Abstract
Introduction
Inhaled corticosteroid (ICS) is the most widely used and effective treatment of asthma. However, some patients do not respond to ICS, which might be due to various genetic factors. Hence, understanding the genetic factors involved in the ICS response could help physicians to individualize their treatment decision and action plans for given patients. This study aimed to analyze the characteristics of corticotropin-releasing hormone receptor 1 (CRHR1) genotypes in children with asthma and the correlation between rs242941 polymorphism of CRHR1 gene and ICS responsiveness.
Methods
This prospective study included children with uncontrolled asthma, assessing their eosinophil count, IgE concentration, lung function, and fractional concentration of nitric oxide in exhaled breath (FENO) and performing CRHR1 polymorphism sequencing. The level of asthma control was assessed by asthma control test (ACT); the responsiveness of asthma treatment with ICS was evaluated by measuring the change of ACT and forced expiratory volume in 1 s (FEV1) after treatment versus at inclusion.
Results
In total, 107 patients were analyzed for CRHR1 at rs242941. Among these, 86 (80.3%) had homozygous wild-type GG, 20 (18.7%) had heterozygous GT genotypes, and 1 (1.0%) had a homozygous variant for TT. Children with personal and family history of atopy were more likely to have GT and TT genotypes. The severity of asthma was similar between children with asthma in the three groups of GG, GT, and TT genotypes of CRHR1 at rs242941. FENO level, total IgE concentration, and eosinophilic count in children with asthma were not significantly different between GG and GT genotypes. The patient with a TT homozygous variant genotype had a higher level of FENO. There was no correlation between CRHR1 polymorphism at rs242941 and asthma control evaluated by asthma control test and lung function parameters.
Conclusion
TT genotype of rs242941 in the CRHR1 gene is not frequent. Clinical and functional characteristics of children with asthma with rs242941 polymorphism of CRHR1 gene remain homogeneously similar. There is no correlation between rs242941 polymorphism and ACT or FEV1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Why carry out the study? |
The links between clinical phenotypes, relevant biomarkers, gene polymorphism, and treatment responsiveness are essential for children with asthma. |
Inhaled corticosteroids are the most widely used drugs for children with asthma, but the inhaled corticosteroid responsiveness varies between individuals. |
There are numerous factors, including genetic factors, related to corticosteroid responsiveness. |
What was learned from the study? |
TT genotype of rs242941 in corticotropin-releasing hormone receptor-1 gene is not frequently seen in children with asthma. |
There is no correlation between rs242941 polymorphism in corticotropin-releasing hormone receptor-1 gene and inhaled corticosteroid responsiveness. |
Introduction
Inhaled corticosteroid (ICS) is the most effective drug used in asthma to reduce airway inflammation, improve lung function and/or asthma symptoms, and reduce the risk of acute asthma exacerbation; it also improves quality of life in patients with asthma [1]. It has been recommended that, to achieve the best outcomes, ICS-containing controller treatment should be started after confirming the diagnosis of asthma [1,2,3]. Treatment with ICS avoids the use of short-acting beta-agonists (SABA) alone as the first-line asthma treatment; however, adherence to ICS treatment is usually poor [1]. Thus, the poor adherence to ICS with as-needed SABA in children with asthma who have infrequent symptoms should be taken into serious consideration [1].
However, more than 20% of patients with asthma are classified as having difficult-to-treat asthma and severe asthma due to different causes [4,5,6]; genetic factors are among the causes related to ICS responsiveness [7,8,9,10,11]. Genetic studies on corticosteroid responsiveness in asthma have focused on different asthma phenotypes with more than 20 target genes. Among these genes, corticotropin-releasing hormone receptor 1 (CRHR1) is a well-established candidate gene in which replacement of G by T at rs242941 in might be associated with a better response to ICS treatment [12, 13].
CRHR1 gene encodes a G-protein receptor. It plays a major role in the mechanism of stress through the synthesis of endogenous glucocorticoids and the production of catecholamines. CRHR1 might be involved in the response to exogenous glucocorticoids because its rs242941 polymorphism may be associated with improved FEV1 respiratory function after ICS treatment [13]. However, a previous study showed that, if the change of FEV1 used to define the cutoff value of a change in lung function after ICS use was less than 7.5%, rs242941 polymorphism of the gene CRHR1 was associated with a low improvement in FEV1 after ICS treatment [14].
