endophytic fungi, these genes are analyzed in supplementation to ITS marker study which reveals the taxonomic lineage at a lower level (genus and species) and to detect novelty. Morakotkarn and his associate could identify 71 strains from host bamboos belonging to Phyllostachys and Sasa species under Sordariomycetes and Dothideomycetes order by employing ITS and 18S, respectively [30]. Similar protocols were followed by other workers for taxonomic diversity analysis of Theobroma cacao and Pinus halepensis, respectively [31, 32].
The abovementioned cultivable methods and techniques have been limited to identification of only those isolates that could be cultivable in artificial media, the establishment of their novelty, and the understanding of diversity existing among the community. Nevertheless, outcome of these protocols fails to throw sufficient light on deciphering the relationship between the host plant and the endophyte and the molecular basing of intricacies behind endophytism probably because these methods do not truly encourage the growth of all the members of endophyte present in the community in the plant tissue in a defined artificial media in vitro (cultivation-dependent method).
3.2 Metagenomic Analysis by Cultivation-Independent Method
Metagenomics is the genomic analysis of total DNA of all the members of the microbial community in an environment which is otherwise called as community genomics or environmental genomics bypassing the detection and in vitro cultivation of every single organism present in any microbiome. Metabolic implications and factors associated with host-endophyte interactions, due to non-cultivable microbes whose population is reported to be much higher (90–99%) in any environmental sample than the in vitro cultivable isolates, can thus be better realized following this protocol [33]. The size of metagenomic DNA (DNA of the entire microbiome present in the sample) is generally of huge size and warrants a fast and efficient high-throughput method to handle and analyze the large-sized genome and suitable pipelines and software to translate into understandable information. Next-generation sequencing (NGS) is the most recent intervention for metagenomic analysis which brings in the unrecorded unprecedented information of the microorganisms present in any host-endophyte association much beyond the data generated from individual cultivable taxa. It is further supported by several numbers of tools that make fat data into information explicable to the analyzer.
With the recent discovery and intervention of alternate omics tools since the last two decades, the above inherent disadvantages of culturable methods can be overcome where total community genomic DNA of the sample (both host plant and the endophyte) is subjected to molecular analysis. Non-cultivable or cultivation-independent methods involve a sequence of molecular reaction steps as shown in Fig. 2: (i) community DNA (genomic DNA of host plant and all the members of endophyte present) isolation; (ii) ITS, 28S, and 18S gene amplification for fungal and 16S for bacterial endophyte; (iii) electrophoretic separation and excision of bands generated from DGGE (denaturing gradient gel electrophoresis); (iv) cloning into vector and transforming into heterologous host E.coli DH5α and sequencing; and (v) phylogenetic analysis using NCBI database for identification of the taxa.
Flow chart showing the cultivation-independent analysis (metagenomic and predicted functional) of endophyte, Source: [65]
The outcome from the genome analysis through uncultivable method employing ITS could unravel novel taxa never been reported through cultivation-dependent method which are YJ4-61, YJ4-9, and YJ4-70 from H. japonica tissues [34], 1 unidentifiable clone from L. chinensis [35], and 14 novel taxonomic units from Magnolia liliifera [36]. The novelty attained by this method of exploring the endophyte community diversity could be possible due to the ability to overcome the technical biases of traditional protocols that did not allow scoring all the genomes present but could not be grown in vitro and the high-resolution ability of DGGE coupled with sequencing covering the whole genome.
