Abstract
Halophilic archaea are multi-stress resistant organisms and their enzymes are of special interest as they are generally stable and functional under extreme conditions of temperature and low water activity. The search for novel extremozymes is an ongoing one because of their superior functionality in extreme conditions encountered in various industries. In this study, extremely halophilic archaea were isolated from two salterns (Marakkanam and Vedaranyam) in Tamil Nadu and from three salterns (Sinquetim, Siridao and Ribandar) in Goa, India. All isolates were Gram negative and their pigmentation ranged from light pink to bright orange. Characterization of the ten halophilic archaeal isolates was carried out by morphological, biochemical and molecular techniques. They spanned 6 different genera; Haloferax, Halorubrum, Halococcus, Haloarcula, Halogeometricum, and Haloterrigena. These extremely halophilic archaeal strains were screened for production of hydrolytic enzymes like amylase, esterase, lipase, protease, pectinase and cellulase. Amylase production by Halogeometricum sp. is being reported for the first time. All isolates showed at least one enzyme activity. Halococcus sp., Haloarcula sp., and Haloferax sp. were capable of producing three extracellular enzymes each. Cellulolytic activity was not observed in any of the isolates. Since these enzymes are inherently salt tolerant, they are very promising from the industrial point of view.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The industrial enzymes market is estimated to be valued at $4.61 billion in 2016 and is projected to grow at a Compound Annual Growth Rate (CAGR) of 5.8% from 2017 to 2022 (Markets and Markets 2016). The global market for industrial enzymes is segmented into cleaning agents, animal feed, food and beverages and biofuel production, and was valued at $3.1 billion in 2010 (BBC research 2012). The major enzymes involved in these applications can be classified broadly as carbohydrases (amylase, pectinase, cellulase, xylanase, etc.), proteases, lipases and others. Carbohydrases represent the fastest growing section in terms of growth rate. With newer applications such as egg processing, protein fortifying being developed, the search for novel and exotic enzymes from unexplored sources is gaining ground (Markets and Markets 2014). Most industrial processes are carried out in harsh physiological conditions such as extremes of pH, temperature, pressure, low water activity, etc. Extremozymes or enzymes from extremophilic microorganisms which have their optima at various extremes are the ideal candidates (Kakhki et al. 2011).
Halophiles form a relatively unexplored group of extremophiles compared to thermophiles, alkaliphiles and acidophiles. Though they have been used traditionally for solar salt production and even for processing of foodstuffs, few products from halophiles (for example, beta carotene from Dunaliella and ectoine from moderately halophilic bacteria) have been able to penetrate the global market (Oren 2010). Halotek, a German company established in 2016, has started marketing products like bacteriorhodopsin, membrane lipids and carotenoids from Halobacterium salinarium (http://halotek.de). However, no haloarchaeal enzyme has reached the level of commercial production and distribution till date.
Halophilic archaea cope with several stresses simultaneously such as, high salinity of up to 30%, high temperature, ultra-violet radiations and pH variations. Time and again, it has been proven that, the enzymes from these organisms display polyextremophilicity, i.e., the ability to function optimally in more than one of these extreme conditions (Karan et al. 2012). The halophilic enzymes or ‘halozymes’ function optimally under ‘dry’ or low water activity condition; the condition most frequently encountered in synthetic chemistry as well as in various chemical purification methods. It has been proved that the high salt required to maintain the function and structure of the halozymes may be partially replaced by organic solvents such as DMSO (Karan et al. 2012; Alsafadi and Paradisi 2013). Hence, it is very important to identify new sources and study these unique enzymes as well as their producers in detail to fine tune the catalytic activity in order to make an entry in the industrial level.
The enzymatic potential of the various halophilic organisms isolated from hypersaline niches of the Indian subcontinent has hardly been explored (Kumar et al. 2012). This study focuses mainly on the isolation of extremely halophilic archaea from various salt pans of India, their characterization by morphological, biochemical and molecular techniques as well as screening for the production of various hydrolytic enzymes.
Materials and methods
Sample collection and growth conditions
Samples were collected from three salt pans in Goa, i.e., Sinquetim (15°14′N; 73°57′E), Siridao (15°26′N, 73°52′E), Ribandar (15°30′N, 73°51′E), as well as from Marakkanam (12°12′N; 79°52′E) and Vedaranyam (10°35′N; 79°78′E) salt pans in Tamil Nadu. Samples obtained were plated out directly on extremely Halophilic (EHM) agar plates containing (g/l) NaCl 250.0; MgSO4.7H2O 20.0; CaCl2 2H2O 0.36; KCl 2.0; NaHCO3 0.06; NaBr 0.23; peptone 5.0; yeast extract 10.0; FeCl3 6H2O in traces, modified Moderately Halophilic Medium (MHM) containing (g/l) NaCl 178.0; MgSO4 7H2O 1.0; CaCl2 2H2O 2.0; KCl 2.0; NaHCO3 0.06; NaBr 0.23; peptone 5.0; yeast extract 10.0; FeCl3 6H2O in traces and JCM 168 containing (g/l) NaCl 200.0; casamino acid 5.0; yeast extract 5.0; trisodium citrate 3.0; sodium glutamate 1.0; MgSO4 7H2O 20.0; KCl 2.0; FeCl2 4H2O 0.03 and MnCl2 4H2O 0.36 mg. The pH of all media was adjusted to 7.0–7.2 using 1 M NaOH. The plates were incubated for 10–15 days at room temperature. The colonies obtained were purified by successive re-streaking till single colonies were obtained. The pure cultures obtained were grown in the respective nutrient media in which they were isolated. The isolates were stored at room temperature and sub-cultured once a month.
Identification of the isolates
The isolates were identified and characterized using various morphological (Gram staining and scanning electron microscope (SEM) analysis), biochemical (polar lipid analysis, carbohydrate utilization and pigment studies) and molecular techniques (16S rRNA gene sequencing).