Although ICSs are effective in treating asthma, side effects of high-dose ICS are always of concern to the patient’s family and the physician in clinical practice. Therefore, it remains necessary to study predictive genes related to clinical asthma phenotype and corticosteroid responsiveness to help physicians manage patients with asthma effectively.
The present study aimed to analyze CRHR1 genotypes in children with asthma and determine the relationship between rs242941 polymorphism of the CRHR1 gene and ICS treatment for asthma control in children.
Methods
Subjects
This study included children aged 6–17 years old who were consulted and diagnosed with asthma in the Clinical Immunology-Allergology and Asthma Department of National Children’s Hospital (Hanoi, Vietnam) from May 2015 to May 2018. The parents/guardians signed consent forms after they received an explanation of the study objectives and the intent to publish the data. Institutional review board (IRB) approval was given by the Ethics Council of National Children’s Hospital (no. 954B/BVNTW-VNCSKTE). This study was performed in accordance with the Helsinki Declaration of 1964 and its later amendments.
Inclusion Criteria
Children > 5 years old whose asthma was uncontrolled or who did not follow their daily asthma treatment for at least 1 month were included in the present study. They were examined by spirometry, exhaled NO measurements (FENO), and skin prick test (SPT). Their parents/guardians signed the IRB-approved consent forms.
The criteria for asthma diagnosis and asthma control were based on Global Initiative for Asthma (GINA) guidelines for children > 5 years old to diagnose asthma [15].
Exclusion Criteria
Children with asthma who met one of the following criteria were excluded: unable to undergo spirometry and/or exhaled NO measurement; having other chronic or acute diseases in addition to asthma; acute asthma exacerbation needing systemic corticosteroid therapy at inclusion; lack of written consent form from parent or guardian.
Study Design
This was a prospective study. All data on anthropometry, allergy status, asthma severity, blood eosinophils, total IgE concentration, skin prick test (SPT), spirometry, and fractional exhaled nitric oxide (FENO) were recorded for analysis (Fig. 1). Fluticasone propionate was used as ICS treatment for study subjects. Patients could use SABA (albuterol) as-needed bronchodilator. Asthma control test (ACT) was used to assess asthma control in study subjects (ACT < 20 points: asthma is not controlled; ACT 20–24 points: asthma is well controlled; ACT 25–27 points: asthma is fully controlled). The evaluation of ICS treatment responsiveness was based on the modifications of ACT and FEV1, and the level of asthma control was evaluated according to GINA (controlled asthma, partially controlled asthma, and uncontrolled asthma).
Laboratory Tests
Peripheral blood analysis with eosinophil count percentage and absolute value was performed by automated machines (XT-4000i-SYSMEX; Kope, Japan). The chemical luminescence technique was used to measure total IgE in peripheral blood (COBASC 501; Hitachi, Japan).
Skin prick test (SPT) with standardized respiratory allergens (Stallergenes, London, UK) was done for all included children with asthma. The negative control was 0.9% saline solution, and the positive control was 1 mg/1 mL histamine solution. Respiratory allergens were tested with Dermatophagoides pteronyssinus (Dp), Dermatophagoides farinae (Df), Blomia tropicalis (Blo), dog and cat hair, and cockroaches. SPT was considered positive when the wheal size was > 3 mm larger than the negative control.
To assess respiratory function, spirometry was performed with Koko (Inspire, Hertford, UK). Bronchial reversibility was defined as an increase in forced expiratory volume in 1 s (FEV1) ≥ 12% and > 200 mL after 15 min of using 200 μg salbutamol [15]. All spirometry parameters, including forced vital capacity (FVC), FEV1, FEV1/FVC, peak expiratory flow (PEF), and forced expiratory flow at 25–75% (FEF25–75), were recorded for analysis (Fig. 2).
FENO measurement was performed with Hypair NO (Medisoft, Sorinnes, Belgium) according to the manufacturer’s instructions. Expiratory air flow of 50 mL/s was used as recommended for FENO and classified by the following levels: FENO ≤ 20 ppb: normal; 20 < FENO ≤ 35 ppb: increased; > 35 ppb: highly increased [16].