3.3 Predicted Functional Analysis of Metagenome
In a sequence-based analysis, genomic information is assessed from microbes without culturing them and can be used to identify microorganisms and genes and compare organisms of different communities. Sequence-based metagenomics can also be used to establish the diversity, enumeration of bacterial species present in the sample, ecophysiological relationship with the microflora dwelling in it with prevailing physiochemical parameters of that environment, and predicted genes and metabolic pathways. Analyzing microbial diversity can provide valuable information at less cost of experimentation and also predict the metabolism prevailing and the ecology of microbes. Recent developments on different efficient cloning vectors, along with newer methods of DNA isolation and sequencing, have been possible to clone and express bigger-sized DNA into large-sized metagenomics clone library for functional analysis. Over the past 10 years, shotgun sequencing technology used in metagenomics has gradually shifted from classical Sanger sequencing to NGS methods [37]. Although Sanger sequencing technology is the best sequencing technology because of its low error rate for sequencing maximum 30 Kb insert size [37], the main disadvantages are the labor-intensive cloning process and cost-intensive factor for giga base pair sequencing (approximately 400,000 USD) [37]. In next-generation sequencing technology, 454/Roche and Illumina/Solexa systems are widely used for analyzing the sequence of the microbial community and functional analysis of metagenomic samples. The sequence reads generated from NGS methods are generally shorter than Sanger’s sequencing read. In 454/Roche technology, average read length is 600–800 bp and produces ~500 Mbp in single run, whereas in Illumina/Solexa, the read length is 150–300 bp and produces ~6 Gbp in single run sequencing [37]. After NGS sequencing, post-sequencing analysis such as assembly, annotation, binning, ORF prediction, taxonomic profiling, and metabolic reconstruction is the most challenging step which decides the output of any metagenomic sample. Several bioinformatic tools and data storage pipelines have been developed to simplify the post-sequencing analysis, such as MEGAN [38], MG-RAST [39], GALAXY [40], CAMERA [13], and MetAMOS [41]. PICRUSt [42] and TAX4FUN [43] tools are used to analyze the predicted functional activity by using 16S rRNA gene sequences, the details of which presented in Fig. 2. However, so far as our knowledge goes, many attempts are taken to predict the functional genes and their possible activities from any community DNA of any endophytic niche.
3.4 Multigenomic Analysis
Whole genome analysis of an individual endophyte harboring a plant may not be able to completely establish their lifestyle, the kind of which may vary from mutualistic symbionts to commensalistic symbionts or saprotropism to biotropism. They can also behave as latent pathogens and latent saprotrophs [44]. Therefore, comparing the genome of isolates having endophyte association and the non-endophyte complement can help realize the controlling factors responsible for their adaptation to host plant, their evolutionary trajectory, and genetic basis of endophytism exhibited by them [45].
Endophyte adaptation, potential to promote the growth of host plant, as well as the tolerance to stress and production of protection metabolites could be understood from metagenomic analysis of rice plant root tissue associated with the endophyte [46]. Dinsdale and his groups presented the differential functional characters of nine endophyte microbiome following comparative metagenomic analysis [33]. Comparative genome analysis using Illumina platform for Cadophora sp. and Periconia macrospinosa with their 32 close relatives with different lifestyles could reveal the functional differences with respect to the presence of a number of genes for aquaporins, melanin synthesis, enzyme proteases, and lipases, despite their common origin. The insight into basic biological and evolutional understandings has been made available through comparative genomic study in several endophyte species, M. bolleyi (37), P. subalpine (29), S. indica (34), X. heveae, (31), P. scopiformis (33), and C. trifolium (35), originating from different habitats [47]. The detailed community diversity of fungal endophyte and its composition in a Japanese forest have been analyzed [48]. Large-scale functional characterization of fungal communities using 454 genomes employing metagenomic protocol could accumulate a surplus of information of ecophysiology of the endophyte community which reported the existence of fungi of both mycorrhizal and endophytic origin [49].
Since the metagenomic analysis has been recently the intervened approach in understanding endophytism as a whole, attention must be given in making the public database more furnished with genome/reference genome sequence information for the target plant/endophyte species. However, the intervention of proteomic analysis in supplementation with the metagenomic analysis in a non-cultivable approach can help further to understand the existing interaction of these two ecotypes.
3.5 Transcriptomics and Metatranscriptomic Analysis
Although whole genome analysis or metagenomic analysis could provide the existence of genes in a community, their functionality in terms of whether the gene is expressed in that particular environment could not be accounted which is very much key in realizing the endophytism in any endophyte-host plant association. The environmental parameters present in and around of any ecological niche determine the expression of a character in any organism irrespective of the presence of the gene that controls it. Therefore, understanding differentially expressed genes with respect to the altered environment through isolation and characterization of all the RNA present in a community (transcriptomics and metatranscriptomics) can be a better way of knowing the response of interacting endophyte species with the host and the environment. Comparative expression analysis of the transcriptome of plants with and without endophyte infestation and of endophytes in and outside of host can help to understand the interactive factors responsible for endophytism, production of secondary metabolites, and plant growth-promoting substances. Ambrose and Belanger and their associates successfully revealed the differential expression of 200 genes associated with host plant Epichloe festucae infested with endophyte named Festuca rubra. However, these transcriptome data correlated with the data generated from their respective genomes can complete the understanding about the facts [50].
