Abstract
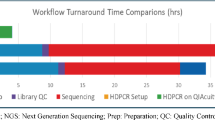
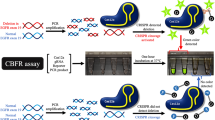
The profiling of EGFR mutations, the most common genetic alterations in non-small cell lung cancer (NSCLC) predictive of targeted therapy efficacy, is crucial to anticipate the patient response to EGFR tyrosine kinase inhibitors. Here, we introduce the naica® system for 6-color Crystal Digital PCRTM and describe in detail a standardized workflow for the multiplexed, single-assay detection of the 19 most prevalent sensitizing and resistance EGFR mutations in both tumor and circulating tumor DNA (ctDNA) samples. Two major advantages of the 6-color multiplexing system over current digital PCR systems are the rapid time to results, and the large quantity of mutational information obtained per patient sample, rendering the 6-color system highly cost effective. The 6-color Crystal Digital PCRTM technology enables highly sensitive and efficient therapeutic monitoring through liquid biopsy, resulting in the early detection of treatment resistance. While the assay presented here specifically addresses EGFR mutation status monitoring in NSCLC patients, 6-color Crystal Digital PCRTM assays are flexible and evolutive in design. As such, 6-color detection assays can be optimized to monitor mutations associated with a range of cancers and other genetic diseases, as well as to detect genetic changes beyond the oncology and human health domains.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Barlesi F, Mazieres J, Merlio J et al (2016) Routine molecular profiling of patients with advanced non-small-cell lung cancer: results of a 1-year nationwide programme of the French cooperative thoracic intergroup (IFCT). Lancet 387(10026):1415–1426
Reguart N, Remon J (2015) Common EGFR-mutated subgroups (Del19/L858R) in advanced non-small-cell lung cancer: chasing better outcomes with tyrosine kinase inhibitors. Future Oncol 11(8):1245–1257
Lee CK, Davies L, Wu YL et al (2017) Gefitinib or erlotinib vs chemotherapy for EGFR mutation-positive lung cancer: individual patient data meta-analysis of overall survival. J Natl Cancer Inst 109(6)
Helena AY, Arcila ME, Rekhtman N et al (2013) Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin Cancer Res 19(8):2240–2247
Wang S, Tsui ST, Liu C et al (2016) EGFR C797S mutation mediates resistance to third-generation inhibitors in T790M-positive non-small cell lung cancer. J Hematol Oncol 9(1):59
Jovelet C, Ileana E, Le Deley MC et al (2016) Circulating cell-free tumor DNA analysis of 50 genes by next-generation sequencing in the prospective MOSCATO trial. Clin Cancer Res 22(12):2960–2968
Madic J, Jovelet C, Lopez J et al (2019) Correction: EGFR C797S, EGFR T790M and EGFR sensitizing mutations in non-small cell lung cancer revealed by six-color crystal digital PCR. Oncotarget 10(13):1345
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Madic, J., Jovelet, C., Dehri, I., Mallory, A.C. (2021). 6-Color Crystal Digital PCRTM for the High-Plex Detection of EGFR Mutations in Non-Small Cell Lung Cancer. In: Santiago-Cardona, P.G. (eds) Lung Cancer. Methods in Molecular Biology, vol 2279. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1278-1_10
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1278-1_10
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1277-4
Online ISBN: 978-1-0716-1278-1
eBook Packages: Springer Protocols