Abstract
Genetic material derived from tumours is constantly shed into the circulation of cancer patients both in the form of circulating free nucleic acids and within circulating cells or extracellular vesicles. Monitoring cancer-specific genomic alterations, particularly mutant allele frequencies, in circulating nucleic acids allows for a non-invasive liquid biopsy for detecting residual disease and response to therapy. The advent of molecular targeted treatments and immunotherapies with increasing effectiveness requires corresponding effective molecular biology methods for the detection of biomarkers such as circulating nucleic acid to monitor and ultimately personalise therapy. The use of polymerase chain reaction (PCR)-based methods, such as droplet digital PCR, allows for a very sensitive analysis of circulating tumour DNA, but typically only a limited number of gene mutations can be detected in parallel. In contrast, next-generation sequencing allows for parallel analysis of multiple mutations in many genes. The development of targeted next-generation sequencing cancer gene panels optimised for the detection of circulating free DNA now provides both the flexibility of multiple mutation analysis coupled with a sensitivity that approaches or even matches droplet digital PCR. In this review, we discuss the advantages and disadvantages of these current molecular technologies in conjunction with how this field is evolving in the context of melanoma diagnosis, prognosis, and monitoring of response to therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Circulating free DNA can provide non-invasive, real-time information about a patient’s tumour burden and subsequent response to therapy. |
Development of sensitive, targeted, next-generation sequencing technologies that detect tumour mutations in circulating free DNA is revolutionising the monitoring of cancer patients. |
1 Introduction
Liquid biopsy involves the extraction, detection, and analysis of DNA, RNA, proteins, vesicles, or cells derived from biofluids such as blood, urine, saliva, pleural effusions, and cerebrospinal fluid (CSF). The development of liquid biopsies for genomic profiling of solid tumours as a means of detecting actionable mutations, monitoring cancer progression/evolution, and predicting response to therapy in a non-invasive manner is a rapidly growing field (reviewed in [1,2,3,4,5,6]).
A major biomarker of tumour-related genetic changes is circulating free DNA (cfDNA), specifically, the tumour-derived circulating tumour DNA (ctDNA) fraction (reviewed in [7,8,9,10,11]). cfDNA is highly fragmented DNA with a size distribution of ~ 130–170 bp [12,13,14], which is equivalent to the size of nuclease-cleaved nucleosomes, and may arise from multiple mechanisms, including apoptosis, necrosis, and active secretion (reviewed in [15,16,17]). cfDNA can be found in many biofluids, including blood, urine, CSF, saliva, and stool. Levels of ctDNA, which often increase with tumour volume, can predict response to targeted and immunotherapies, can be used to monitor residual disease and tumour heterogeneity, and can reveal expanding therapy-resistant tumour clones (reviewed in [18,19,20,21]).
One major challenge associated with the clinical application of ctDNA arises from the variability in patient ctDNA levels that is associated with cancer stage, tumour burden and location, response to therapy, vascularity and cellular turnover. Thus, levels of ctDNA relative to the total cfDNA pool of an individual can vary from < 0.01 to > 50% [22]. Coupled with the fact that cfDNA is not abundant and has a short half-life of only a few hours, the reliable and sensitive detection of ctDNA remains challenging, especially in patients with early-stage cancer (reviewed in [23]). For blood-derived cfDNA, a number of studies have sought to optimise the yield and stability of cfDNA by comparing a range of collection tubes and other factors during blood collection [24,25,26,27,28,29,30] and a range of commercial cfDNA purification kits [31,32,33,34,35,36,37].
The methodology used to detect and analyse the genomic alterations in cfDNA, along with the strengths and weaknesses of these various approaches are the subject of a number of recent reviews [4, 10, 20, 38,39,40]. Cancer-associated alterations in cfDNA include single nucleotide variations (SNVs), rearrangement of genomic sequences, copy number variations (CNVs), microsatellite instability, loss of heterozygosity and DNA methylation (reviewed in [11]). The most common methods used to detect cfDNA can be classified into standard polymerase chain reaction (PCR)-based, digital PCR, whole-exome or targeted deep sequencing. The primary focus of this review is the application of digital PCR and targeted next-generation sequencing (NGS) gene panels to identify and monitor tumour-associated genetic changes in cfDNA isolated from cancer patients with an emphasis on cutaneous melanoma (Fig. 1). Other sources of circulating nucleic acids that have the potential to complement detection of cfDNA are also highlighted.
Comparison of ddPCR and NGS platforms for the monitoring of ctDNA in melanoma. cfDNA circulating free DNA, CNV copy number variation, ctDNA circulating tumour DNA, CSF cerebrospinal fluid, ddPCR droplet digital polymerase chain reaction, indels insertions or deletions, NGS next-generation sequencing, SNV single nucleotide variation
2 Biomarkers and Genetics of Melanoma
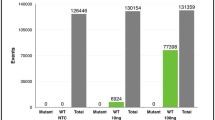
The high tumour mutation burden (TMB; i.e. the number of somatic mutations found in the genome of a single tumour) of cutaneous melanoma has been well documented, with the most significant driver mutations in BRAF, NRAS, NF1, and KIT genes [41,42,43,44,45,46,47,48,49,50]. Prior to 2010, the typical 1-year survival for stage IV melanoma patients was only 25% [51]. With the introduction of tyrosine kinase inhibitors targeting the mitogen-activated protein kinase (MAPK) pathway in patients with BRAFV600-mutated melanoma, and antibodies against immune checkpoints such as programmed cell death protein 1 (PD-1) or cytotoxic T-lymphocyte-associated antigen 4 (CTLA-4), the 1-year survival for patients receiving combination dabrafenib and trametinib, combination encorafenib and binimetinib, combination vemurafenib and cobimetinib, single-agent PD-1 antibodies, and a combination of PD-1 and CTLA-4 antibodies has improved to 72%, 75%, 75%, 73%, and 73%, respectively [52,53,54,55,56]. However, major limitations exist for both MAPK and immune checkpoint inhibitors, including the emergence of drug resistance in the majority of patients within 12 months for MAPK inhibitors and low but durable response rates for single-agent immune checkpoint inhibitors at 10–40% [5, 57,58,59,60,61,62,63]. Thus, the identification of biomarkers that can predict and monitor patient responses remains a critical unmet need, and the use of liquid biopsies and specifically cfDNA to inform patient selection and monitor patient response to treatment is a rapidly evolving area of research (reviewed in [4, 5, 10, 64, 65]).
3 PCR-Based Methods to Detect ctDNA in Melanoma
The use of PCR-based methods, particularly droplet digital (dd) PCR and BEAMing (beads, emulsion, amplification and magnetics), for the detection of ctDNA as a biomarker in metastatic melanoma and other cancers has been well documented (reviewed in [10, 66]). The potential clinical applications of PCR-based analysis of ctDNA in melanoma as a predictive biomarker, measuring tumour heterogeneity/dynamics, identifying resistance driver mutations for targeted therapy, evaluating early response to therapy, and monitoring the development of resistance to therapy, have been highlighted in many studies (reviewed in [64]). These allele-specific PCR-based methods mainly detect single driver mutations in key genes such as BRAF and NRAS. They require prior knowledge of the driver mutation, which typically comes from sequencing of the primary solid tissue biopsy using targeted pan-cancer panels that analyse mutations in genes such as BRAF and NRAS [48].
The use of ddPCR-based detection of BRAF or NRAS mutations in ctDNA from metastatic melanoma patients has illustrated an inverse correlation between ctDNA copy number and response to either targeted or immunotherapy [67,68,69,70,71,72,73,74,75,76]. A direct correlation between ctDNA levels, based on ddPCR-detection, and tumour burden has also been reported [77, 78]. The potential for ddPCR-based detection of ctDNA to replace tumour genotyping of melanomas has been postulated both for detecting major driver mutations in BRAF and NRAS [79] as well as other mutations, such as in the TERT promoter [80], which can be found in up to 70% of all melanomas [81].