Morphological characterization
For Gram staining, cell suspensions were prepared on clean glass slides in a drop of 15% (w/v) NaCl solution and air-dried. The cells were desalted with 2% acetic acid (Dussault 1955) followed by standard Gram staining and observed using phase contrast microscope (100× objective; Olympus BX41) (Mani et al. 2012).
For Scanning Electron Microscopic (SEM) analysis, 50-100 µl of cell suspension in 15% (w/v) NaCl solution was mounted onto glass coverslips, air dried and desalted with 2% acetic acid (Dussault 1955). The cells were fixed with 2% gluteraldehyde. The cells were dehydrated using a series of increasing concentrations (10–100%) of acetone (Mani et al. 2012). These dehydrated samples were dried further using critical point dryer (CPD) (Leica EM CPD 300, Austria). The sample was coated with Gold-Palladium using Leica EM ACE-200 sputter coater and mounted onto aluminum stubs for analysis using SEM (Quanta FEG 250, The Netherland).
Biochemical characterization
Polar lipid analysis
Haloarchaeal isolates in their exponential phase of growth (5–6 days old culture) were centrifuged at 10,000 rpm for 10 min to separate the cells. The cell pellets obtained were suspended in a mixture of chloroform: methanol in the ratio 1:2 (v/v). The extraction process was carried out for 4–6 h. The suspension was centrifuged (10,000 rpm, 20 min) and the supernatant was stored in a dark container. The pellet was vortexed with methanol: chloroform: water in the ratio 2:1:0.8 (v/v). The supernatant obtained after centrifugation (10,000 rpm, 20 min) of this suspension was pooled with the stored supernatant and a mixture of chloroform: water (1:1 v/v) was added to achieve phase separation. The organic phase containing the lipids was separated completely by centrifugation, collected in a clean dry glass container, and allowed to dry by evaporation. Thin layer chromatography (TLC) was carried out on silica gel plates (Silica gel 60 F254, Merck). The dried lipids were resuspended in chloroform and spotted on the TLC plates. Chloroform:methanol:acetic acid:water (85:22.5:10:4 v/v) was used to separate the lipids (Oren et al. 1996; Litchfield et al. 2000; Elevi et al. 2004). Glycolipids were visualized by spraying with 0.5% α-napthol in 50% methanol–water followed by 5% H2SO4 in ethanol and spots were detected by heating the TLC plate at 100 °C. Phospholipids were visualized by spraying molybdenum blue spray reagent (Sigma-Aldrich) (Bligh and Dyer 1959; Kates 1977).
Carbohydrate utilization
A stock (10% w/v) of various sugars (viz., glucose, sucrose, fructose, mannitol, maltose, sorbitol, and lactose) were prepared and sterilized separately at 121 °C for 15 min. For qualitatively analyzing the carbohydrate utilization, each of this sugar solution was added to sterilized Norberg and Hofstein medium to obtain a final concentration of 1%. Growth was observed by turbidity. The results were noted after 10–15 days of growth at 37 °C (Balkrishna 2015).
Pigment studies
The cells were pelleted by centrifugation at 10,000 rpm, 4 °C for 15 min. Chloroform: methanol in the ratio 1:2 (v/v) was added to the wet cell pellet and vortexed for 5 min. The suspension was centrifuged and the supernatant was taken for pigment analysis. The pigment was analyzed by spectrophotometric scan in the range of 200–800 nm using a UV–visible double beam spectrophotometer (Shimadzu, Japan, UV-2450) with chloroform:methanol (1:2) as the blank. The extraction procedure was carried out in dim light and the pigment extracts were collected in tubes covered with aluminum foil (as the pigments tend to get photobleached).
Molecular characterization
Genomic DNA extraction
One ml of culture broth (5–6 days old) was centrifuged at 5000 rpm for 5 min. The pellet was resuspended in 200 µl of sterile distilled water. To this, 200 µl of buffer-saturated phenol was added and incubated at 60 °C for 60 min. This suspension was centrifuged at 8000 rpm for 5 min. Four hundred µl of cold ethanol was added to the aqueous phase for DNA precipitation. The precipitated DNA pellet was washed with 70% ethanol and resuspended in nuclease free water (Dyall-Smith 2008).
Amplification of 16S rRNA gene using PCR
The extracted genomic DNA was used as template for the amplification of 16S rRNA gene fragment with primers A109(F) AC(G/T) GCTCAGTAACACGT and 1510 (R) GGTTACCTTGTTACGACTT (Mani et al. 2012; Birbir et al. 2007). Each PCR reaction mixture contained 10× Taq buffer, 2 mM MgCl2, 10 mM of dNTPs (Sigma), 10 µM of each primer (IDT technologies, Singapore), 2U Taq Polymerase and 1 µl of template DNA. The reaction was initiated by denaturation for 5 min at 94 °C followed by 35 cycles of Denaturation (30 s at 94 °C), Annealing (40 s at 53.5 °C) and Elongation (60 s at 68 °C) and reaction was terminated after a final elongation for 5 min at 68 °C.
Amplified product was subjected to electrophoresis on a 1.5% agarose gel and was found to be approximately 1.4 kb in size. The purified product was sequenced bidirectionally, using an automated DNA sequencer (Applied Biosystems, Foster City, CA, USA). The 16S rRNA sequences were submitted to DDBJ (DNA Databank of Japan). Multiple sequence alignment was done using MUSCLE and phylogenetic tree was constructed with MEGA X by neighbor-joining method with bootstrap analysis using 1000 replicates and displayed for 100 (Tamura et al. 2011; Wright 2006; Mani et al. 2012).
