Abstract
Pracinostat, an emerging hydroxamate histone deacetylase (HDAC) inhibitor has shown better efficacy than approved inhibitor suberoylanilide hydroxamic acid (SAHA). Apart from haematological malignancies, this inhibitor has shown promising results in preclinical models of solid tumours. Being pan-inhibitor pracinostat targets various classical HDACs and has demonstrated antiproliferative properties in a series of cancer cell lines. Currently, no energetic and structural studies are available about the pracinostat against four HDAC isozymes of Class I. Taking this into account, the current study involved flexible molecular docking for gaining insights regarding pracinostat-HDAC isozyme interactions, molecular mechanics generalized born surface area (MM-GBSA) for estimating binding affinity of this inhibitor towards these isozymes and energetically optimized pharmacophores (e-Pharmacophores) technique for delineating the critical e-pharmacophoric features of pracinostat in its least energy state in the binding pocket of these HDACs. The outcome from this study will help in further optimization of pracinostat towards better therapeutic and the e-Pharmacophores generated will serve as queries in e-Pharamcophores guided virtual screening.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
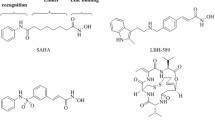
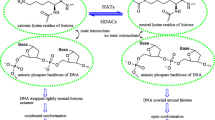
Various cancers are associated with deviated expression of zinc-dependent classical histone deacetylase (HDACs) (Sanaei and Kavoosi 2019). Due to promising clinical trial results, four HDAC inhibitors (HDACi) have been labelled as FDA approved. Among the approved HDACi, suberoylanilide hydroxamic acid (SAHA) and belinostat come under the jurisdiction of hydroxamate group HDAC inhibitors (Ho et al. 2020; Tu et al. 2020). However, due to its poor pharmacokinetics, SAHA which is effective against haematological malignancies does not prove to be effective against solid tumours (Gryder et al. 2012). Pracinostat, an emerging hydroxamate group HDAC inhibitor has demonstrated strong anticancer activity against both haematological and solid malignancies (Ganai 2016). Compared to SAHA, pracinostat has shown efficient pharmaceutical, pharmacokinetic and physicochemical properties. Importantly, this inhibitor unlike SAHA has manifested above 100-fold affinity towards classical HDACs in comparison to other zinc-reliant metalloenzymes (Ganai 2016; Novotny-Diermayr et al. 2010). For ease in understanding the structures of pracinostat and SAHA have been shown in Fig. 1.
Pracinostat in association with azacytidine was found to be better tolerated by patients having newly confirmed acute myeloid leukaemia (AML) (Garcia-Manero et al. 2019). Pracinostat exhibited synergistic effects in AML (pre-clinical models) in collaboration with pacritinib, a JAK2 inhibitor (Novotny-Diermayr et al. 2012). Pracinostat, a strong inhibitor targetting multiple HDACs has shown extremely good druglike properties and has proved to be highly effective in inhibiting tumour growth in, in vivo models (Wang et al. 2011). Although this inhibitor inhibits HDACs in nanomolar concentration, the antiproliferative IC50 values of pracinostat range from 0.34 to 0.56 µM depending on the cancer cell lines (Novotny-Diermayr et al. 2012; Novotny-Diermayr et al. 2010; Wang et al. 2011). However, no structural studies are available regarding the pracinostat against clinically relevant targets (HDAC1-3 and HDAC8). Thus, this study focussed on exploring the binding affinity and interactions of pracinostat with defined HDACs through extra-precision (XP) - flexible molecular docking and a relatively superior alternative to MM/PBSA termed as MM-GBSA. Most importantly the e-Pharmacophoric features of pracinostat in docked state with these inhibitors were generated.
Methodology followed
Pracinostat preparation
The coordinate files of pracinostat and SAHA were fetched from PubChem (PubChem CID: 49,855,250 and 5311 respectively). Using the LigPrep tool pracinostat was minimized and protonation states were generated through epik. Following this, metal binding states were generated and additionally specific chiralities were retained during its (pracinostat) preparation (Ganai et al. 2017; Madhavi Sastry et al. 2013).