The majority of studies involving ddPCR-based detection of ctDNA have been based on the isolation of ctDNA from plasma. One of the major limitations is tracking patients with brain metastases, which occur in 50–75% of all stage IV melanoma patients [82,83,84]. The low detectability of ctDNA seen in patients with predominant brain metastases suggests that the blood–brain barrier may significantly impact the release of ctDNA into the circulation (reviewed in [10]), and in fact, plasma ctDNA has been shown not to be a reliable indicator of melanoma brain metastases [71]. There are two studies which were able to successfully detect ctDNA using ddPCR in the CSF of melanoma patients with metastasis to the central nervous system [85, 86]. Detection of CSF ctDNA may provide greater sensitivity than conventional cytology and appears to reflect central nervous system-tumour burden and is an indicator of response to therapy [85, 86].
The limitations of only being able to detect one to two mutations simultaneously and the need for specialised equipment encountered with ddPCR and more traditional fluorescence-based PCR have been addressed in a recent report [87]. Alternative approaches to ddPCR have also been investigated. The combination of surface-enhanced Raman spectroscopy (SERS) nanotags combined with PCR was shown to provide similar sensitivity to ddPCR in simultaneously detecting BRAF, NRAS, and KIT mutations in ctDNA from melanoma patients [87]. The potential of nanoparticle-based sensing of mutations in ctDNA, without the need for PCR amplification, has also been highlighted [88]. Whether these approaches truly approach the same sensitivity, specificity, reproducibility, and accuracy in ctDNA quantification as ddPCR requires further validation.
A fully automated integrated platform from Biocartis (Belgium), designated Idylla, for multiplex real-time PCR-based detection of major NRAS, BRAF, KRAS, or EGFR mutations in either ctDNA from blood or genomic DNA from tissue has been developed. This platform has been validated using mainly tissue from melanoma patients, and showed > 95% concordance with other methods of detection, and as such, represents a feasible diagnostic platform given the ease of use, automation, and fast 1-day turnaround [89,90,91,92,93].
These targeted ctDNA mutation detection methods, which rely on screening previously identified tumour mutations, can be used for the real-time monitoring of patient response to therapy. However, they provide no information on tumour evolution and cannot identify newly acquired somatic mutations that may confer resistance to therapy and which may be targetable with alternative therapies. Furthermore, 20–30% of melanoma patients who have rare mutations or no identifiable mutations on standard tissue mutation testing platforms provide additional challenges regarding ctDNA mutation detection. The identification of a broader number of mutations with the detection of acquired resistance markers requires the use of NGS approaches, as discussed in the next section.
4 NGS-Based Methods to Detect ctDNA in Melanoma
The use of NGS in the context of melanoma and other cancers, whether in the form of whole-genome, whole-exome, or targeted gene panels, is well documented for tissue-derived samples [38, 49, 94,95,96,97,98,99,100]. These NGS approaches are able to interrogate mutations in many genes and provide increasing sensitivity as the sequencing becomes more targeted. The drive to complement or replace tissue-based sequencing with sequencing of ctDNA for both initial diagnosis and longitudinal assessment of cancer patients on therapy is now a major focus in the field. The less invasive sampling required for ctDNA has obvious advantages over the more invasive tissue collection, with which serial sampling may not be possible in many cases.
The use of fragmented and low copy number ctDNA as a template for NGS, compared with the more abundant higher molecular weight genomic DNA used in tissue-based NGS, has been a major limiting factor, and this has been largely overcome with advancements in sequencing technology (reviewed in [40]). All of these advancements have aimed to improve the detection sensitivity and detection specificity of monitoring rare ctDNA mutations within the larger pool of total cfDNA. However, there are continuing limitations due to varying cfDNA quality and quantity arising from non-standardised pre-analytical workflows used for both blood collection and subsequent extraction of cfDNA (reviewed in [40]).
The feasibility of whole-genome and whole-exome sequencing using ctDNA as a template to identify somatic mutations, which can subsequently be used to guide choice and response to therapy, has been demonstrated in a number of cancers, including melanoma [76, 101,102,103,104,105,106,107,108,109]. The value of using whole-genome and whole-exome sequencing, given their turnaround time, cost, and the actual need for such a broad approach, has come into question with the advent of more targeted NGS gene panels compatible with cfDNA (Table 1). The whole-genome or whole-exome sequencing approach can still be used initially to design patient-specific targeted NGS gene panels for longitudinal monitoring of therapy (reviewed in [19]).
4.1 Targeted NGS Gene Panels to Guide and Monitor Response to Therapy
Targeted NGS gene panels are typically based on proprietary technology for sequencing library preparation and are compatible with one of the three current NGS platforms: Thermofisher Ion Torrent (USA); Illumina (USA); or Roche (Switzerland). Whether pan cancer (covering major solid tumour cancers, including lung, breast, prostate, colorectal and melanoma) or cancer specific, NGS cancer panels generally include common somatic driver mutations such as those in EGFR, BRAF, NRAS, KRAS, PDGFRA, and KIT. In the case of melanoma, this can guide the choice of initial targeted MAPK inhibitor therapy, i.e. patients with BRAFV600-mutant melanoma can be effectively treated with combination inhibitors targeting mutant BRAF and its downstream effector MEK [110]. Further inclusion of known somatic resistance mutations plus oncogenic targets in downstream signalling pathways [63] can identify pathways of resistance to first-line therapy and guide the choice of alternative therapies. A practical application of NGS targeted gene panels for the parallel analysis of several genes in genotyping primary melanoma patients has been documented using genomic DNA [111]. Such a targeted panel could also be applied in the analysis of ctDNA, to provide diagnostic/prognostic information and monitor resistance during treatment of patients with stage III/IV melanoma.
4.1.1 Targeted NGS Gene Panels Developed for Genomic DNA and Applied to cfDNA
The application of targeted NGS gene panels in the analysis of ctDNA can identify dominant and therapeutically targetable genetic mutations. Initially, targeted NGS gene panels based on library preparation and sequencing technology developed for tissue-derived genomic DNA templates were used in studies employing ctDNA as a template (Table 1, rows 1–5). These consisted of off-the-shelf and custom-designed pan-cancer panels [86, 112,113,114,115] as well as a custom-designed melanoma panel [109].
In one study, a custom melanoma-specific NGS gene panel was designed based on whole-genome sequencing analysis of cfDNA (and tissue) from pre- and post-treatment (targeted or immune checkpoint inhibition) samples to identify a signature of SNVs in non-coding and coding regions associated with progression on therapy (Table 1, row 2) [109]. The targeted NGS gene panel was then used to monitor response to therapy based on increases in mutant allele frequency of the signature SNVs within cfDNA over several time points post-therapy. Interestingly, a TERT promoter mutation identified by whole-exome sequencing could not be incorporated into the targeted NGS gene panel based on the repetitive sequence of this region [109].
For one of the custom pan-cancer NGS gene panels used for melanoma (Table 1, row 4) [114], the panel design attempted to incorporate known melanoma driver and BRAF inhibitor resistance mutations, including mutations in BRAF, NRAS, KRAS, MAP2K1, and CDKN2A [63]. The feasibility of this panel was then highlighted using ctDNA from melanoma patients, who had previously received (or were still receiving) targeted or immune checkpoint therapy, to measure mutant allele frequency of the targeted genes. This showed a high concordance with tissue samples even though the tissue samples had been collected earlier than the plasma and during this interval patients had received one or more therapies [114].