In the metatranscriptomic analysis, the transcripts or RNAs are directly isolated from environment or community. This type of analysis brings the direct connection between the genetic makeup of the community and the respective functionality in situ through the profiling of the expressed transcripts and linking them with the prevailing ecophysiological conditions. Such metatranscriptomic analysis is accomplished by either cDNA clonal libraries derived from mRNA as given in Fig. 3.
Flow chart showing the metatranscriptomic analysis of endophyte, Source: [65]
Using dual RNA-sequencing technology for comparative transcriptional profiling, the differential regulation of genes meant for nutrient availability was observed in wheat roots infested with bacterial endophyte A. brasilense [51]. This helped him to interpret the basic mutualistic relationship existing between them. The occurrence of transcripts foreign to host soybean genome system through comparative metatranscriptomic analysis helped in tracing the infestation of endophytes and free-living microbes in different soybean host plants [52].
Although these recent methods of analyzing community RNA provide a considerable amount of information and insight, they are not free from limitations. First, extraction of RNA directly from an environmental sample is often problematic and the concentration is often low. For this reason, previous studies have used additional amplification steps to increase the concentration of initial transcripts [45, 53]. Second, separation of mRNA from the abundant non-mRNA (e.g., ribosomal or transfer RNA) is also problematic, and, as a result, the gene expression profile of the sample often remains limited. Consequently, the low gene expression profile may not result in statistically meaningful transcription patterns or may not provide sufficient coverage for most of the genes of a complex community. Thus, in earlier studies, the focus was only on the most dominant members present in a respective community.
3.6 Proteomics and Metaproteomic Analysis
Soon after the realization that genomic and metagenomic analysis is still unable to unravel the real-time in situ functional information about the community, post-genomic analysis like proteomics and metaproteomics is gaining expedition. Proteomics involves the scale analysis of total proteins present in an organism, and on the other hand metaproteomics is basically the analysis of functional expression of the community genes and interpretation of activities at the time of sampling. Metaproteomics is the process of direct identification and assessment of the prevailing functionality of the microbial community of an environmental sample. It directly assesses the microbial functional profile. In addition, the developments of computing and bioinformatic tools provide a more solid source of protein identification [54].
The metaproteomic analyses include four important steps, the process flow of which is given in (Fig. 4): (i) extraction and purification concentration of protein; (ii) denaturation and reduction; (iii) protein separation, digestion, and analysis by MS; and (iv) protein identification basing on spectroscopic data [54]. In metaproteomics it is vital that the sample protein should be a characteristic one in terms of both quality and quality [55]. The first metaproteomic analysis conducted was the AMD biofilm system [56]. Metaproteomic analysis of endophytes has been either done by direct lysis method which involves extraction of total protein of the endosphere (the microenvironment where the plant and endophyte association is established) under different environmental conditions or comparative analysis of their fingerprinting two-dimensional gel electrophoresis to understand the effect of any parameter on secondary metabolite production, etc. [55]. On the other hand, the indirect method of lysis involves extraction of total protein of isolated endophytes subjected to different treatments or stress environments [57]. However, going another step further, similar protein analysis protocol may be followed for host plants with and without the association of endophytes in order to ascertain some particular proteins responsible for bringing in the possible interactions between the host and the endophyte. One such metaproteomic report in sugarcane associated with endophyte Gluconacetobacter reveals 78 differentially expressed proteins using mass spectrometry-based analysis.
Flow chart showing the metaproteomic analysis of endophyte, Source: [65]
The most common methodological bottlenecks in this type of analysis could be the on-site interference of large quantity of secondary metabolites and other cell contents (organic acids, lipids, and polysaccharides) present in the sample tissue. In addition, lack of sufficient amount of information on the microbial community from varied possible ecological niches to characterize these endophytes adds insufficiency of this technique. However, the metaproteomic study needs to be supplemented with its genomic information to make the analysis complete.