Preparation of target enzymes
It is well known that precise structure based modelling highly depends on correct starting structures. The crystal structure coordinates of four Class I HDAC isozymes were retrieved from pandemically famous ‘Protein Data Bank’ (www.rcsb.org) (Berman et al. 2000). The structures of these isozymes were individually prepared using the Protein Preparation Wizard (Madhavi Sastry et al. 2013; Shankaran et al. 2017). This was done as structural correctness is among the main requisites of accurate molecular docking. To these structures the missing hydrogen atoms were added. Sometimes crystal structures possess missing residues, missing side chains and missing loops. These missing side chains and loops were filled using the Prime incorporated in this wizard. Following this, the water molecules over 5 Å were removed (Ganai et al. 2017). The junk chains and irrelevant heteroatoms in these crystal structures were expunged. However, the cocrystallized ligands wherever present were kept as such. As HDACs are reliant on zinc this heteroatom was retained in all the four enzymes under consideration (Ganai et al. 2017). These structures were further processed to make them suitable for the upcoming procedure. During the defined process the structures of HDACs were optimized and water molecules were deleted as per certified guidelines. All the structures were finally minimized using the default parameters (Kalyaanamoorthy and Chen 2013; Madhavi Sastry et al. 2013).
Specifying the binding pocket through glide
HDAC2 and HDAC8 were having native ligands which were kept as such during protein preparation. These ligands were used for demarcating active site in these HDACs. However, for HDAC1 and HDAC3 which were devoid of experimental ligand, grid was generated by selecting the active site residues of these enzymes. Grid was specified in all the four cases using the grid generation option of the Glide program (Friesner et al. 2004; Halgren et al. 2004; Shankaran et al. 2017).
Molecular docking of pracinostat against class I HDACs
Pracinostat was docked against the individual grid-specified HDAC isozymes using the globally accepted molecular docking tool Glide. Molecular docking was carried out in extra-precision flexible mode and identical parameters were set for all the isozymes. Docking score of the most appropriate pose was only taken into consideration (Friesner et al. 2006; Ganai et al. 2018; Halgren et al. 2004). Further, the SAHA was compared with pracinostat against HDAC2 and HDAC8 using the AutoDock Vina based CB-Dock. This server first predicts five active sites and then docks ligands flexibly into them using the Vina. Based on information of co-crystallized ligand, the most correct pose was finally selected in all cases (Liu et al. 2020; Pettersen et al. 2004; Trott and Olson 2010).
Binding energy estimation though MM/PBSA variant, MM-GBSA
The binding energy of pracinostat-HDAC docked complexes was estimated by using pandemically accepted method Prime/MMGBSA (Molecular Mechanics Generalized Born Surface Area. During the calculations no flexibility was imparted to receptor (HDACs) and default parameters were not altered (Balaji and Ramanathan 2012). Prime MMGBSA method relies on molecular mechanics and implicit solvation. Its implicit solvent models estimate the free energies of solute-solvent interactions while molecular mechanics component measures the enthalpic contributions of receptor-ligand interactions (Ganai et al. 2017; Kalyaanamoorthy and Chen 2013). Out of various energy calculations done by MMGBSA, the ultimate calculation was done through below mentioned equation;
ΔG (bind) = complex − receptor − ligand
Favourable binding affinity is positively related to more negative values of ΔG (Bind). In other words more negative the value of ΔG (Bind) more is the pracinostat-HDAC affinity.
Binding affinity (kcal/mol) indicated by Vina Score was estimated for SAHA and pracinostat against two representatives of Class I HDAC isozymes. The isozymes selected were HDAC2 and HDAC8 as they were integrated with native ligands and it was easy to cross-check the correct pose generated by AutoDock Vina algorithm.
Pracinostat-HDAC 3D-interaction profile generation through PLIP
Characterizing ligand-receptor interactions has crucial significance in drug discovery. The interaction profile of pracinostat in complex with individual isozyme was generated at atomistic level using the protein-ligand interaction profiler (PLIP). Pracinostat-HDAC docked complexes were separately given as an input and then the ligand was selected for generating 3D interaction profile between pracinostat and HDACs. Seven types of non-covalent interactions including hydrophobic and hydrogen bonding interactions existing between the pracinostat and HDACs were explored through PLIP (Salentin et al. 2015).
e-Pharmacophoric feature generation
The energetically optimized structure based pharmacophores (e-Pharmacophores)—method merges the fruitful aspects of both the structure and ligand based approaches. The e-pharmacophores for pracinostat-HDAC1-3 and pracinostat-HDAC8 were generated using the auto e-Pharmacophore tool of Phase of Schrödinger. For generating these features the pose of pracinostat with more negative value of docking score was selected in each case (Kalyaanamoorthy and Chen 2013; Salam et al. 2009).
