Abstract
Nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH) are becoming the dominant liver diseases in the US and Western World. Extensive work is being done to diagnose, understand, and explore the pathogenesis of these multivariable complex diseases. Recently, a new avenue of biologic regulation is being explored. MicroRNAs are noncoding RNAs that modulate the expression of multiple genes and have been implicated in multiple diseases. Recently, there is a growing body of evidence supporting a significant role of microRNAs in NAFLD pathogenesis and progression to NASH, and hinting at their use as targets, biomarkers, and potential therapeutic tools. This review is designed to highlight some of the recent work on a few of the key microRNAs involved in the pathogenesis of NAFLD and NASH.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nonalcoholic fatty liver disease (NAFLD) encompasses a spectrum of disease that spans from isolated fatty liver (IFL) to nonalcoholic steatohepatitis (NASH) to cirrhosis and hepatocellular carcinoma (HCC). NASH can be defined by necro-inflammation with hepatocyte injury and ballooning, along with a variable amount of fibrosis and potential for progression to cirrhosis. NALFD is considered by most to be the hepatic manifestation of the metabolic syndrome, which is defined by the presence of three or more of the following: visceral obesity, elevated fasting plasma glucose, hypertension, hypertriglyceridemia, or low high-density lipoprotein levels [1].
Global NAFLD prevalence rates ranges from 2.8 to 46 % [2, 3•]. With time, NAFLD has become the most common cause of chronic liver disease in the United States and other western countries, and its prevalence has mirrored the rising obesity and diabetes mellitus (DM) epidemics. Recent global analysis estimates that 1.6 billion adults are overweight with a body mass index (BMI) of 25 kg/m2 or greater, and 500 million adults are obese with a BMI of 30 kg/m2 or greater [4]. In the United States, approximately 33.8 % of the populations are obese and 10.6 % have type 2 DM [5].
The National Health and Nutrition Examination Surveys (NHANES) performed in the United States from 1988 to 2008 showed stable prevalence rates for particular chronic liver diseases, such as hepatitis B, hepatitis C, and alcoholic liver disease. Recent advances in the treatment of HCV will likely lessen its prevalence and impact on public health [6]. However, the prevalence of NAFLD dramatically increased from 5.5 to 11 %, in the same time period. Overall, NAFLD represented approximately 47 % of all chronic liver disease in 1988, rising to 75 % by 2008 [7].
Estimating the true incidence and prevalence of NAFLD, specifically NASH, remains a challenge due both to the asymptomatic presentation of the disease as well as the lack of accurate and noninvasive diagnostic measurements. The prevalence of NAFLD was first estimated to be 36 % in lean patients and 72 % in obese patients in 1990 via autopsy. The prevalence of NASH in this same cohort was found to be 2.7 % in lean patients and 18.5 % in obese patients [8]. The evaluation of patients undergoing bariatric surgery confirmed the high prevalence rates among obese populations in which the prevalence of NAFLD and NASH was found to be 91 % and 37 %, respectively [9]. Of note, although NAFLD is strongly associated with obesity and other metabolic syndrome components, it can occur in patients without obesity and obesity does not portend the presence of NAFLD.
What is especially concerning, NASH may progress to cirrhosis from any stage [10]. After the development of cirrhosis, the patient is at risk for development of hepatocellular carcinoma (HCC) [11] and liver-related mortality [12]. Even without the development of NASH, obesity alone is associated to the risk of developing HCC. Large meta-analysis of cohort studies investigating nonviral causes of HCC shows a 90 % increased risk of HCC in obese patients [13, 14]. This has been suggested as a major cause of the growing rates of HCC in developed countries. In particular, the annual incidence of HCC over the last two decades in the United States has increased by 80 % [15–17]. The association between NASH and HCC represents a growing area of study and concern as metabolic syndrome and obesity rates continue to rise.
