Abstract
This study proposes the treatment and valorization of denim textile effluents through a fermentative hydrogen production process. Also, the study presents the decolorizing capabilities of bacterial and fungal isolates obtained from the fermented textile effluents. The maximum hydrogen production rate was 0.23 L H2/L-d, achieving at the same time color removal. A total of thirty-five bacteria and one fungal isolate were obtained from the fermented effluents and screened for their abilities to decolorize indigo dye, used as a model molecule. From them, isolates identified as Bacillus BT5, Bacillus BT9, Lactobacillus BT20, Lysinibacillus BT32, and Aspergillus H1T showed notable decolorizing capacities. Lactobacillus BT20 reached 90% of decolorization using glucose as co-substrate after 11 days of incubation producing colorless metabolites. Bacillus BT9 was able to utilize the indigo dye as the sole carbon source achieving a maximum decolorization of 60% after 9 days of incubation and producing a red-colored metabolite. In contrast, Bacillus BT5 and Lysinibacillus BT32 exhibited the lowest percentages of decolorization, barely 33% after 16 and 11 days of incubation, respectively. When Aspergillus H1T was grown in indigo dye supplemented with glucose, 96% of decolorization was reached after 2 days. This study demonstrates the valorization of denim textile effluents for the production of hydrogen via dark fermentation with concomitant color removal.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Textile industry demands large volumes of water for its dying and washing processes. In Mexico, this industry consumes almost 73 million m3 water per year, and 45% of these effluents are discharged without treatment [1]. Textile effluents have a mixture of contaminants such as synthetic dyes, surfactants, acids or bases, heavy metals (Cr, Ni, Cu), chlorinated organic compounds, starch, salts, fats and oils, and phenols with an alkaline pH [2], causing considerable pollution to water-receiving bodies. In general terms, the textile effluents are characterized by a high biochemical oxygen demand (from 210 to 5500 mg/L), chemical oxygen demand (COD, from 340 to 17,900 mg/L), total suspended solids from 50 to 24,000 mg/L, and a visible color from 300 to 3500 color units. Also, dyes present in textile effluents result in the abnormal coloration of surface waters affecting the sunlight entering the water [3]. The necessity of treatment for textile effluents is evident. Biological treatment is considered a competitive alternative in comparison with physicochemical pretreatments [3]. Fungi, such as Aspergillus sp., Penicillium sp., and Hypocrea koningii [4, 5], and several bacteria such as Enterococcus faecalis and Bacillus spp. have the ability to decolorize various types of industrial dyes [6,7,8].
A recent trend is the development of biological treatment systems coupled with the generation of value-added products, such as biohydrogen, creating a sustainable alternative in the context of a circular bioeconomy [9]. Recent efforts to valorize textile effluents for producing biohydrogen via dark fermentation have been already reported. Lay et al. [10] studied the effect of pH (4.0–8.0), substrate concentration (5–25 g COD/L), and type of inoculum (sewage sludge, soil, and cow dung) on the hydrogen production from textile wastewater in batch fermentations, observing the maximum hydrogen production rate with the cow dung inoculum, pH of 7.0, and substrate concentration of 20 g COD/L. With a different approach, the pretreatment of textile effluents has been successful in improving the fermentative hydrogen production. Li et al. [11] evaluated the hydrogen production from textile effluents pretreated with amylases, activated carbon, and cation resins. The amylase-activated carbon treatment yielded the best performance in terms of hydrogen production. In this study, the microbial community that achieved the decolorization was molecularly identified as Clostridium butyricum and Klebsiella oxytoca. The coagulation process was also used as pretreatment of textile desizing wastewater for batch [12] and continuous hydrogen production [13]. In batch reactors, the coagulation pretreatment improved by 120% the hydrogen yield compared with the untreated effluent. In the continuous bioreactor, the coagulation pretreatment reached a maximum hydrogen production rate similar to those reported previously with substrates like sugarcane vinasses.
The traditional approach for the biological treatment of textile effluents does not consider valorization of organic matter contained in such effluents for obtaining value-added products. At the other extreme, studies on hydrogen production from textile effluents do not address the decolorizing abilities of the microbial communities involved in the process. This study is more in line with the valorization of textile effluents for obtaining biohydrogen, but at the same time, to address the decolorizing abilities of the involved microorganisms. Therefore, the goals of this study were to valorize denim textile effluents for hydrogen production and characterize the decolorizing capacities of cultivable microorganisms.
