Abstract
With more than 500 species, most of them occurring in the Neotropical region, the genus Passiflora is of key importance when one considers exploring biodiversity for food and medicinal purposes. Plants from the genus Passiflora produce fruits with economical importance, the passionfruits. Passionfruit breeding programs are leaving their infancy and molecular markers are increasingly being used to assist selection for desirable traits in novel passionfruit commercial varieties. However, the molecular and genetic basis of many of the selected characters affecting passionfruit production and quality cannot be studied in model species such as Arabidopsis thaliana, due to developmental particularities of Passiflora species. Therefore, in this review we comment on the development of genetic, genomic and transcriptomic tools that are recently becoming available for passionfruits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Passionfruit is the common name for many species belonging to the genus Passiflora. This genus is the largest one of the Passifloraceae family that encompasses more than 500 wild species distributed especially in the Neotropical region (Ulmer and MacDougal 2004). The commercial production of passionfruits is an activity of crucial economical importance for Brazil, mainly based on the yellow or sour passionfruit, Passiflora edulis. With around 50 thousand hectares producing around one million tons of fruits per year, Brazil is the largest passionfruit producer in the world, but it is also the highest consumer, absorbing near all its own production of fresh fruits, leaving insignificant amounts for export (IBGE 2017).
Passionfruit productivity in Brazil is very low and has stagnated around 14 ton per hectare for over a decade (Gonçalves and Souza 2006; Cavichioli et al. 2018). Nonetheless, there is a large discrepancy in productivity between areas where technology is incorporated by producers (more than 20 ton per hectare in Southeast Brazil) and areas where there is less investment in the orchards (around 11 ton per hectare in Northeast Brazil) (Gonçalves and Souza 2006; Cavichioli et al. 2018). Among the technologies employed for successful passionfruit commercial production is obviously genetic breeding for the development of improved varieties. Passionfruit breeders traditionally look after genotypes that produce more and larger fruits, but also that are resistant and/or tolerant to soil pathogens, viruses and abiotic stresses such as drought (Cerqueira-Silva et al. 2014; Freitas et al. 2016; Rosado et al. 2017). But more recently, there is a quest for products of aggregated valued such as fruits with different colors and flavors, ornamental varieties and varieties developed for the production of medicinal or cosmetic extracts (Abreu et al. 2009; Cavichioli et al. 2018; Cazarin 2013; Cerqueira-Silva et al. 2014; Giovannini et al. 2012; Santos et al. 2012). Therefore, understanding the genetic and molecular mechanisms that regulate these traits is not just a fascinating question of basic passionfruit physiology, but will also help to understand how these traits might be selected during passionfruit breeding and may also have important implications for the rational improvement of passionfruit quality and productivity.
2 The origin of passionfruits
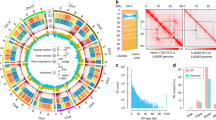
The genus Passiflora L. is the largest of the Passifloraceae family, and the majority of these are herbaceous lianas but some passionfruits are also produced by shrubs and trees. Killip (1938) and MacDougal (1994) described a high morphological diversity among species, especially when flower structures are concerned, as Passiflora flowers display a wide variation in size and color, with synapomorphic structures such as the corona and the androgynophore (Fig. 1). Coevolution with pollinators has been suggested as an explanation for this observed diversity (MacDougal 1994).
Frontal view (a, e, i) and longitudinal sections (b, f, j) of flowers at anthesis of Passiflora edulis (a, b), P. alata (e, f) and P. caerulea (i, j). External aspect (c, g, k) and longitudinal sections (d, h, l) of mature fruits of P. edulis (c, d), P. alata (g, h) and P. caerulea (k, l). Bars: a, b, e and f: 1.5 cm; c and d: 2.5 cm; g and h: 3.5 cm; i and j: 1 cm; k and l: 2 cm
Although early works suggested that the genus Passiflora should be divided into 22 or 23 subgenera (Killip 1938; Escobar 1989), there is a general agreement that only four subgenera (Passiflora, Decaloba, Astrophea and Deidamioides) are supported, as suggested by Feuillet and MacDougal (2004) and corroborated by more recent morphological, molecular and ecological data (Hansen et al. 2006; Muschner et al. 2003, 2012; Yotoko et al. 2011). The first molecular phylogeny of Passiflora, by Muschner et al. (2003), used plastidial and nuclear genome markers and clearly defined three major clades (Passiflora, Decaloba, Astrophea) with the fourth one (Deidamioides) less defined due to the small number of species classified in it. Accordingly, the available cytogenetic information on passion fruit species describes four different chromosome numbers distributed among the four subgenera of Passiflora: x = 6 in Decaloba, x = 9 and x = 10 or 11 in Passiflora, x = 10 in Astrophaea, and x = 12 in Deidamiodes (Melo et al. 2001; Melo and Guerra 2003). Although there is no agreement on the estimation of the Passiflora basic number of chromosomes, studies have suggested 6 or 12 as the basic number (Hansen et al. 2006; Melo et al. 2001). Although some species might be considered tetra- hexa- or even octoploid (Hansen et al. 2006; Melo et al. 2001), most of Passiflora species are considered diploid, with 2n = 12, 18, or 20 chromosomes. The most commercially important species, such as P. alata and P. edulis, are diploid (2n = 18), which generally allows interspecific hybrids to be obtained for genetic breeding purposes (Abreu et al. 2009; Santos et al. 2012; Souza et al. 2008).