3.7 Metaproteogenomic Analysis
It is much well known to the scientific community that not all the genes present in any ecological niche (individual organism or community) are functional at any point of time under a specific environmental condition which makes the analysis of DNA, RNA, or protein (individual organism or community) incomplete in isolation. Metaproteogenomics is a study which deals with the combined exploration of metaproteome as well as metagenome of the same sample linking the genome and proteome of any environmental sample. One outstanding experimental analysis following metaproteogenomic approach has been done in rice where they analyzed the microbial communities of both rhizosphere and phyllosphere and reported that the expression of nifH genes was restricted to rhizosphere only, although present in both [58]. Similarly using metaproteomic approach, a group of workers could mine out certain distinctive traits that were restricted only to phyllospheric bacteria but in the rhizosphere [59]. This approach has the potentiality to correlate the genetic and functional diversity of any community. With time there has been the advent of newer tools and establishment of suitable specific proteogenomic pipelines which urges application of such techniques for more insight study of endosphere and endophytism. The functional proteins are involved in establishing the plant-endophyte interactions, the endophyte protein secretion systems and their identification [60, 61].
3.8 Microarray-Based Analysis
Microarray is basically a laboratory tool where two-dimensional ordered array of microscopic amount of DNA of entire genome of any organism is immobilized onto a solid surface (slide/chip/membrane) so as to measure the simultaneous expressions of all these genes/genetic material or to genotype (polymorphism and mutation) multiple regions of entire genome together. Microarray-based analysis has been attempted to understand the mystery lying behind the endophytism, gene profiling and expression studies of endophytes, unravelling the facts behind the possible interaction between the host plant and the associated endophytes. The advantage of the use of Symbiosis Chip in this technique has been the unique consecrations to study the expressional analysis of both the partners to understand the exchange of signals between them in terms of differential coordinated differential expressions [62]. He specially designed a dual-genome Symbiosis Chip to reveal the physiology behind the nodule development host legume plant Medicago truncatula and the bacterial host Sinorhizobium meliloti using its complete genome. Another advantage of this method is its ability to characterize an unknown species if the genome sequencing of its allied species is done by following genomic interspecies microarray hybridization technique [63]. One such successful accomplishment could be the efficient discovery of genes of unknown endophyte K. pneumoniae 342 by hybridizing its DNA associated with those of Escherichia coli K12. Thus this technique became popular and got very fast applications in endophyte genome analysis. Identification of genes in host plants responsible for initiation of endophyte infestations could be possible in Epichloe-Neotyphodium endophyte [64] and differential regulation of genes in Arabidopsis-Pseudomonas endophyte [25] through microarray studies of induced transcriptional changes. The limitations of this advanced technique are restricted access to the specific gene profiling databases and absence of a specific reference.
4 Conclusion
Profound knowledge of endophytism is inevitable to utilize the enormous potential of endophytes for human welfare in many different valuable means employing multidisciplinary omics science and techniques. This will sure help the better realization of the establishment of such symbiosis between plant and endophyte, tolerance exhibited by endophytes, and their role in growth promotion of host plants. Omics study-based generation of information when supplemented with other disciplinary approaches related to systems biology, several myths behind the total physiological and biochemical processes involved in host-endophyte interaction can be busted and most expectedly predicted models can be established to further expedite the process of understanding. This can ultimately pave a path to sustainable bioprospecting through several biotechnological means.
