Abstract
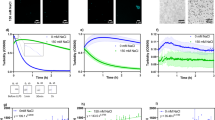
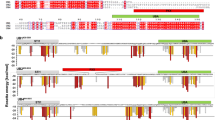
Liquid–liquid phase separation (LLPS) is hypothesized to be the underlying mechanism for how membraneless organelles or biomolecular condensates form inside both prokaryotic and eukaryotic cells. Protein LLPS is a biophysical process during which proteins demix from homogeneous solution to form protein-dense droplets with liquid-like properties. Disruptions to LLPS, such as changes to material properties of condensates or physicochemical parameters for LLPS onset, are implicated in neurodegenerative diseases and cancer. Therefore, it is essential to determine the physicochemical parameters that promote protein LLPS. Here, we present our UV-Vis spectrophotometric turbidity assay to characterize the temperature and concentration dependence of LLPS for UBQLN2, a protein that undergoes LLPS via homotypic interactions in vitro and forms stress-induced condensates in cells. Mutations in UBQLN2 cause amyotrophic lateral sclerosis (ALS) and disrupt UBQLN2 LLPS. We present a detailed expression and purification protocol for a C-terminal construct of UBQLN2 and how we use microscopy to image UBQLN2 LLPS. We use our UV-Vis assay to construct temperature–concentration phase diagrams for wild-type and mutant UBQLN2 constructs to determine the effects of domain deletions and/or mutations on UBQLN2 phase separation.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Brangwynne CP, Eckmann CR, Courson DS et al (2009) Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324:1729–1732
Brangwynne CP, Mitchison TJ, Hyman AA (2011) Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc Natl Acad Sci U S A 108:4334–4339
Wippich F, Bodenmiller B, Trajkovska MG et al (2013) Dual specificity kinase DYRK3 couples stress granule condensation/dissolution to mTORC1 signaling. Cell 152:791–805
Mitrea DM, Cika JA, Guy CS et al (2016) Nucleophosmin integrates within the nucleolus via multi-modal interactions with proteins displaying R-rich linear motifs and rRNA. elife 5:e13571
Conicella AE, Zerze GH, Mittal J et al (2016) ALS mutations disrupt phase separation mediated by α-helical structure in the TDP-43 low-complexity C-terminal domain. Structure 24:1537–1549
Alberti S, Dormann D (2019) Liquid–liquid phase separation in disease. Annu Rev Genet 53:171–194
Kato M, Han TW, Xie S et al (2012) Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149:753–767
Bouchard JJ, Otero JH, Scott DC et al (2018) Cancer mutations of the tumor suppressor SPOP disrupt the formation of active, phase-separated compartments. Mol Cell 72:19–36.e8
Ambadipudi S, Biernat J, Riedel D et al (2017) Liquid–liquid phase separation of the microtubule-binding repeats of the Alzheimer-related protein tau. Nat Commun 8:275
Murakami T, Qamar S, Lin JQ et al (2015) ALS/FTD mutation-induced phase transition of FUS liquid droplets and reversible hydrogels into irreversible hydrogels impairs RNP granule function. Neuron 88:678–690
Banani SF, Rice AM, Peeples WB et al (2016) Compositional control of phase-separated cellular bodies. Cell 166:651–663
Li P, Banjade S, Cheng H-C et al (2012) Phase transitions in the assembly of multivalent signalling proteins. Nature 483:336–340
Jonas S, Izaurralde E (2013) The role of disordered protein regions in the assembly of decapping complexes and RNP granules. Genes Dev 27:2628–2641
Martin EW, Holehouse AS, Peran I et al (2020) Valence and patterning of aromatic residues determine the phase behavior of prion-like domains. Science 367:694–699
Dignon GL, Best RB, Mittal J (2020) Biomolecular phase separation: from molecular driving forces to macroscopic properties. Annu Rev Phys Chem 71:53–75