Genotyping of CRHR1 gene rs242941 polymorphism was performed at the Laboratory for Molecular Biomedicine of the School of Medicine and Pharmacy at Vietnam National University, Hanoi, Vietnam. DNA was extracted from white blood cells collected from the peripheral blood of the study patients, as described previously [17]. Polymerase chain reaction method and DNA sequencing were used to determine single-nucleotide polymorphism (SNP) rs242941 in the CRHR1 gene using the BigDye Terminator v3.1 Cycle Sequencing Kit onABI 3500 Series Genetic Analyzer (Thermo Fisher Scientific, Waltham, MA, USA). SNP alleles were identified by employing the BioEdit Version 7.1.9 (Abbott Laboratories, Carlbad, CA, USA) for sequence alignment and analyses. Population allele and genotype frequencies were estimated on the basis of the Hardy–Weinberg principles [18].
Statistical Analyses
Sample size was calculated using sample size formula for prospective study by estimating the absolute error rate:
n is the number of asthma patients; p is the proportion of patients not responding to medication (40%, according to some studies); Δ: absolute error, 0.1; Z (1 − α/2): coefficient of confidence; α: with a probability of 95%, the index is 1.96. The minimum sample size needed for the present study was 93 patients.
SPSS software 22.0 (Chicago, IL, USA) was used to analyze recorded parameters. Qualitative variables are expressed in terms of numbers and percentages. Categorical variables are presented as mean ± standard deviation (SD) or median (min–max). Quantitative variables are presented as mean and SD. The distribution (standard, nonstandard) of variables was assessed. The chi-square test was used to compare two categorical variables. Pair comparison of mean was performed using Mann–Whitney U test.
Results
Clinical and Functional Characteristics of Children with Asthma
This study included 107 children with asthma aged > 5 years old, with a mean age of 9.2 ± 2.6 years and a predominance of males (67.3%; Table 1). In total, 76.7% of patients had a personal atopic history and 67.3% had a family atopic history (Table 1). The severity of asthma was mild and moderate (52.3% and 47.7%, respectively). Spirometry results demonstrated an average reduction of FEV1 at inclusion (80.1 ± 19.9%; Table 1). The mean level of FENO was 25.4 ± 20.1 ppb.
Frequency of Genotypes and Alleles of CRHR1 Gene in rs242941
rs242941 polymorphism in CRHR1 of 107 children with asthma was analyzed. Eighty-six (80.3%) of children had homozygous wild-type GG, 20 (18.7%) had heterozygous GT genotypes, and 1 (1%) had a homozygous variant for TT (Table 2 and Fig 2). The frequency of alleles G and T was 89.7% and 10.3%, respectively (Table 2).
Phenotype Asthma Characteristics Classified by rs242941 in CRHR1 Gene
rs242941 in the CRHR1 gene was analyzed in 107 children with asthma, including 86 patients for GG, 20 for GT, and 1 for TT, respectively (Table 3). There were no gender or BMI differences across these three genotypes (p > 0.05; Table 3). There was no difference in the percentage of personal atopic history between the three groups (GG, GT, and TT), but there were significant differences between the three groups regarding the percentage of family atopic history (GG versus GT and TT; Table 3). The asthma level was not significantly different between the three groups. However, the percentage of asthma crisis at admission (inclusion) and self-withdrawn treatment was significantly different between GG and GT groups (p = 0.019 and p = 0.029; Table 3).
There were no significant differences between the three groups for spirometry parameters, FENO, total IgE, and blood eosinophil count (Table 3). Although the level of FENO and total IgE in children with asthma with TT genotype was higher than in the two other groups, statistical significance could not be achieved owing to the limited number of study patients in the TT group (n = 1; Table 3).
ACT Scores, Asthma Control, and Modification of FEV1 after 3 Months Classified by rs242941—CRHR1 Genotype
The percentage of ACT scores (< 20 or ≥ 20) and asthma control (controlled or uncontrolled) in each genotype group (GG, GT, and TT) is presented in Table 4.
