Abstract
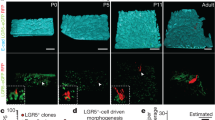
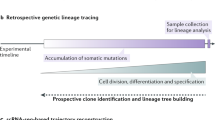
Lineage-tracing experiments aim to identify and track the progeny and/or fate of cells. The use of inducible recombinases and fluorescent reporters has been instrumental in defining cellular hierarchies and allowing for the identification of stem cells in an unperturbed in vivo setting. The refinement of these approaches, labeling single cells, and the subsequent quantitative analysis of the clonal dynamics have allowed the comparison of different stem cell populations as well as establishing different mechanisms of cellular replenishment during steady-state homeostasis as well as during morphogenesis and disease. Utilizing this approach, it is now possible to establish the cellular hierarchy in a given tissue and the frequency of cell fate decisions on a population basis, thus providing a comprehensive analysis of cellular behavior in vivo. Although in this chapter we describe a protocol for lineage tracing of cells from fetal intestinal epithelium to the adult intestine, this approach can be widely applied to quantitatively assess the cell fate of any fetal cell during morphogenesis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kretzschmar K, Watt FM (2012) Lineage tracing. Cell 148(1–2):33–45. https://doi.org/10.1016/j.cell.2012.01.002
Cheng H, Leblond CP (1974) Origin, differentiation and renewal of the four main epithelial cell types in the mouse small intestine. I. Columnar cell. Am J Anat 141(4):461–479. https://doi.org/10.1002/aja.1001410403
Barker N, van Es JH, Kuipers J, Kujala P, van den Born M, Cozijnsen M, Haegebarth A, Korving J, Begthel H, Peters PJ, Clevers H (2007) Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449(7165):1003–1007. https://doi.org/10.1038/nature06196
Lopez-Garcia C, Klein AM, Simons BD, Winton DJ (2010) Intestinal stem cell replacement follows a pattern of neutral drift. Science 330(6005):822–825. https://doi.org/10.1126/science.1196236
Ritsma L, Ellenbroek SIJ, Zomer A, Snippert HJ, de Sauvage FJ, Simons BD, Clevers H, van Rheenen J (2014) Intestinal crypt homeostasis revealed at single-stem-cell level by in vivo live imaging. Nature 507(7492):362–365. https://doi.org/10.1038/nature12972
Snippert HJ, van der Flier LG, Sato T, van Es JH, van den Born M, Kroon-Veenboer C, Barker N, Klein AM, van Rheenen J, Simons BD, Clevers H (2010) Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143(1):134–144. https://doi.org/10.1016/j.cell.2010.09.016
Guiu J, Hannezo E, Yui S, Demharter S, Ulyanchenko S, Maimets M, Jorgensen A, Perlman S, Lundvall L, Mamsen LS, Larsen A, Olesen RH, Andersen CY, Thuesen LL, Hare KJ, Pers TH, Khodosevich K, Simons BD, Jensen KB (2019) Tracing the origin of adult intestinal stem cells. Nature 570(7759):107–111. https://doi.org/10.1038/s41586-019-1212-5
Andersen MS, Hannezo E, Ulyanchenko S, Estrach S, Antoku Y, Pisano S, Boonekamp KE, Sendrup S, Maimets M, Pedersen MT, Johansen JV, Clement DL, Feral CC, Simons BD, Jensen KB (2019) Tracing the cellular dynamics of sebaceous gland development in normal and perturbed states. Nat Cell Biol 21(8):924–932. https://doi.org/10.1038/s41556-019-0362-x
Clayton E, Doupe DP, Klein AM, Winton DJ, Simons BD, Jones PH (2007) A single type of progenitor cell maintains normal epidermis. Nature 446(7132):185–189. https://doi.org/10.1038/nature05574
Latil M, Nassar D, Beck B, Boumahdi S, Wang L, Brisebarre A, Dubois C, Nkusi E, Lenglez S, Checinska A, Vercauteren Drubbel A, Devos M, Declercq W, Yi R, Blanpain C (2017) Cell-type-specific chromatin states differentially prime squamous cell carcinoma tumor-initiating cells for epithelial to mesenchymal transition. Cell Stem Cell 20(2):191–204. e195. https://doi.org/10.1016/j.stem.2016.10.018
Han S, Fink J, Jorg DJ, Lee E, Yum MK, Chatzeli L, Merker SR, Josserand M, Trendafilova T, Andersson-Rolf A, Dabrowska C, Kim H, Naumann R, Lee JH, Sasaki N, Mort RL, Basak O, Clevers H, Stange DE, Philpott A, Kim JK, Simons BD, Koo BK (2019) Defining the identity and dynamics of adult gastric isthmus stem cells. Cell Stem Cell 25(3):342–356. e347. https://doi.org/10.1016/j.stem.2019.07.008
Lilja AM, Rodilla V, Huyghe M, Hannezo E, Landragin C, Renaud O, Leroy O, Rulands S, Simons BD, Fre S (2018) Clonal analysis of Notch1-expressing cells reveals the existence of unipotent stem cells that retain long-term plasticity in the embryonic mammary gland. Nat Cell Biol 20(6):677–687. https://doi.org/10.1038/s41556-018-0108-1
Tetteh PW, Basak O, Farin HF, Wiebrands K, Kretzschmar K, Begthel H, van den Born M, Korving J, de Sauvage F, van Es JH, van Oudenaarden A, Clevers H (2016) Replacement of lost Lgr5-positive stem cells through plasticity of their enterocyte-lineage daughters. Cell Stem Cell 18(2):203–213. https://doi.org/10.1016/j.stem.2016.01.001
Acknowledgments
We thank Dr. Kim B. Jensen and Jensen lab for his support, advice, and helpful discussion. We acknowledge Biorender.com as Figs. 1, 3, 4, and 6 were created with Biorender.com and exported under a paid subscription. This work was supported by MSCA-IF-2014-EF-656099. We thank CERCA Programme / Generalitat de Catalunya for institutional support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ulyanchenko, S., Guiu, J. (2021). A Quantitative Lineage-Tracing Approach to Understand Morphogenesis in Gut. In: Ebrahimkhani, M.R., Hislop, J. (eds) Programmed Morphogenesis. Methods in Molecular Biology, vol 2258. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1174-6_3
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1174-6_3
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1173-9
Online ISBN: 978-1-0716-1174-6
eBook Packages: Springer Protocols