Abstract
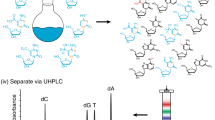
5-Hydroxymethyluracil (5hmU) and 5-formyluracil (5fU) are mutagenic and major oxidized nucleobases in DNA that are derived from thymine (T) or 5-methylcytosine (5mC). 5hmU, and possibly 5fU as well, in DNA may also assume an epigenetic role. The analysis of 5hmU and 5fU in DNA has suffered from multiple drawbacks such as a lack of specificity, artifacts generated in sample preparation, as well as poor applicability to analysis of nucleoside forms of DNA base damage. To overcome these problems, we developed a capillary liquid chromatography–electrospray ionization–tandem mass spectrometry (LC-ESI-MS3) method that is combined with the stable isotope-dilution technique, which allows quantification of 5hmU, 5fU, and several other oxidation products in DNA with high sensitivity, specificity, and accuracy. In this chapter, we describe in detail the protocols from DNA extraction, through DNA digestion and off-line HPLC enrichment, to LC-MS3 analysis. The method is readily applicable to other modified nucleosides in DNA.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Marnett LJ (2000) Oxyradicals and DNA damage. Carcinogenesis 21:361–370
Lindahl T (1999) DNA lesions generated in vivo by reactive oxygen species, their accumulation and repair. NATO ASI Ser Ser A 302:251–257
Finkel T, Holbrook NJ (2000) Oxidants, oxidative stress and the biology of ageing. Nature 408:239–247
Cadet J, Bellon S, Douki T et al (2004) Radiation-induced DNA damage: formation, measurement, and biochemical features. J Environ Pathol Toxicol Oncol 23:33–43
Kasai H, Iida A, Yamaizumi Z et al (1990) 5-Formyldeoxyuridine: a new type of DNA damage induced by ionizing radiation and its mutagenicity to salmonella strain TA102. Mutat Res 243:249–253
Olinski R, Starczak M, Gackowski D (2016) Enigmatic 5-hydroxymethyluracil: oxidatively modified base, epigenetic mark or both? Mutat Res Rev Mutat Res 767:59–66
Teebor GW, Boorstein RJ, Cadet J (1988) The repairability of oxidative free radical mediated damage to DNA: a review. Int J Radiat Biol 54:131–150
Levy DD, Teebor GW (1991) Site directed substitution of 5-hydroxymethyluracil for thymine in replicating phi X-174am3 DNA via synthesis of 5-hydroxymethyl-2′-deoxyuridine-5′-triphosphate. Nucleic Acids Res 19:3337–3343
Herrala AM, Vilpo JA (1989) Template-primer activity of 5-(hydroxymethyl)uracil-containing DNA for prokaryotic and eukaryotic DNA and RNA polymerases. Biochemistry 28:8274–8277
Mellac S, Fazakerley GV, Sowers LC (1993) Structures of base pairs with 5-(hydroxymethyl)-2′-deoxyuridine in DNA determined by NMR spectroscopy. Biochemistry 32:7779–7786
Rusmintratip V, Sowers LC (2000) An unexpectedly high excision capacity for mispaired 5-hydroxymethyluracil in human cell extracts. Proc Natl Acad Sci U S A 97:14183–14187
Hashimoto H, Hong S, Bhagwat AS et al (2012) Excision of 5-hydroxymethyluracil and 5-carboxylcytosine by the thymine DNA glycosylase domain: its structural basis and implications for active DNA demethylation. Nucleic Acids Res 40:10203–10214
Morera S, Grin I, Vigouroux A et al (2012) Biochemical and structural characterization of the glycosylase domain of MBD4 bound to thymine and 5-hydroxymethyuracil-containing DNA. Nucleic Acids Res 40:9917–9926
Cortellino S, Xu J, Sannai M et al (2011) Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 146:67–79
Wibley JE, Waters TR, Haushalter K et al (2003) Structure and specificity of the vertebrate anti-mutator uracil-DNA glycosylase SMUG1. Mol Cell 11:1647–1659
Modrzejewska M, Gawronski M, Skonieczna M et al (2016) Vitamin C enhances substantially formation of 5-hydroxymethyluracil in cellular DNA. Free Radic Biol Med 101:378–383
Kawasaki F, Martinez Cuesta S, Beraldi D et al (2018) Sequencing 5-hydroxymethyluracil at single-base resolution. Angew Chem Int Ed Engl 57:9694–9696
Kawasaki F, Beraldi D, Hardisty RE et al (2017) Genome-wide mapping of 5-hydroxymethyluracil in the eukaryote parasite Leishmania. Genome Biol 18:23
Pfaffeneder T, Spada F, Wagner M et al (2014) Tet oxidizes thymine to 5-hydroxymethyluracil in mouse embryonic stem cell DNA. Nat Chem Biol 10:574–581
Klungland A, Paulsen R, Rolseth V et al (2001) 5-Formyluracil and its nucleoside derivatives confer toxicity and mutagenicity to mammalian cells by interfering with normal RNA and DNA metabolism. Toxicol Lett 119:71–78
Kamiya H, Murata-Kamiya N, Karino N et al (2002) Induction of T --> G and T --> A transversions by 5-formyluracil in mammalian cells. Mutat Res 513:213–222
Terato H, Masaoka A, Kobayashi M et al (1999) Enzymatic repair of 5-formyluracil. II. Mismatch formation between 5-formyluracil and guanine during dna replication and its recognition by two proteins involved in base excision repair (AlkA) and mismatch repair (MutS). J Biol Chem 274:25144–25150