Abbreviations
- BLAST:
-
Basic local alignment search tool
- BLAT:
-
BLAST-like alignment tool
- Bp:
-
Base pairs
- Brenda:
-
Braunschweig enzyme database
- CAMERA:
-
Community cyberinfrastructure for advanced microbial ecology research and analysis
- COGs:
-
Clusters of orthologous groups
- DGGE:
-
Denaturing gradient gel electrophoresis
- Gbp:
-
Giga base pairs
- ITS:
-
Intertranscribing regions
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- LSU:
-
Large subunit
- LTQ:
-
Linear trap quadrupole
- MALDI:
-
Matrix-assisted laser desorption/ionization
- MALDI ToF:
-
Matrix-assisted laser desorption/ionization time of flight
- Mbp:
-
Mega base pair
- MEGAN:
-
MEtaGenome ANalyzer
- MetAMOS:
-
Open source and modular metagenomic assembly and analysis pipeline
- MG-RAST:
-
Metagenomic rapid annotations using subsystems technology
- MS:
-
Mass spectroscopy
- NCBI:
-
National center for biotechnology information
- NGS:
-
Next-generation sequencing
- NOGs:
-
Non-supervised orthologous groups
- NR:
-
Negative regulatory domain
- Pfam:
-
Protein families
- PICRUSt:
-
Phylogenetic investigation of communities by reconstruction of unobserved states
- PRINTS:
-
Protein fingerprints
- Q-ToF:
-
Quadruple time-of-flight mass spectrometer
- RDP:
-
Ribosomal database project
- SMART:
-
Simple modular architecture research tool
- SRTINGS:
-
Search tool for the retrieval of interacting genes/proteins
- SSU:
-
Small subunit
References
Premjanu N, Jayanthy C (2012) Endophytic fungi a repository of bioactive compounds-a review. Int J Inst Pharm Life Sci 2:135–162
Mousa WK, Raizada MN (2013) The diversity of anti-microbial secondary metabolites produced by fungal endophytes: an interdisciplinary perspective. Front Microbiol. https://doi.org/10.3389/fmicb.2013.00065
Johnson LJ, Johnson RD, Schardl CL, Panaccione DG (2003) Identification of differentially expressed genes in the mutualistic association of tall fescue with Neotyphodium coenophialum. Physiol Mol Plant Pathol 63:305–317. https://doi.org/10.1016/j.pmpp.2004.04.001
Rodriguez RJ, Henson J, Van Volkenburgh E et al (2008) Stress tolerance in plants via habitat-adapted symbiosis. ISME J 2:404–416. https://doi.org/10.1038/ismej.2007.106
Suryanarayanan TS (2013) Endophyte research: going beyond isolation and metabolite documentation. Fungal Ecol 6:561–568. https://doi.org/10.1016/j.funeco.2013.09.007
Deckert RJ, Melville LH, Peterson RL (2001) Structural features of a Lophodermium endophyte during the cryptic life-cycle phase in the foliage of Pinus strobus. Mycol Res 105:991–997. https://doi.org/10.1016/S0953-7562(08)61957-7
Lucero ME, Unc A, Cooke P et al (2011) Endophyte microbiome diversity in micropropagated Atriplex canescens and Atriplex torreyi var griffithsii. PLoS One. https://doi.org/10.1371/journal.pone.0017693
Hallmann J, Berg G, Schulz B (2006) Isolation procedures for endophytic microorganisms. Soil Biol 9:299–319. https://doi.org/10.1007/3-540-33526-9_17
Wang Y, Guo L (2007) A comparative study of endophytic fungi in needles, bark, and xylem of Pinus tabulaeformis. Can J Bot 85:911–917. https://doi.org/10.1139/B07-084
Li W, Guo JZS, Guo L (2007) Endophytic fungi associated with lichens in Baihua mountain of Beijing, China. Fungal Divers 25:69–80
Guo LD, Huang GR, Wang Y (2008) Seasonal and tissue age influences on endophytic fungi of Pinus tabulaeformis (Pinaceae) in the Dongling Mountains, Beijing. J Integr Plant Biol 50:997–1003. https://doi.org/10.1111/j.1744-7909.2008.00394.x