Material and methods
Hydrogen production from denim textile effluents
Denim textile effluents were obtained from a local textile industry located in Guanajuato, Mexico, with a chemical oxygen demand of 2260 mg/L and pH of 8.7. These denim textile effluents were used as a substrate in 500 mL-bioreactors with a working volume of 300 mL (run in duplicate). The inoculum consisted of an anaerobic sludge with a total solid content of 2.8 ± 0.9 mg/L [14]. The anaerobic sludge received a heat-shock treatment in a water bath at 105 °C during 1 h to select sporulating H2-producing bacteria [15]. During the start-up, the bioreactors were inoculated with 20% v/v of the pretreated inoculum and 80% v/v of denim textile effluents supplemented with 2 g/L of glucose and 5 g/L of peptone to accelerate the bacterial growth [16]. After 5 days, the bioreactors were operated in continuous mode with a hydraulic and cellular retention times of 3 days, 35 °C, pH of 5.5 ± 0.3, and 100 rpm. During the continuous operation, the denim textile effluents were supplemented with 2.0 g/L of peptone as the nitrogen source.
Isolation and screening for decolorizing microorganisms
On day 30, a 10-mL sample of fermented denim effluents was used for screening for decolorizing microorganisms. The fermented effluents were diluted 1:10 in saline solution and 0.1 mL was spread on the surface of agar plates containing 25% v/v of unfermented denim textile effluents. Three plates were incubated at 28 °C for fungal growth, and three plates were incubated at 37 °C for bacterial growth. Then, monocultures were obtained and microscopically verified. All the isolates were screened during a decolorization test performed in tubes containing 12 mL of the sterilized medium with the following composition in gram per liter: glucose 2.5, peptone 5.0, and 0.05 of indigo dye supplied by a local textile industry (Guanajuato, Mexico). The tubes were inoculated by the loop technique and incubated at 100 rpm for 15 days. Bacterial monocultures were maintained in Luria Bertani medium (LB, in g/L: tryptone 5, yeast extract 5, NaCl 10), while fungal monocultures were maintained in potato dextrose agar (PDA).
Decolorization kinetics of indigo dye
Inoculum preparation
Inocula from bacterial monocultures were prepared in tubes with 30 mL of sterile LB medium inoculated by the loop technique and incubated at 37 ± 1 °C and 100 rpm for 48 h until to reach an optical density (OD600 nm) of 0.8–1.0. These monocultures were centrifuged at 4000×g for 15 min and the cell pellet was resuspended in LB broth for using as the inoculum source. A fungal isolate was grown in potato dextrose broth at 28 ± 1 °C for 2 days, and conidia were recovered and used as the inoculum source.
Procedure
The experiments were performed in 250-mL flasks containing 70 mL of sterile medium (2.0 g/L of peptone, 1.0 g/L of NaCl) with glucose (gl+, 2.5 g/L) or without glucose (gl−), and 50 mg/L of indigo dye. The initial pH was adjusted to 7.0 with NaOH. The flasks were inoculated with 2 mL of the bacterial inoculum and incubated at 37 °C, 100 rpm for 21 days. Samples were drawn for analysis on days 0, 2, 4, 7, 9, 11, 14, 16, 18, and 21. The flasks for fungal decolorization (gl+ and gl−) were inoculated with the conidial suspension and incubated at 28 ± 1 °C and 100 rpm. Samples were drawn on days 0, 1, 2, and 3. A control treatment without inoculum was always included. All the tests were performed in triplicate, and the results were expressed as the mean ± standard deviation. The percentage of decolorization was determined as follows:
Analytical methods
The biogas volume was measured by liquid displacement of a saturated solution of sodium chloride with pH 3.0. The biogas composition (H2, CO2, CH4) was determined by gas chromatography as previously reported [17]. Hydrogen production was reported at standard temperature and pressure (STP). The decolorization was assessed using supernatants after centrifugation at 5500×g/10 min by measuring the absorbance with a UV-visible spectrophotometer (Eppendorf, BioSpectrometer®) following the procedure reported by Ramya et al. [18], where the non-inoculated control was taken as 100%. The bacterial growth in the decolorization kinetics of indigo dye was determined using a Bio-Rad Bradford Protein Assay (Bio-Rad, Hercules, CA, USA). Fungal growth in the decolorization kinetics of indigo dye was determined gravimetrically from a known volume. Glucose consumption was measured as reducing sugars using the dinitrosalicylic acid method [19].
