Abstract
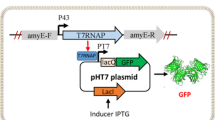
Bacterial proteases are important enzymes used in several technical applications where controlled cleavage of proteins is needed. They are challenging enzymes to express recombinantly as parts of the proteome can be hydrolyzed by their activity. The eukaryotic model organism Saccharomyces cerevisiae is potentially a good expression host as it tolerates several stress conditions and is known to better express insoluble proteins compared to bacterial systems. In this chapter we describe how the protease IdeS from Streptococcus pyogenes can be expressed in S. cerevisiae. The expression of IdeS was followed by constructing a fused protein with GFP and measuring the fluorescence with flow cytometry. The protease presence was confirmed with a Western blot assay and activity was measured with an in vitro assay. To reduce potentially toxic effect on the host cell, the growth and production phases were separated by using the inducible promoter GAL1p to control recombinant gene expression. The protocol provided may be adopted for other bacterial proteases through minor modifications of the fused protein.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Ward OP (2011) 3.49 – proteases. In: Moo-Young M (ed) Comprehensive biotechnology, 2nd edn. Academic, Burlington, pp 571–582
Nelson AD et al (2012) IgG Fab fragments forming bivalent complexes by a conformational mechanism that is reversible by osmolytes. J Biol Chem 287(51):42936–42950
Komai T et al (1997) Development of HIV-1 protease expression methods using the T7 phage promoter system. Appl Microbiol Biotechnol 47(3):241–245
Kwon K et al (2011) Recombinant expression and functional analysis of proteases from Streptococcus pneumoniae, Bacillus anthracis, and Yersinia pestis. BMC Biochem 12:17
Xie Y, Han X, Miao Y (2018) An effective recombinant protein expression and purification system in Saccharomyces cerevisiae. Curr Protoc Mol Biol 123(1):e62
Liu ZH et al (2012) Different expression systems for production of recombinant proteins in Saccharomyces cerevisiae. Biotechnol Bioeng 109(5):1259–1268
Borodina I, Nielsen J (2014) Advances in metabolic engineering of yeast Saccharomyces cerevisiae for production of chemicals. Biotechnol J 9(5):609–620
Hahn-Hägerdal B et al (2007) Towards industrial pentose-fermenting yeast strains. Appl Microbiol Biotechnol 74(5):937–953
Fraczek MG, Naseeb S, Delneri D (2018) History of genome editing in yeast. Yeast 35(5):361–368
Alexander WG (2018) A history of genome editing in Saccharomyces cerevisiae. Yeast 35(5):355–360
Jessop-Fabre MM et al (2016) EasyClone-MarkerFree: a vector toolkit for marker-less integration of genes into Saccharomyces cerevisiae via CRISPR-Cas9. Biotechnol J 11(8):1110–1117
Mikkelsen MD et al (2012) Microbial production of indolylglucosinolate through engineering of a multi-gene pathway in a versatile yeast expression platform. Metab Eng 14(2):104–111
Li J et al (2000) Green fluorescent protein in Saccharomyces cerevisiae: real-time studies of the GAL1 promoter. Biotechnol Bioeng 70(2):187–196
von Pawel-Rammingen U, Johansson BP, Bjorck L (2002) IdeS, a novel streptococcal cysteine proteinase with unique specificity for immunoglobulin G. EMBO J 21(7):1607–1615
Johansson BP, Shannon O, Björck L (2008) IdeS: a bacterial proteolytic enzyme with therapeutic potential. PLoS One 3(2):e1692
Collin M, Olsén A (2003) Extracellular enzymes with immunomodulating activities: variations on a theme in Streptococcus pyogenes. Infect Immun 71(6):2983–2992
Vincents B et al (2004) Enzymatic characterization of the streptococcal endopeptidase, IdeS, reveals that it is a cysteine protease with strict specificity for IgG cleavage due to exosite binding. Biochemistry 43(49):15540–15549
Heins A-L et al (2019) Quantitative flow cytometry to understand population heterogeneity in response to changes in substrate availability in Escherichia coli and Saccharomyces cerevisiae chemostats. Front Bioeng Biotechnol 7:187
Davey HM, Hexley P (2011) Red but not dead? Membranes of stressed Saccharomyces cerevisiae are permeable to propidium iodide. Environ Microbiol 13(1):163–171
Gietz RD, Schiestl RH (2007) High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat Protoc 2(1):31–34
Grote A et al (2005) JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res 33(Web Server issue):W526–W531
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Lindh, T., Collin, M., Lood, R., Carlquist, M. (2023). Expression of the Bacterial Enzyme IdeS Using a GFP Fusion in the Yeast Saccharomyces cerevisiae. In: Nordenfelt, P., Collin, M. (eds) Bacterial Pathogenesis. Methods in Molecular Biology, vol 2674. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3243-7_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3243-7_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3242-0
Online ISBN: 978-1-0716-3243-7
eBook Packages: Springer Protocols