4.1.2 Targeted NGS Gene Panels Developed for cfDNA
The subsequent development of cfDNA-optimised targeted NGS gene panels for deep sequencing of ctDNA looks to revolutionise the use of liquid biopsies as a diagnostic tool in cancer therapy (Table 1, rows 6–13). Many of these panels have the reported ability to detect mutant allele frequencies as low as 0.01%. All recommend around 20–30 ng of starting cfDNA for high-quality library preparation, and this amount of cfDNA can be commonly obtained from 10 ml of blood (4–5 ml of plasma). Less cfDNA can be used as the input, but will result in a subsequent decrease in the limit of detection. For instance, if the limit of mutation detection was a 0.01% frequency in 100 ng input cfDNA, 10 ng input cfDNA would only allow for detection of mutations occurring at a 0.1% frequency.
Several targeted NGS gene panels developed for cfDNA are now available for research and include those from Thermofisher, Roche, and ArcherDX (USA) (Table 1). Thermofisher is the only company which currently provides the option of custom-designing cfDNA-optimised NGS gene panels in the form of their new AmpliSeq HD technology (Table 1). The AmpliSeq HD technology can also be used for genomic DNA from tissue sources. Thermofisher also offer cancer-type–specific off-the-shelf cfDNA panels for breast, colorectal, and lung cancer in their Oncomine range. To date, no melanoma-specific cfDNA-optimised NGS gene panels are available, though the off-the-shelf panels do include a subset of melanoma somatic driver and resistance mutations.
The potential of targeted ctDNA sequencing is evident from a growing number of clinically focused companies who have embraced this approach, including Guardant Health (USA), CellMax Life (USA), and Foundation Medicine (USA), who have all developed off-the-shelf pan-cancer cfDNA-optimised NGS gene panels (Table 1). Of particular note is the use of the Guardant360 platform (Table 1) to analyse ctDNA from a cohort of > 20,000 cancer patients across several cancers, including melanoma, illustrating the ability of this platform to distinguish primary driver from secondary emerging clonal resistance mutations [116]. Another clinically focused company, Natera (USA), offers an individualised NGS panel, known as Signatera, which is custom-designed after whole-exome sequencing of cancer tissue (Table 1). The additional tissue sequencing step of Signatera adds significantly to the turnaround time and diagnostic cost, but provides patient specificity.
NGS of cfDNA can detect low frequency variants, but the clinical relevance of these variants, which may reflect clonal hematopoiesis, remains unclear [117]. In addition, detection of low frequency variants in NGS of cfDNA can lead to discordance when comparing different NGS platforms [118]. Therefore, although the concept of identifying multiple actionable mutations in the circulation to guide therapeutic decisions is an attractive concept, the results from these tests must be used with caution and comparison of NGS tests across large numbers of patients with cancer needs to be done to improve clinical utility.
4.2 Analysis of ctDNA as a Predictor of Response to Immunotherapy
The use of ctDNA as a predictor of response to immunotherapy has largely relied on detection using ddPCR. Several studies involving metastatic melanoma patients, based on ddPCR detection of a single mutation in BRAF or NRAS, have shown that elevated ctDNA levels at baseline and on immunotherapy correlate with a poor prognosis [71, 78]. Lee et al. used a combination of baseline and early on treatment ctDNA levels to predict response to immunotherapy, where undetectable ctDNA early during treatment was associated with improved objective response and overall survival. Furthermore, ctDNA was able to accurately and rapidly differentiate between melanoma patients receiving immune checkpoint inhibitors displaying pseudoprogression, defined as initial growth followed by eventual tumour response, and true disease progression [72].
An alternative to ddPCR-based approaches is the measure of TMB based on targeted NGS gene panels. High TMB within a particular tumour often correlates with a greater number of neoepitopes, leading to greater immunogenicity and a likely greater chance of responding to immunotherapy (reviewed in [119, 120]). The use of whole-exome sequencing and targeted NGS gene panels, such as the Guardant360 panel (Table 1), have shown promising results for a range of cancers, including melanoma, in assessing response to immunotherapies based on measuring TMB [121, 122]. A recent retrospective analysis of two large clinical trials in non-small cell lung cancer demonstrated concordance between tumour and blood TMB, with high mutation burden in plasma associated with clinically significant improvement in progression-free survival from anti-PD-L1 [123]. However, the assays used in matching tumour and plasma, despite overlap, identified non-identical variants, which likely originated from the samples themselves as opposed to technical variation.
Therefore, several factors in the use of cfDNA and targeted NGS gene panels in quantifying TMB still need to be addressed in more detail. Such factors include whether the depth of sequencing coverage of targeted NGS gene panels is sufficient for an accurate determination of TMB especially for mutations of low allelic frequency present in the tumour, and whether the cfDNA reflects the TMB of an individual or a number of tumours.
4.3 Factors to Consider when Choosing Allele-Specific PCR or NGS
One important consideration when selecting a molecular test is the cost effectiveness and turnaround time. In one study, the detection of BRAFV600E in melanoma using allele-specific PCR-based monitoring of ctDNA was compared to NGS targeted gene panel-based monitoring of genomic DNA [124]. The turnaround time was reported to be faster at 2.9 ± 1.1 days for PCR compared to 4.7 ± 1.6 days for NGS [124]. The cost was also significantly higher for NGS, at US$270 per sample compared to US$40 per sample for allele-specific PCR [124]. The advantage of NGS lies in the ability to capture much greater mutational information compared to allele-specific PCR techniques such as ddPCR (Fig. 1). Thus, although NGS can provide significant additional mutation data, the value of this data, which may constitute non-clinically relevant mutations, to the clinical management of cancer patients needs to be considered relative to the increase in cost and extensive bioinformatic resources needed for NGS analysis.
Allele-specific PCR, such as ddPCR, is a quantitative technique, whereas NGS is only semiquantitative (Fig. 1). Quantitating ctDNA can be influenced by multiple factors which are likely to induce an increased release of non-tumour DNA into the plasma. These factors include physiopathological factors, such as inflammation, autoimmune diseases, pregnancy and physical exercise (reviewed in [125]), or preanalytical factors primarily during blood collection (reviewed in [126]). This needs to be taken into consideration when using NGS or ddPCR to measure mutant allele frequency, since it depends on the number of wild-type cfDNA copies derived from normal cells.
5 Other Sources of Circulating Nucleic Acid
In an effort to increase both the sensitivity of detection and to capture the evolution of cancer-related genomic changes in the circulation, other sources of nucleic acid, both DNA and RNA, are being investigated. These alternative sources of DNA and RNA include circulating free RNA (cfRNA), extrachromosomal circular DNA, circulating tumour cells (CTCs), circulating endothelial cells, tumour educated platelets and extracellular vesicles such as exosomes (reviewed in [8, 127,128,129,130,131,132]).
Given that there is a requirement for specialised workflows to isolate cells or vesicles prior to the extraction of DNA or RNA and the fact that isolation methods are yet to be standardised (reviewed in [132, 133]), these alternate sources of tumour markers have not been thoroughly explored. The particular difficulty of isolating and assessing the genomic profile of melanoma-derived CTCs has been addressed [134, 135]. These issues stem from the apparent diversity of melanoma CTCs, which require further characterisation of specific surface markers to enable immunoaffinity-based purification of these CTC subpopulations. A subsequent report appears to have overcome some of these issues in identifying an RNA signature from CTCs isolated from melanoma patients on immunotherapy which may be a predictor of early response [136]. A recent observation has highlighted that prostate tumour-derived large extracellular vesicles (oncosomes) contain significant levels of circulating tumour genomic DNA with identifiable genomic alterations [137]. Given the limitations of detecting ctDNA in early-stage cancers, the search for other diagnostic/prognostic DNA/RNA-based biomarkers from these alternative sources is certainly worth further investigation.