Molecular identification
Genomic DNA was extracted according to the protocol established by Burbano et al. [20] from the bacterial monocultures. The primer pair 9F (5′-GAGTTTGATCCTGGCTCAG-3′) and 1542R (5′-AGAAAGGAGGTGATCCAGCC-3′) was used to amplify a region of the 16S rDNA gene, and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) and ITS5 (5′-GGAAGTAAAAGTCGTAACAAGG-3′) for a region of the 5.8S rDNA gene. DNA amplification as performed using the Quick Load Taq2x Master Mix kit (New England BioLabs, M0271S) using a thermal cycler (C1000 Touch, Bio-Rad, USA) following the thermal profile of Yoon et al. [21] and Luo and Mitchell [22] for bacteria and fungi, respectively. The PureLink PCR Purification kit (Invitrogen, K3100-01) served to purify the PCR products that then were sequenced at the National Laboratory of Genomics for Biodiversity (LANGEBIO Irapuato, Mexico). The sequences were aligned using the National Center for Biotechnology Information (NCBI) database with the BLAST. Finally, a phylogenetic analysis of the identified strains and selected sequences of the GenBank was performed. Evolutionary analyses were conducted in MEGA v7.0 software [23]. The multiple sequence alignments were performed using MUSCLE [24], and to identify the best nucleotide substitution model for the sequences, an algorithm was used which is integrated in MEGA. For bacteria, the best substitution pattern was determined by maximum likelihood with the Kimura 2-parameter nucleotide substitution model using a discrete gamma distribution (+G). Meanwhile, for fungi, the best substitution pattern was determined by maximum likelihood with the Tamura 3-parameter nucleotide substitution model. The maximum likelihood trees were inferred from bootstrap method with 1000 replicates and were constructed applying the nearest-neighbor-join interchange containing all the sites [25,25,26,28]. The sequences were deposited into the GenBank database under accession numbers MG241238–MG241241 for the bacterial isolates and MG256171 for the fungal isolate.
Results and discussion
Hydrogen production from denim textile effluents
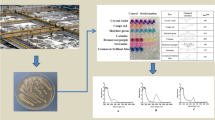
During the batch operation (first 5 days in Fig. 1), the bioreactor fed with denim textile effluents produced biogas with a negligible content of hydrogen. When the continuous operation mode was established, the hydrogen percentage increased to 55%. On average, the bioreactor achieved a stable hydrogen production for 20 days with a maximum hydrogen rate of 0.23 L H2/L-d (Fig. 1a).
In literature, only a few studies have reported production of hydrogen from the dark fermentation of textile effluents. From them, batch reactors yielded hydrogen production rates ranging from 1.1 to 4.3 L H2/L-d (Table 1). For intermittent or continuous bioreactors, the hydrogen production rate averaged 3.2 L H2/L-d. The hydrogen production rate reached in this study was lower in comparison with the previous reports due probably the low concentration of substrate in the denim textile effluents (chemical oxygen demand of 2.2 g COD/L). For instance, Lay et al. [29] reported a hydrogen production rate almost 10-fold higher than the value reached in this study, since the authors fed a textile effluent with a higher substrate concentration of 33 g/L. This higher substrate concentration represents higher organic loading rates that directly improve the hydrogen production rate but also could be related to the substrate or product inhibition if some threshold is surpassed [30].
In parallel to the hydrogen production, the distinctive blue color of the denim textile effluents changed to light transparent yellow (Fig. 1a). Anaerobic bacteria decompose azo dyes into potential carcinogenic amines considered a treatment less effective and safer than that performed by aerobic bacteria [31, 32]. Under certain conditions of microaerophilic/aerobic operation, the formation of aromatic amines could be avoided [33]. In this study, the acclimated microbial consortium received denim textile effluents with soluble oxygen that could be enough to maintain the growth of facultative bacteria. Therefore, the study of cultivable members responsible for decolorization was focused on aerobic members that could remain active in the H2-producing bioreactor cooperating with anaerobic members for the biodegradation of dyes.