Although divergence time studies suggest a post-Gondwanic origin of the Passifloraceae (Muschner et al. 2012), they also indicated that when Passiflora ancestors arrived in Central America they diversified quickly from there. The very ancient (~ 40 Mya) separation of Astrophea from the other Passiflora clades indicates why this subgenus contains species that present the most unusual morphological traits within Passiflora. Nonetheless the diversification age within subgenus Passiflora (~ 16.8 Mya) was much more recent than the diversification in Decaloba (~ 29 Mya; Muschner et al. 2012). Most of the cultivated passionfruit species are not considered completely domesticated, although the first reports of passionfruit cultivation (mostly for ornamental purposes) date back from the XVII century when living plants were taken from central America to Europe (Ulmer and MacDougal 2004). The Passiflora species that is mostly cultivated world-wide nowadays is certainly P. edulis, wich is a semi-woody perennial climbing vine producing round to oval fruits containing hundreds of seeds, each surrounded by an aromatic juicy aril. Two major varieties, with purple (P. edulis Sims f. edulis) and yellow (P. edulis Sims f. flavicarpa Deg.) fruits, are grown mainly for fresh fruit and/or juice production, in mild subtropical (purple variety) to warm tropical (yellow variety) climates (Menzel et al. 1987). The purple variety occurs wildly in southeast Brazil, Paraguay and northern Argentina, but the origin of the yellow variety (most commonly cultivated in Brazil) is unclear. Hybrids of these two varieties are commercially grown in Hawaii and Israel (Nave et al. 2010).
3 Approaches and tools used to study important passionfruit traits
From a commercial point of view, there are several very important passionfruit traits, depending on the purpose of cultivation. These traits include plant architecture, flower/fruit development, fruit seasonality and resistance to biotic and abiotic stresses (Cerqueira-Silva et al. 2014; Freitas et al. 2016; Rosado et al. 2017; Scorza et al. 2017). But let us not forget that Passiflora species are also highly valued for their use as ornamental and medicinal (or cosmetic) plants (Abreu et al. 2009; Cavichioli et al. 2018; Cazarin 2013; Cerqueira-Silva et al. 2014; Giovannini et al. 2012; Santos et al. 2012). During the past decade, molecular and genetic approaches started to be used to identify and functionally characterize genes and gene networks associated with important passionfruit traits (Aizza and Dornelas 2011; Rocha et al. 2015, 2016; Rosa et al. 2013a, b, c). However, until early 2000, very little information was available on gene expression in passionfruits. The very first report on gene cloning and gene expression analysis in passionfruits is the work of Mita et al. (1998) that studied fruit ripening processes and the influence of ethylene and the involvement of genes related to its biosynthesis and perception. The findings of Mita et al. (1998) could have biotechnological interest since manipulation of ethylene biosynthesis might be of interest to control fruit ripening. This is probably the only reported study on any molecular aspect of fruit development in Passiflora, although recently some groups started to be interested on the molecular basis of aril development (Silveira et al. 2016). More recently, a thorough physiological work was performed to clearly establish the role for etilene was established during the ripening process of passionfruits (Goldenberg et al. 2012) and the early molecular work by Mita et al. (1998) might contribute to elucidate the molecular aspects of this process.
Nonetheless, fruit production starts with the transition of juvenile plants to the reproductive stage followed by successful flowering. There is a great interest in understanding the molecular basis of Passiflora reproductive development, as the manipulation of flowering pathways might allow controlling fruit production. The control of flowering in Passiflora is based on the interplay between photoperiod and the balance between biosynthesis and perception of gibberellins and cytokinins (Chayut et al. 2014; Cutri et al. 2013; Nave et al. 2010; Sobol et al. 2014). Indeed, it has been shown that doubling the number of fruits per node might be a simple matter of increasing the production of cytokinins by the adult plant (Cutri et al. 2013). The molecular pathways involved in the interplay between photoperiod and hormonal balance is a key issue to be explored by Passiflora molecular biology studies in the near future as the outcome of this interplay, in model plants such as Arabidopsis thaliana, is the activation of inflorescence and flower meristem identity genes (see the review by Bloomer and Dean 2018). Regrettably, most of what is known of the molecular regulation of arabidopsis flowering might only partially be applicable to Passiflora. The study of the flowering processes in passionfruits is complicated by the shared common origins of flowers and tendrils (Cutri et al. 2013; Nave et al. 2010). Actually, Passiflora tendrils are thought to be modified flowers or inflorescences (Krosnick and Freudenstein 2005). Recent molecular characterization of floral development in Passiflora provided new molecular evidence that tendrils are indeed part of the Passiflora inflorescence and express putative orthologs of the Arabidopsis APETALA1, and FRUITFULL genes (Scorza et al. 2017). Besides pointing to the convergence of similar developmental processes involving the recruitment of genes related to flower identity in the origin of tendrils, this work clearly indicates that molecular resources are urgently needed in Passiflora as the traditional candidate gene approach using comparisons to plant models such as arabidopsis might not be useful for many passionfruit important traits.