Screening of isolates for hydrolytic enzyme production
The various strains were screened for production of various extracellular hydrolytic enzymes by plate assay method. Activity was checked in different types of minimal media like Norberg Hofstein (NH) media containing (g/l) NaCl 200.0, MgSO4 7H2O 10.0, KCl 5.0, yeast extract 1.0 (pH adjusted to 7.0 using 1 M KOH) and NaCl synthetic media (NSM) containing NaCl (g/l) 200.0, MgCl2 6H2O 13.0, KCl 4.0, CaCl2 2H2O 1.0, NaHCO3 0.2, NH4Cl 2.0, FeCl3 6H2O 0.005, KH2PO4 0.5 and NSM with 0.1% yeast extract (pH adjusted to 7.0 using 1 M NaOH). The media was supplemented with the substrate of the enzyme which is to be checked. Log phase culture (50 µl) was spotted on all plates except the ones for checking lipase and esterase activity where 50 µl of the culture was added to wells bored in the solid media (Kakhki et al. 2011).
Amylase activity
Plate assay was carried out in the three media described above, supplemented with 0.5, 1 and 2% (w/v) soluble starch. The different cultures were spotted on the solid medium. The plates were incubated at room temperature for 20 days and then flooded with 0.3% I2-0.6% KI solution. Amylase production is indicated by the formation of a clear zone around the culture against a blue-black background.
Protease activity
Different volumes of sterile skimmed milk was separately added to the three media (NH, NSM and NSM + 0.1% yeast extract) to get the final substrate concentrations of 0.5%/1%/2% (w/v). Cultures were spotted on the plates. The plates were incubated at room temperature for 15 days. Formation of a halo/clearance around the culture indicates the production of protease.
Lipase activity
Olive oil (0.5%/1%/2% (v/v)) was supplemented in the three media respectively. A 50 µl of culture was added into wells bored in the solid medium. The plates were observed for the presence of white sediment around the culture growth which indicates lipase production after 15 days incubation at room temperature.
Cellulase activity
Cellulase activity was checked in the minimal media, supplemented with 0.5%/1%/2% (w/v) CM cellulose. The plates were incubated at room temperature for 20 days and then flooded with 0.1% (w/v) Congo red solution. Cellulase production is indicated by the formation of a clear zone around the culture.
Pectinase activity
Plate assay was carried out in the minimal media, supplemented with 0.5%/1%/2% (w/v) pectin. The plates were incubated at room temperature for 20 days and then flooded with 0.3% I2-0.6% KI solution. Pectinase production is indicated by the formation of a clear zone around the culture against a blue-black background.
Esterase activity
Esterase activity was determined on plates supplemented with 0.5%/1%/2% (v/v) of Tween 20 and Tween 80 in minimal medium. A 50 µl of culture was added into the wells bored in the solid medium. Formation of white sediment around the culture growth indicates esterase production.
Results and discussion
Sampling sites
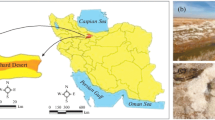
The sampling sites are from the east and west coast of India; Siridao, Ribandar and Sinquetin in Goa as well as Vedaranyam and Marakkanam in Tamil Nadu (Fig. 1). Variation in geographical locations are known to play a role in the diversity as well as characteristics of halophilic archaea (Zafrilla et al. 2010; Mani et al. 2012). Goa is lined by Arabian Sea whereas Tamil Nadu has its coastal area facing Bay of Bengal in the east and Indian Ocean in the south. The rainfall patterns are considerably different in these two states and this plays an important part in salt production pattern and the resident microbial community (Litchfield et al. 2000). Goa experiences higher seasonal fluctuation with heavy rainfall in the monsoon season (approximately 2916.7 mm in 2012) accounting for over 95% of the total annual rainfall (3048.9 mm in 2012). Hence, salt production cycle has three stages (i) monsoon period, (ii) pre-salt harvesting phase, (iii) salt harvesting phase. In this period (spanning the entire year), the salinity will increase from ~ 3% in the monsoon period to ~ 30% in the salt harvesting phase. Total annual rainfall in Tamil Nadu is less than 25% of the rainfall received in Goa (708.3 mm in 2012), and the rainfall follows a more distributed pattern with the monsoon period contributing towards 30–40% of the annual rainfall. Therefore, salt production occurs throughout the year without a drastic variation in the salinity (Mani et al. 2012; Kaur and Purohit 2012).
Sampling sites: Siridao (1), Ribandar (2), Sinquetim (3) in Goa and Marakkanam (4), Vedaranyam (5) in Tamil Nadu. Annual rainfall obtained in various parts of India is depicted in the central map (www.mapsofindia.com)
Isolation, growth and maintenance of haloarchaeal isolates
The samples obtained from various sites were plated out and visually distinct cultures were purified by repeated streaking. E1, E2, E3, E4 and E5 were isolated and maintained in EHM media, M1, M2, M3 and M5 isolated and maintained in MHM media; J1 in JCM168 media. Table 1 shows the details of each isolate with respect to the respective sampling sites, growth media, pigmentation and colony morphology. The pigmentation of most of the isolates was in different shades of orange ranging from pink, to orange and orangish red colour. The orange colouration is a common attribute associated with halophilic archaea due to the presence of bacterioruberin, a C50 carotenoid present in the membrane of the organism. The colony of all isolates were circular in shape with entire margin. Haloarchaea and extremely halophilic bacteria obtained from neutral thalassohaline environment generally have an optimum pH requirement of 6.0–8.0 (Grant 2004). Na+ and/or K+ is an obligate requirement as it helps the haloarchaea maintain osmolarity (Martin et al. 1999). Mg2+ is required for the activity of many haloarchaeal enzymes (Madern et al. 2000). Visually different cultures were purified by repeated streaking and started appearing in 7–10 days after acclimatization to the growth medium. The growth was faster if the plates were incubated at 37 °C but this caused the salts in the media to crystallize out more rapidly. Hence, room temperature was preferred for the long-term maintenance of cultures. Figure 2 shows some of the pure isolates used in the study.