Isolation of microorganisms and screening
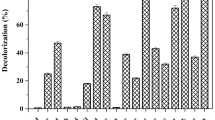
Thirty-five aerobic isolates were recovered from the fermented effluents using raw denim textile effluents as the sole carbon source. Screening for decolorizing microorganisms was performed using indigo dye, used as a model molecule. Indigo dye is one of the most used dyes in the blue jeans manufacturing, and its biological removal has been previously reported [34]. The screening decolorization assay showed that nineteen isolates showed some degree of decolorization. From them, the isolates called BT5, BT9, BT20, and BT32 had the highest decolorizing activities (Fig. 1b). These isolates were molecularly identified as Bacillus BT5, Bacillus BT9, Lactobacillus BT20, and Lysinibacillus BT32 (Fig. 2a), as well as a fungal isolate identified as Aspergillus H1T (Fig. 2b).
Decolorization kinetics of indigo dye
During the decolorization kinetic of indigo dye in presence of Lactobacillus BT20, the glucose consumption and color removal showed a parallel behavior achieving up to 90% of decolorization under aerobic growth and after 264 h of incubation (Fig. 3). In the absence of a co-substrate (glucose), Lactobacillus BT20 was unable to decolorize, a phenomenon previously reported for this genus [32, 35]. The Lactobacillus species express azoreductases (EC 1.7.1.6) to degrade to some extent azo dyes such as acid black under aerobic conditions [32], as well as methyl orange, methyl red, ponceau BS, orange G, amaranth, orange II, direct blue, Sudan III, and Sudan IV under anaerobic growth with accumulation of toxic aromatic amines [35, 36]. In this study, Lactobacillus BT20 reached a high decolorization of indigo dye under aerobic growth possibly an azoreductase-mediated decolorization.
Bacillus BT9 showed a poor decolorization in the presence of glucose, decolorizing barely 20% (Fig. 3). In contrast in the absence of glucose, the cell growth correlated well with the color removal reaching the highest decolorization of 59%. Bacillus BT9 modified the blue-colored dye to red-colored metabolites, a behavior previously reported for laccase-catalyzed decolorization [34, 37]. In literature, expression of laccases (EC. 1.10.3.2) by the Bacillus genus has been extensively reported with application in decolorization of azo dyes such as indigo carmine, reactive black 5, Congo red, reactive red 11, and reactive blue 171 [21, 37,37,38,40]. The Bacillus species also perform a reductive decolorization of indigo carmine with the formation of indoline sulfonic acid [41]. Bacillus BT5 and Lysinibacillus BT32 exhibit a similar growth behavior, decolorizing barely 10% of indigo dye in the presence of glucose (Fig. 3). This poor performance could indicate a removal mechanism by physical adsorption [42]. In the absence of glucose, Bacillus BT5 and Lysinibacillus BT32 increased the percentages of removal to 30 and 36%, respectively.
Most of the previous studies demonstrated that Bacillus is an efficient degrader of indigo dye with percentages of decolorization up to 87% (Table 2). In contrast, in this study, the Bacillus spp. showed lower decolorizing capacities than Lactobacillus BT20. It is important to stress that all these isolates, Bacillus, Lysinibacillus, and Lactobacillus, belonged to an acclimated consortium that together with other bacteria produced hydrogen and decolorized the denim textile effluents. However, further studies should confirm if this acclimated consortium degraded the azo dyes into amines.
