Abstract
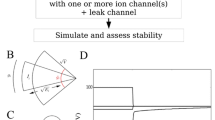
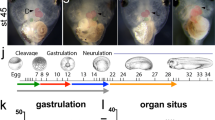
Embryogenesis, as well as regeneration, is increasingly recognized to be orchestrated by an interplay of transcriptional and bioelectric networks. Spatiotemporal patterns of resting potentials direct the size, shape, and locations of numerous organ primordia during patterning. These bioelectrical properties are established by the function of ion channels and pumps that set voltage potentials of individual cells, and gap junctions (electrical synapses) that enable physiological states to propagate across tissue networks. Functional experiments to probe the roles of bioelectrical states can be carried out by targeting endogenous ion channels during development. Here, we describe protocols, optimized for the highly tractable Xenopus laevis embryo, for molecular genetic targeting of ion channels and connexins based on CRISPR, and monitoring of resting potential states using voltage-sensing fluorescent dye. Similar strategies can be adapted to other model species.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Sundelacruz S, Levin M, Kaplan DL (2008) Membrane potential controls adipogenic and osteogenic differentiation of mesenchymal stem cells. PLoS One 3:e3737
Sundelacruz S, Levin M, Kaplan DL (2009) Role of membrane potential in the regulation of cell proliferation and differentiation. Stem Cell Rev Rep 5:231–246
Funk RH (2015) Endogenous electric fields as guiding cue for cell migration. Front Physiol 6:143
Levin M, Martyniuk CJ (2018) The bioelectric code: an ancient computational medium for dynamic control of growth and form. Biosystems 164:76–93. https://doi.org/10.1016/j.biosystems.2017.08.009
Adams DS et al (2006) Early, H+-V-ATPase-dependent proton flux is necessary for consistent left-right patterning of non-mammalian vertebrates. Development 133:1657–1671
Levin M (2017) Seminars in cell & developmental biology. Elsevier, Amsterdam, pp 543–556
McLaughlin KA, Levin M (2017) Bioelectric signaling in regeneration: mechanisms of ionic controls of growth and form. Dev Biol 433(2):177–189
Mathews J, Levin M (2017) Gap junctional signaling in pattern regulation: physiological network connectivity instructs growth and form. Dev Neurobiol 77:643–673. https://doi.org/10.1002/dneu.22405
Bates E (2015) Ion channels in development and cancer. Annu Rev Cell Dev Biol 31:231–247. https://doi.org/10.1146/annurev-cellbio-100814-125338
Adams DS, Levin M (2013) Endogenous voltage gradients as mediators of cell-cell communication: strategies for investigating bioelectrical signals during pattern formation. Cell Tissue Res 352:95–122. https://doi.org/10.1007/s00441-012-1329-4
Chernet BT, Adams DS, Lobikin M, Levin M (2016) Use of genetically encoded, light-gated ion translocators to control tumorigenesis. Oncotarget 7:19575–19588. https://doi.org/10.18632/oncotarget.8036
Adams DS et al (2016) Bioelectric signalling via potassium channels: a mechanism for craniofacial dysmorphogenesis in KCNJ2-associated Andersen-Tawil Syndrome. J Physiol 594:3245–3270. https://doi.org/10.1113/JP271930
Adams DS, Lemire JM, Kramer RH, Levin M (2014) Optogenetics in developmental biology: using light to control ion flux-dependent signals in Xenopus embryos. Int J Dev Biol 58:851–861. https://doi.org/10.1387/ijdb.140207ml
Adams DS, Tseng AS, Levin M (2013) Light-activation of the Archaerhodopsin H(+)-pump reverses age-dependent loss of vertebrate regeneration: sparking system-level controls in vivo. Biol Open 2:306–313. https://doi.org/10.1242/bio.20133665
Adams DS, Levin M (2006) Inverse drug screens: a rapid and inexpensive method for implicating molecular targets. Genesis 44:530–540
Sullivan KG, Levin M (2018) Inverse drug screening of bioelectric signaling and neurotransmitter roles: illustrated using a xenopus tail regeneration assay. Cold Spring Harb Protoc 2018:pdb.prot099937. https://doi.org/10.1101/pdb.prot099937
Belus MT et al (2018) Kir2.1 is important for efficient BMP signaling in mammalian face development. Dev Biol 444(Suppl 1):S297–S307. https://doi.org/10.1016/j.ydbio.2018.02.012
Ardissone A, Sansone V, Colleoni L, Bernasconi P, Moroni I (2017) Intrafamilial phenotypic variability in Andersen-Tawil syndrome: a diagnostic challenge in a potentially treatable condition. Neuromuscul Disord 27:294–297. https://doi.org/10.1016/j.nmd.2016.11.006