Identification and characterization of haloarchaeal isolates
Morphological characterization
All halophilic isolates stained Gram negative and most appeared to be cocci in morphology when observed using a phase contrast microscope (1000× magnification). SEM imaging (Fig. 3) revealed that the size of the haloarchaeal cells ranged from 0.7 to 1.6 µm arranged as single cells, double cells, in chains or in clusters, cocci, disk shaped or highly pleomorphic morphology (Table 2). Cocci morphology is characteristic of Halococcus sp. whereas Haloarcula and Halogeometricum sp. cells are known to be triangular or even pleomorphic in shape (Montalvo-Rodriguez et al. 1998). Rod shaped and pleomorphic Halorubrum cells have also been reported (Cui et al. 2007; Trigui et al. 2011).
Biochemical characterization
Polar lipid analysis
TLC was performed to separate the polar lipids extracted from the halophilic isolates (Fig. 4). The polar lipids of haloarchaea consists of one major phospholipid and one major glycolipid accompanied by a few minor phospholipids and glycolipids. A combination of phospholipid and glycolipid structures are recognized as molecular markers for the taxonomic classification of family Halobacteriaceae (Kamekura and Kates 1999). Isolates M1, M2, E1 and E5 showed the presence of monomethylated derivative of phosphatidyl glycerophosphate (PGP-Me) as the major phospholipid and phospatidyl glycerol (PG) as a minor phospholipid accompanied by sulfated diglycosyl diether (S DGD) as the major glycolipid which suggests that they could belong to the genera Haloarcula, Haloferax, Halorubrum or Halococcus (Kamekura 1999; Kamekura and Kates 1999; Moldoveanu et al. 1990).
TLC of the total polar lipids of various haloarchaeal isolates spotted on silica gel plates and separated using chloroform: methanol: acetic acid: water (85:22.5:10:4 v/v). Phospholipids were visualized by spraying molybdenum blue spray reagent and glycolipids were visualized using α-naphthol spray followed by 5% H2SO4 in ethanol
Isolates E4 and M3 exhibited nearly identical profiles and the presence of PGP-Me, PG, S DGD, sulfated triglycosyl/tetraglycosyl diether (S TGD/S TeGD) and also a glycolipid X at an Rf of 0.01. A similar profile and an unnamed glycolipid, Glycolipid X, from Halococcus hamelinensis was described by Goh and coworkers in 2006 (Goh et al. 2006; Kamekura and Kates 1999). In isolate E3 a lack of glycolipids was noted which point towards the possibility of it belonging to the genus Natronococcus or Halogeometricum. The lipid profile of E2 showed the presence of phospholipid PGP-Me and glycolipid S DGD. Since this is found in many genera, E2 could belong to Haloarcula, Halorubrum, Haloferax, Halococcus or Haloterrigena (Kamekura and Kates 1999). Isolate M5 showed the presence of PGP- Me only and glycolipid X; very faint spots of PGP-Me and PG were detected in M4. Since these did not correspond to any known pattern, an inference regarding the identity could not be drawn for isolate M4 and M5 from the chromatogram.
The major phospholipid in several haloarchaeal genera (viz., Halobacteria, Haloarcula, Haloferax, Halococci, Natronobacteria and Natronococci) is identified to be PGP-Me which is found in all the isolates tested (Kates 1993).
Archaeal polar lipids or the phospholipid ether lipids, are composed of di- and tetraethers of glycerol (archaeols and caldarchaeols, respectively) or more complex polyols with side chains consisting of C15, C20, C25 or C40 isoprenoids. Extreme halophiles contain only archaeol derived lipids. The polar lipids of archaea are unique and readily distinguished from the acyl phospholipids of bacteria and eukaryotes. Archaeal lipids contain two phytanyl chains ether linked to glycerol or other alcohols (viz. nonitol). The acyl groups are normally fully saturated isoprenoids (De Rosa et al. 1991) containing two fatty acyl chains ester linked to a glycerol molecule with the third –OH of glycerol bound to phospho/glycol containing moieties. (Kates 1993; Gattinger et al. 2003). The ether lipids of archaea are more stable and less permeable than ester linked lipids of other forms of life. This forms one more adaptation of archaeal cells to survive in extreme conditions (Elferink et al. 1992; Thompson et al. 1992; Konings et al. 2002).
Carbohydrate utilization
The isolates were found to utilize various sugars. Fructose was metabolized by seven of the ten isolates tested, whereas, glucose was used up by only five. Lactose was utilized by three isolates and sucrose by two. Sorbitol and mannitol were utilized only by isolate E2 and maltose was not metabolized by any of the isolates. Isolate E2 was able to utilize ribose, glucose, sorbitol, mannitol, fructose, sucrose and lactose. Isolate M5 could not utilize any of the sugars tested. The results have been tabulated in Table 3.
Haloarchaea are either aerobes or facultatively anaerobic heterotrophs. Some are capable of growing in a wide range of energy sources whereas others such as Haloquadratum walsbyi have specific demands for growth (Burns et al. 2007; Bardavid and Oren 2008). The Embden–Meyerhof pathway seems to be incomplete in halophiles as inferred from an absence of phosphofructokinase. The enzymes of semi phosholylated Entner–Doudonoff pathway for glycolysis are present and are highly conserved in most sequenced haloarchaea suggesting a common ancestry. However, genome comparative analysis of several haloarchaeal genera reveal that they prefer amino acids over sugar as energy and carbon source. One exception is seen in the case of Halorhabdus utahensis which seems to be well adapted to grow on carbohydrates (Anderson et al. 2011).
