Abstract
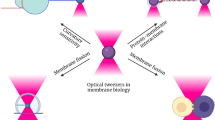
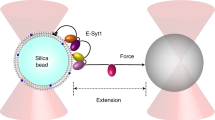
Numerous proteins directly or indirectly bind membranes to exert their roles in a wide variety of biological processes. Such membrane binding often occurs in the presence of an external mechanical force. It remains challenging to quantify these interactions using traditional experimental approaches based on a large number of molecules, due to ensemble averaging or the lack of mechanical force. Here we described a new single-molecule approach based on high-resolution optical tweezers to characterize protein–membrane interactions. A single membrane binding protein is attached to the lipid bilayer coated on a silica bead via a flexible polypeptide linker, tethered to another bead via a long DNA handle, and pulled away from the bilayer using optical tweezers. Dynamic protein binding and unbinding is detected by the corresponding changes in the extension of the protein-DNA tether with high spatiotemporal resolution, which reveals the membrane binding affinity, kinetics, and intermediates. We demonstrated the method using C2 domains of extended synaptotagmin 2 (E-Syt2) with a detailed protocol. The method can be widely applied to investigate complex protein–membrane interactions under well-controlled experimental conditions.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Das DK, Feng Y, Mallis RJ, Li X, Keskin DB, Hussey RE, Brady SK, Wang JH, Wagner G, Reinherz EL, Lang MJ (2015) Force-dependent transition in the T-cell receptor beta-subunit allosterically regulates peptide discrimination and pMHC bond lifetime. Proc Natl Acad Sci U S A 112:1517–1522
Cho WH, Stahelin RV (2005) Membrane-protein interactions in cell signaling and membrane trafficking. Annu Rev Biophys Biomol Struct 34:119–151
Wang XF, Ha T (2013) Defining single molecular forces required to activate integrin and notch signaling. Science 340:991–994
Brunger AT, Choi UB, Lai Y, Leitz J, White KI, Zhou Q (2019) The pre-synaptic fusion machinery. Curr Opin Struct Biol 54:179–188
Brose N, Brunger A, Cafiso D, Chapman ER, Diao J, Hughson FM, Jackson MB, Jahn R, Lindau M, Ma C, Rizo J, Shin YK, Sollner TH, Tamm L, Yoon TY, Zhang YL (2019) Synaptic vesicle fusion: today and beyond. Nat Struct Mol Biol 26:663–668
Baker RW, Hughson FM (2016) Chaperoning SNARE assembly and disassembly. Nat Rev Mol Cell Biol 17:465–479
Yu HJ, Liu YH, Gulbranson DR, Paine A, Rathore SS, Shen JS (2016) Extended synaptotagmins are Ca2+-dependent lipid transfer proteins at membrane contact sites. Proc Natl Acad Sci U S A 113:4362–4367
Saheki Y, De Camilli P (2017) Endoplasmic reticulum-plasma membrane contact sites. Annu Rev Biochem 86:659–684
Reinisch KM, De Camilli P (2016) SMP-domain proteins at membrane contact sites: structure and function. Biochim Biophys Acta 1861:924–927
Bian X, Zhang Z, Xiong QC, De Camilli P, Lin CX (2019) A programmable DNA-origami platform for studying lipid transfer between bilayers. Nat Chem Biol 15:830
Yu H, Siewny MGW, Edwards DT, Sanders AW, Perkins TT (2017) Hidden dynamics in the unfolding of individual bacteriorhodopsin proteins. Science 355:945–949
Choi HK, Min D, Kang H, Shon MJ, Rah SH, Kim HC, Jeong H, Choi HJ, Bowie JU, Yoon TY (2019) Watching helical membrane proteins fold reveals a common N-to-C-terminal folding pathway. Science 366:1150–1156
Popot JL, Engelman DM (2016) Membranes do not tell proteins how to fold. Biochemistry 55:5–18
Lemmon MA (2008) Membrane recognition by phospholipid-binding domains. Nat Rev Mol Cell Biol 9:99–111
Corbalan-Garcia S, Gomez-Fernandez JC (2014) Signaling through C2 domains: more than one lipid target. Biochim Biophys Acta 1838:1536–1547
Saheki Y, De Camilli P (2017) The extended-synaptotagmins. Biochim Biophys Acta 1864:1490–1493
Ma L, Cai Y, Li Y, Jiao J, Wu Z, O'Shaughnessy B, De Camilli P, Karatekin E, Zhang YL (2017) Single-molecule force spectroscopy of protein-membrane interactions. eLife 6:e30493
Perez-Lara A, Thapa A, Nyenhuis SB, Nyenhuis DA, Halder P, Tietzel M, Tittmann K, Cafiso DS, Jahn R (2016) PtdInsP2 and PtdSer cooperate to trap synaptotagmin-1 to the plasma membrane in the presence of calcium. eLife 5:e15886