Su YY, Guo LD, Hyde KD (2010) Response of endophytic fungi of Stipa grandis to experimental plant function group removal in Inner Mongolia steppe, China. Fungal Divers 43:93–101. https://doi.org/10.1007/s13225-010-0040-6
Sun X, Guo LD, Hyde KD (2011) Community composition of endophytic fungi in Acer truncatum and their role in decomposition. Fungal Divers 47:85–95. https://doi.org/10.1007/s13225-010-0086-5
Petrini O, Stone J, Carroll FE (1982) Endophytic fungi in evergreen shrubs in Western Oregon: a preliminary study. Can J Bot 60:789–796. https://doi.org/10.1139/b82-102
Rodrigues KF, Samuels GJ (1990) Preliminary study of endophytic fungi in a tropical palm. Mycol Res 94:827–830. https://doi.org/10.1016/S0953-7562(09)81386-5
Guo LD, Hyde KDLE (2000) Identification of endophytic fungi from Livistona chinensis (Palmae) using morphological and molecular techniques. New Phytol 147:617–630
de Souza Vieira PD, de Souza Motta CM, Lima D et al (2011) Endophytic fungi associated with transgenic and non-transgenic cotton. Mycology 2:91–97. https://doi.org/10.1080/21501203.2011.584390
Ding G, Zheng Z, Liu S et al (2009) Photinides A-F, cytotoxic benzofuranone-derived γ-lactones from the plant endophytic fungus Pestalotiopsis photiniae. J Nat Prod 72:942–945. https://doi.org/10.1021/np900084d
Wang Y, Zheng Z, Liu S et al (2010) Oxepinochromenones, furochromenone, and their putative precursors from the endolichenic fungus Coniochaeta sp. J Nat Prod 73:920–924. https://doi.org/10.1021/np100071z
Li J, Li L, Si Y et al (2011) Virgatolides A – C, benzannulated spiroketals from the plant endophytic fungus Pestalotiopsis virgatula. Org Lett 13:2670–2673. https://doi.org/10.1021/ol200770k
Tejesvi MV, Kajula M, Mattila S, Pirttilä AM (2011) Bioactivity and genetic diversity of endophytic fungi in Rhododendron tomentosum Harmaja. Fungal Divers 47:97–107. https://doi.org/10.1007/s13225-010-0087-4
Gamboa MA, Laureano S, Bayman P (2003) Measuring diversity of endophytic fungi in leaf fragments: does size matter? Mycopathologia 156:41–45. https://doi.org/10.1023/A:1021362217723.
Petrini O, Sieber TN, Toti L, Viret O (1993) Ecology, metabolite production, and substrate utilization in endophytic fungi. Nat Toxins 1:185–196. https://doi.org/10.1002/nt.2620010306
Guo L, Hyde KD, Liew E (1998) A method to promote sporulation in palm endophytic fungi. Fungal Divers 1:109–113
Wang Y, Ohara Y, Nakayashiki H et al (2005) Microarray analysis of the gene expression profile induced by the endophytic plant growth-promoting Rhizobacteria, Pseudomonas fluorescens FPT 9601-T 5 in Arabidopsis. Mol Plant-Microbe Interact 18:385–396
Guo LD, Huang GR, Wang Y, He WH, Zheng WH, Hyde KD (2003) Molecular identification of white morphotype strains of endophytic fungi from Pinus tabulaeformis. Mycol Res 107:680–688
González V, Tello ML (2011) The endophytic mycota associated with Vitis vinifera in Central Spain. Fungal Divers 47:29–42. https://doi.org/10.1007/s13225-010-0073-x
Hoff JA, Klopfenstein NB, McDonald GI et al (2004) Fungal endophytes in woody roots of Douglas-fir(Pseudotsuga menziesii) and ponderosa pine (Pinus ponderosa). Forest Pathology 34(4):255–271
Ghimire SR, Charlton ND, Bell JD et al (2011) Biodiversity of fungal endophyte communities inhabiting switchgrass (Panicum virgatum L.) growing in the native tallgrass prairie of northern Oklahoma. Fungal Divers 47:19–27. https://doi.org/10.1007/s13225-010-0085-6
Morakotkarn D, Kawasaki H, Seki T (2007) Molecular diversity of bamboo-associated fungi isolated from Japan. FEMS Microbiol Lett 266:10–19. https://doi.org/10.1111/j.1574-6968.2006.00489.x