Masaoka A, Terato H, Kobayashi M et al (1999) Enzymatic repair of 5-formyluracil. I. Excision of 5-formyluracil site-specifically incorporated into oligonucleotide substrates by alka protein (Escherichia coli 3-methyladenine DNA glycosylase II). J Biol Chem 274:25136–25143
Bjelland S, Birkeland NK, Benneche T et al (1994) DNA glycosylase activities for thymine residues oxidized in the methyl group are functions of the AlkA enzyme in Escherichia coli. J Biol Chem 269:30489–30495
Knaevelsrud I, Slupphaug G, Leiros I et al (2009) Opposite-base dependent excision of 5-formyluracil from DNA by hSMUG1. Int J Radiat Biol 85:413–420
Matsubara M, Masaoka A, Tanaka T et al (2003) Mammalian 5-formyluracil-DNA glycosylase. 1. Identification and characterization of a novel activity that releases 5-formyluracil from DNA. Biochemistry 42:4993–5002
Masaoka A, Matsubara M, Hasegawa R et al (2003) Mammalian 5-formyluracil-DNA glycosylase. 2. Role of SMUG1 uracil-DNA glycosylase in repair of 5-formyluracil and other oxidized and deaminated base lesions. Biochemistry 42:5003–5012
Liu P, Burdzy A, Sowers LC (2003) Repair of the mutagenic DNA oxidation product, 5-formyluracil. DNA Repair 2:199–210
Schiesser S, Pfaffeneder T, Sadeghian K et al (2013) Deamination, oxidation, and C-C bond cleavage reactivity of 5-hydroxymethylcytosine, 5-formylcytosine, and 5-carboxycytosine. J Am Chem Soc 135:14593–14599
Cadet J, Odin F, Mouret JF et al (1992) Chemical and biochemical postlabeling methods for singling out specific oxidative DNA lesions. Mutat Res 275:343–354
Teebor GW, Frenkel K, Goldstein MS (1984) Ionizing radiation and tritium transmutation both cause formation of 5-hydroxymethyl-2′-deoxyuridine in cellular DNA. Proc Natl Acad Sci U S A 81:318–321
Frenkel K, Cummings A, Solomon J et al (1985) Quantitative determination of the 5-(hydroxymethyl)uracil moiety in the DNA of gamma-irradiated cells. Biochemistry 24:4527–4533
Bjelland S, Eide L, Time RW et al (1995) Oxidation of thymine to 5-formyluracil in DNA: mechanisms of formation, structural implications, and base excision by human cell free extracts. Biochemistry 34:14758–14764
Dizdaroglu M, Jaruga P, Birincioglu M et al (2002) Free radical-induced damage to DNA: mechanisms and measurement. Free Radic Biol Med 32:1102–1115
Douki T, Delatour T, Paganon F et al (1996) Measurement of oxidative damage at pyrimidine bases in gamma-irradiated DNA. Chem Res Toxicol 9:1145–1151
Mori T, Dizdaroglu M (1994) Ionizing radiation causes greater DNA base damage in radiation-sensitive mutant M10 cells than in parent mouse lymphoma L5178Y cells. Radiat Res 140:85–90
Dizdaroglu M (1998) Facts about the artifacts in the measurement of oxidative DNA base damage by gas chromatography-mass spectrometry. Free Radic Res 29:551–563
Cadet J, Douki T, Ravanat JL (1997) Artifacts associated with the measurement of oxidized DNA bases. Environ Health Perspect 105:1034–1039
Cadet J, Douki T, Frelon S et al (2002) Assessment of oxidative base damage to isolated and cellular DNA by HPLC-MS/MS measurement. Free Radic Biol Med 33:441–449
Frelon S, Douki T, Ravanat JL et al (2000) High-performance liquid chromatography--tandem mass spectrometry measurement of radiation-induced base damage to isolated and cellular DNA. Chem Res Toxicol 13:1002–1010
Hua Y, Wainhaus SB, Yang Y et al (2001) Comparison of negative and positive ion electrospray tandem mass spectrometry for the liquid chromatography tandem mass spectrometry analysis of oxidized deoxynucleosides. J Am Soc Mass Spectrom 12:80–87
Wang J, Yuan B, Guerrero C et al (2011) Quantification of oxidative DNA lesions in tissues of Long-Evans Cinnamon rats by capillary high-performance liquid chromatography-tandem mass spectrometry coupled with stable isotope-dilution method. Anal Chem 83:2201–2209
Tretyakova N, Goggin M, Sangaraju D et al (2012) Quantitation of DNA adducts by stable isotope dilution mass spectrometry. Chem Res Toxicol 25:2007–2035
Hong H, Cao H, Wang Y et al (2006) Identification and quantification of a guanine-thymine intrastrand cross-link lesion induced by Cu(II)/H2O2/ascorbate. Chem Res Toxicol 19:614–621
Acknowledgments
The development of the methodology used on which this procedure was prepared was supported by the National Institutes of Health (Grants R01 CA101864, R01 DK082779, R01 DK071111, and P30-DK41296). The preparation of this procedure was supported by Inner Mongolia University with a grant from its “Outstanding Young Talents” Program (to YL) and a grant from its “Steed Plan” High-Level Talents Program (to JW).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Liu, Y., Wang, J. (2022). Isotope-Dilution Liquid Chromatography–Tandem Mass Spectrometry for Detection of 5-Hydroxymethyluracil and 5-Formyluracil in DNA. In: Yuan, BF. (eds) DNA Modification Detection Methods . Springer Protocols Handbooks. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1229-3_12
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1229-3_12
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1228-6
Online ISBN: 978-1-0716-1229-3
eBook Packages: Springer Protocols