Pigment analysis
Each purified halophilic isolate showed a characteristic colour when grown in nutrient media. This is attributed to the presence of pigments in them. The pigments from the haloarchaeal isolates were analyzed using UV–Visible spectrophotometer. The pigment profiles of the cultures were seen to have their λmax at 395, 472, 506 and 539 (Fig. 5) which indicates the presence of bacterioruberin class of pigments, a characteristic pigment present in halophilic archaea. The carotenoid pigments serve to protect the cells from the harmful effect of UV radiation and assists in photoreactivation (Rodrigo-Baños et al. 2015). There are very few known non-pigmented haloarchaea, for example, H. utahensis (Antunes et al. 2008) and Halarchaeum acidiphilum (Minegishi et al. 2010).
Molecular characterization
Phylogenic analysis of all the halophilic isolates was carried out using 16S rRNA gene sequencing. Among the isolates obtained from the solar salterns of India, ten cultures phylogenetically affiliated to Archaea were screened for enzyme production. These ten isolates were grouped under six different genera. Phylogenetic tree analysis (Fig. 6) and sequence comparison with the GenBank database showed that isolates M1 and E5 were most closely related to Haloferax prahovense strain TL6 (100 and 99.85% similarity), M2 and M5 belonged to the genus Halorubrum. Isolate M3 showed 99.18 similarity to Halococcus agarilyticus strain 62E whereas E4 showed 99.86% similarity to Halococcus saccharolyticus strain DSM 5350.
Phylogenetic tree indicating the positions of haloarchaeal isolates. The evolutionary history was inferred using the neighbour-joining method with bootstrap values for 100 replicates. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the maximum composite likelihood method and are in the units of the number of base substitutions per site. Evolutionary analyses were conducted in MEGA X
Isolates E1, E2 belonged to the genus Haloarcula and J1 belonged to the genus Haloterrigena. Isolate E3 was most closely related to Halogeometricum borinquense strain DSM 11551 (99.69%).
The obtained 16S rRNA sequences were submitted in DDBJ/GenBank under the accession numbers (given in brackets): E1 (AB904831), E2 (AB904832), E3 (AB904833), E4 (AB904834), E5 (AB904835), M1 (AB904836), M2 (MK810744), M3 (AB904838), M5 (AB904840), J1 (AB971350) (Table 4).
Screening of isolates for extracellular hydrolytic enzyme production
Ten haloarchaeal cultures namely, Haloferax sp. M1 and E5, Halorubrum sp. M2 and M5, Halococcus sp. M3 and E4, Haloarcula sp. E1 and E2, Halogeometricum sp. E3 and Haloterrigena sp. J1 were screened for the production of amylase, protease, lipase, esterase, pectinase and cellulase using plate assays with media containing 20% NaCl (Fig. 7).
Plate assays exhibiting amylase activity (a), protease activity (b), lipase activity (c), esterase activity (d) and pectinase activity (e). The zones of clearance indicate enzyme activity of amylase, protease and pectinase around the spotted cultures in their respective plates whereas lipase and esterase activity were indicated by the formation of white precipitate
The producers for amylase were Halorubrum sp. M2 and M5, Halococcus sp. M3 and E4, Haloarcula sp. E1 and E2, and Halogeometricum sp. E3. Out of these, E3, M5 and E1 were found to be the best producers as indicated by the zone of clearance around the colonies after adding iodine solution (Fig. 7a). Among the three media used, NH and NSMY showed maximal activity compared to NSM without yeast extract. α-Amylases (1,4-α-d-glucan glucanohydrolase, EC 3.2.1.1) are a class of industrial enzymes that find application in diverse areas such as food, laundry, brewing, textile and even in pharmaceutical industry. Many sources have been identified among halophilic and non-halophilic bacteria such as Nesterenkonia sp. and Bacillus sp., as well as eukarya such as Trametes trogii and Aspergillus niger but very few have been found among halophilic archaea. (Shafiei et al. 2010; Liu and Xu 2008; Levin et al. 2008; Mahadik et al. 2002). The amylase produced by Halorubrum sp. (Moshfegh et al. 2013) and Haloarcula sp. have been characterized but we report for the first time, the production of amylase from Halococcus sp. and Halogeometricum sp.
Protease activity was observed in E3, E4 and J1. E3 showed the best activity (Fig. 7b). Yeast extract did not have much effect on the production of protease. NSM and NSM with 0.1% yeast extract showed comparable activities while in NH media, the activity was less. Proteases find application in leather, detergent and other industries, where its stability and activity in the presence of salts and in alkaline conditions is highly desirable (Sinha and Khare 2012). In this study, we found three genera viz., Halogeometricum sp., Halococcus sp, and Haloterrigena sp. with extracellular protease activity. It was seen that the presence of yeast extract in the growth medium did not have much effect on the protease activity. This could be because the protein components of skimmed milk which was supplied as the carbon source could double up as a nitrogen supplement as well reducing the necessity of yeast extract.
Lipase activity was seen only in one of the isolates, Haloterrigena sp. J1 (Fig. 7c). It was observed that NH media having 0.5% olive oil showed maximum activity seen as a white precipitate of calcium salts of the fatty acids released. Growth of the organism and subsequent production of lipase was profoundly affected by the absence of yeast extract as seen in NS medium where lipase production was negligible. There are very few reports of true lipase activity from haloarchaea. Recently, organic solvent-tolerant lipase from Haloarcula sp. G41 and organic solvent-tolerant, detergent-stable lipase from Haloferax sp. was isolated and characterized (Li and Yu 2014; Akmoussi-Toumi et al. 2018). Though Haloferax sp. and Haloarcula sp. were screened in our study, lipase activity was not observed.
