Abstract
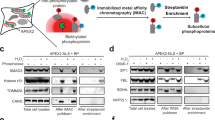
Enzyme clustering is a phenomenon that involves partitioning of proteins that function together in a common subcellular or sub-organellar compartment. Traditional genetic, biochemical, and biophysical approaches for studying protein–protein interactions in complexes with defined stoichiometry yield inconclusive results when applied to clustered proteins. This chapter describes a combination of approaches to study clustered proteins including co-immunoprecipitation, biochemical co-localization in purified mitochondria, and super resolution imaging of endogenous proteins in situ. These approaches can be used to study interactions among proteins that form clusters. We will illustrate this approach by using the urea cycle enzymes that localize in the mitochondrial matrix, and form clusters at the inner mitochondrial membrane.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Battesti A, Bouveret E (2012) The bacterial two-hybrid system based on adenylate cyclase reconstitution in Escherichia coli. Methods 58:325–334
Young KH (1998) Yeast two-hybrid: so many interactions, (in) so little time. Biol Reprod 58:302–331
Rich RL, Myszka DG (2007) Higher-throughput, label-free, real-time molecular interaction analysis. Anal Biochem 361:1–6
Attri AK, Minton AP (2005) Composition gradient static light scattering: a new technique for rapid detection and quantitative characterization of reversible macromolecular hetero-associations in solution. Anal Biochem 346:132–138
Martin SF, Tatham MH, Hay RT, Samuel ID (2008) Quantitative analysis of multi-protein interactions using FRET: application to the SUMO pathway. Protein Sci 17:777–784
Matsumoto S, Hammes GG (1975) Fluorescence energy transfer between ligand binding sites on aspartate transcarbamylase. Biochemistry 14(2):214–224
Pollok BA, Heim R (1999) Using GFP in FRET-based applications. Trends Cell Biol 9:57–60
Sekar RB, Periasamy A (2003) Fluorescence resonance energy transfer (FRET) microscopy imaging of live cell protein localizations. J Cell Biol 160(5):629–633
Vinogradova O, Qin J (2012) NMR as a unique tool in assessment and complex determination of weak protein-protein interactions. Top Curr Chem 326:35–45
Englander SW (2006) Hydrogen exchange and mass spectrometry: a historical perspective. J Am Soc Mass Spectrom 17:1481–1489
Keilhauer EC, Hein MY, Mann M (2015) Accurate protein complex retrieval by affinity enrichment mass spectrometry (AE-MS) rather than affinity purification mass spectrometry (AP-MS). Mol Cell Proteomics 14:120–135
Morris JH, Knudsen GM, Verschueren E, Johnson JR, Cimermancic P, Greninger AL, Pico AR (2014) Affinity purification-mass spectrometry and network analysis to understand protein-protein interactions. Nat Protoc 9:2539–2554
Rigaut G, Shevchenko A, Rutz B, Wilm M, Mann M, Seraphin B (1999) A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol 17:1030–1032
Herzog F, Kahraman A, Boehringer D, Mak R, Bracher A, Walzthoeni T, Leitner A, Beck M, Hartl FU, Ban N, Malmstrom L, Aebersold R (2012) Structural probing of a protein phosphatase 2A network by chemical cross-linking and mass spectrometry. Science 337:1348–1352
Wu F, Minteer S (2015) Krebs cycle metabolon: structural evidence of substrate channeling revealed by cross-linking and mass spectrometry. Angew Chem Int Ed Engl 54:1851–1854
Rhee HW, Zou P, Udeshi ND, Martell JD, Mootha VK, Carr SA, Ting AY (2013) Proteomic mapping of mitochondria in living cells via spatially restricted enzymatic tagging. Science 339:1328–1331
Floyd BJ, Wilkerson EM, Veling MT, Minogue CE, Xia C, Beebe ET, Wrobel RL, Cho H, Kremer LS, Alston CL, Gromek KA, Dolan BK, Ulbrich A, Stefely JA, Bohl SL, Werner KM, Jochem A, Westphall MS, Rensvold JW, Taylor RW, Prokisch H, Kim JP, Coon JJ, Pagliarini DJ (2016) Mitochondrial protein interaction mapping identifies regulators of respiratory chain function. Mol Cell 63:621–632