Results and discussion
Pracinostat manifested differential docking score against class I HDACs
Molecular docking provides atomistic details of ligand-protein interactions (Meng et al. 2011). Pracinostat showed differential docking score towards HDAC1-3 and HDAC8. While more negative value of docking score was shown by pracinostat against HDAC1 (− 9.8) followed by HDAC2 (− 9.65) and HDAC8 (− 8.93) respectively, the least negative value of this score was seen against HDAC3 (− 8.89) (Fig. 2). This differential docking score of single inhibitor pracinostat against four Class I HDACs may be attributed to differences in amino acid residues at the active sites of these HDACs. Small differences in amino acid residues at the active sites of these HDACs have discernible effect on docking score (Ganai et al. 2018).
Docking score of pracinostat against three different Class I isozymes. Pracinostat was docked against members of Class I HDACs using the extra-precision flexible docking method and the docking scores were obtained from respective output files. Pracinostat showed more negative value of docking score with HDAC1 (− 9.8) and HDAC2 (− 9.65). Comparatively lesser negative values were obtained with HDAC8 (− 8.93) and HDAC3 (− 8.89). GlideScore + Epik energy together constitute docking score
Pracinostat exhibited distinct binding free energy values against HDAC isozymes
Binding free energy values designate the binding affinity. It is globally accepted that more negative values of this energy suggests stronger binding. The more negative value of binding energy was shown by pracinostat against HDAC2, followed by HDAC8 and HDAC3 respectively. While pracinostat demonstrated binding free energy value of − 51.46 (kcal/mol) towards HDAC2, relatively lower value was observed for HDAC3 (− 38 kcal/mol) and the lowest binding energy value (− 47.30 kcal/mol) was seen in case of pracinostat-HDAC8 (Fig. 3). More negative binding free energy value is directly related to stronger binding tendency. The docking score of HDAC2-3 and HDAC8 values correlated well with binding free energy values further authenticated the accuracy of results.
Binding free energy values of pracinostat against predefined HDACs. The binding free energy of pracinostat-HDAC2, pracinostat-HDAC3 and pracinostat-HDAC8 docked complexes were calculated through alternative but superior method than MM/PBSA, Prime MM-GBSA. This method performs several energy calculations from which the ΔGbind values are calculated using the globally accepted equation:
ΔGbind = Complex – (Receptor + Ligand)
The binding free energy was calculated by imparting no receptor flexibility (frozen condition). More negative the value of ΔGbind stronger is the ligand-enzyme binding. Thus from the above graph it is quite visible that pracinostat shows more binding inclination towards HDAC2 (− 51.46), followed by HDAC8 (− 47.31 kcal/mol) and HDAC3 (− 38.0 kcal/mol)
Further, the binding affinity calculated by CB-Dock as indicated by Vina Score showed similar trend in case of pracinostat. While SAHA showed the binding affinity of − 8.1 and − 6.6 kcal/mol against HDAC2 and HDAC8 respectively, the corresponding values of pracinostat were found to be − 7.2 and − 6.9 kcal/mol respectively. Thus SAHA demonstrated more negative value against HDAC2 compared to pracinostat and these values correlated with their in vitro IC50 values (4 nM and 96 nM for SAHA and pracinostat respectively) (Bradner et al. 2010a; Bradner et al. 2010b). In case of HDAC8, pracinostat demonstrated relatively more negative value of affinity − 6.9 in comparison to SAHA − 6.6 kcal/mol. This was expected as IC50 of pracinostat is lesser as compared to SAHA against HDAC8 (140 nM for pracinostat and 1100 nM for SAHA) (Bradner et al. 2010a). This has been put in simplified in Table 1.
Pracinostat targeted aspartate, glycine and phenylalanine and tyrosine residues of these HDACs
.