Halorubrum sp. M1, Halococcus sp. M3, Haloarcula sp. E1 and Haloferax sp. E5 showed positive result for esterase in Tween 20 while in Tween 80, Halorubrum sp. M1, Halorubrum sp. M2, Haloarcula sp. E1, Haloferax sp. E5 showed esterase activity observed as a white precipitate formed around the colonies (Fig. 7d). Haloferax sp. E5 was found to be the best producer as it produced the maximum amount of white precipitate around the colonies. Esterase activity is common among halophilic archaea like Haloarcula sp. (Camacho et al. 2009; Müller-Santos et al. 2009), Natronococcus sp. (Singh and Singh 2017), Haloferax sp. (Rathod et al. 2016), Halococcus sp. (Legat et al. 2013), Halorubrum sp. (Cui et al. 2006; Zhang and Cui 2014). Active microbial lipolytic enzymes have been discovered from marine sediments (Li and Yu 2014) as well as surface and deep-sea water (Chu et al. 2008; Fang et al. 2014), implying their potential role in marine organic carbon degradation and recycling (Zhang et al. 2017).
Moderate pectinase activity was observed in NSM medium having 0.5% pectin as the sole carbon source by three different genera (Haloferax, Halococcus and Haloarcula) and cellulase activity was shown by none of the isolates (Fig. 7e).
More than one type of hydrolytic activity was observed in many of the haloarchaeal isolates. Similar findings have been previously reported (Karray et al. 2018; Moreno et al. 2007; Rohban et al. 2009; Sánchez-Porro et al. 2003). There are several unique adaptations in their sequence and folding towards this effect (Reed et al. 2013). Halophilic proteins in general, have more of acidic residues than the basic ones with the ion pair or salt bridge being one of the most important stabilizing factors for maintaining their native conformation (Nayek et al. 2014). The drawback of using halophilic archaea for the production of these enzymes is their slow growth and production rate (Ventosa et al. 1998). But once the novel halophilic enzymes are identified and characterized, they can be overexpressed in mesophilic organisms (Promchai et al. 2018) for utilization in several industries as green and sustainable technology.
References
Akmoussi-Toumi S, Khemili-Talbi S, Ferioune I, Kebbouche-Gana S (2018) Purification and characterization of an organic solvent-tolerant and detergent-stable lipase from Haloferax mediterranei CNCMM 50101. Int J Biol Macromol 116:817–830. https://doi.org/10.1016/j.ijbiomac.2018.05.087
Alsafadi D, Paradisi F (2013) Effect of organic solvents on the activity and stability of halophilic alcohol dehydrogenase (ADH2) from Haloferax volcanii. Extremophiles 17(1):115–122
Anderson I, Scheuner C, Göker M, Mavromatis K, Hooper SD, Porat I, Klenk HP, Ivanova N, Kyrpides N (2011) Novel insights into the diversity of catabolic metabolism from ten haloarchaeal genomes. PLoS One 6(5):e20237
Antunes A, Taborda M, Huber R, Moissl C, Nobre MF, da Costa MS (2008) Halorhabdus tiamatea sp. nov., a non-pigmented, extremely halophilic archaeon from a deep-sea, hypersaline anoxic basin of the Red Sea, and emended description of the genus Halorhabdus. Int J Syst Evolut Microbiol 58(1):215–220
Balkrishna SB (2015) Synthesis of polyhydroxyalkanoates by halophilic archaea and bacteria and their osmoadaptation. Doctoral dissertation, Birla Institute of Technology and Science, Pilani)
Bardavid RE, Oren A (2008) Dihydroxyacetone metabolism in Salinibacter ruber and in Haloquadratum walsbyi. Extremophiles 12(1):125–131
BBC Research (2012) In Report BIO030G—Global markets for enzymes in industrial applications. https://www.bccresearch.com/market-research/biotechnology/enzymes-industrial-applications-markets-bio030g.html
Birbir M, Calli B, Mertoglu B, Bardavid RE, Oren A, Ogmen MN, Ogan A (2007) Extremely halophilic Archaea from Tuz Lake, Turkey, and the adjacent Kaldirim and Kayacik salterns. World J Microbiol Biotechnol 23(3):309–316
Bligh EG, Dyer WJ (1959) A rapid method of total lipid extraction and purification. Can J Biochem Physiol 37(8):911–917
Burns DG, Janssen PH, Itoh T, Kamekura M, Li Z, Jensen G, Dyall-Smith ML (2007) Haloquadratum walsbyi gen. nov., sp. nov., the square haloarchaeon of Walsby, isolated from saltern crystallizers in Australia and Spain. Int J Syst Evolut Microbiol 57(2):387–392
Camacho RM, Mateos JC, González-Reynoso O, Prado LA, Córdova J (2009) Production and characterization of esterase and lipase from Haloarcula marismortui. J Ind Microbiol Biotechnol 36(7):901–909
Chu X, He H, Guo C, Sun B (2008) Identification of two novel esterases from a marine metagenomic library derived from South China Sea. Appl Microbiol Biotechnol 80(4):615–625
Cui HL, Tohty D, Zhou PJ, Liu SJ (2006) Halorubrum lipolyticum sp. nov. and Halorubrum aidingense sp. nov., isolated from two salt lakes in Xin-Jiang, China. Int J Syst Evolut Microbiol 56(7):1631–1634
Cui HL, Lin ZY, Dong Y, Zhou PJ, Liu SJ (2007) Halorubrum litoreum sp. nov., an extremely halophilic archaeon from a solar saltern. Int J Syst Evolut Microbiol 57(10):2204–2206
De Rosa M, Trincone A, Nicolaus B, Gambacorta A (1991) Archaebacteria: lipids, membrane structures, and adaptation to environmental stresses. Life under extreme conditions. Springer, Berlin, Heidelberg, pp 61–87