Huttlin EL, Bruckner RJ, Paulo JA, Cannon JR, Ting L, Baltier K, Colby G, Gebreab F, Gygi MP, Parzen H, Szpyt J, Tam S, Zarraga G, Pontano-Vaites L, Swarup S, White AE, Schweppe DK, Rad R, Erickson BK, Obar RA, Guruharsha KG, Li K, Artavanis-Tsakonas S, Gygi SP, Harper JW (2017) Architecture of the human interactome defines protein communities and disease networks. Nature 545:505–509
Schweppe DK, Huttlin EL, Harper JW, Gygi SP (2018) BioPlex display: an interactive suite for large-scale AP-MS protein-protein interaction data. J Proteome Res 17:722–726
Jie J, Lohr F, Barbar E (2015) Interactions of yeast dynein with dynein light chain and dynactin: general implications for intrinsically disordered duplex scaffolds in multiprotein assemblies. J Biol Chem 290:23863–23874
Nyarko A, Song Y, Barbar E (2012) Intrinsic disorder in dynein intermediate chain modulates its interactions with NudE and dynactin. J Biol Chem 287:24884–24893
Nyarko A, Song Y, Novacek J, Zidek L, Barbar E (2013) Multiple recognition motifs in nucleoporin Nup159 provide a stable and rigid Nup159-Dyn2 assembly. J Biol Chem 288:2614–2622
Velot C, Mixon MB, Teige M, Srere PA (1997) Model of a quinary structure between Krebs TCA cycle enzymes: a model for the metabolon. Biochemistry 36:14271–14276
Bulutoglu B, Garcia KE, Wu F, Minteer SD, Banta S (2016) Direct evidence for metabolon formation and substrate channeling in recombinant TCA cycle enzymes. ACS Chem Biol 11:2847–2853
Castellana M, Wilson MZ, Xu Y, Joshi P, Cristea IM, Rabinowitz JD, Gitai Z, Wingreen NS (2014) Enzyme clustering accelerates processing of intermediates through metabolic channeling. Nat Biotechnol 32:1011–1018
De La Fuente IM, Martinez L, Perez-Samartin AL, Ormaetxea L, Amezaga C, Vera-Lopez A (2008) Global self-organization of the cellular metabolic structure. PLoS One 3:e3100
Brusilow SW, Horwich AL (2001) Urea cycle enzymes. In: Scriver CR, Beaudet AL, Sly WS, Valle D (eds) The metabolic & molecular bases of inherited disease, vol 2. McGraw-Hill, New York, NY, pp 1909–1963
Grisolia S, Cohen PP (1952) The catalytic role of carbamyl glutamate in citrulline biosynthesis. J Biol Chem 198:561–571
Grisolia S, Cohen PP (1953) Catalytic role of glutamate derivatives in citrulline biosynthesis. J Biol Chem 204:753–757
Bradford NM, McGivan JD (1980) Evidence for the existence of an ornithine/citrulline antiporter in rat liver mitochondria. FEBS Lett 113:294–298
Kobayashi K, Sinasac DS, Iijima M, Boright AP, Begum L, Lee JR, Yasuda T, Ikeda S, Hirano R, Terazono H, Crackower MA, Kondo I, Tsui LC, Scherer SW, Saheki T (1999) The gene mutated in adult-onset type II citrullinaemia encodes a putative mitochondrial carrier protein. Nat Genet 22:159–163
Cheung CW, Cohen NS, Raijman L (1989) Channeling of urea cycle intermediates in situ in permeabilized hepatocytes. J Biol Chem 264:4038–4044
Cohen NS, Cheung CW, Sijuwade E, Raijman L (1992) Kinetic properties of carbamoyl-phosphate synthase (ammonia) and ornithine carbamoyltransferase in permeabilized mitochondria. Biochem J 282:173–180
Tuchman M (1989) Persistent acitrullinemia after liver transplantation for carbamylphosphate synthetase deficiency. N Engl J Med 320:1498–1499
Cohen NS, Cheung CW, Kyan FS, Jones EE, Raijman L (1982) Mitochondrial carbamyl phosphate and citrulline synthesis at high matrix acetylglutamate. J Biol Chem 257:6898–6907
Raijman L (1976) Enzyme and reactant concentrations and the regulation of urea synthesis. In: Grisolia S, Baguena R, Mayor F (eds) The urea cycle. John Wiley & Sons, New York, pp 243–259