6 Conclusions and Future Directions
Conventional tissue biopsy for genotyping in cancer diagnosis is still considered to be the gold standard. However, this approach has its limitations when faced with limited tumour tissue and often depends on sample tissue collected from a section of a single tumour and thus does not provide a true representation of the heterogeneous tumour burden. Furthermore, the availability of tissue for monitoring treatment over time is an issue. Liquid biopsy-based genotyping makes it possible to assess both tumour heterogeneity and to provide an accurate assessment of TMB [123]. In particular, the use of cfDNA from biofluids in conjunction with molecular technologies such as ddPCR or targeted NGS can provide non-invasive real-time information about a patient’s tumour burden and subsequent response to therapy.
The increasing sensitivity (detecting as low as 0.01% mutant allele frequency) of targeted NGS gene panels for the detection of cfDNA in cancer patients is now comparable with the sensitivity of such common digital PCR approaches as ddPCR. Given the capacity for multiplexing, currently not possible with ddPCR, and for designing custom targeted NGS gene panels optimised for cfDNA (as offered by Thermofisher’s new AmpliSeq HD technology), the likelihood is that targeted NGS gene panels will supersede ddPCR both in a research and diagnostic setting. This is now feasible in a diagnostic context given that a typical workflow for targeted NGS gene panels can be completed in 2–3 days. Importantly, the increasing availability of cfDNA standards for NGS (Seracare, USA, and Horizon Discovery, UK) containing cancer-relevant somatic mutations of known allele frequencies allows validation and standardisation of targeted NGS gene panels.
One of the main issues which still needs to be resolved to fully incorporate NGS technology into ctDNA analysis is the need to address whether NGS methods are truly as specific as PCR-based methods such as ddPCR and BEAMing. Applications such as upfront mutational profiling [80], prediction of early response [67, 70, 71], pseudoprogression [72] and early detection of relapse (during treatment and in stage III melanoma) [69, 73] have important clinical implications for the management of patients. To date, optimised ctDNA-based NGS targeted gene panels for melanoma remain to be fully evaluated for their effectiveness for future use in the clinic.
Finally, several important considerations that need to be addressed with cfDNA analysis include: the limited ability to detect cfDNA in early-stage cancers, including melanoma; and the fact that ctDNA arising from a tumour may not be detectable in cfDNA because of the site of the tumour [138, 139]. Therefore, analysis of other biofluids for cfDNA and new technologies that enrich for cfDNA is warranted. The rapid development of sensitive technologies that accurately detect tumour mutations in circulating nucleic acids is revolutionising the monitoring of cancer patients, and although tissue biopsies still provide essential diagnostic information, new targeted and immune-based therapies require real-time, longitudinal monitoring of tumour evolution via non-invasive and serial liquid biopsies.
References
Siravegna G, Marsoni S, Siena S, Bardelli A. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol. 2017;14(9):531–48.
Perakis S, Speicher MR. Emerging concepts in liquid biopsies. BMC Med. 2017;15(1):75.
Thompson JR, Menon SP. Liquid biopsies and cancer immunotherapy. Cancer J. 2018;24(2):78–83.
Gaiser MR, von Bubnoff N, Gebhardt C, Utikal JS. Liquid biopsy to monitor melanoma patients. J Dtsch Dermatol Ges. 2018;16(4):405–14.
Lim SY, Lee JH, Diefenbach RJ, Kefford RF, Rizos H. Liquid biomarkers in melanoma: detection and discovery. Mol Cancer. 2018;17(1):8.
Cheung AH, Chow C, To KF. Latest development of liquid biopsy. J Thorac Dis. 2018;10(Suppl 14):S1645–51.
Sumbal S, Javed A, Afroze B, Zulfiqar HF, Javed F, Noreen S, et al. Circulating tumor DNA in blood: future genomic biomarkers for cancer detection. Exp Hematol. 2018;65:17–28.
Pos O, Biro O, Szemes T, Nagy B. Circulating cell-free nucleic acids: characteristics and applications. Eur J Hum Genet. 2018;26(7):937–45.
Lu L, Bi J, Bao L. Genetic profiling of cancer with circulating tumor DNA analysis. J Genet Genomics. 2018;45(2):79–85.
Calapre L, Warburton L, Millward M, Ziman M, Gray ES. Circulating tumour DNA (ctDNA) as a liquid biopsy for melanoma. Cancer Lett. 2017;28(404):62–9.
Marzese DM, Hirose H, Hoon DS. Diagnostic and prognostic value of circulating tumor-related DNA in cancer patients. Expert Rev Mol Diagn. 2013;13(8):827–44.
Underhill HR, Kitzman JO, Hellwig S, Welker NC, Daza R, Baker DN, et al. Fragment length of circulating tumor DNA. PLoS Genet. 2016;12(7):e1006162.
Thierry AR, Mouliere F, Gongora C, Ollier J, Robert B, Ychou M, et al. Origin and quantification of circulating DNA in mice with human colorectal cancer xenografts. Nucleic Acids Res. 2010;38(18):6159–75.
Lo YM, Chan KC, Sun H, Chen EZ, Jiang P, Lun FM, et al. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci Transl Med. 2010;2(61):61ra91.
Thierry AR, El Messaoudi S, Gahan PB, Anker P, Stroun M. Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev. 2016;35(3):347–76.
Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, et al. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61(4):1659–65.
Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001;313(1–2):139–42.
Donaldson J, Park BH. Circulating tumor DNA: measurement and clinical utility. Annu Rev Med. 2018;29(69):223–34.
Wan JCM, Massie C, Garcia-Corbacho J, Mouliere F, Brenton JD, Caldas C, et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat Rev Cancer. 2017;17(4):223–38.
Oellerich M, Schutz E, Beck J, Kanzow P, Plowman PN, Weiss GJ, et al. Using circulating cell-free DNA to monitor personalized cancer therapy. Crit Rev Clin Lab Sci. 2017;54(3):205–18.
Cabel L, Riva F, Servois V, Livartowski A, Daniel C, Rampanou A, et al. Circulating tumor DNA changes for early monitoring of anti-PD1 immunotherapy: a proof-of-concept study. Ann Oncol. 2017;28(8):1996–2001.
Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14(9):985–90.
Heitzer E, Perakis S, Geigl JB, Speicher MR. The potential of liquid biopsies for the early detection of cancer. NPJ Precis Oncol. 2017;1(1):36.
Alidousty C, Brandes D, Heydt C, Wagener S, Wittersheim M, Schafer SC, et al. Comparison of blood collection tubes from three different manufacturers for the collection of cell-free DNA for liquid biopsy mutation testing. J Mol Diagn. 2017;19(5):801–4.
van Dessel LF, Beije N, Helmijr JC, Vitale SR, Kraan J, Look MP, et al. Application of circulating tumor DNA in prospective clinical oncology trials—standardization of preanalytical conditions. Mol Oncol. 2017;11(3):295–304.
El Messaoudi S, Rolet F, Mouliere F, Thierry AR. Circulating cell free DNA: preanalytical considerations. Clin Chim Acta. 2013;23(424):222–30.
Nikolaev S, Lemmens L, Koessler T, Blouin JL, Nouspikel T. Circulating tumoral DNA: preanalytical validation and quality control in a diagnostic laboratory. Anal Biochem. 2018;11(542):34–9.
Parpart-Li S, Bartlett B, Popoli M, Adleff V, Tucker L, Steinberg R, et al. The effect of preservative and temperature on the analysis of circulating tumor DNA. Clin Cancer Res. 2017;23(10):2471–7.