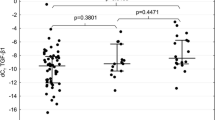
The percentage of ACT < 20 was not significant difference between GG genotype versus GT and TT (14.5% versus 20.0% and 0%; p = 0.296 and p = 0.340, respectively; Table 4). The percentage of ACT ≥ 20 also showed no significant difference between the three genotypes (Table 4). The percentage of controlled and uncontrolled asthma was not significantly different between the GG genotype versus GT and TT, and between the GT genotype versus TT (Table 4).
Discussion
The classification of asthma into phenotypes is essential for managing patients with asthma [19]. Currently, with the advances in molecular biology in emerging countries, the study of gene polymorphism in subjects with asthma is crucial to expand our knowledge of asthma pathogenesis and endotype [20, 21]. In particular, the link between clinical phenotypes, relevant biomarkers, gene polymorphism, and asthma treatment responsiveness, especially in children with asthma, should be investigated [22].
In the present study, we analyzed the rs242941 polymorphism of the CRHR1 gene in children with asthma because this gene might be related to the corticosteroid responsiveness in asthma. CRHR1 gene is located on the long chromosome 17 of position q21.31 and 13-exonsin structure. It encodes protein molecules of 415 amino acids: the G-protein receptors. These G-protein receptors combine with their neuropeptide pair (corticotropin-releasing hormone, CRH), the primary hormone controlling the hypothalamic–pituitary–adrenal axis for regulating the adrenal glands’ production of cortisol. CRHR1 might significantly influence stress-induced endogenous glucocorticoid synthesis and catecholamine production. CRHR1 gene might be also involved in exogenous glucocorticoid responses, especially in the ICS treatment of patients with asthma [13].
Analysis of rs242941 polymorphism of the CRHR1 gene in 107 patients showed genotypic and allele rates consistent with the Hardy–Weinberg equilibrium (p > 0.05). The analysis of rs242941 polymorphism revealed that 86 patients were GG homozygous and 20 patients were GT heterozygous, accounting for 80.3% and 18.7%, respectively (Table 2). In these two genotype groups, the clinical, functional, and biological parameters were not significantly different (Table 3). The GG genotype was more common in patients with a lower frequency of acute asthma exacerbation than the GT group (p < 0.05; Table 3). The present study also demonstrated only one patient with TT mutation. This patient was a 6-year-old boy with occasional urticaria. His level of eosinophilia was comparable to that of the other groups. Moreover, the level of FENO and total IgE of these children with asthma was higher than the mean FENO and the median total IgE of the two other groups (47 ppb versus 22.9 ± 18.8 ppb and 885.0 UI/mL versus 705.4 UI/mL; Table 2). There was no difference in lung function parameters at inclusion and after 3 months of treatment in children with asthma with TT mutation compared with the two other groups (Table 3).
Previous studies have demonstrated FEV1 improvements in adults and children with asthma using ICS for 6–8 weeks, in both adults (patients with TT genotype of rs242941: ΔFEV1 13.28 ± 3.11% in the homozygous TT versus 5.49 ± 1.40% in wild-type GG homozygous) and children (CAMP study: ΔFEV1 17.80 ± 6.77% in the homozygous TT versus 7.57 ± 1.5% in wild-type homozygotes) [13]. However, other studies have shown the opposite results. Firstly, Mougey et al. showed that, after 16 weeks of ICS treatment, ΔFEV1 improvement in patients with the homozygous minor allele of rs242941 was lower than in those with the homozygous major allele, in 65 patients with asthma (LOCCS study) [23]. Secondly, Roger et al. also found that the mutant allele at rs242941 of CRHR1 was associated with poorly improved respiratory function [odds ratio (OR) 1.9] and was not associated with acute asthma (OR 0.95) during follow-up for 4 years [14]. Finally, in another study focused on the long-term ICS response (20 years), Dijkstra found no association between CRHR1 polymorphisms at rs242941, rs242939, and rs1876828 and improved respiratory function in patients with asthma [24]. We suggest that these results might be due to the different criteria for evaluating FEV1 changes after treatment with ICS and the mutant group’s follow-up duration.