On the other hand, very little is known about how passionfruits cope with abiotic and biotic stresses. Although some potyvirus and some aphid-borne mosaic viruses have been isolated and shown to affect passionfruit production (Barros et al. 2011; Chen et al. 2018; Yang et al. 2018), nothing is known about the molecular or physiological mechanisms involved. The key publication by Lopes et al. (2006) mapped resistance genes to Xanthomonas axonopodis pv. passiflorae in yellow passionfruit. They found quantitative resistance loci (QRLs) which explained a great deal of the total phenotypic variation. This work pointed to the importance of fine-tuning genetic mapping data for marker-assisted selection in passionfruit breeding programs. Additionally, Abreu and Aragão (2007) cloned the PeMIPS1 gene from P. edulis, that is supposed to code for an enzyme involved in the biosynthesis of all inositol-containing compounds, and showed that it is differentially regulated under cold and heat stress, presenting a light-responsive transcription, indicating that this gene might be involved with environmental stress response.
4 Use of molecular markers as tools for Passiflora breeding and for the study of genotypic diversity
The first Passiflora linkage maps were available in the early 2000s. The aim of the groups that developed these maps was to locate or tag genomic regions associated with important passionfruit traits, most of the time quantitative trait loci (QTLs). Nonetheless, linkage analysis in passionfruit has been complicated by the difficulty of obtaining inbred lines, due to self-incompatibility and high heterozygosity (Cerqueira-Silva et al. 2014). The early passionfruit linkage maps were based on the characterization of a segregant population of P. edulis using random amplified polimofic DNA (RAPD) markers (Carneiro et al. 2002). Later, the use of amplified fragment length polymorphism (AFLP) markers, allowed the detection of QRLs potentially related to passionfruit genes conferring resistance to Xanthomonas (Lopes et al. 2006). One of the first steps to produce higher quality molecular marker-based maps for passionfruits was the introduction of microsatellite markers for map construction in addition to AFLP (Cazé et al. 2012; Cerqueira-Silva et al. 2012; Padua et al. 2005; Oliveira et al. 2005). This allowed the building of a single P. edulis integrated and highly saturated map that lead to the identification of the first QTLs in passionfruit (Oliveira et al. 2008). It did not take too long for other passionfruit species, such as P. alata to have its integrated genetic map (Penha et al. 2013; Pereira et al. 2013). This map also included the use of single sequence repeats (SSRs) and, for the first time, single nucleotide polimorphisms (SNPs) with potential practical application were detected in passionfruits (Costa et al. 2017; Pereira et al. 2017).
Recently, research dedicated to the construction of the first passionfruit physical map was published (Santos et al. 2014), that lead to the sequencing of the first gene-rich regions of a commercially important passionfruit species (Munhoz et al. 2018). Together with data on expressed sequence tags (ESTs) that first approached transcriptomics in Passiflora (Cutri and Dornelas 2012), the genomic age has clearly begun for passionfruits.
5 Transcriptomics and genomics as tools for the study of Passiflora biology and for Passiflora breeding
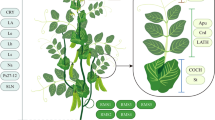
Studies dealing with transcriptomic arrived first in passionfruit literature, before actual genomic analysis. Cutri and Dornelas (2012) used the analysis of expressed sequence tags (ESTs) as an approach for the characterization of genes expressed during the reproductive development of P. edulis and P. suberosa. Combining both species, 10,272 assembled Passiflora sequences, assumed as proxy for representative gene (cDNA) sequences, were obtained. Among these sequences, many showed significant similarity to genes known to be involved in reproductive development in model plants (Cutri and Dornelas 2012). RT-PCR and in situ hybridization were used to confirm the expression patterns of some of the sequences during Passiflora flower development. Among these sequences there are P. edulis putative orthologs to the Arabidopsis KANADI and YABBY genes. In Arabidopsis, the YABBY and KANADI genes interact to establish and maintain the abaxial-adaxial polarity of all plant organs (Bowman 2000; Eshed et al. 2001). Manipulation of these gene expression patterns might alter organ size, including fruit size (Bowman 2000; Eshed et al. 2001). A second group of conserved genes that were found to express during P. edulis flower development belong to the now classical ABC model (Coen and Meyerowitz 1991). Putative Passiflora orthologs to the Arabidopsis C-class AGAMOUS gene and B-class PISTILLATA genes were found to be expressed in restricted domains of the floral whorls and were linked to the differentiation of the Passifloraceae-specific floral organ, the corona (Hemingway et al. 2011; Scorza et al. 2017). Understanding how floral morphology affects pollination might lead to knowledge that would allow incrementing pollination efficiency and thus, fruit formation. Additionally, putative orthologs to genes related to meristem behavior, important for the activity of the axillary meristems, were flowers are produced in Passiflora, were also found (Cutri and Dornelas 2012). These include members of the TCP gene family (see review by Martín-Trillo and Cubas 2010). It is known that members of the TCP family are closely linked to the process of formation of tendrils in Cucurbitaceae (Mizuno et al. 2015; Wang et al. 2015).
