Abstract
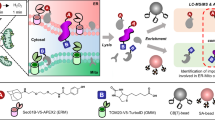
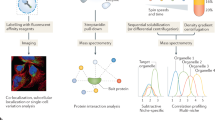
The ability to purify cell organelles and protein complexes on a large scale, combined with advances in protein identification using mass spectrometry, has provided a wealth of information regarding protein localization and function. A major challenge in these studies has been the ability to identify bona fide organelle components from a background of co-purifying contaminants because none of the available biochemical purification protocols afford pure preparations. Since this situation is unlikely to change alternative strategies have been devised to meet this challenge by making use of the information inherent in the fractionation profile of organelles isolated by density gradient centrifugation. In this chapter we describe strategies based on protein correlation profiling and quantitative mass spectrometry to sort out likely candidates. The organelle inventories defined by these methods are suitable to guide future functional experiments.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Aebersold, R., and Mann, M. (2003) Mass spectrometry-based proteomics. Nature 422, 198–207.
Yates, J. R., 3rd, Gilchrist, A., Howell, K. E., and Bergeron, J. J. (2005) Proteomics of organelles and large cellular structures. Nat Rev Mol Cell Biol 6, 702–714.
Andersen, J. S., and Mann, M. (2006) Organellar proteomics: turning inventories into insights. EMBO Rep 7, 874–879.
Yan, W., Aebersold, R., and Raines, E. W. (2008) Evolution of organelle-associated protein profiling. J Proteomics 15, 4–11.
Sadowski, P. G., Dunkley, T. P., Shadforth, I. P., Dupree, P., Bessant, C., Griffin, J. L., and Lilley, K. S. (2006) Quantitative proteomic approach to study subcellular localization of membrane proteins. Nat. Protoc. 1, 1778–1789.
Andersen, J. S., Wilkinson, C. J., Mayor, T., Mortensen, P., Nigg, E. A., and Mann, M. (2003) Proteomic characterization of the human centrosome by protein correlation profiling. Nature 426, 570–574.
Gilchrist, A., Au, C. E., Hiding, J., Bell, A. W., Fernandez-Rodriguez, J., Lesimple, S., Nagaya, H., Roy, L., Gosline, S. J., Hallett, M., Paiement, J., Kearney, R. E., Nilsson, T., and Bergeron, J. J. (2006) Quantitative proteomics analysis of the secretory pathway. Cell 127, 1265–1281.
Dunkley, T. P., Hester, S., Shadforth, I. P., Runions, J., Weimar, T., Hanton, S. L., Griffin, J. L., Bessant, C., Brandizzi, F., Hawes, C., Watson, R. B., Dupree, P., and Lilley, K. S. (2006) Mapping the Arabidopsis organelle proteome. Proc. Natl. Acad. Sci. USA 103, 6518–6523.
Dunkley, T. P., Watson, R., Griffin, J. L., Dupree, P., and Lilley, K. S. (2004) Localization of organelle proteins by isotope tagging (LOPIT). Mol. Cell Proteomics 3, 1128–1134.
Ong, S. E., Blagoev, B., Kratchmarova, I., Kristensen, D. B., Steen, H., Pandey, A., and Mann, M. (2002) Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell Proteomics 1, 376–386.
Foster, L. J., de Hoog, C. L., Zhang, Y., Zhang, Y., Xie, X., Mootha, V. K., and Mann, M. (2006) A mammalian organelle map by protein correlation profiling. Cell 125, 187–199.
Yan, W., Hwang, D., and Aebersold, R. (2008) Quantitative proteomic analysis to profile dynamic changes in the spatial distribution of cellular proteins. Methods Mol. Biol. 432, 389–401.
Blagoev, B., and Mann, M. (2006) Quantitative proteomics to study mitogen-activated protein kinases. Methods 40, 243–250.
Ong, S. E., and Mann, M. (2007) Stable isotope labeling by amino acids in cell culture for quantitative proteomics. Methods Mol. Biol. 359, 37–52.
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V., and Mann, M. (2006) In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860.
Rappsilber, J., Mann, M., and Ishihama, Y. (2007) Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906.
Cox, J., and Mann, M. (2008) MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372.
Acknowledgments
The research leading to these results has received funding from the European Commission’s 7th Framework Programme and grant agreements HEALTH-F4-2008-201648/PROSPECTS and HEALTH-F4-2007-200767/APO-SYS. JD was supported by the European Molecular Biology Organization. We thank all CEBI group members for helpful discussions and support.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2010 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Dengjel, J., Jakobsen, L., Andersen, J.S. (2010). Organelle Proteomics by Label-Free and SILAC-Based Protein Correlation Profiling. In: Cutillas, P., Timms, J. (eds) LC-MS/MS in Proteomics. Methods in Molecular Biology, vol 658. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-60761-780-8_15
Download citation
DOI: https://doi.org/10.1007/978-1-60761-780-8_15
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-60761-779-2
Online ISBN: 978-1-60761-780-8
eBook Packages: Springer Protocols