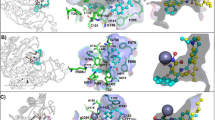
Interaction profile study of pracinostat in docked state with the above mentioned isozymes revealed various interactions. Seven hydrophobic interactions, one salt bridge and two hydrogen bonds were manifested by pracinostat-HDAC1. While salt bridge was formed with ASP 99, hydrogen bonds were seen with HIS 178 and GLY 149. LEU 271, TYR 204, PHE 150 and PHE 205 participated in hydrophobic interactions with pracinostat (Fig. 4; Table 2). On the other hand six hydrophobic interactions (PHE 155, PHE 210, and LEU 276), three hydrogen bonds (TYR 308, GLY 154) one salt bridge and one π-Stacking were observed between pracinostat-HDAC2 (Fig. 5; Table 2). Pracinostat-HDAC3 displayed two salt bridges (ASP 92, ASP 93) besides showing multiple hydrophobic and hydrogen bonding interactions (Fig. 5; Table 2). Further, in case of Pracinostat-HDAC8 two hydrogen bonds, seven hydrophobic interactions one π-Stacking (PHE 208) and one salt bridge (ASP 101) were noticed (Fig. 5; Table 2).
Pracinostat interacts with HDAC1 through multiple interactions. While first figure is the Chimera generated view of pracinostat-HDAC1 docked complex, the second figure represents the 3D interaction of pracinostat and HDAC1 residues. As per analysis through PLIP, in addition to seven hydrophobic interactions, one salt bridge and two hydrogen bonding interactions were observed between pracinostat and the residues of HDAC1. GLY 149, HIS 178 were involved in hydrogen bonding interaction, ASP 99 formed salt bridge where as PHE 150, TYR 204, LEU 271 and PHE 205 showed hydrophobic interaction with the inhibitor pracinostat
3D interaction of pracinostat with HDAC2, HDAC3 and HDAC8. Pracinostat (brown colour) interacts will all these isozymes through various interactions. Majority of the interactions are hydrophobic, followed by hydrogen bonding and salt bridges. In case of HDAC2 and HDAC8, π-stacking is also seen with pracinostat
E-Pharmacophoric features of pracinostat against HDAC1/2/3/8
These features are generated from the ligand-protein complex. Docked complexes of pracinostat with these HDACs were taken individually for e-Pharmacophoric feature generation. Thus, during generation of these features receptor is also taken into consideration. For each docked complex, the pose of ligand with more negative value of docking score is considered (Salam et al. 2009). This method takes the advantage of scoring function of Glide XP for specifying receptor-ligand interactions precisely (Ganai et al. 2017; Kalyaanamoorthy and Chen 2013). While pracinostat displayed 3 e-Pharmacophoric features against HDAC2 and HDAC8 only two features were observed against HDAC3. One hydrogen bond donor, one aromatic ring and one hydrogen bond acceptor were noted both in case of HDAC2 and HDAC8. In case of HDAC3, pracinostat showed one aromatic ring and one hydrogen bond donor (Fig. 6).
E-Pharmacophoric features of pracinostat against Class I HDACs. Pracinostat showed three e-Pharmacophoric features against HDAC1 including hydrogen bond donor (D5), aromatic ring (R9) and positive ionisable group (P7). With HDAC2 this inhibitor showed three features including one hydrogen bond donor (D5), one hydrogen bond acceptor (A5) and one aromatic ring (R9). Regarding HDAC3, pracinostat displayed only two features, one hydrogen bond donor (D4) and one aromatic ring (R9). Like HDAC2, pracinostat showed three features against HDAC8 including hydrogen bond donor (D5), hydrogen bond acceptor (A3) and aromatic ring
Aromatic ring in linker component of pracinostat proved to be the most critical
Like other hydroxamates, pracinostat has three components in its structure including zinc-binding group, cap region and the linker connecting the two (Noureen et al. 2010). The score of these e-Pharmacophoric features was estimated and the location of these features in pracinostat components was also delineated. Pracinostat exhibited three features positive ionizable group in cap region, aromatic ring residing in linker region and hydrogen bond donor in the zinc chelating region of this inhibitor. Among the defined features the first feature proved to be critical in energy terms (− 3.43 kcal/mol). Pracinostat showed aromatic ring (located in linker region) as the most crucial feature in terms of score against all the isozymes. The score of this feature was calculated to be − 1.49 against HDAC2, − 2.07 for HDAC3 and − 1.34 in case of HDAC8. These findings align well with the previous findings where it has been reported that inhibitors with minimum one aromatic ring in the linker component bind strong whereas the inhibitors devoid of aromatic rings are feeble binders (Kalyaanamoorthy and Chen 2013). Hydrogen bond donor located in the ZBG of pracinostat was found to be another crucial feature after aromatic ring. This feature showed a score of – 0.7 in case of all the concerned HDAC isozymes. Another feature namely hydrogen bond acceptor in ZBG of pracinostat was seen only in case of HDAC2 and HDAC8. This feature was not seen in case of pracinostat-HDAC3 docked complex (Table 3).