On the other hand, the use of suppression subtractive hybridization allowed Munhoz et al. (2015) to construct two cDNA libraries enriched for transcripts induced and repressed by the interaction of P. edulis with the passionfruit pathogen Xanthomonas axonopodis. Sixty-three out of the total of 998 obtained transcripts were similar to Arabidopsis defense-related proteins. From these, 76% (48/63) had their expression profiles changed in response to the pathogen as demonstrated by quantitative PCR data. The gene that responded most strongly to the pathogen attack encodes a lipoxygenase 2 (Munhoz et al. 2015). The detection of these novel sequences can contribute to the development of gene-based markers for important agronomic traits as well as to the establishment of genomic tools to study the naturally occurring diversity among Passiflora species.
The seminal work of Santos et al. (2014) provides an initial overview of the P. edulis genome using BAC-end sequence (BES) data. The ‘IAPAR-123’ accession was used as a donor of genomic DNA to construct the BAC library. Around 9.6% of all BES were found to have high levels of similarity to plant genes and identities could be assigned to 940 of all the analyzed sequences. The results of comparative mapping between the BES of P. edulis and the reference genomes of Arabidopsis thaliana, Populus trichocarpa and Vitis vinifera were used to choose BAC clones for full sequencing (Munhoz et al. 2018). Over 100 large-inserts were completely sequenced and structural sequence annotation resulted in the prediction of about 1900 genes. Most of them were annotated as coding proteins involved in metabolic and cellular processes and biochemically involved in binding and catalytic activity (Munhoz et al. 2018). Noteworthy, almost all of the predicted encoded proteins showed sthe highest levels of similarity to proteins from Jatropha curcas, Populus trichocarpa, Populus euphratica and Ricinus communis. The authors mention that these results were expected, since all these represent species belonging to the Malpighiales order for which fully sequenced genomes are available and they are phylogenetically close to P. edulis. Accordingly, there is a high degree of comparative microsynteny between P. edulis and both P. trichocarpa and M. esculenta. These results are the foundation of a novel era of comparative genomics in Passiflora but further studies are still necessary to access the abundance of repetitive DNA associated with gene-poor regions and the annotation of a larger number of Passiflora genes, since the (already large) number of available annotated P. edulis genes represent only 1% of the expected number of genes in the genome.
Finally the complete P. edulis chloroplast genome became available recently (Cauz-Santos et al. 2017). It is characterized by the presence of two copies of an inverted repeat sequence, each separating a two single copy regions of 13,378 bp and 85,720 bp. The annotation of the P. edulis chloroplast genome identified 105 unique genes, including 30 tRNAs, 4 rRNAs, and 71 protein coding genes (Cauz-Santos et al. 2017). These sequences might be useful as genetic markers, taking into account that chloroplast inheritance in Passiflora might be either paternal or maternal, depending on the species (Muschner et al. 2006).
6 Conclusions and future perspectives
The last decade has presented the Passiflora literature with a large number of papers that indicate that passionfruit species have entered the “omics” era where large amounts of gene and protein sequence are becoming available for functional studies and/or for their potential use as genetic markers to assist breeding programs. It will be of great interest that we will expect the unfolding of novel available genomic resources for Passiflora species, such as the complete sequencing of the P. organensis genome (unpublished results from our group) the smallest Passiflora genome with around 300 Mbp (Yotoko et al. 2011). Also as the techniques for obtaining transgenic plants is already available for passionfruits (Manders et al. 1994; Monteiro-Hara et al. 2011; Trevisan et al. 2006) genetic manipulation of candidate genes is already possible.