Dahal GR et al (2012) An inwardly rectifying K+ channel is required for patterning. Development 139:3653–3664. https://doi.org/10.1242/dev.078592
Marrus SB, Cuculich PS, Wang W, Nerbonne JM (2011) Characterization of a novel, dominant negative KCNJ2 mutation associated with Andersen-Tawil syndrome. Channels (Austin) 5:500–509. https://doi.org/10.4161/chan.5.6.18524
George LF et al (2019) Ion channel contributions to wing development in Drosophila melanogaster. G3 (Bethesda) 9(4):999–1008. https://doi.org/10.1534/g3.119.400028
Smith RS et al (2018) Sodium channel SCN3A (NaV1.3) regulation of human cerebral cortical folding and oral motor development. Neuron 99:905–913 .e907. https://doi.org/10.1016/j.neuron.2018.07.052
Masotti A et al (2015) Keppen-Lubinsky syndrome is caused by mutations in the inwardly rectifying K(+) channel encoded by KCNJ6. Am J Hum Genet 96:295–300. https://doi.org/10.1016/j.ajhg.2014.12.011
Kortum F et al (2015) Mutations in KCNH1 and ATP6V1B2 cause Zimmermann-Laband syndrome. Nat Genet 47(6):661–667. https://doi.org/10.1038/ng.3282
Veale EL, Hassan M, Walsh Y, Al-Moubarak E, Mathie A (2014) Recovery of current through mutated TASK3 potassium channels underlying Birk Barel syndrome. Mol Pharmacol 85:397–407. https://doi.org/10.1124/mol.113.090530
Curran J, Mohler PJ (2015) Alternative paradigms for ion channelopathies: disorders of ion channel membrane trafficking and posttranslational modification. Annu Rev Physiol 77:505–524
Perathoner S et al (2014) Bioelectric signaling regulates size in zebrafish fins. PLoS Genet e1004080:10
Hoptak-Solga AD, Klein KA, DeRosa AM, White TW, Iovine MK (2007) Zebrafish short fin mutations in connexin43 lead to aberrant gap junctional intercellular communication. FEBS Lett 581:3297–3302
La Russa MF, Qi LS (2015) The new state of the art: Cas9 for gene activation and repression. Mol Cell Biol 35:3800–3809
Beck CW, Slack JM (2001) An amphibian with ambition: a new role for Xenopus in the 21st century. Genome Biol 2:reviews1029
Blum M, Ott T (2018) Xenopus: an undervalued model organism to study and model human genetic disease. Cells Tissues Organs 205(5–6):303–313. https://doi.org/10.1159/000490898
Tseng AS (2017) Seeing the future: using Xenopus to understand eye regeneration. Genesis 55:1–2. https://doi.org/10.1002/dvg.23003
Getwan M, Lienkamp SS (2017) Toolbox in a tadpole: Xenopus for kidney research. Cell Tissue Res 369:143–157. https://doi.org/10.1007/s00441-017-2611-2
Dubey A, Saint-Jeannet JP (2017) Modeling human craniofacial disorders in Xenopus. Curr Pathobiol Rep 5:79–92. https://doi.org/10.1007/s40139-017-0128-8
Adams DS, Levin M (2012) Measuring resting membrane potential using the fluorescent voltage reporters DiBAC4(3) and CC2-DMPE. Cold Spring Harb Protoc 2012:459–464. https://doi.org/10.1101/pdb.prot067702
Adams DS, Levin M (2012) General principles for measuring resting membrane potential and ion concentration using fluorescent bioelectricity reporters. Cold Spring Harb Protoc 2012:385–397. https://doi.org/10.1101/pdb.top067710
Kim EZ, Vienne J, Rosbash M, Griffith LC (2017) Non-reciprocal homeostatic compensation in Drosophila potassium channel mutants. J Neurophysiol 117(6):2125–2136. https://doi.org/10.1152/jn.00002.2017
Pietak A, Levin M (2017) Bioelectric gene and reaction networks: computational modelling of genetic, biochemical and bioelectrical dynamics in pattern regulation. J R Soc Interface 14(134):20170425. https://doi.org/10.1098/rsif.2017.0425
Pietak A, Levin M (2016) Exploring instructive physiological signaling with the bioelectric tissue simulation engine (BETSE). Front Bioeng Biotechnol 4:55. https://doi.org/10.3389/fbioe.2016.00055
Adams DS, Levin M (2012) Measuring resting membrane potential using the fluorescent voltage reporters DiBAC4 (3) and CC2-DMPE. Cold Spring Harb Protoc 2012:pdb.prot067702
Oviedo NJ, Nicolas CL, Adams DS, Levin M (2008) Live imaging of planarian membrane potential using DiBAC4 (3). Cold Spring Harb Protoc 2008:pdb.prot5055
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Nanos, V., Levin, M. (2021). Rewiring Endogenous Bioelectric Circuits in the Xenopus laevis Embryo Model. In: Ebrahimkhani, M.R., Hislop, J. (eds) Programmed Morphogenesis. Methods in Molecular Biology, vol 2258. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1174-6_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1174-6_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1173-9
Online ISBN: 978-1-0716-1174-6
eBook Packages: Springer Protocols