Conclusions
From about past 30 years HDACi are emerging as prosperous molecules for neuroregeneration and antineoplastic therapy. An oral HDAC inhibitor namely pracinostat has shown multiple benefits over approved drug SAHA. This has diverted the attention of scientific community towards this promising hydroxamate pracinostat. Preclinical and clinical benefits of this inhibitor especially in combinatorial therapy have been certified by various research groups. Despite these studies pracinostat was not studied at atomistic level against therapeutically relevant Class I HDACs. Keeping these facts in view a combined approach involving different techniques was utilized for studying pracinostat-HDAC2-3/HDAC8 interactions, binding energy and importantly e-Pharmacophoric features of pracinostat in binding state with these isozymes. Pracinostat showed distinct docking scores and binding free energies against these three Class I members expectedly. Docking scores and their corresponding binding energy values showed excellent correlation which further stamps the accuracy of results. While two e-Pharmacophoric features were displayed by pracinostat in case of isozyme HDAC3, an additional feature as hydrogen bond acceptor was shown by this inhibitor in case of HDAC2 and HDAC8. Further, calculating the scores of e-Pharmacophoric features revealed aromatic ring in linker as the key feature in terms of energy in all the four isozymes excluding HDAC1. Taken together, this study provides insights for further optimization of pracinostat and the differences in features of pracinostat in HDAC3 compared to HDAC2/HDAC8 will help in identification of novel HDAC3 selective inhibitors through e-Pharmacophore based virtual screening.
References
Balaji B, Ramanathan M (2012) Prediction of estrogen receptor β ligands potency and selectivity by docking and MM-GBSA scoring methods using three different scaffolds. J Enzyme Inhib Med Chem 27:832–844
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE, 2000. The Protein Data Bank. Nucleic acids research 28, 235–242
Bradner JE, Mak R, Tanguturi SK, Mazitschek R, Haggarty SJ, Ross K, Chang CY, Bosco J, West N, Morse E, Lin K, Shen JP, Kwiatkowski NP, Gheldof N, Dekker J, DeAngelo DJ, Carr SA, Schreiber SL, Golub TR, Ebert BL (2010a) Chemical genetic strategy identifies histone deacetylase 1 (HDAC1) and HDAC2 as therapeutic targets in sickle cell disease. Proc Natl Acad Sci U S A 107:12617–12622
Bradner JE, West N, Grachan ML, Greenberg EF, Haggarty SJ, Warnow T, Mazitschek R (2010b) Chemical phylogenetics of histone deacetylases. Nat Chem Biol 6:238–243
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, Repasky MP, Knoll EH, Shelley M, Perry JK, Shaw DE, Francis P, Shenkin PS (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med chem 47:1739–1749
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J Med Chem 49:6177–6196
Ganai SA (2016) Histone deacetylase inhibitor pracinostat in doublet therapy: a unique strategy to improve therapeutic efficacy and to tackle herculean cancer chemoresistance. Pharmaceutical Biol 54:1926–1935
Ganai SA, Abdullah E, Rashid R, Altaf M (2017) Combinatorial In Silico Strategy towards Identifying Potential Hotspots during Inhibition of Structurally Identical HDAC1 and HDAC2 Enzymes for Effective Chemotherapy against Neurological Disorders. Frontiers in molecular neuroscience 10, 357
Ganai SA, Farooq Z, Banday S, Altaf M (2018) silico approaches for investigating the binding propensity of apigenin and luteolin against class I HDAC isoforms. Future Med Chem 10:1925–1945
Garcia-Manero G, Abaza Y, Takahashi K, Medeiros BC, Arellano M, Khaled SK, Patnaik M, Odenike O, Sayar H, Tummala M, Patel P, Maness-Harris L, Stuart R, Traer E, Karamlou K, Yacoub A, Ghalie R, Giorgino R, Atallah E (2019) Pracinostat plus azacitidine in older patients with newly diagnosed acute myeloid leukemia: results of a phase 2 study. Blood Adv 3:508–518