Crozier J, Thomas SE, Aime MC et al (2006) Molecular characterization of fungal endophytic morphospecies isolated from stems and pods of Theobroma cacao. Plant Pathol 55:783–791. https://doi.org/10.1111/j.1365-3059.2006.01446.x
Botella L, Javier Diez J (2011) Phylogenic diversity of fungal endophytes in Spanish stands of Pinus halepensis. Fungal Divers 47:9–18. https://doi.org/10.1007/s13225-010-0061-1
Dinsdale EA, Edwards RA, Hall D et al (2008) Functional metagenomic profiling of nine biomes. Nature 452:629–632. https://doi.org/10.1038/nature06810
Liang Y, Guo LD, Ma KP (2005) Population genetic structure of an ectomycorrhizal fungus Amanita manginiana in a subtropical forest over two years. Mycorrhiza 15:137–142
Guo LD, Hyde KD, Liew ECY (2001) Detection and taxonomic placement of endophytic fungi within frond tissues of Livistona chinensis based on rDNA sequences. Mol Phylogenet Evol 20:1–13. https://doi.org/10.1006/mpev.2001.0942
Duong LM, Jeewon R, Lumyong S, Kevin D (2006) DGGE coupled with ribosomal DNA gene phylogenies reveal uncharacterized fungal phylotypes. Fungal Divers 23:121–138
Thomas T, Gilbert J, Meyer F (2012) Metagenomics – a guide from sampling to data analysis. Microb Inform Exp 2:3. https://doi.org/10.1186/2042-5783-2-3
Markowitz VM, Ivanova NN, Szeto E et al (2008) IMG/M: a data management and analysis system for metagenomes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkm869
Meyer F, Paarmann D, D’Souza M et al (2008) The metagenomics RAST server – a public resource for the automatic phylo- genetic and functional analysis of metagenomes. BMC Bioinformatics 9:386. https://doi.org/10.1186/1471-2105-9-386
Blankenberg D, Kuster GV, Coraor N et al (2010) Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol. https://doi.org/10.1002/0471142727.mb1910s89
Treangen TJ, Koren S, Sommer DD et al (2013) MetAMOS: a modular and open source metagenomic assembly and analysis pipeline. Genome Biol. https://doi.org/10.1186/gb-2013-14-1-r2
Langille MG, Zaneveld J, JG C et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821
Aßhauer KP, Wemheuer B, Daniel R, Meinicke P (2015) Tax4Fun: predicting functional profiles from metagenomic 16S rRNA data. Bioinformatics 31:2882–2884. https://doi.org/10.1093/bioinformatics/btv287
Zuccaro A, Lahrmann U, Güldener U et al (2011) Endophytic life strategies decoded by genome and transcriptome analyses of the mutualistic root symbiont Piriformospora indica. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1002290
Frias-Lopez J, Shi Y, Tyson GW et al (2008) Microbial community gene expression in ocean surface waters. Proc Natl Acad Sci USA 105:3805–3810. https://doi.org/10.1073/pnas.0708897105
Sessitsch A, Hardoim P, Döring J et al (2012) Functional characteristics of an endophyte community colonizing Rice roots as revealed by metagenomic analysis. Mol Plant-Microbe Interact 25:28–36. https://doi.org/10.1094/MPMI-08-11-0204
Knapp DG, Németh JB, Barry K et al (2018) Comparative genomics provides insights into the lifestyle and reveals functional heterogeneity of dark septate endophytic fungi. Sci Rep. https://doi.org/10.1038/s41598-018-24686-4
Toju H, Yamamoto S, Sato H, Tanabe AS, Gilbert GS, Kadowaki K (2013) Community composition of root-associated fungi in a Quercus dominated temperate forest:“codominance”of mycorrhizal and root- endophytic fungi. Ecol Evol 3:1281–1293
Jumpponen A, Jones KL, Mattox JD, Yaege C (2010) Massively parallel 454-sequencing of fungal communities in Quercus spp. ectomycorrhizas indicates seasonal dynamics in urban and rural sites. Mol Ecol 19(Suppl 1):41–53. https://doi.org/10.1111/j.1365-294X.2009.04483.x.
