Abstract
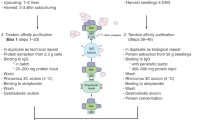
Defining protein complexes is critical to virtually all aspects of cell biology because most cellular processes are regulated by stable or more dynamic protein interactions. Elucidation of the protein–protein interaction network around transcription factors is essential to fully understand their function and regulation. In the last decade, new technologies have emerged to study protein–protein interactions under near-physiological conditions. We have developed a high-throughput tandem affinity purification (TAP)/mass spectrometry (MS) platform for cell suspension cultures to analyze protein complexes in Arabidopsis thaliana. This streamlined platform follows an integrated approach comprising generic Gateway-based vectors with high cloning flexibility, the fast generation of transgenic suspension cultures, TAP adapted for plant cells, and tandem matrix-assisted laser desorption ionization MS for the identification of purified proteins. Recently, we evaluated the GS tag, originally developed to study mammalian protein complexes, that combines two IgG-binding domains of protein G with a streptavidin-binding peptide, separated by two tobacco etch virus cleavage sites. We found that this GS tag outperforms the traditional TAP tag in plant cells, regarding both specificity and complex yield. Here, we provide detailed protocols of the GS-based TAP platform that allowed us to characterize transcription factor complexes involved in signaling in response to the plant phytohormone jasmonate.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Lorenzo, O., Chico, J. M., Sánchez-Serrano, J. J., and Solano, R. (2004) JASMONATE INSENSITIVE1 encodes a MYC transcription factor essential to discriminate between different jasmonate-regulated defense responses in Arabidopsis. Plant Cell 16, 1938–1950.
Katsir, L., Schilmiller, A. L., Staswick, P. E., He, S. Y., and Howe, G. A. (2008) COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc. Natl. Acad. Sci. USA 105, 7100–7105.
Chini, A., Fonseca, S., Fernández, G., Adie, B., Chico, J. M., Lorenzo, O., García-Casado, G., López-Vidriero, I., Lozano, F. M., Ponce, M. R., Micol, J. L., and Solano, R. (2007) The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448, 666–671.
Thines, B., Katsir, L., Melotto, M., Niu, Y., Mandaokar, A., Liu, G., Nomura, K., He, S. Y., Howe, G. A., and Browse, J. (2007) JAZ repressor proteins are targets of the SCFCOI1 complex during jasmonate signalling. Nature 448, 661–665.
Rigaut, G., Shevchenko, A., Rutz, B., Wilm, M., Mann, M., and Séraphin, B. (1999) A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 17, 1030–1032.
Gavin, A.-C., Aloy, P., Grandi, P., Krause, R., Boesche, M., Marzioch, M., Rau, C., Jensen, L. J., Bastuck, S., Dümpelfeld, B., Edelmann, A., Heurtier, M.-A., Hoffman, V., Hoefert, C., Klein, K., Hudak, M., Michon, A.-M., Schelder, M., Schirle, M., Remor, M., Rudi, T., Hooper, S., Bauer, A., Bouwmeester, T., Casari, G., Drewes, G., Neubauer, G., Rick, J. M., Kuster, B., Bork, P., Russell, R. B., and Superti-Furga, G. (2006) Proteome survey reveals modularity of the yeast cell machinery. Nature 440, 631–636.
Gavin, A.-C., Bösche, M., Krause, R., Grandi, P., Marzioch, M., Bauer, A., Schultz, J., Rick, J. M., Michon, A.-M., Cruciat, C.-M., Remor, M., Höfert, C., Schelder, M., Brajenovic, M., Ruffner, H., Merino, A., Klein, K., Hudak, M., Dickson, D., Rudi, T., Gnau, V., Bauch, A., Bastuck, S., Huhse, B., Leutwein, C., Heurtier, M.-A., Copley, R. R., Edelmann, A., Querfurth, E., Rybin, V., Drewes, G., Raida, M., Bouwmeester, T., Bork, P., Seraphin, B., Kuster, B., Neubauer, G., and Superti-Furga, G. (2002) Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147.