The challenge ahead is that many of the traits that might be interesting for passionfruit breeding will have to be studied in Passiflora species and will have limited profit from studies conducted in model plants such as Arabidopsis. This is because contrary to what is observed in Arabidopsis, passionfruits are semi-perenial plants and flowers (and thus fruits) are produced by the axillary meristems and not by the conversion of the apical shoot meristem into an inflorescence. Additionally, Arabidopsis does not form tendrils (which have a shared morphological origin with flowers in passionfruits) and produces dried dehiscent fruits (siliques) opposed to Passiflora species that produce (at least in the species of commercial interest) indehiscent fruits whose seeds are involved with a juice sac, the aril. The aril being a structure that is also absent in Arabidopsis. As for reasons of space restrictions, we intentionally neglected aspects of Passiflora biology related to the production of medicinal compounds (most of them glycosides, alakloids, flavonoids and phenolic compounds, Cazarin 2013) that, although some of them might also be present in Arabidopsis and thus share a conserved biosynthetic pathway, the final steps in their biosynthesis might involve proteins present exclusively in the Passiflora genome, as it was observed for compounds involved in the composition of rose and citrus scents (Amrad et al. 2016).
Based on the observations above, it has become clear that the model species suited for the study of passionfruit physiology and the source of gain of knowledge that will benefit genetic breeding programs are not those already available such as Arabidopsis, but must come from the genus Passiflora itself. Therefore this is the reason why passionfruit genomic resources will continue to be prized and will largely contribute to the understanding of Passiflora physiology and to breeding programs in the decades to come.
References
Abreu EF, Aragão FJ (2007) Isolation and characterization of a myo-inositol-1-phosphate synthase gene from yellow passion fruit (Passiflora edulis f. flavicarpa) expressed during seed development and environmental stress. Ann Bot 99:285–292. https://doi.org/10.1093/aob/mcl256
Abreu PP, Magalhães-Souza M, Santos E, Pires M, Pires M, Almeida AA (2009) Passion flower hybrids and their use in the ornamental plant market: perspectives for sustainable development with emphasis on Brazil. Euphytica 166:307–315. https://doi.org/10.1007/s10681-008-9835-x
Aizza LCB, Dornelas MC (2011) A genomic approach to study anthocyanin synthesis and flower pigmentation in passionflowers. J Nucleic Acids 2011:1–17
Amrad A, Moser M, Mandel T, de Vries M, Schuurink RC, Freitas LB, Kuhlemeier C (2016) Gain and loss of floral scent production through changes in structural genes during pollinator-mediated speciation. Curr Biol 26:3303–3312. https://doi.org/10.1016/j.cub.2016.10.023
Barros DR, Alfenas-Zerbini P, Beserra JE Jr, Antunes TF, Zerbini FM (2011) Comparative analysis of the genomes of two isolates of cowpea aphid-born mosaic virus (CABMB) obtained from different hosts. Arch Virol 156:1085–1091. https://doi.org/10.1007/s00705-011-0962-7
Bloomer RH, Dean C (2018) Fine-tuning timing: natural variation informs the mechanistic basis of the switch to flowering in Arabidopsis thaliana. J Exp Bot 68:5439–5452. https://doi.org/10.1093/jxb/erx270
Bowman JL (2000) Axial patterning in leaves and other lateral organs. Curr Opin Genet Dev 10:399–404
Carneiro MS, Camargo LEA, Coelho ASG, Vencovsky R, Leite RP, Stenzel NM, Vieira MLC (2002) RAPD-based genetic linkage maps of yellow passion fruit (Passiflora edulis Sims. f. flavicarpa Deg.). Genome 35:670–678
Cauz-Santos LA, Munhoz CF, Rodde N, Cauet S, Santos AA, Penha H, Dornelas MC, Varani AM, Oliveira GCX, Bergès H, Vieira MLC (2017) The chloroplast genome of Passiflora edulis (Passifloraceae) assembled from long sequence reads: structural organization and phylogenomic studies in Malpighiales. Front Plant Sci 8:334
Cavichioli JC, Meletti LMM, Narita N (2018) Aspectos da cultura do maracujazeiro no Brasil. http://www.todafruta.com.br/wp-content/uploads/2018/05/MARACUJA.pdf. Accessed 25 Sept 2018
Cazarin C (2013) Antioxidant activity of aqueous extract of passion fruit (Passiflora edulis) leaves: in vitro and in vivo study. Food Res Int 52:882–890