Gryder BE, Sodji QH, Oyelere AK (2012) Targeted cancer therapy: giving histone deacetylase inhibitors all they need to succeed. Future Med Chem 4:505–524
Halgren TA, Murphy RB, Friesner RA, Beard HS, Frye LL, Pollard WT, Banks JL (2004) Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. Journal of medicinal chemistry 47:1750–1759
Ho TCS, Chan AHY, Ganesan A (2020) Thirty Years of HDAC Inhibitors: 2020 Insight and Hindsight. Journal of medicinal chemistry
Kalyaanamoorthy S, Chen Y-PP (2013) Energy based pharmacophore mapping of HDAC inhibitors against class I HDAC enzymes. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 1834, 317–328
Liu Y, Grimm M, Dai W-t, Hou M-c, Xiao Z-X, Cao Y (2020) CB-Dock: a web server for cavity detection-guided protein–ligand blind docking. Acta Pharmacol Sin 41:138–144
Madhavi Sastry G, Adzhigirey M, Day T, Annabhimoju R, Sherman W (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27:221–234
Meng X-Y, Zhang H-X, Mezei M, Cui M (2011) Molecular docking: a powerful approach for structure-based drug discovery. Curr Comput Aided Drug Des 7:146–157
Noureen N, Rashid H, Kalsoom S (2010) Identification of type-specific anticancer histone deacetylase inhibitors: road to success. Cancer Chemother Pharmacol 66:625–633
Novotny-Diermayr V, Hart S, Goh KC, Cheong A, Ong LC, Hentze H, Pasha MK, Jayaraman R, Ethirajulu K, Wood JM (2012) The oral HDAC inhibitor pracinostat (SB939) is efficacious and synergistic with the JAK2 inhibitor pacritinib (SB1518) in preclinical models of AML. Blood Cancer J 2:e69–e69
Novotny-Diermayr V, Sangthongpitag K, Hu CY, Wu X, Sausgruber N, Yeo P, Greicius G, Pettersson S, Liang AL, Loh YK, Bonday Z, Goh KC, Hentze H, Hart S, Wang H, Ethirajulu K, Wood JM (2010) SB939, a novel potent and orally active histone deacetylase inhibitor with high tumor exposure and efficacy in mouse models of colorectal cancer. Mol Cancer Ther 9:642–652
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Salam NK, Nuti R, Sherman W (2009) Novel method for generating structure-based pharmacophores using energetic analysis. J Chem Inf Model 49:2356–2368
Salentin S, Schreiber S, Haupt VJ, Adasme MF, Schroeder M (2015) PLIP: fully automated protein-ligand interaction profiler. Nucleic acids research 43, W443-W447
Sanaei M, Kavoosi F (2019) Histone Deacetylases and Histone Deacetylase Inhibitors: Molecular Mechanisms of Action in Various Cancers. Adv Biomed Res 8:63–63
Shankaran KS, Ganai SA, K PA, Mahadevan PB, V (2017) In silico and In vitro evaluation of the anti-inflammatory potential of Centratherum punctatum Cass-A. J Biomol Struct Dyn 35:765–780
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Tu B, Zhang M, Liu T, Huang Y (2020) Nanotechnology-Based Histone Deacetylase Inhibitors for Cancer Therapy. Front Cell Developmental Biol 8
Wang H, Yu N, Chen D, Lee KCL, Lye PL, Chang JWW, Deng W, Ng MCY, Lu T, Khoo ML, Poulsen A, Sangthongpitag K, Wu X, Hu C, Goh KC, Wang X, Fang L, Goh KL, Khng HH, Goh SK, Yeo P, Liu X, Bonday Z, Wood JM, Dymock BW, Kantharaj E, Sun ET (2011) Discovery of (2E)-3-{2-Butyl-1-[2-(diethylamino)ethyl]-1H-benzimidazol-5-yl}-N-hydroxyacrylamide (SB939), an Orally Active Histone Deacetylase Inhibitor with a Superior Preclinical Profile. J Med Chem 54, 4694–4720
Acknowledgements
Ganai SA acknowledges monetary support from Science and Engineering Research Board. The author used the power facility of SDAU palanpur for writing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ganai, S.A. Characterizing binding intensity and energetic features of histone deacetylase inhibitor pracinostat towards class I HDAC isozymes through futuristic drug designing strategy. In Silico Pharmacol. 9, 18 (2021). https://doi.org/10.1007/s40203-021-00077-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40203-021-00077-y