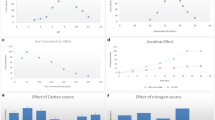
Figure 4 displays the growth kinetic during the decolorization process of indigo dye in the presence of Aspergillus H1T. Parallel to the glucose consumption, this fungus formed blue-colored spherical masses of mycelium during the first 2 days, the time during which it reached 96% of decolorization. In the absence of glucose, this phenomenon took more time, a behavior previously reported for Aspergillus ochraceus NCIM-1146 [47]. The Aspergillus genus expresses lignin-peroxidases and laccases for decolorizing indigo dye [46,46,48]. Recently, Silva et al. [45] reported the decolorization of textile wastewater containing indigo dye using Aspergillus niger AN 400 in a sequential batch bioreactor. Also, ligninolytic basidiomycetes fungi such as Phellinus, Pleurotus, Pycnoporus, and Trametes showed notable decolorizing capacities of indigo dye, the expression of laccases responsible for decolorizing are associated with the secondary metabolism (Table 2; [31, 44]).
In this study, Aspergillus H1T was isolated from fermented textile effluents; similarly, Aspergillus lentulus was isolated from textile effluents [49]. It is uncertain if Aspergillus H1T was an active decolorizing microorganism during the operation of the H2-producing bioreactor. However, previous evidence suggests that Aspergillus is ubiquitous in anaerobic environments such as the gastrointestinal tract of beef cattle [50], a similar source to the seed used for the H2-producing bioreactor. The relevance of this finding is that Aspergillus has important cellulolytic activity convenient for treating textile effluents containing cellulose fibers. Next-generation sequencing technologies for transcriptome assembly could elucidate if filamentous fungi are responsible for hydrolyzing and decolorizing of textile effluents.
Conclusions
Biological treatment of textile effluents is moving towards the reuse and recovery of value-added products. Here, denim textile effluents served as a substrate for producing bioenergy in the form of hydrogen, with the simultaneous removal of color. Different microorganisms were isolated from the fermented effluents where the decolorizing took place. From these isolates, Lactobacillus sp. BT20 and Aspergillus sp. H1T had the highest decolorizing capacities in the presence of glucose. However, further studies are necessary to test the toxicity of the treated denim effluents.
References
Comisión Nacional del Agua (CONAGUA) (2011) Situación del subsector de agua potable, alcantarillado y saneamiento, Edición 2011 (situation of the subsector of drinking water, sewerage and sanitation, edition 2011), Mexico
Verma AK, Dash RR, Bhunia P (2012) A review on chemical coagulation/flocculation technologies for removal of colour from textile wastewaters. J Environ Manag 93(1):154–168
Rai HS, Bhattacharyya MS, Singh J, Bansal TK, Vats P, Banerjee UC (2005) Removal of dyes from the effluent of textile and dyestuff manufacturing industry: a review of emerging techniques with reference to biological treatment. Crit Rev Environ Sci Technol 35(3):219–238
Gomaa OM, Momtaz OA, El Kareem HA, Fathy R (2011) Isolation, identification, and biochemical characterization of a brown rot fungus capable of textile dye decolorization. World J Microbiol Biotechnol 27(7):1641–1648
Gajera HP, Bambharolia RP, Hirpara DG, Patel SV, Golakiya BA (2015) Molecular identification and characterization of novel Hypocrea koningii associated with azo dyes decolorization and biodegradation of textile dye effluents. Process Saf Environ 98:406–416
Sahasrabudhe MM, Saratale RG, Saratale GD, Pathade GR (2014) Decolorization and detoxification of sulfonated toxic diazo dye CI Direct Red 81 by Enterococcus faecalis YZ 66. J Environ Health Sci Eng 12(1):151. https://doi.org/10.1186/s40201-014-0151-1
Wang J, Lu L, Feng F (2017) Improving the indigo carmine decolorization ability of a Bacillus amyloliquefaciens laccase by site-directed mutagenesis. Cat 7(9):275