Posey AE, Holehouse AS, Pappu RV (2018) Phase separation of intrinsically disordered proteins. In: Methods in enzymology. Academic Press, 611:1–30
Bracha D, Walls MT, Wei M-T et al (2018) Mapping local and global liquid phase behavior in living cells using photo-oligomerizable seeds. Cell 175:1467–1480.e13
Riback JA, Zhu L, Ferrolino MC et al (2020) Composition-dependent thermodynamics of intracellular phase separation. Nature 581:209–214
Peran I, Martin EW, Mittag T (2020) Walking along a protein phase diagram to determine coexistence points by static light scattering. In: Kragelund BB, Skriver K (eds) Intrinsically disordered proteins: methods and protocols. Springer US, New York, pp 715–730
Milkovic NM, Mittag T (2020) Determination of protein phase diagrams by centrifugation. In: Kragelund BB, Skriver K (eds) Intrinsically disordered proteins: methods and protocols. Springer, US, New York, NY, pp 685–702
Holland J, Crabtree MD, Nott TJ (2020) In vitro transition temperature measurement of phase-separating proteins by microscopy. In: Kragelund BB, Skriver K (eds) Intrinsically disordered proteins: methods and protocols. Springer US, New York, pp 703–714
Riback JA, Katanski CD, Kear-Scott JL et al (2017) Stress-triggered phase separation is an adaptive, evolutionarily tuned response. Cell 168:1028–1040.e19
Dao TP, Kolaitis R-M, Kim HJ et al (2018) Ubiquitin modulates liquid-liquid phase separation of UBQLN2 via disruption of multivalent interactions. Mol Cell 69:965–978.e6
Martin EW, Mittag T (2018) Relationship of sequence and phase separation in protein low-complexity regions. Biochemistry 57:2478–2487
Molliex A, Temirov J, Lee J et al (2015) Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163:123–133
Ruff KM, Roberts S, Chilkoti A et al (2018) Advances in understanding stimulus-responsive phase behavior of intrinsically disordered protein polymers. J Mol Biol 430:4619–4635
Yang Y, Jones HB, Dao TP et al (2019) Single amino acid substitutions in stickers, but not spacers, substantially alter UBQLN2 phase transitions and dense phase material properties. J Phys Chem B 123:3618–3629
Renaud L, Picher-Martel V, Codron P et al (2019) Key role of UBQLN2 in pathogenesis of amyotrophic lateral sclerosis and frontotemporal dementia. Acta Neuropathol Commun 7:103
Zheng T, Yang Y, Castañeda CA (2020) Structure, dynamics and functions of UBQLNs: at the crossroads of protein quality control machinery. Biochem J 477:3471–3497
Kleijnen MF, Shih AH, Zhou P et al (2000) The hPLIC proteins may provide a link between the ubiquitination machinery and the proteasome. Mol Cell 6:409–419
Deng H-X, Chen W, Hong S-T et al (2011) Mutations in UBQLN2 cause dominant X-linked juvenile and adult onset ALS and ALS/dementia. Nature 477:211–215
Synofzik M, Maetzler W, Grehl T et al (2012) Screening in ALS and FTD patients reveals 3 novel UBQLN2 mutations outside the PXX domain and a pure FTD phenotype. Neurobiol Aging 33:2949.e13–2949.e17
Dao TP, Martyniak B, Canning AJ et al (2019) ALS-linked mutations affect UBQLN2 oligomerization and phase separation in a position- and amino acid-dependent manner. Structure 27:937–951
Acknowledgments
This material is based on work supported by NSF MCB grant 1750462 to C.A.C.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Raymond-Smiedy, P., Bucknor, B., Yang, Y., Zheng, T., Castañeda, C.A. (2023). A Spectrophotometric Turbidity Assay to Study Liquid-Liquid Phase Separation of UBQLN2 In Vitro. In: Cieplak, A.S. (eds) Protein Aggregation. Methods in Molecular Biology, vol 2551. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2597-2_32
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2597-2_32
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2596-5
Online ISBN: 978-1-0716-2597-2
eBook Packages: Springer Protocols