Dussault HP (1955) An improved technique for staining red halophilic bacteria. J Bacteriol 70(4):484
Dyall-Smith M (2008) The halohandbook: protocols for halobacterial genetics. http://www.haloarchaea.com/resources/halohandbook/index.html. Accessed 31 Aug 2018
Elevi R, Assa P, Birbir M, Ogan A, Oren A (2004) Characterization of extremely halophilic Archaea isolated from the Ayvalik Saltern, Turkey. World J Microbiol Biotechnol 20(7):719–725
Elferink MGL, De Wit JG, Demel R, Driessen AJM, Konings WN (1992) Functional reconstitution of membrane proteins in monolayer liposomes from bipolar lipids of Sulfolobus acidocaldarius. J Biol Chem 267:1375–1381
Fang Z, Li J, Wang Q, Fang W, Peng H, Zhang X, Xiao Y (2014) A novel esterase from a marine metagenomic library exhibiting salt tolerance ability. World J Microbiol Biotechnol 24:771–780
Gattinger A, Günthner A, Schloter M, Munch JC (2003) Characterization of archaea in soils by polar lipid analysis. Eng Life Sci 23(1):21–28
Goh F, Leuko S, Allen MA, Bowman JP, Kamekura M, Neilan BA, Burns BP (2006) Halococcus hamelinensis sp. nov., a novel halophilic archaeon isolated from stromatolites in Shark Bay, Australia. Int J Syst Evolut Microbiol 56(6):1323–1329
Grant WD (2004) Life at low water activity. Philos Trans R Soc Lond B Biol Sci 359(1448):1249–1267
Kakhki AM, Amoozegar MA, Khaledi EM (2011) Diversity of hydrolytic enzymes in haloarchaeal strains isolated from salt lake. Int J Environ Sci Technol 8(4):705–714
Kamekura M (1999) Diversity of members of the family Halobacteriaceae. Microbiol Biogeochem Hypersaline Environ 13:26
Kamekura M, Kates M (1999) Structural diversity of membrane lipids in members of Halobacteriaceae. Biosci Biotechnol Biochem 63(6):969–972
Karan R, Capes MD, Das Sarma S (2012) Function and biotechnology of extremophilic enzymes in low water activity. Aquat Biosyst 8:4
Karray F, Abdallah MB, Kallel N, Hamza M, Fakhfakh M, Sayadi S (2018) Extracellular hydrolytic enzymes produced by halophilic bacteria and archaea isolated from hypersaline lake. Molecular Biology Reports 45(5):1297–1309
Kates M (1977) The phytanyl ether-linked polar lipids and isoprenoid neutral lipids of extremely halophilic bacteria. Prog Chem Fats Other Lipids 15(4):301–342
Kates M (1993) Membrane lipids of archaea. New comprehensive biochemistry, vol 26. Elsevier, Amsterdam, pp 261–295
Kaur S, Purohit MK (2012) Rainfall Statistics of India, Indian Meteorological Department, Ministry of earth sciences. Report number: ESSO/IMD/HS/R.F. REP/02 (2013)/16
Konings WN, Albers SV, Koning S, Driessen AJ (2002) The cell membrane plays a crucial role in survival of bacteria and archaea in extreme environments. Antonie Van Leeuwenhoek 81(1–4):61–72
Kumar S, Karan R, Kapoor S, Singh SP, Khare SK (2012) Screening and isolation of halophilic bacteria producing industrially important enzymes. Braz J Microbiol 43(4):1595–1603
Legat A, Denner E, Dornmayr-Pfaffenhuemer M, Pfeiffer P, Knopf B, Claus H, Stan-Lotter H (2013) Properties of Halococcus salifodinae, an isolate from Permian rock salt deposits, compared with halococci from surface waters. Life 3(1):244–259
Levin L, Herrmann C, Papinutti VL (2008) Optimization of lignocellulolytic enzyme production by the white-rot fungus Trametes trogii in solid-state fermentation using response surface methodology. Biochem Eng J 39(1):207–214
Li X, Yu HY (2014) Characterization of an organic solvent-tolerant lipase from Haloarcula sp. G41 and its application for biodiesel production. Folia Microbiol 59(6):455–463
Litchfield CD, Irby A, Kis-Papo T, Oren A (2000) Comparisons of the polar lipid and pigment profiles of two solar salterns located in Newark, California, USA, and Eilat, Israel. Extremophiles 4(5):259–265
Liu XD, Xu Y (2008) A novel raw starch digesting α-amylase from a newly isolated Bacillus sp. YX-1: purification and characterization. Bioresour Technol 99(10):4315–4320
Madern D, Ebel C, Zaccai G (2000) Halophilic adaptation of enzymes. Extremophiles 4(2):91–98
Mahadik ND, Puntambekar US, Bastawde KB, Khire JM, Gokhale DV (2002) Production of acidic lipase by Aspergillus niger in solid state fermentation. Process Biochem 38(5):715–721
Mani K, Salgaonkar BB, Braganca JM (2012) Culturable halophilic archaea at the initial and crystallization stages of salt production in a natural solar saltern of Goa, India. Aquat Biosyst 8(1):15
Markets and Markets (2014) In Report Code: FB 2277 industrial enzymes market by Types (Carbohydrase, Protease, Lipase), applications (food & beverages, cleaning agents, bio-fuel, animal feed), & Geography—Global Trends & Forecasts to 2018
Markets and Markets (2016) In Report Code: FB 2277 industrial enzymes market by type (Amylases, Cellulases, Proteases, Lipases, and Phytases), application (food and beverages, cleaning agents, and animal feed), source (microorganism, plant, and animal), and Region—Global Forecast to 2022
Martin DD, Ciulla RA, Roberts MF (1999) Osmoadaptation in archaea. Appl Environ Microbiol 65(5):1815–1825
Minegishi H, Echigo A, Nagaoka S, Kamekura M, Usami R (2010) Halarchaeum acidiphilum gen. nov., sp. nov., a moderately acidophilic haloarchaeon isolated from commercial solar salt. Int J Syst Evolut Microbiol 60(11):2513–2516