Cazé ALR, Kriedt RA, Beheregaray LB, Bonatto S, Freitas LB (2012) Isolation and characterization of microsatellite markers for Passiflora contracta. Int J Mol Sci 13:11343–11348
Cerqueira-Silva CBM, Santos ES, Souza AM, Mori GM, Oliveira EJ, Corrêa RX, Souza AP (2012) Development and characterization of microsatellite markers for the wild South American Passiflora cincinnata (Passifloraceae). Am J Bot 99:170–172. https://doi.org/10.3732/ajb.1100477
Cerqueira-Silva CBM, Jesus ON, Santos ESL, Corrêa RX, Souza AP (2014) Genetic breeding and diversity of the Genus Passiflora: progress and perspectives in molecular and genetic studies. Int J Mol Sci 15:14122–14152. https://doi.org/10.3390/ijms150814122
Chayut N, Sobol S, Nave N, Samach A (2014) Shielding flowers developing under stress: translating theory to field application. Plants 3:304–323. https://doi.org/10.3390/plants3030304
Chen S, Yu N, Yang S, Zhong B, Lan H (2018) Identification of Telosma mosaic virus infection in Passiflora edulis and its impact on phytochemical contents. Virol J 15:168. https://doi.org/10.1186/s12985-018-1084-6
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Costa ZP, Munhoz CF, Vieira MLC (2017) Report on the development of putative functional SSR and SNP markers in passion fruits. BMC Res Notes 10:445
Cutri L, Dornelas MC (2012) PASSIOMA: exploring expressed sequence tags during flower development in Passiflora spp. Comp Funct Genomics 2012:1–11
Cutri L, Nave N, Ami MB, Chayut N, Samach A, Dornelas MC (2013) Evolutionary, genetic, environmental and hormonal-induced plasticity in the fate of organs arising from axillary meristems in Passiflora spp. Mech Dev 130:61–69
Escobar LK (1989) A new subgenus and five new species in Passiflora (Passifloraceae) from South America. Ann Mis Bot Gard 76:877–885
Eshed Y, Baum SF, Perea JV, Bowman JL (2001) Establishment of polarity in lateral organs of plants. Curr Biol 11:1251–1260
Feuillet C, MacDougal JM (2004) A new infrageneric classification of Passiflora. Passiflora 14:1–4
Freitas JCO, Viana AP, Santos E, Paiva C, Silva F, Magalhães-Souza M (2016) Sour passion fruit breeding: strategy applied to individual selection in segregating population of Passiflora resistant to Cowpea aphid-born mosaic virus (CABMV). Sci Hortic 211:241–247. https://doi.org/10.1016/j.scienta.2016.09.002
Giovannini A, Dente F, Benedetti L, Nicoletti F, Braglia L, Gavazzi F, Mercuri A (2012) Interspecific hybridization in ornamental passion flowers. Acta Hortic 953:111–118. https://doi.org/10.17660/actahortic.2012.953.15
Goldenberg L, Feygenberg O, Samach A, Pesis E (2012) Ripening attributes of new passion fruit line featuring seasonal non-climacteric behavior. J Agric Food Chem 60:1810–1821. https://doi.org/10.1021/jf203313r
Gonçalves JS, Souza SAM (2006) Fruta da Paixão: panorama econômico do maracujá no Brasil. Inf Econ 36:29–36
Hansen AK, Gilbert LE, Simpson BB, Downie SR, Cervi AC, Jansen RK (2006) Phylogenetic relationships and chromosome number evolution in Passiflora. Syst Bot 31:138–150
Hemingway CA, Christensen AR, Malcomber ST (2011) B- and C-class gene expression during corona development of the blue passionflower (Passiflora caerulea, Passifloraceae). Am J Bot 98:923–934
IBGE. Instituto Brasileiro de Geografia e Estatística (2017) Levantamento sistemático da produção agrícola V30: 1-81.ftp://ibge.gov.br/Producao_Agricola/Levantamento_Sistematico_da_Producao_Agricola_[mensal]/Fasciculo/2017/lspa_201701.pdf. Accessed 25 Sept 2018
Killip EP (1938) The American species of Passifloraceae. Field Mus Nat Hist, Bot Ser 19:1–613
Krosnick SE, Freudenstein JV (2005) Monophyly and floral character homology of old world Passiflora (Subgenus Decaloba, Supersection Disemma). Syst Bot 30:139–152
Lopes R, Lopes MT, Carneiro MS, Fde PM, Camargo LE, Vieira ML (2006) Linkage and mapping of resistance genes to Xanthomonas axonopodis pv. passiflorae in yellow passion fruit. Genome 49:17–29
MacDougal JM (1994) Revision of Passiflora subgenus Decaloba section Pseudodysosmia (Passifloraceae). Syst Bot Monogr 41:1–46
Manders G, Otoni WC, d’Utra Vaz FB, Blackball NW, Power JB, Davey MR (1994) Transformation of passionfruit (Passiflora edulis fv flavicarpa Degener.) using Agrobacterium tumefaciens. Plant Cell Rep 13:697–702. https://doi.org/10.1007/BF00231627