Krogan, N. J., Cagney, G., Yu, H., Zhong, G., Guo, X., Ignatchenko, A., Li, J., Pu, S., Datta, N., Tikuisis, A. P., Punna, T., Peregrín-Alvarez, J. M., Shales, M., Zhang, X., Davey, M., Robinson, M. D., Paccanaro, A., Bray, J. E., Sheung, A., Beattie, B., Richards, D. P., Canadien, V., Lalev, A., Mena, F., Wong, P., Starostine, A., Canete, M. M., Vlasblom, J., Wu, S., Orsi, C., Collins, S. R., Chandran, S., Haw, R., Rilstone, J. J., Gandi, K., Thompson, N. J., Musso, G., St Onge, P., Ghanny, S., Lam, M. H. Y., Butland, G., Altaf-Ul, A. M., Kanaya, S., Shilatifard, A., O’Shea, E., Weissman, J. S., Ingles, C. J., Hughes, T. R., Parkinson, J., Gerstein, M., Wodak, S. J., Emili, A., and Greenblatt, J. F. (2006) Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643.
Van Leene, J., Stals, H., Eeckhout, D., Persiau, G., Van De Slijke, E., Van Isterdael, G., De Clercq, A., Bonnet, E., Laukens, K., Remmerie, N., Henderickx, K., De Vijlder, T., Abdelkrim, A., Pharazyn, A., Van Onckelen, H., Inzé, D., Witters, E., and De Jaeger, G. (2007) A tandem affinity purification-based technology platform to study the cell cycle interactome in Arabidopsis thaliana. Mol. Cell. Proteomics 6, 1226–1238.
Bürckstümmer, T., Bennett, K. L., Preradovic, A., Schütze, G., Hantschel, O., Superti-Furga, G., and Bauch, A. (2006) An efficient tandem affinity purification procedure for interaction proteomics in mammalian cells. Nat. Methods 3, 1013–1019.
Van Leene, J., Witters, E., Inzé, D., and De Jaeger, G. (2008) Boosting tandem affinity purification of plant protein complexes. Trends Plant Sci. 13, 517–520.
Grunewald, W., Vanholme, B., Pauwels, L., Plovie, E., Inzé, D., Gheysen, G., and Goossens, A. (2009) Expression of the Arabidopsis jasmonate signalling repressor JAZ1/TIFY10A is stimulated by auxin. EMBO Rep. 10, 923–928.
Pauwels, L., Fernández Barbero, G., Geerinck, J., Tilleman, S., Grunewald, W., Cuéllar Pérez, A., Chico, J. M., Vanden Bossche, R., Sewell, J., Gil, E., García-Casado, G., Witters, E., Inzé, D., Long, J. A., De Jaeger, G., Solano, R., and Goossens, A. (2010) NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464, 788–791.
Chini, A., Fonseca, S., Chico, J. M., Fernández-Calvo, P., and Solano, R. (2009) The ZIM domain mediates homo- and heteromeric interactions between Arabidopsis JAZ proteins. Plant J. 59, 77–87.
Menges, M., and Murray, J. A. H. (2002) Synchronous Arabidopsis suspension cultures for analysis of cell-cycle gene activity. Plant J. 30, 203–212.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2011 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Van Leene, J. et al. (2011). Isolation of Transcription Factor Complexes from Arabidopsis Cell Suspension Cultures by Tandem Affinity Purification. In: Yuan, L., Perry, S. (eds) Plant Transcription Factors. Methods in Molecular Biology, vol 754. Humana Press. https://doi.org/10.1007/978-1-61779-154-3_11
Download citation
DOI: https://doi.org/10.1007/978-1-61779-154-3_11
Published:
Publisher Name: Humana Press
Print ISBN: 978-1-61779-153-6
Online ISBN: 978-1-61779-154-3
eBook Packages: Springer Protocols