Moldoveanu N, Kates M, Montero CG, Ventosa A (1990) Polar lipids of non-alkaliphilic Halococci. Biochim Biophys Acta (BBA) Lipids Lipid Metab 1046(2):127–135
Montalvo-Rodriguez RAFAEL, Vreeland RH, Oren A, Kessel M, Betancourt C, López-Garriga JUAN (1998) Halogeometricum borinquense gen. nov., sp. nov., a novel halophilic archaeon from Puerto Rico. Int J Syst Evolut Microbiol 48(4):1305–1312
Moreno ML, Mellado E, Garcia MT, Ventosa A (2007) Diversity of extreme halophiles producing hydrolytic enzymes in hypersaline habitats. Halophiles-2007 booklet, 59–60
Moshfegh M, Shahverdi AR, Zarrini G, Faramarzi MA (2013) Biochemical characterization of an extracellular polyextremophilic α-amylase from the halophilic archaeon Halorubrum xinjiangense. Extremophiles 17:1–11
Müller-Santos M, de Souza EM, Pedrosa FDO, Mitchell DA, Longhi S, Carrière F, Krieger N (2009) First evidence for the salt-dependent folding and activity of an esterase from the halophilic archaea Haloarcula marismortui. Biochim Biophys Acta (BBA) Mol Cell Biol Lipids 1791(8):719–729
Nayek A, Gupta PSS, Banerjee S, Mondal B, Bandyopadhyay AK (2014) Salt-bridge energetics in halophilic proteins. PLoS One 9(4):e93862
Oren A (2010) Industrial and environmental applications of halophilic microorganisms. Environ Technol 31(8–9):825–834
Oren A, Duker S, Ritter S (1996) The polar lipid composition of Walsby’s square bacterium. FEMS Microbiol Lett 138(2–3):135–140
Promchai R, Boonchalearn A, Visessanguan W, Luxananil P (2018) Rapid production of extracellular thermostable alkaline halophilic protease originating from an extreme haloarchaeon, Halobacterium salinarum by recombinant Bacillus subtilis. Biocatal Agric Biotechnol 15:192–198
Rathod BN, Bhatt HH, Upasani VN (2016) Extracellular Hydrolases producing Haloarchaea from Marine Salterns at Okhamadhi, Gujarat, India. Int J Curr Microbiol App Sci 5(11):51–64
Reed CJ, Lewis H, Trejo E, Winston V, Evilia C (2013) Protein adaptations in archaeal extremophiles. Archaea. https://doi.org/10.1155/2013/373275
Rodrigo-Baños M, Garbayo I, Vílchez C, Bonete MJ, Martínez-Espinosa RM (2015) Carotenoids from haloarchaea and their potential in biotechnology. Mar Drugs 13(9):5508–5532
Rohban R, Amoozegar MA, Ventosa A (2009) Screening and isolation of halophilic bacteria producing extracellular hydrolyses from Howz Soltan Lake, Iran. J Ind Microbiol Biot 36(3):333–340
Sánchez-Porro C, Martin S, Mellado E, Ventosa A (2003) Diversity of moderately halophilic bacteria producing extracellular hydrolytic enzymes. J Appl Microbiol 94(2):295–300
Shafiei M, Ziaee AA, Amoozegar MA (2010) Purification and biochemical characterization of a novel SDS and surfactant stable, raw starch digesting, and halophilic α-amylase from a moderately halophilic bacterium, Nesterenkonia sp. strain F. Process Biochem 45(5):694–699
Singh A, Singh AK (2017) Haloarchaea: worth exploring for their biotechnological potential. Biotech Lett 39(12):1793–1800
Sinha R, Khare SK (2012) Characterization of detergent compatible protease of a halophilic Bacillus sp. EMB9: differential role of metal ions in stability and activity. Bioresour Technol 145:357–361
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Thompson DH, Wong KF, Humphry-Baker R, Wheeler JJ, Kim JM, Rananavare SB (1992) Tetraether bolaform amphiphiles as models of archaebacterial membrane lipids: raman spectroscopy, phosphorus-31 NMR, x-ray scattering, and electron microscopy. J Am Chem Soc 114(23):9035–9042
Trigui H, Masmoudi S, Brochier-Armanet C, Maalej S, Dukan S (2011) Characterization of Halorubrum sfaxense sp. nov., a new halophilic archaeon isolated from the solar saltern of Sfax in Tunisia. Int J Microbiol. https://doi.org/10.1155/2011/240191
Ventosa A, Nieto JJ, Oren A (1998) Biology of moderately halophilic aerobic bacteria. Microbiol Mol Biol R 62(2):504–544
Wright AG (2006) Phylogenetic relationships within the order Halobacteriales inferred from 16S rRNA gene sequences. Int J Syst Evol Microbiol 56:1223–1227
Zafrilla B, Martínez-Espinosa RM, Alonso MA, Bonete MJ (2010) Biodiversity of Archaea and floral of two inland saltern ecosystems in the Alto Vinalopó Valley, Spain. Saline Syst 6(1):10
Zhang WJ, Cui HL (2014) Halorubrum salinum sp. nov., isolated from a marine solar saltern. Arch Microbiol 196(6):395–400
Zhang Y, Hao J, Zhang YQ, Chen XL, Xie BB, Shi M, Li PY (2017) Identification and characterization of a novel salt-tolerant esterase from the deep-sea sediment of the South China Sea. Front Microbiol 8:441
Acknowledgements
This work was supported by BITS Pilani Seed Grant 2013 to JB. DD would like to thank BITS Pilani, KK Birla Goa Campus for the fellowship. The authors thank Dr Hiroaki Minegishi, Bio-Nano Electronics Research Center, Tokyo University, Japan for identifying the haloarchaeal isolates.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Das, D., Kalra, I., Mani, K. et al. Characterization of extremely halophilic archaeal isolates from Indian salt pans and their screening for production of hydrolytic enzymes. Environmental Sustainability 2, 227–239 (2019). https://doi.org/10.1007/s42398-019-00077-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42398-019-00077-x