Schauder CM, Wu XD, Saheki Y, Narayanaswamy P, Torta F, Wenk MR, De Camilli P, Reinisch KM (2014) Structure of a lipid-bound extended synaptotagmin indicates a role in lipid transfer. Nature 510:552–555
Min SW, Chang WP, Sudhof TC (2007) E-Syts, a family of membranous Ca2+-sensor proteins with multiple C2 domains. Proc Natl Acad Sci U S A 104:3823–3828
Giordano F, Saheki Y, Idevall-Hagren O, Colombo SF, Pirruccello M, Milosevic I, Gracheva EO, Bagriantsev SN, Borgese N, De Camilli P (2013) PI(4,5)P2-dependent and Ca2+-regulated ER-PM interactions mediated by the extended synaptotagmins. Cell 153:1494–1509
Xu JJ, Bacaj T, Zhou A, Tomchick DR, Sudhof TC, Rizo J (2014) Structure and Ca2+-binding properties of the tandem C2 domains of E-Syt2. Structure 22:269–280
Zhang YL, Sirinakis G, Gundersen G, Xi ZQ, Gao Y (2012) DNA translocation of ATP-dependent chromatin remodelling factors revealed by high-resolution optical tweezers. Methods Enzymol 513:3–28
Zhang YL, Smith CL, Saha A, Grill SW, Mihardja S, Smith SB, Cairns BR, Peterson CL, Bustamante C (2006) DNA translocation and loop formation mechanism of chromatin remodeling by SWI/SNF and RSC. Mol Cell 24:559–568
Ma L, Jiao J, Zhang YL (2019) Single-molecule optical tweezers study of regulated SNARE assembly. Methods Mol Biol 1860:95–114
Sirinakis G, Ren YX, Gao Y, Xi ZQ, Zhang YL (2012) Combined and versatile high-resolution optical tweezers and single-molecule fluorescence microscopy. Rev Sci Instrum 83:093708
Moffitt JR, Chemla YR, Izhaky D, Bustamante C (2006) Differential detection of dual traps improves the spatial resolution of optical tweezers. Proc Natl Acad Sci U S A 103:9006–9011
Gittes F, Schmidt CF (1998) Interference model for back-focal-plane displacement detection in optical tweezers. Opt Lett 23:7–9
Zhang XM, Ma L, Zhang YL (2013) High-resolution optical tweezers for single-molecule manipulation. Yale J Biol Med 86:367–383
Cecconi C, Shank EA, Bustamante C, Marqusee S (2005) Direct observation of the three-state folding of a single protein molecule. Science 309:2057–2060
Cecconi C, Shank EA, Marqusee S, Bustamante C (2011) DNA molecular handles for single-molecule protein-folding studies by optical tweezers. DNA Nanotechnol Method Protoc 749:255–271
Gao Y, Zorman S, Gundersen G, Xi ZQ, Ma L, Sirinakis G, Rothman JE, Zhang YL (2012) Single reconstituted neuronal SNARE complexes zipper in three distinct stages. Science 337:1340–1343
Gao Y, Sirinakis G, Zhang YL (2011) Highly anisotropic stability and folding kinetics of a single coiled coil protein under mechanical tension. J Am Chem Soc 133:12749–12757
Petrache HI, Dodd SW, Brown MF (2000) Area per lipid and acyl length distributions in fluid phosphatidylcholines determined by 2H NMR spectroscopy. Biophys J 79:3172–3192
Jiao JY, Rebane AA, Ma L, Zhang YL (2017) Single-molecule protein folding experiments using high-resolution optical tweezers. Methods Mol Biol 1486:357–390
Zhang YL, Jiao J, Rebane AA (2016) Hidden Markov modeling with detailed balance and its application to single protein folding. Biophys J 111:2110–2124
Rabiner LR (1989) A tutorial on hidden Markov-models and selected applications in speech recognition. Proc IEEE 77:257–286
Rebane AA, Ma L, Zhang YL (2016) Structure-based derivation of protein folding intermediates and energies from optical tweezers. Biophys J 110:441–454
Acknowledgments
This work was supported by NIH grant R35GM131714, R01GM093341, and R01GM120193 to Y. Z. and by the National Natural Science Foundation of China grant 31770812 to L. M.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ma, L., Ge, J., Bian, X., Zhang, Y. (2022). Single-Molecule Optical Tweezers Study of Protein–Membrane Interactions. In: Shen, J. (eds) Membrane Trafficking. Methods in Molecular Biology, vol 2473. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2209-4_23
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2209-4_23
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2208-7
Online ISBN: 978-1-0716-2209-4
eBook Packages: Springer Protocols