Wong D, Moturi S, Angkachatchai V, Mueller R, DeSantis G, van den Boom D, et al. Optimizing blood collection, transport and storage conditions for cell free DNA increases access to prenatal testing. Clin Biochem. 2013;46(12):1099–104.
Warton K, Yuwono NL, Cowley MJ, McCabe MJ, So A, Ford CE. Evaluation of Streck BCT and PAXgene stabilised blood collection tubes for cell-free circulating DNA studies in plasma. Mol Diagn Ther. 2017;21(5):563–70.
Warton K, Graham LJ, Yuwono N, Samimi G. Comparison of 4 commercial kits for the extraction of circulating DNA from plasma. Cancer Genet. 2018. https://doi.org/10.1016/j.cancergen.2018.02.004.
Perez-Barrios C, Nieto-Alcolado I, Torrente M, Jimenez-Sanchez C, Calvo V, Gutierrez-Sanz L, et al. Comparison of methods for circulating cell-free DNA isolation using blood from cancer patients: impact on biomarker testing. Transl Lung Cancer Res. 2016;5(6):665–72.
Sorber L, Zwaenepoel K, Deschoolmeester V, Roeyen G, Lardon F, Rolfo C, et al. A comparison of cell-free DNA isolation kits: isolation and quantification of cell-free DNA in plasma. J Mol Diagn. 2017;19(1):162–8.
Kloten V, Ruchel N, Bruchle NO, Gasthaus J, Freudenmacher N, Steib F, et al. Liquid biopsy in colon cancer: comparison of different circulating DNA extraction systems following absolute quantification of KRAS mutations using Intplex allele-specific PCR. Oncotarget. 2017;8(49):86253–63.
Markus H, Contente-Cuomo T, Farooq M, Liang WS, Borad MJ, Sivakumar S, et al. Evaluation of pre-analytical factors affecting plasma DNA analysis. Sci Rep. 2018;8(1):7375.
Devonshire AS, Whale AS, Gutteridge A, Jones G, Cowen S, Foy CA, et al. Towards standardisation of cell-free DNA measurement in plasma: controls for extraction efficiency, fragment size bias and quantification. Anal Bioanal Chem. 2014;406(26):6499–512.
Diefenbach RJ, Lee JH, Kefford RF, Rizos H. Evaluation of commercial kits for purification of circulating free DNA. Cancer Genet. 2018;228–229:21–7.
Serrati S, De Summa S, Pilato B, Petriella D, Lacalamita R, Tommasi S, et al. Next-generation sequencing: advances and applications in cancer diagnosis. Onco Targets Ther. 2016;9:7355–65.
Groisberg R, Roszik J, Conley A, Patel SR, Subbiah V. The role of next-generation sequencing in sarcomas: evolution from light microscope to molecular microscope. Curr Oncol Rep. 2017;19(12):78.
Postel M, Roosen A, Laurent-Puig P, Taly V, Wang-Renault SF. Droplet-based digital PCR and next generation sequencing for monitoring circulating tumor DNA: a cancer diagnostic perspective. Expert Rev Mol Diagn. 2018;18(1):7–17.
Reddy BY, Miller DM, Tsao H. Somatic driver mutations in melanoma. Cancer. 2017;123(S11):2104–17.
Zhang T, Dutton-Regester K, Brown KM, Hayward NK. The genomic landscape of cutaneous melanoma. Pigment Cell Melanoma Res. 2016;29(3):266–83.
Lu X, Zhang Q, Wang Y, Zhang L, Zhao H, Chen C, et al. Molecular classification and subtype-specific characterization of skin cutaneous melanoma by aggregating multiple genomic platform data. J Cancer Res Clin Oncol. 2018;144(9):1635–47.
Cancer Genome Atlas N. Genomic classification of cutaneous melanoma. Cell. 2015;161(7):1681–96.
Hodis E, Watson IR, Kryukov GV, Arold ST, Imielinski M, Theurillat JP, et al. A landscape of driver mutations in melanoma. Cell. 2012;150(2):251–63.
Guan J, Gupta R, Filipp FV. Cancer systems biology of TCGA SKCM: efficient detection of genomic drivers in melanoma. Sci Rep. 2015;20(5):7857.
Luke JJ, Flaherty KT, Ribas A, Long GV. Targeted agents and immunotherapies: optimizing outcomes in melanoma. Nat Rev Clin Oncol. 2017;14(8):463–82.
Siroy AE, Boland GM, Milton DR, Roszik J, Frankian S, Malke J, et al. Beyond BRAF(V600): clinical mutation panel testing by next-generation sequencing in advanced melanoma. J Invest Dermatol. 2015;135(2):508–15.
Xia J, Jia P, Hutchinson KE, Dahlman KB, Johnson D, Sosman J, et al. A meta-analysis of somatic mutations from next generation sequencing of 241 melanomas: a road map for the study of genes with potential clinical relevance. Mol Cancer Ther. 2014;13(7):1918–28.
Bailey MH, Tokheim C, Porta-Pardo E, Sengupta S, Bertrand D, Weerasinghe A, et al. Comprehensive characterization of cancer driver genes and mutations. Cell. 2018;173(2):371e18–385e18.
Middleton M, Hauschild A, Thomson D, Anderson R, Burdette-Radoux S, Gehlsen K, et al. Results of a multicenter randomized study to evaluate the safety and efficacy of combined immunotherapy with interleukin-2, interferon-{alpha}2b and histamine dihydrochloride versus dacarbazine in patients with stage IV melanoma. Ann Oncol. 2007;18(10):1691–7.
Robert C, Karaszewska B, Schachter J, Rutkowski P, Mackiewicz A, Stroiakovski D, et al. Improved overall survival in melanoma with combined dabrafenib and trametinib. N Engl J Med. 2015;372(1):30–9.
Robert C, Long GV, Brady B, Dutriaux C, Maio M, Mortier L, et al. Nivolumab in previously untreated melanoma without BRAF mutation. N Engl J Med. 2015;372(4):320–30.
Wolchok JD, Chiarion-Sileni V, Gonzalez R, Rutkowski P, Grob JJ, Cowey CL, et al. Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N Engl J Med. 2017;377(14):1345–56.
Dummer R, Ascierto PA, Gogas HJ, Arance A, Mandala M, Liszkay G, et al. Encorafenib plus binimetinib versus vemurafenib or encorafenib in patients with BRAF-mutant melanoma (COLUMBUS): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2018;19(5):603–15.
Ascierto PA, McArthur GA, Dreno B, Atkinson V, Liszkay G, Di Giacomo AM, et al. Cobimetinib combined with vemurafenib in advanced BRAF(V600)-mutant melanoma (coBRIM): updated efficacy results from a randomised, double-blind, phase 3 trial. Lancet Oncol. 2016;17(9):1248–60.
Helgadottir H, Rocha Trocoli Drakensjo I, Girnita A. Personalized medicine in malignant melanoma: towards patient tailored treatment. Front Oncol. 2018;8:202.
Hogan SA, Levesque MP, Cheng PF. Melanoma immunotherapy: next-generation biomarkers. Front Oncol. 2018;8:178.
Verykiou S, Ellis RA, Lovat PE. Established and emerging biomarkers in cutaneous malignant melanoma. Healthcare (Basel). 2014;2(1):60–73.
Buder-Bakhaya K, Hassel JC. Biomarkers for clinical benefit of immune checkpoint inhibitor treatment—a review from the melanoma perspective and beyond. Front Immunol. 2018;9:1474.
Lim SY, Menzies AM, Rizos H. Mechanisms and strategies to overcome resistance to molecularly targeted therapy for melanoma. Cancer. 2017;123(S11):2118–29.