Martín-Trillo M, Cubas P (2010) TCP genes: a family snapshot ten years later. Trends Plant Sci 15:31–39. https://doi.org/10.1016/j.tplants.2009.11.003
Melo NF, Guerra M (2003) Variability of the 5S and rDNA sites in Passiflora L. with species with distinct base chromosome numbers. Ann Bot 92:309–316
Melo NF, Cervi AC, Guerra M (2001) Kariology and citotaxonomy of the genus Passiflora L. Plant Syst Evol 226:68–84
Menzel CM, Simpson DR, Winks CW (1987) Effect of temperature on growth, flowering and nutrient uptake of three passionfruit cultivars under low irradiance. Sci Hortic 31:259–268
Mita S, Kawamura S, Yamawaki K, Nakamura K, Hyodo H (1998) Differential expression of genes involved in the biosynthesis and perception of ethylene during ripening of passion fruit (Passiflora edulis Sims). Plant Cell Physiol 39:1209–1217
Mizuno S, Sonoda M, Tamura Y, Eisho N, Suzuki H, Sato T, Oizumi T (2015) Chiba Tendril-Less locus determines tendril organ identity in melon (Cucumis melo L.) and potentially encodes a tendril-specific TCP homolog. J Plant Res. https://doi.org/10.1007/s10265-015-0747-2
Monteiro-Hara ACBA, Jadão AS, Mendes MBJ, Rezende JAM, Trevisan F, Mello APOA, Vieira MLC, Meletti LMM, Piedade SMS (2011) Genetic transformation of passionflower and evaluation of R1 and R2 generations for resistance to cowpea aphid borne mosaic virus. Plant Dis 95:1021–1025. https://doi.org/10.1094/PDIS-12-10-0873
Munhoz CF, Santos AA, Arenhart RA, Santini L, Monteiro-vitorello CB, Vieira MLC (2015) Analysis of plant gene expression during passion fruit-Xanthomonas axonopodis interaction implicates lipoxygenase 2 in host defence. Ann Appl Biol 167:135–155. https://doi.org/10.1111/aab.12215
Munhoz CF, Costa ZP, Cauz-Santos LA, Reátegui ACE, Rodde N, Cauet S, Dornelas MC, Leroy P, Varani AM, Bergès H, Vieira MLC (2018) A gene-rich fraction analysis of the Passiflora edulis genome reveals highly conserved microsyntenic regions with two related Malpighiales species. Sci Rep 8:13024. https://doi.org/10.1038/s41598-018-31330-8
Muschner VC, Lorenz AP, Cervi AC, Bonatto SL, Souza-Chies TT, Salzano FM, Freitas LB (2003) A first molecular phylogenetic analysis of Passiflora (Passifloracae). Am J Bot 90:1229–1238
Muschner VC, Lemke APL, Vecchia M, Bonatto S, Salzano FM, Freitas LB (2006) Differential organellar inheritance in Passiflora’s (Passifloraceae) subgenera. Genetica 128:449–453
Muschner VC, Zamberlan PM, Bonatto SL, Freitas LB (2012) Phylogeny, biogeography and divergence times in Passiflora (Passifloraceae). Genet Mol Biol 35:1036–1043
Nave N, Katz E, Chayut N, Gazit S, Samach A (2010) Flower development in the passion fruit Passiflora edulis requires a photoperiod-induced systemic graft-transmissible signal. Plant, Cell Environ 33:2065–2083. https://doi.org/10.1111/j.1365-3040.2010.02206.x
Oliveira EJ, Pádua JG, Zucchi MI, Camargo LEA, Fungaro MHP, Vieira MLC (2005) Development and characterization of microsatellite markers from the yellow passion fruit (Passiflora edulis f. flavicarpa). Mol Ecol Notes 5:331–333
Oliveira EJ, Vieira MLC, Garcia AAF, Munhoz CEF, Margarido GRA, Consoli L, Matta FP, Moraes MC (2008) An integrated molecular map of yellow passion fruit based on simultaneous maximum-likelihood estimation of linkage and linkage phases. J Am Soc Hortic Sci 133:35–41
Padua JG, Oliveira EJ, Zucchi MI, Oliveira GCX, Camargo LEA, Vieira MLC (2005) Isolation and characterization of microsatellite markers from the sweet passion fruit (Passiflora alata Curtis: Passifloraceae). Mol Ecol Notes 5:863–865
Penha HA, Pereira GS, Zucchi MI, Diniz AL, Carneiro ML (2013) Development of microsatellite markers in sweet passion fruit, and identification of length and conformation polymorphisms within repeat sequences. Plant Breed 132:731–735
Pereira GS, Nunes ES, Laperuta LDC, Braga MF, Penha HÁ, Diniz AL, Munhoz CF, Gazaffi R, Garcia AAF, Vieira MLC (2013) Molecular polymorphism and linkage analysis in sweet passion fruit, an outcrossing species. Ann Appl Biol 162:347–361
Pereira GS, Laperuta LC, Nunes ES, Chavarría L, Pastina MM, Gazaffi R, Geraldi IO, Garcia AAF, Vieira MLC (2017) The sweet passion fruit (Passiflora alata) crop: genetic and phenotypic parameter estimates and QTL mapping for fruit traits. Trop Plant Biol 10:18–29