Sonoda T, Tatibana M (1983) Purification of N-acetyl-L-glutamate synthetase from rat liver mitochondria and substrate and activator specificity of the enzyme. J Biol Chem 258:9839–9844
Wang Y, Palmfeldt J, Gregersen N, Makhov AM, Conway JF, Wang M, McCalley SP, Basu S, Alharbi H, St Croix C, Calderon MJ, Watkins S, Vockley J (2019) Mitochondrial fatty acid oxidation and the electron transport chain comprise a multifunctional mitochondrial protein complex. J Biol Chem 294:12380–12391
An S, Kumar R, Sheets ED, Benkovic SJ (2008) Reversible compartmentalization of de novo purine biosynthetic complexes in living cells. Science 320(5872):103–106
Chan CY, Zhao H, Pugh RJ, Pedley AM, French J, Jones SA, Zhuang X, Jinnah H, Huang TJ, Benkovic SJ (2015) Purinosome formation as a function of the cell cycle. Proc Natl Acad Sci U S A 112:1368–1373
Lapuente-Brun E, Moreno-Loshuertos R, Acin-Perez R, Latorre-Pellicer A, Colas C, Balsa E, Perales-Clemente E, Quiros PM, Calvo E, Rodriguez-Hernandez MA, Navas P, Cruz R, Carracedo A, Lopez-Otin C, Perez-Martos A, Fernandez-Silva P, Fernandez-Vizarra E, Enriquez JA (2013) Supercomplex assembly determines electron flux in the mitochondrial electron transport chain. Science 340:1567–1570
Subramanian K, Jochem A, Le Vasseur M, Lewis S, Paulson BR, Reddy TR, Russell JD, Coon JJ, Pagliarini DJ, Nunnari J (2019) Coenzyme Q biosynthetic proteins assemble in a substrate-dependent manner into domains at ER-mitochondria contacts. J Cell Biol 218:1353–1369
Haskins N, Bhuvanendran S, Anselmi C, Gams A, Kanholm T, Kocher KM, LoTempio J, Krohmaly KI, Sohai D, Stearrett N, Bonner E, Tuchman M, Morizono H, Jaiswal JK, Caldovic L (2020) Mitochondrial enzymes of the urea cycle cluster at the inner mitochondrial membrane. Front Physiol 11:542950
Makris G, Lauber M, Rufenacht V, Gemperle C, Diez-Fernandez C, Caldovic L, Froese DS, Haberle J (2021) Clinical and structural insights into potential dominant negative triggers of proximal urea cycle disorders. Biochimie 183:89–99
Powers-Lee SG, Mastico RA, Bendayan M (1987) The interaction of rat liver carbamoyl phosphate synthetase and ornithine transcarbamoylase with inner mitochondrial membranes. J Biol Chem 262:15683–15688
Thul PJ, Akesson L, Wiking M, Mahdessian D, Geladaki A, Ait Blal H, Alm T, Asplund A, Bjork L, Breckels LM, Backstrom A, Danielsson F, Fagerberg L, Fall J, Gatto L, Gnann C, Hober S, Hjelmare M, Johansson F, Lee S, Lindskog C, Mulder J, Mulvey CM, Nilsson P, Oksvold P, Rockberg J, Schutten R, Schwenk JM, Sivertsson A, Sjostedt E, Skogs M, Stadler C, Sullivan DP, Tegel H, Winsnes C, Zhang C, Zwahlen M, Mardinoglu A, Ponten F, von Feilitzen K, Lilley KS, Uhlen M, Lundberg E (2017) A subcellular map of the human proteome. Science 356:aal3321
Uhlen M, Fagerberg L, Hallstrom BM, Lindskog C, Oksvold P, Mardinoglu A, Sivertsson A, Kampf C, Sjostedt E, Asplund A, Olsson I, Edlund K, Lundberg E, Navani S, Szigyarto CA, Odeberg J, Djureinovic D, Takanen JO, Hober S, Alm T, Edqvist PH, Berling H, Tegel H, Mulder J, Rockberg J, Nilsson P, Schwenk JM, Hamsten M, von Feilitzen K, Forsberg M, Persson L, Johansson F, Zwahlen M, von Heijne G, Nielsen J, Ponten F (2015) Proteomics. Tissue-based map of the human proteome. Science 347:1260419
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Caldovic, L., Bhuvanendran, S., Jaiswal, J. (2022). Assessing Protein Interactions for Clustering of Mitochondrial Urea Cycle Enzymes. In: Stamatis, H. (eds) Multienzymatic Assemblies. Methods in Molecular Biology, vol 2487. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2269-8_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2269-8_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2268-1
Online ISBN: 978-1-0716-2269-8
eBook Packages: Springer Protocols