Wilmott JS, Rizos H, Scolyer RA, Long GV. The “tricky business” of identifying mechanisms of resistance to anti-PD-1. Clin Cancer Res. 2017;23(12):2921–3.
Shi H, Hugo W, Kong X, Hong A, Koya RC, Moriceau G, et al. Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy. Cancer Discov. 2014;4(1):80–93.
Busser B, Lupo J, Sancey L, Mouret S, Faure P, Plumas J, et al. Plasma circulating tumor DNA levels for the monitoring of melanoma patients: landscape of available technologies and clinical applications. Biomed Res Int. 2017;2017:5986129.
Huynh K, Hoon DS. Liquid biopsies for assessing metastatic melanoma progression. Crit Rev Oncog. 2016;21(1–2):141–54.
Perkins G, Lu H, Garlan F, Taly V. Droplet-based digital PCR: application in cancer research. Adv Clin Chem. 2017;79:43–91.
Ashida A, Sakaizawa K, Uhara H, Okuyama R. Circulating tumour DNA for monitoring treatment response to anti-PD-1 immunotherapy in melanoma patients. Acta Derm Venereol. 2017;97(10):1212–8.
Chang GA, Tadepalli JS, Shao Y, Zhang Y, Weiss S, Robinson E, et al. Sensitivity of plasma BRAFmutant and NRASmutant cell-free DNA assays to detect metastatic melanoma in patients with low RECIST scores and non-RECIST disease progression. Mol Oncol. 2016;10(1):157–65.
Gray ES, Rizos H, Reid AL, Boyd SC, Pereira MR, Lo J, et al. Circulating tumor DNA to monitor treatment response and detect acquired resistance in patients with metastatic melanoma. Oncotarget. 2015;6(39):42008–18.
Herbreteau G, Vallee A, Knol AC, Theoleyre S, Quereux G, Varey E, et al. Quantitative monitoring of circulating tumor DNA predicts response of cutaneous metastatic melanoma to anti-PD1 immunotherapy. Oncotarget. 2018;9(38):25265–76.
Lee JH, Long GV, Boyd S, Lo S, Menzies AM, Tembe V, et al. Circulating tumour DNA predicts response to anti-PD1 antibodies in metastatic melanoma. Ann Oncol. 2017;28(5):1130–6.
Lee JH, Long GV, Menzies AM, Lo S, Guminski A, Whitbourne K, et al. Association between circulating tumor DNA and pseudoprogression in patients with metastatic melanoma treated with anti-programmed cell death 1 antibodies. JAMA Oncol. 2018;4(5):717–21.
Lee RJ, Gremel G, Marshall A, Myers KA, Fisher N, Dunn JA, et al. Circulating tumor DNA predicts survival in patients with resected high-risk stage II/III melanoma. Ann Oncol. 2018;29(2):490–6.
Sanmamed MF, Fernandez-Landazuri S, Rodriguez C, Zarate R, Lozano MD, Zubiri L, et al. Quantitative cell-free circulating BRAFV600E mutation analysis by use of droplet digital PCR in the follow-up of patients with melanoma being treated with BRAF inhibitors. Clin Chem. 2015;61(1):297–304.
Tsao SC, Weiss J, Hudson C, Christophi C, Cebon J, Behren A, et al. Monitoring response to therapy in melanoma by quantifying circulating tumour DNA with droplet digital PCR for BRAF and NRAS mutations. Sci Rep. 2015;22(5):11198.
Girotti MR, Gremel G, Lee R, Galvani E, Rothwell D, Viros A, et al. Application of sequencing, liquid biopsies, and patient-derived xenografts for personalized medicine in melanoma. Cancer Discov. 2016;6(3):286–99.
McEvoy AC, Warburton L, Al-Ogaili Z, Celliers L, Calapre L, Pereira MR, et al. Correlation between circulating tumour DNA and metabolic tumour burden in metastatic melanoma patients. BMC Cancer. 2018;18(1):726.
Valpione S, Gremel G, Mundra P, Middlehurst P, Galvani E, Girotti MR, et al. Plasma total cell-free DNA (cfDNA) is a surrogate biomarker for tumour burden and a prognostic biomarker for survival in metastatic melanoma patients. Eur J Cancer. 2018;88:1–9.
Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O’Connell A, Messineo MM, et al. Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res. 2014;20(6):1698–705.
McEvoy AC, Calapre L, Pereira MR, Giardina T, Robinson C, Khattak MA, et al. Sensitive droplet digital PCR method for detection of TERT promoter mutations in cell free DNA from patients with metastatic melanoma. Oncotarget. 2017;8(45):78890–900.
Horn S, Figl A, Rachakonda PS, Fischer C, Sucker A, Gast A, et al. TERT promoter mutations in familial and sporadic melanoma. Science. 2013;339(6122):959–61.
de la Monte SM, Moore GW, Hutchins GM. Patterned distribution of metastases from malignant melanoma in humans. Cancer Res. 1983;43(7):3427–33.
Patel JK, Didolkar MS, Pickren JW, Moore RH. Metastatic pattern of malignant melanoma. A study of 216 autopsy cases. Am J Surg. 1978;135(6):807–10.
Ajithkumar T, Parkinson C, Fife K, Corrie P, Jefferies S. Evolving treatment options for melanoma brain metastases. Lancet Oncol. 2015;16(13):e486–97.
Momtaz P, Pentsova E, Abdel-Wahab O, Diamond E, Hyman D, Merghoub T, et al. Quantification of tumor-derived cell free DNA(cfDNA) by digital PCR (DigPCR) in cerebrospinal fluid of patients with BRAFV600 mutated malignancies. Oncotarget. 2016;7(51):85430–6.
Ballester LY, Glitza Oliva IC, Douse DY, Chen MM, Lan C, Haydu LE, et al. Evaluating circulating tumor DNA from the cerebrospinal fluid of patients with melanoma and leptomeningeal disease. J Neuropathol Exp Neurol. 2018;77(7):628–35.
Wee EJ, Wang Y, Tsao SC, Trau M. Simple, sensitive and accurate multiplex detection of clinically important melanoma DNA mutations in circulating tumour DNA with SERS nanotags. Theranostics. 2016;6(10):1506–13.
Hu P, Zhang S, Wu T, Ni D, Fan W, Zhu Y, et al. Fe–Au nanoparticle-coupling for ultrasensitive detections of circulating tumor DNA. Adv Mater. 2018;22:e1801690.
Barel F, Guibourg B, Lambros L, Le Flahec G, Marcorelles P, Uguen A. Evaluation of a rapid, fully automated platform for detection of BRAF and NRAS mutations in melanoma. Acta Derm Venereol. 2018;98(1):44–9.
Bisschop C, Ter Elst A, Bosman LJ, Platteel I, Jalving M, van den Berg A, et al. Rapid BRAF mutation tests in patients with advanced melanoma: comparison of immunohistochemistry, droplet digital PCR, and the Idylla mutation platform. Melanoma Res. 2018;28(2):96–104.
Harle A, Salleron J, Franczak C, Dubois C, Filhine-Tressarieu P, Leroux A, et al. Detection of BRAF mutations using a fully automated platform and comparison with high resolution melting, real-time allele specific amplification, immunohistochemistry and next generation sequencing assays, for patients with metastatic melanoma. PLoS One. 2016;11(4):e0153576.
Seremet T, Planken S, Schreuer M, Jansen Y, Delaunoy M, El Housni H, et al. Illustrative cases for monitoring by quantitative analysis of BRAF/NRAS ctDNA mutations in liquid biopsies of metastatic melanoma patients who gained clinical benefits from anti-PD1 antibody therapy. Melanoma Res. 2018;28(1):65–70.