Neetha JN, Ujwal P, Sandesh K, Santhosh H, Girish K (2018) Aerobic biodegradation of acid blue-9 dye by Bacillus fermus isolated from Annona reticulata. Environ Technol Innov 11:253–261
Pensupa N, Leu SY, Hu Y, Du C, Liu H, Jing H, Wang H, Sze C, Lin K (2017) Recent trends in sustainable textile waste recycling methods: current situation and future prospects. Top Curr Chem 375(5):76
Lay CH, Kuo SY, Sen B, Chen CC, Chang JS, Lin CY (2012) Fermentative biohydrogen production from starch-containing textile wastewater. Int J Hydrogen Energ 37(2):2050–2057
Li YC, Chu CY, Wu SY, Tsai CY, Wang CC, Hung CH, Lin CY (2012) Feasible pretreatment of textile wastewater for dark fermentative hydrogen production. Int J Hydrogen Energ 37(20):15511–15517
Lin CY, Chiang CC, Nguyen TML, Lay CH (2017) Enhancement of fermentative biohydrogen production from textile desizing wastewater via coagulation-pretreatment. Int J Hydrogen Energ 42(17):12153–12158
Lin CY, Chiang CC, Nguyen TML, Lay CH (2017) Continuous biohydrogen production from coagulation-pretreated textile desizing wastewater. Int J Hydrogen Energ 42(49):29159–29165
Pérez-Rangel M, Quiroz-Figueroa FR, González-Castañeda J, Valdez-Vazquez I (2015) Microscopic analysis of wheat straw cell wall degradation by microbial consortia for hydrogen production. Int J Hydrogen Energ 40(1):151–160
Valdez-Vazquez I, Poggi-Varaldo HM (2009) Hydrogen production by fermentative consortia. Renew Sust Energ Rev 13(5):1000–1013
Valdez-Vazquez I, Sparling R, Risbey D, Rinderknecht-Seijas N, Poggi-Varaldo HM (2005) Hydrogen generation via anaerobic fermentation of paper mill wastes. Bioresour Technol 96(17):1907–1913
Lara-Vázquez A R, Sánchez A, Valdez-Vazquez I (2014) Hydration treatments increase the biodegradability of native wheat straw for hydrogen production by a microbial consortium. Int J Hydrogen Energy39(35):19899–19904
Ramya M, Anusha B, Kalavathy S (2008) Decolorization and biodegradation of indigo carmine by a textile soil isolate Paenibacillus larvae. Biodegradation 19(2):283–291
Miller GL (1959) Use of determination of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428
Burbano E, Sierra S, Torres K, Mercado M, Carrascal A, Poutou R (2006) Rapid DNA extraction and PCR validation for direct detection of Listeria monocytogenes in raw milk. Rev MVZ Córdova11(1):715–724
Yoon JH, Lee ST, Kim SB, Kim WY, Goodfellow M, Park YH (1997) Restriction fragment length polymorphism analysis of PCR amplified 16S ribosomal DNA for rapid identification of Saccharomonospora strains. Int J Syst Bacteriol 47(1):111–114
Luo G, Mitchell TG (2002) Rapid identification of pathogenic fungi directly from cultures by using multiplex PCR. J Clin Microbiol 40(8):2860–2865
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Tamura K (1992) Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol Biol Evol 9:678–687
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Lay CH, Sen B, Kuo SY, Chen CC, Lin CY (2014) Biohydrogen production from textile wastewater by mixed microflora in an intermittent-flow, stirred tank reactor: effect of feeding frequency. J Chin Chem Soc 61(7):791–796
Arimi MM, Knodel J, Kiprop A, Namango SS, Zhang Y, Geißen SU (2015) Strategies for improvement of biohydrogen production from organic-rich wastewater: a review. Biomass Bioenergy 75:101–118
Wong Y, Yu J (1999) Laccase-catalyzed decolorization of synthetic dyes. Water Res 33:3512–3520
Huang G, Wang W, Liu G (2015) Simultaneous chromate reduction and azo dye decolourization by Lactobacillus paracase CL1107 isolated from deep sea sediment. J Environ Manag 157:297–302
Lade H, Kadam A, Paul D, Govindwar S (2015) Biodegradation and detoxification of textile azo dyes by bacterial consortium under sequential microaerophilic/aerobic processes. EXCLI J 14:158–174
Younes SB, Sayadi S (2013) Detoxification of indigo carmine using a combined treatment via a novel trimeric thermostable laccase and microbial consortium. J Mol Catal B Enzym 87:62–68