Rocha DI, Pinto DLP, Vieira LM, Tanaka FAO, Dornelas MC, Otoni WC (2015) Cellular and molecular changes associated with competence acquisition during passion fruit somatic embryogenesis: ultrastructural characterization and analysis of SERK gene expression. Protoplasma 253:595–609. https://doi.org/10.1007/s00709-015-0837-y
Rocha DI, Monte-Bello CC, Aizza LCN, Dornelas MC (2016) A passion fruit putative ortholog of the somatic embryogenesis receptor kinase1 gene is expressed throughout the in vitro de novo shoot organogenesis developmental program. Plant Cell, Tissue Org Cult 125:107–117. https://doi.org/10.1007/s11240-015-0933-x
Rosa YBCJ, Aizza LCB, Armanhi JSL, Dornelas MC (2013a) A Passiflora homolog of a D-type cyclin gene is differentially expressed in response to sucrose, auxin, and cytokinin. Plant Cell, Tissue Org Cult 115:233–242
Rosa YBCJ, Aizza LCB, Monte-Bello CC, Dornelas MC (2013b) PmTCP1 encodes a putative TCP transcription factor and is differentially expressed during in vitro organogenesis in Passiflora. In Vitro Cell Dev Biol-Plant 50:36–44
Rosa YBCJ, Aizza LCB, Monte-Bello CC, Dornelas MC (2013c) The PmNAC1 gene is induced by auxin and expressed in differentiating vascular cells in callus cultures of Passiflora. Plant Cell, Tissue Org Cult 115:275–283
Rosado RDS, Rosado LDS, Cremasco JPG, dos Santos CEM, Dias DCFS, Cruz CD (2017) Genetic divergence between passion fruit hybrids and reciprocals based on seedling emergence and vigor. J Seed Sci 39:417–425. https://doi.org/10.1590/2317-1545v39n4183293
Santos EA, Souza MM, Abreu PP, Conceição LDHCS, Araujo IS, Viana AP, Almeida AF, Freitas JCO (2012) Confirmation and characterization of interspecific hybrids of Passiflora L. (Passifloraceae) for ornamental use. Euphytica 184:389–399. https://doi.org/10.1007/s10681-011-0607-7
Santos AA, Penha HA, Arnaud B, Munhoz C, Pedrosa-Harand A, Bergès H, Vieira MLC (2014) Begin at the beginning: a BAC-end view of the passion fruit (Passiflora) genome. BMC Genomics 15:816
Scorza LCT, Hernandes-Lopes J, Melo-de-Pinna GFA, Dornelas MC (2017) Expression patterns of Passiflora edulis APETALA1/FRUITFULL homologues shed light onto tendril and corona identities. EvoDevo 8:3. https://doi.org/10.1186/s13227-017-0066-x
Silveira SR, Dornelas MC, Martinelli AP (2016) Perspectives for a framework to understand aril initiation and development. Front Plant Sci 7:1912
Sobol S, Chayut N, Nave N, Kafle D, Hegele M, Kaminetsky R, Wünsche JN, Samach A (2014) Genetic variation in yield under hot ambient temperatures spotlights a role for cytokinin in protection of developing floral primordia. Plant, Cell Environ 37:643–657. https://doi.org/10.1111/pce.12184
Souza MM, Pereira TNS, Vieira MLC (2008) Cytogenetic studies in some species of Passiflora L. (Passifloraceae): a review emphasizing Brazilian species. Braz Arch Biol Technol 51:247–258
Trevisan F, Mendes BMJ, Maciel SC, Vieira MLC, Meletti LM, Rezende JA (2006) Resistance to passion fruit woodiness virus in transgenic passionflower expressing the virus coat protein gene. Plant Dis 90:1026–1030
Ulmer T, MacDougal JM (2004) Passiflora: passionflowers of the world. Timber Press, Portland, p 430
Wang S, Yang X, Xu M, Lin X, Lin T, Qui J, Shao JQ, Tian N, Yang Q, Zhang Z, Huang S (2015) A rare SNP identified a TCP transcription factor essential for tendril development in cucumber. Mol Plant 8:1795–1808. https://doi.org/10.1016/j.molp.2015.10.005
Yang K, Yan H, Song L, Jin P, Miao W, Cui H (2018) Analysis of the complete genome sequence of a potyvirus from passion fruit suggests its taxonomic classification as a member of a new species. Arch Virol 163:2583–2586. https://doi.org/10.1007/s00705-018-3885-8
Yotoko KSC, Dornelas MC, Togni PD, Fonseca TC, Salzano FM, Bonatto SL, Freitas LB (2011) Does variation in genome sizes reflect adaptive or neutral processes? New clues from Passiflora. PLoS ONE 6:e18212
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gioppato, H.A., da Silva, M.B., Carrara, S. et al. Genomic and transcriptomic approaches to understand Passiflora physiology and to contribute to passionfruit breeding. Theor. Exp. Plant Physiol. 31, 173–181 (2019). https://doi.org/10.1007/s40626-018-0134-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40626-018-0134-1