Ambrose KV, Belanger FC (2012) SOLiD-SAGE of endophyte-infected red fescue reveals numerous effects on host transcriptome and an abundance of highly expressed fungal secreted proteins. PLoS One. https://doi.org/10.1371/journal.pone.0053214
Camilios-Neto D, Bonato P, Wassem R et al (2014) Dual RNA-seq transcriptional analysis of wheat roots colonized by Azospirillum brasilense reveals up-regulation of nutrient acquisition and cell cycle genes. BMC Genomics. https://doi.org/10.1186/1471-2164-15-378
Molina LG, da Fonseca GC, de Morais GL et al (2012) Metatranscriptomic analysis of small RNAs present in soybean deep sequencing libraries. Genet Mol Biol 35:292–303. https://doi.org/10.1590/S1415-47572012000200010
Gilbert JA, Meyer F, Bailey MJ (2011) The future of microbial metagenomics (or is ignorance bliss). ISME J 5:777–779. https://doi.org/10.1038/ismej.2010.178
Schneider T, Riedel K (2009) Environmental proteomics: analysis of structure and function of microbial communities. Proteomics 10:785–798. https://doi.org/10.1002/pmic.200900450.
Maron PA, Ranjard L, Mougel C, Lemanceau P (2007) Metaproteomics: a new approach for studying functional microbial ecology. Microb Ecol 53:486–493. https://doi.org/10.1007/s00248-006-9196-8
Ram RJ, Verberkmoes NC, Thelen MP et al (2013) Community proteomics of a natural microbial biofilm. Science 308:1915–1920. https://doi.org/10.1126/science
Yadava P, Bhuyan SK, Bandyopadhyay P, Yadava PK (2015) Extraction of proteins for two-dimensional gel electrophoresis and proteomic analysis from an endophytic fungus. Protoc Exch. https://doi.org/10.1038/protex.2015.084
Knief C, Delmotte N, Chaffron S et al (2012) Metaproteogenomic analysis of microbial communities in the phyllosphere and rhizosphere of rice. ISME J 6:1378–1390. https://doi.org/10.1038/ismej.2011.192
Delmotte N, Knief C, Chaffron S et al (2009) Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc Natl Acad Sci 106:16428–16433. https://doi.org/10.1073/pnas.0905240106
Downie JA (2010) The roles of extracellular proteins, polysaccharides and signals in the interactions of rhizobia with legume roots. FEMS Microbiol Rev 34:150–170. https://doi.org/10.1111/j.1574-6976.2009.00205.x
Uszkoreit J, Plohnke N, Rexroth S et al (2014) The bacterial proteogenomic pipeline. BMC Genomics. https://doi.org/10.1186/1471-2164-15-S9-S19
Barnett MJ, Toman CJ, Fisher RF, Long SR (2004) A dual-genome Symbiosis Chip for coordinate study of signal exchange and development in a prokaryote-host interaction. Proc Natl Acad Sci 101:16636–16641. https://doi.org/10.1073/pnas.0407269101
Dong Y, Glasner JD, Blattner FR, Triplett EW (2001) Genomic interspecies microarray hybridization: rapid discovery of three thousand genes in the maize endophyte, Klebsiella pneumoniae 342, by microarray hybridization with Escherichia coli K-12 open reading frames. Appl Environ Microbiol 67:1911–1921. https://doi.org/10.1128/AEM.67.4.1911-1921.2001
Felitti S, Shields K, Ramsperger M et al (2006) Transcriptome analysis of Neotyphodium and Epichloe grass endophytes. Fungal Genet Biol 43:465–475. https://doi.org/10.1016/j.fgb.2006.01.013
Sahoo RK, Gaur M, Subudhi E (2017) Function profiling of microbial community, published in New and Future Development in Microbial Biotechnology and Bioengineering-Microbial genes, Elsevier (In press)
Acknowledgments
We gratefully acknowledge the infrastructure and support provided by Siksha O Anusandhan University, deemed to be university located at Bhubaneswar, for completing this work.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this entry
Cite this entry
Subudhi, E., Sahoo, R.K., Dey, S., Das, A., Sahoo, K. (2019). Unraveling Plant-Endophyte Interactions: An Omics Insight. In: Jha, S. (eds) Endophytes and Secondary Metabolites. Reference Series in Phytochemistry. Springer, Cham. https://doi.org/10.1007/978-3-319-90484-9_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-90484-9_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-90483-2
Online ISBN: 978-3-319-90484-9
eBook Packages: Chemistry and Materials ScienceReference Module Physical and Materials ScienceReference Module Chemistry, Materials and Physics