Seesuriyachan P, Takenaka S, Kuntiya A, Klayraung S, Murakami S, Aoki K (2007) Metabolism of azo dyes by Lactobacillus casei TISTR 1500 and effects of various factors on decolorization. Water Res 41(5):985–992
Chen H, Xu H, Heinze TM, Cerniglia CE (2009) Decolorization of water and oil-soluble azo dyes by Lactobacillus acidophilus and Lactobacillus fermentum. J Ind Microbiol Biotechnol 36(12):1459–1466
Reiss R, Ihssen J, Thöny-Meyer L (2011) Bacillus pumilus laccase: a heat stable enzyme with a wide substrate spectrum. BMC Biotechnol 11(1):9
Lu L, Zhao M, Li GF, Li J, Wang TN, Li DB, Xu TF (2012) Decolorization of synthetic dyes by immobilized spore from Bacillus amyloliquefaciens. Catal Commun 26:58–62
Lončar N, Gligorijević N, Božić N, Vujčić Z (2014) Congo red degrading laccases from Bacillus amyloliquefaciens strains isolated from salt spring in Serbia. Int Biodeterior Biodegradation 91:18–23
Cho EA, Seo J, Lee DW, Pan G (2011) Decolorization of indigo carmine by laccase displayed on Bacillus subtilis spores. Enzym Microb Technol 49(1):100–104
Li HX, Xu B, Tang L, Zhang JH, Mao ZG (2015) Reductive decolorization of indigo carmine dye with Bacillus sp. MZS10. Int Biodeterior Biodegradation 103:30–37
Tony BD, Goyal D, Khanna S (2009) Decolorization of textile azo dyes by aerobic bacterial consortium. Int Biodeterior Biodegradation 63(4):462–469
Manu B, Chaudhari S (2003) Decolorization of indigo and azo dyes in semicontinuous reactors with long hydraulic retention time. Process Biochem 38(8):1213–1221
Balan DSL, Monteiro RTR (2001) Decolorization of textile indigo dye by ligninolytic fungi. J Biotechnol 89:141–145
Silva KML, Wanderley CRP, Marinho G, Oliveira JCD, Santos ADOD, Rodrigues K (2015) Influence of excess nitrogen in the treatment of the textile effluent in sequential batch reactors with Aspergillus niger AN 400. Eng Sanit Ambient 20(4):635–643
Khelifi E, Ayed L, Bouallagui H, Touhami Y, Hamdi M (2009) Effect of nitrogen and carbon sources on indigo and Congo red decolourization by Aspergillus alliaceus strain 121C. J Hazard Mater 163:1056–1062
Parshetti GK, Kalme SD, Gomare SS, Govindwar SP (2007) Biodegradation of reactive blue-25 by Aspergillus ochraceus NCIM-1146. Bioresour Technol 98(18):3638–3642
Jin X, Liu G, Xu Z, Wen Y (2007) Decolorization of a dye industry effluent by Aspergillus fumigatus XC6. Appl Microbiol Biotechnol 74:239–243
Kaushik P, Malik A (2011) Process optimization for efficient dye removal by Aspergillus lentulus FJ172995. J Hazard Mater 185(2–3):837–843
Abrão FO, Duarte ER, Pessoa MS, Santos VL, Freitas Júnior LF, Barros KO, Hughes AFDS, Silva TD, Rodriguez NM (2017) Notable fibrolytic enzyme production by Aspergillus spp. isolates from the gastrointestinal tract of beef cattle fed in lignified pastures. PLoS One 12(8):e0183628
Funding
This work was supported by the PROMEP/103.5/12/3680 and UGto-DAIP 000105/11. K.M. Muñoz-Páez acknowledges the support from CONACYT through the CÁTEDRAS program (Researcher ID 6407, Project 265).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Responsible Editor: Cyntia Canedo Silva
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
- Hydrogen production and decolorization of denim effluents were achieved.
- Four bacteria and one fungus were isolated from the fermented effluent.
- Lactobacillus BT20 removed indigo dye completely using glucose as co-substrate.
- Bacillus BT9 used the dye as the sole carbon source and removed 60% of color.
- Aspergillus H1T decolorized indigo dye faster than the bacteria isolated.
Rights and permissions
About this article
Cite this article
Valdez-Vazquez, I., Robledo-Rizo, J.G., Muñoz-Páez, K.M. et al. Simultaneous hydrogen production and decolorization of denim textile wastewater: kinetics of decolorizing of indigo dye by bacterial and fungal strains. Braz J Microbiol 51, 701–709 (2020). https://doi.org/10.1007/s42770-019-00157-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-019-00157-4