The main limitation of our study is its low number of recruited study patients. Thus, we found only one child with asthma who was CRHR1 mutant homozygous with the mutant allele at rs242941 of CRHR1. Although this patient’s asthma was partially controlled and had an ACT score of 21 after 3 months of treatment, we did not find a significant correlation between CRHR1 polymorphism at rs242941 and ACT or FEV1change. Therefore, it is crucial to perform a long-term study with a larger sample size to reevaluate the association of CRHR1 polymorphism with the corticosteroid response.
Conclusion
The frequency of the TT genotype of rs242941 of the CRHR1 gene is low. The clinical and functional characteristics of this genotype are similar to those of other genotypes. However, there is no significant correlation between CRHR1 polymorphism at rs242941 and ACT scores and improvement of FEV1. Studies with long-term follow-up and large sample size should be conducted to demonstrate the correlation between CRHR1 gene polymorphism and the responsiveness of asthma treatment with ICS and asthma control.
References
Global Strategy for Asthma Management and Prevention (GINA) 2022. Available at www.ginasthma.org. Last accessed: Aug 5th 2022.
Busse WW, Pedersen S, Pauwels RA, Tan WC, Chen YZ, Lamm CJ, O’Byrne PM. The Inhaled Steroid Treatment As Regular Therapy in Early Asthma (START) study 5-year follow-up: effectiveness of early intervention with budesonide in mild persistent asthma. J Allergy Clin Immunol. 2008;121:1167–74.
Selroos O. Effect of disease duration on dose-response of inhaled budesonide in asthma. Respir Med Dis. 2008;102:1065–72.
Chung KF, et al. International ERS/ATS guidelines on definition, evaluation and treatment of severe asthma. Eur Respir J. 2014;43:343–73.
Hekking PP, et al. The prevalence of severe refractory asthma. J Allergy Clin Immunol. 2015;135:896–902.
Difficult-to-treat & Severe Asthma(GINA) 2019. Available at www.ginasthma.org. Last accessed: July 7th 2022.
Tse SM, TantisiraWeiss KST. The pharmacogenetics and pharmacogenomics of asthma therapy. Pharmacogenomics J. 2011;11(6):383–92.
Drazen JM, Silverman EK, Lee TH. Heterogeneity of therapeutic responses in asthma. Br Med Bull. 2000;56(4):1054–70.
Lima JJ, Blake KV, Tantisira KG, et al. Pharmacogenetics of asthma. Curr Opin Pulm Med. 2009;15(1):57–62.
Duong-Thi-Ly H, Nguyen-Thi-Thu H, Nguyen-Hoang L, et al. Effects of genetic factors to inhaled corticosteroid response in children with asthma: a literature review. J Int Med Res. 2017;45(6):1818–30.
Nguyen ThiBich H, Duong Thi Ly H, Vu Thi T, Phan Dinh L, Le Thi Minh H, Craig TJ, Duong-Quy S. Study of the correlations between FENO in exhaled breath and atopic status, blood eosinophils, FCER2 mutation, and asthma control in Vietnamese children. J Asthma Allergy. 2016;9:163–70.
Tantisira KG, Damask A, Szefler SJ, et al. Genome-wide association identifies the T gene as a novel asthma pharmacogenetic locus. Am J Respir Crit Care Med. 2012;185:1286–91.
Tantisira KG, Lake S, Silverman ES, et al. Corticosteroid pharmacogenetics: association of sequence variants in CRHR1 with improved lung function in asthmatics treated with inhaled corticosteroids. Hum Mol Genet. 2004;13(13):1353–9.
Rogers AJ, Tantisira KG, Fuhlbrigge AL, et al. Predictors of poor response during asthma therapy differ with definition of outcome. Pharmacogenomics. 2009;10(8):1231–42.
Global Strategy for Asthma Management and Prevention (GINA) 2015. Difficult-to-Treat and Severe Asthma. Available at www.ginasthma.org. Last accessed: Sep 8th 2019.
Dweik RA, Boggs PB, Erzurum SC, et al. American Thoracic Society Committee on Interpretation of Exhaled Nitric Oxide Levels (FENO) for Clinical Applications. An official ATS clinical practice guideline: interpretation of exhaled nitric oxide levels (FENO) for clinical applications. Am J Respir Crit Care Med. 2011;184:602–15.