Uguen A, Troncone G. A review on the Idylla platform: towards the assessment of actionable genomic alterations in one day. J Clin Pathol. 2018;71(9):757–62.
Griewank KG, Schilling B. Next-generation sequencing to guide treatment of advanced melanoma. Am J Clin Dermatol. 2017;18(3):303–10.
Reiman A, Kikuchi H, Scocchia D, Smith P, Tsang YW, Snead D, et al. Validation of an NGS mutation detection panel for melanoma. BMC Cancer. 2017;17(1):150.
Miraflor AP, de Abreu FB, Peterson JD, Turner SA, Amos CI, Tsongalis GJ, et al. Somatic mutation analysis in melanoma using targeted next generation sequencing. Exp Mol Pathol. 2017;103(2):172–7.
Giardina T, Robinson C, Grieu-Iacopetta F, Millward M, Iacopetta B, Spagnolo D, et al. Implementation of next generation sequencing technology for somatic mutation detection in routine laboratory practice. Pathology. 2018;50(4):389–401.
Carlson JA, Caldeira Xavier JC Jr, Tarasen A, Sheehan CE, Otto G, Miller VA, et al. Next-generation sequencing reveals pathway activations and new routes to targeted therapies in cutaneous metastatic melanoma. Am J Dermatopathol. 2017;39(1):1–13.
Mullauer L. Next generation sequencing: clinical applications in solid tumours. Memo. 2017;10(4):244–7.
Kamps R, Brandao RD, Bosch BJ, Paulussen AD, Xanthoulea S, Blok MJ, et al. Next-generation sequencing in oncology: genetic diagnosis, risk prediction and cancer classification. Int J Mol Sci. 2017;18(2):308.
Chicard M, Colmet-Daage L, Clement N, Danzon A, Bohec M, Bernard V, et al. Whole-exome sequencing of cell-free DNA reveals temporo-spatial heterogeneity and identifies treatment-resistant clones in neuroblastoma. Clin Cancer Res. 2018;24(4):939–49.
Klevebring D, Neiman M, Sundling S, Eriksson L, Darai Ramqvist E, Celebioglu F, et al. Evaluation of exome sequencing to estimate tumor burden in plasma. PLoS One. 2014;9(8):e104417.
Luo H, Li H, Hu Z, Wu H, Liu C, Li Y, et al. Noninvasive diagnosis and monitoring of mutations by deep sequencing of circulating tumor DNA in esophageal squamous cell carcinoma. Biochem Biophys Res Commun. 2016;471(4):596–602.
Manier S, Park J, Capelletti M, Bustoros M, Freeman SS, Ha G, et al. Whole-exome sequencing of cell-free DNA and circulating tumor cells in multiple myeloma. Nat Commun. 2018;9(1):1691.
Olmedillas-Lopez S, Garcia-Olmo DC, Garcia-Arranz M, Peiro-Pastor R, Aguado B, Garcia-Olmo D. Liquid biopsy by NGS: differential presence of exons (DPE) in cell-free DNA reveals different patterns in metastatic and nonmetastatic colorectal cancer. Cancer Med. 2018;7(5):1706–16.
Vandekerkhove G, Todenhofer T, Annala M, Struss WJ, Wong A, Beja K, et al. Circulating tumor DNA reveals clinically actionable somatic genome of metastatic bladder cancer. Clin Cancer Res. 2017;23(21):6487–97.
Annala M, Vandekerkhove G, Khalaf D, Taavitsainen S, Beja K, Warner EW, et al. Circulating tumor DNA genomics correlate with resistance to abiraterone and enzalutamide in prostate cancer. Cancer Discov. 2018;8(4):444–57.
Dietz S, Schirmer U, Merce C, von Bubnoff N, Dahl E, Meister M, et al. Low input whole-exome sequencing to determine the representation of the tumor exome in circulating DNA of non-small cell lung cancer patients. PLoS One. 2016;11(8):e0161012.
Cutts A, Venn O, Dilthey A, Gupta A, Vavoulis D, Dreau H, et al. Characterisation of the changing genomic landscape of metastatic melanoma using cell free DNA. NPJ Genom Med. 2017;4(2):25.
Carlino MS, Long GV, Kefford RF, Rizos H. Targeting oncogenic BRAF and aberrant MAPK activation in the treatment of cutaneous melanoma. Crit Rev Oncol Hematol. 2015;96(3):385–98.
de Unamuno Bustos B, Murria Estal R, Perez Simo G, de Juan Jimenez I, Escutia Munoz B, Rodriguez Serna M, et al. Towards personalized medicine in melanoma: implementation of a clinical next-generation sequencing panel. Sci Rep. 2017;7(1):495.
Kaisaki PJ, Cutts A, Popitsch N, Camps C, Pentony MM, Wilson G, et al. Targeted next-generation sequencing of plasma DNA from cancer patients: factors influencing consistency with tumour DNA and prospective investigation of its utility for diagnosis. PLoS One. 2016;11(9):e0162809.
Malapelle U, Mayo de-Las-Casas C, Rocco D, Garzon M, Pisapia P, Jordana-Ariza N, et al. Development of a gene panel for next-generation sequencing of clinically relevant mutations in cell-free DNA from cancer patients. Br J Cancer. 2017;116(6):802–10.
Gangadhar TC, Savitch SL, Yee SS, Xu W, Huang AC, Harmon S, et al. Feasibility of monitoring advanced melanoma patients using cell-free DNA from plasma. Pigment Cell Melanoma Res. 2018;31(1):73–81.
Du J, Wu X, Tong X, Wang X, Wei J, Yang Y, et al. Circulating tumor DNA profiling by next generation sequencing reveals heterogeneity of crizotinib resistance mechanisms in a gastric cancer patient with MET amplification. Oncotarget. 2017;8(16):26281–7.
Zill OA, Banks KC, Fairclough SR, Mortimer SA, Vowles JV, Mokhtari R, et al. The landscape of actionable genomic alterations in cell-free circulating tumor DNA from 21,807 advanced cancer patients. Clin Cancer Res. 2018;24(15):3528–38.
Coombs CC, Zehir A, Devlin SM, Kishtagari A, Syed A, Jonsson P, et al. Therapy-related clonal hematopoiesis in patients with non-hematologic cancers is common and associated with adverse clinical outcomes. Cell Stem Cell. 2017;21(3):374e4–382e4.
Kuderer NM, Burton KA, Blau S, Rose AL, Parker S, Lyman GH, et al. Comparison of 2 commercially available next-generation sequencing platforms in oncology. JAMA Oncol. 2017;3(7):996–8.
Cabel L, Proudhon C, Romano E, Girard N, Lantz O, Stern MH, et al. Clinical potential of circulating tumour DNA in patients receiving anticancer immunotherapy. Nat Rev Clin Oncol. 2018;15(10):639–50.
Khagi Y, Kurzrock R, Patel SP. Next generation predictive biomarkers for immune checkpoint inhibition. Cancer Metastasis Rev. 2017;36(1):179–90.
Koeppel F, Blanchard S, Jovelet C, Genin B, Marcaillou C, Martin E, et al. Whole exome sequencing for determination of tumor mutation load in liquid biopsy from advanced cancer patients. PLoS One. 2017;12(11):e0188174.
Khagi Y, Goodman AM, Daniels GA, Patel SP, Sacco AG, Randall JM, et al. Hypermutated circulating tumor DNA: correlation with response to checkpoint inhibitor-based immunotherapy. Clin Cancer Res. 2017;23(19):5729–36.
Gandara DR, Paul SM, Kowanetz M, Schleifman E, Zou W, Li Y, et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat Med. 2018;24(9):1441–8.
Zhu ML, Zhou L, Sadri N. Comparison of targeted next generation sequencing (NGS) versus isolated BRAF V600E analysis in patients with metastatic melanoma. Virchows Arch. 2018;473(3):371–7.
Khier S, Lohan L. Kinetics of circulating cell-free DNA for biomedical applications: critical appraisal of the literature. Future Sci OA. 2018;4(4):FSO295.
Trigg RM, Martinson LJ, Parpart-Li S, Shaw JA. Factors that influence quality and yield of circulating-free DNA: a systematic review of the methodology literature. Heliyon. 2018;4(7):e00699.
Best MG, Wesseling P, Wurdinger T. Tumor-educated platelets as a noninvasive biomarker source for cancer detection and progression monitoring. Cancer Res. 2018;78(13):3407–12.
Mader S, Pantel K. Liquid biopsy: current status and future perspectives. Oncol Res Treat. 2017;40(7–8):404–8.
Wang J, Chang S, Li G, Sun Y. Application of liquid biopsy in precision medicine: opportunities and challenges. Front Med. 2017;11(4):522–7.
Khetrapal P, Lee MWL, Tan WS, Dong L, de Winter P, Feber A, et al. The role of circulating tumour cells and nucleic acids in blood for the detection of bladder cancer: a systematic review. Cancer Treat Rev. 2018;66:56–63.
Zhu J, Chen S, Zhang F, Wang L. Cell-free eccDNAs: a new type of nucleic acid component for liquid biopsy? Mol Diagn Ther. 2018;22(5):515–22.
Momen-Heravi F, Getting SJ, Moschos SA. Extracellular vesicles and their nucleic acids for biomarker discovery. Pharmacol Ther. 2018. https://doi.org/10.1016/j.pharmthera.2018.08.002.
Mansilla C, Soria E, Ramirez N. The identification and isolation of CTCs: a biological Rubik’s cube. Crit Rev Oncol Hematol. 2018;126:129–34.
Marsavela G, Aya-Bonilla CA, Warkiani ME, Gray ES, Ziman M. Melanoma circulating tumor cells: benefits and challenges required for clinical application. Cancer Lett. 2018;28(424):1–8.
Reid AL, Freeman JB, Millward M, Ziman M, Gray ES. Detection of BRAF-V600E and V600K in melanoma circulating tumour cells by droplet digital PCR. Clin Biochem. 2015;48(15):999–1002.
Hong X, Sullivan RJ, Kalinich M, Kwan TT, Giobbie-Hurder A, Pan S, et al. Molecular signatures of circulating melanoma cells for monitoring early response to immune checkpoint therapy. Proc Natl Acad Sci USA. 2018;115(10):2467–72.
Vagner T, Spinelli C, Minciacchi VR, Balaj L, Zandian M, Conley A, et al. Large extracellular vesicles carry most of the tumour DNA circulating in prostate cancer patient plasma. J Extracell Vesicles. 2018;7(1):1505403.
Chae YK, Davis AA, Jain S, Santa-Maria C, Flaum L, Beaubier N, et al. Concordance of genomic alterations by next-generation sequencing in tumor tissue versus circulating tumor DNA in breast cancer. Mol Cancer Ther. 2017;16(7):1412–20.
Yang N, Li Y, Liu Z, Qin H, Du D, Cao X, et al. The characteristics of ctDNA reveal the high complexity in matching the corresponding tumor tissues. BMC Cancer. 2018;18(1):319.
Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014;20(5):548–54.
Agarwal N, Pal SK, Hahn AW, Nussenzveig RH, Pond GR, Gupta SV, et al. Characterization of metastatic urothelial carcinoma via comprehensive genomic profiling of circulating tumor DNA. Cancer. 2018;124(10):2115–24.
Barata PC, Koshkin VS, Funchain P, Sohal D, Pritchard A, Klek S, et al. Next-generation sequencing (NGS) of cell-free circulating tumor DNA and tumor tissue in patients with advanced urothelial cancer: a pilot assessment of concordance. Ann Oncol. 2017;28(10):2458–63.
Lanman RB, Mortimer SA, Zill OA, Sebisanovic D, Lopez R, Blau S, et al. Analytical and clinical validation of a digital sequencing panel for quantitative, highly accurate evaluation of cell-free circulating tumor DNA. PLoS One. 2015;10(10):e0140712.
McCoach CE, Blakely CM, Banks KC, Levy B, Chue BM, Raymond VM, et al. Clinical utility of cell-free DNA for the detection of ALK fusions and genomic mechanisms of ALK inhibitor resistance in non-small cell lung cancer. Clin Cancer Res. 2018;24(12):2758–70.
Rossi G, Mu Z, Rademaker AW, Austin LK, Strickland KS, Costa RLB, et al. Cell-free DNA and circulating tumor cells: comprehensive liquid biopsy analysis in advanced breast cancer. Clin Cancer Res. 2018;24(3):560–8.
Yang M, Topaloglu U, Petty WJ, Pagni M, Foley KL, Grant SC, et al. Circulating mutational portrait of cancer: manifestation of aggressive clonal events in both early and late stages. J Hematol Oncol. 2017;10(1):100.
Barata PC, Mendiratta P, Heald B, Klek S, Grivas P, Sohal DPS, et al. Targeted next-generation sequencing in men with metastatic prostate cancer: a pilot study. Target Oncol. 2018;13(4):495–500.
Thompson JC, Yee SS, Troxel AB, Savitch SL, Fan R, Balli D, et al. Detection of therapeutically targetable driver and resistance mutations in lung cancer patients by next-generation sequencing of cell-free circulating tumor DNA. Clin Cancer Res. 2016;22(23):5772–82.
Schwaederle M, Chattopadhyay R, Kato S, Fanta PT, Banks KC, Choi IS, et al. Genomic alterations in circulating tumor DNA from diverse cancer patients identified by next-generation sequencing. Cancer Res. 2017;77(19):5419–27.
Clark TA, Chung JH, Kennedy M, Hughes JD, Chennagiri N, Lieber DS, et al. Analytical validation of a hybrid capture-based next-generation sequencing clinical assay for genomic profiling of cell-free circulating tumor DNA. J Mol Diagn. 2018;20(5):686–702.
Jamal-Hanjani M, Wilson GA, Horswell S, Mitter R, Sakarya O, Constantin T, et al. Detection of ubiquitous and heterogeneous mutations in cell-free DNA from patients with early-stage non-small-cell lung cancer. Ann Oncol. 2016;27(5):862–7.
Kirkizlar E, Zimmermann B, Constantin T, Swenerton R, Hoang B, Wayham N, et al. Detection of clonal and subclonal copy-number variants in cell-free DNA from patients with breast cancer using a massively multiplexed PCR methodology. Transl Oncol. 2015;8(5):407–16.
Abbosh C, Birkbak NJ, Wilson GA, Jamal-Hanjani M, Constantin T, Salari R, et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature. 2017;545(7655):446–51.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Russell J. Diefenbach was supported in part by a donation to Melanoma Institute Australia from the Clearbridge Foundation. This work was also supported in part by the National Health and Medical Research Council (APP1093017 and APP1128951). Helen Rizos is supported by a National Health and Medical Research Council Research Fellowship.
Conflict of interest
Russell J. Diefenbach, Jenny H. Lee, and Helen Rizos declare that they have no conflicts of interest that might be relevant to the contents of this manuscript.
Rights and permissions
About this article
Cite this article
Diefenbach, R.J., Lee, J.H. & Rizos, H. Monitoring Melanoma Using Circulating Free DNA. Am J Clin Dermatol 20, 1–12 (2019). https://doi.org/10.1007/s40257-018-0398-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40257-018-0398-x