Nguyen-Thi-Bich H, Duong-Thi-Ly H, Thom VT, et al. Study of the correlations between fractional exhaled nitric oxide in exhaled breath and atopic status, blood eosinophils, FCER2 mutation, and asthma control in Vietnamese children. J Asthma Allergy. 2016;9:163–70.
Guo SW, Thompson EA. Performing the exact test of Hardy–Weinberg proportion for multiple alleles. Biometrics. 1992;48(2):361–72.
Wenzel SE. Asthma phenotypes: the evolution from clinical to molecular approaches. Nat Med. 2012;18:716–25.
Zhu Y, Yan X, Zhai C, Yang L, Li M. Association between risk of asthma and gene polymorphisms in CHI3L1 and CHIA: a systematic meta-analysis. BMC Pulm Med. 2017;17(1):193.
Shi F, Zhang Y, Qiu C. Gene polymorphisms in asthma: a narrative review. Ann Transl Med. 2022;10(12):711.
Dahlin A, Sordillo JE, McGeachie M, Kelly RS, Tantisira KG, Lutz SM, Lasky-Su J, Wu AC. Genome-wide interaction study reveals age-dependent determinants of responsiveness to inhaled corticosteroids in individuals with asthma. PLoS ONE. 2020;15(3): e0229241.
Mougey EB, Chen C, Tantisira KG, et al. Pharmacogenetics of asthma controller treatment. Pharmacogenomics J. 2013;13(3):242–50.
Dijkstra A, Koppelman GH, Vonk JM, et al. Pharmacogenomics and outcome of asthma: no clinical application for long-term steroid effects by CRHR1 polymorphisms. J Allergy Clin Immunol. 2008;121(6):1510–3.
Acknowledgements
The authors would like to thank the Hanoi National University–Vietnamfor the grant contributed to gene analyses (Project QG.14.59).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
No funding or sponsorship was received for this study or publication of this article.
Author Contributions
Conceptualization, H. Nguyen-Thi-Bich, H. Le-Thi-Minh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Methodology, H. Nguyen-Thi-Bich, H. Le-Thi-Minh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Validation, H. Nguyen-Thi-Bich, H. Le-Thi-Minh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Formal Analysis, H. Nguyen-Thi-Bich, L. Nguyen-Ngoc-Quynh, S. Duong-Quy; Investigation, H. Nguyen-Thi-Bich, L. Nguyen-Ngoc-Quynh, H. Le-Thi-Minh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Resources, H. Nguyen-Thi-Bich, S. Duong-Quy; Data Curation, H. Nguyen-Thi-Bich, L. Nguyen-Ngoc-Quynh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Writing – Original Draft Preparation, H. Nguyen-Thi-Bich, L. Nguyen-Ngoc-Quynh, S. Duong-Quy; Writing – Review & Editing, H. Nguyen-Thi-Bich, L. Nguyen-Ngoc-Quynh, H. Le-Thi-Minh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Visualization, H. Le-Thi-Minh, T. Nguyen-Thi-Dieu, S. Duong-Quy; Supervision,H. Le-Thi-Minh, S. Duong-Quy; Project Administration, H. Nguyen-Thi-Bich, H. Le-Thi-Minh, S. Duong-Quy.
Disclosures
H. Nguyen-Thi-Bich, T. Nguyen-Thi-Dieu, L. Nguyen-Ngoc-Quynh, H. Le-Thi-Minh, S. Duong-Quyhave nothing to disclose.
Compliance with Ethics Guidelines
This study followed the Declaration of Helsinki – Ethical Principles for Medical Research involving human subjects. This study was approved by the Ethics Council of National Children’s Hospital (954B/BVNTW-VNCSKTE, approved on 23 May 2014).
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Nguyen-Thi-Bich, H., Nguyen-Thi-Dieu, T., Nguyen-Ngoc-Quynh, L. et al. Responsiveness of Inhaled Corticosteroid Treatment in Children with Asthma: The Role of rs242941 Polymorphism of CRHR1 Gene. Pulm Ther 9, 127–137 (2023). https://doi.org/10.1007/s41030-022-00205-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41030-022-00205-9