The pathogenesis of NASH may be inherently involved with the metabolic syndrome and its associated insulin resistance, both of which are thought to be proinflammatory processes [18]. For example, obesity is frequently linked to type-II diabetes and consequently, to hyperinsulinemia, which leads to increased insulin-like factor 1 (IGF-1) and elevated production of sex steroids and cytokines. In 2010, it was reported that obesity is associated with a chronic inflammatory state characterized by the release of IL-6 and TNFα, two well-known inflammation promoting cytokines [19]. However, the cause(s) of NAFLD and the progression of NAFLD to NASH have yet to be defined.
Further hindering the study of NAFLD and NASH is the difficulty in determining the presence of NASH by noninvasive means. To that end, multiple avenues investigating different markers and pathways associated with the development of NASH are being investigated. One of the promising possible mechanisms in the development of NASH is altered expression of microribonucleic acids (microRNAs). This review will discuss the current breakthroughs in the investigation of microRNAs and their relation to the development of NASH. After covering the background of how microRNAs function, we will discuss new data relating to miR-122, miR-33, miR-34a, and miR-24.
The Significance of MicroRNA in Human disease
Investigations of the human transcriptome following completion of the Human Genome Project in 2003 revealed that our genome encodes noncoding RNAs. First described in 1993 [20], microRNAs play an important role in posttranscriptional gene-expression regulation [21]. MicroRNAs are thought to fine-tune entire intracellular molecular cascades such as intracellular signaling [22]. MicroRNA networks are differentially expressed: microRNA expression is organ/tissue specific [23] and is differentially altered during disease states. Thus, induction or inhibition of microRNA expression may provide unique therapeutic tools [24, 25]. Aberrant expression of microRNAs in liver tissue has been implicated in the progression of liver fibrosis and hepatocarcinogenesis [26–28].
MicroRNAs are small noncoding RNAs 21–25 nucleotides in size that play an important role in the regulation of gene expression. They are produced as a pre-microRNA and exported from the nucleus to the cytoplasm where they are cleaved by the endoribonuclease called Dicer. During this process, the cleaved portion(s) are unwound to single strands and loaded into the RNA-induced silencing complex (RISC). This microRNA–RISC complex can interact with the 3′-end untranslated region (3′ UTR) of the target gene’s messenger-RNA resulting in the suppression of mRNA translation or direct degradation of target mRNA [29]. It has been found that a single microRNA can have several target genes, allowing it to play a broad role in gene regulation.
MicroRNAs and mRNA can be released from cells and carried by exosomes produced from the endoplasmic reticulum (ER) [30]. Recent work has demonstrated that intercellular signaling performed by the exosome is through the action of microRNA. From the donor cell, microRNA-containing exosomes are released through ceramide-dependent secretory machinery and transferred to a receiving target cell [31]. The transfer can occur through actively secreted micro vesicles, as demonstrated in human blood and cell cultures [32]. Viruses such as EBV may produce microRNAs themselves that are then brought to target cells by exosomes where then modulate the immune response [33]. In the setting of liver disease, HCC has been demonstrated to use microRNA exchange via exosomes in intercellular communication [34].
Since their discovery, microRNAs have been investigated for possible connections in human disease. Alterations in their expression patterns have been seen in multiple disease states compared to normal tissues and serum. In particular, changes in microRNA expression in heart disease, sepsis, malignancies, and autoimmune diseases (reviewed in [35]) have suggested a new avenue of research and targets for treating human disease. This promising new field has propelled research and opened the possibility that stable microRNAs which are detectable in serum and plasma may serve as biomarkers for early disease states or be noninvasive means of determining the severity of disease [36, 37]. In particular, miR-122, miR-33a, miR-34a, and miR-24 have shown recent promise as markers for hepatic inflammation in the setting of NASH/NAFLD or as factors contributing to or involved in the development of NASH/NAFLD.
MiR-122
In 2008, expression profiling studies of patients identified miR-122 as differentially expressed after comparing normal patients and NASH patients’ microRNA profiles, indicating a possible role in the pathogenesis of NASH [38]. MiR-122 has received considerable attention as a biomarker of liver disease because it comprises approximately 70 % of microRNAs in the liver [39, 40]. In fact compared to the rest of the body, miR-122 has been found to be the most abundant in the liver, and is released into circulation during hepatocyte damage [41, 42•].
In the liver, increased expression of miR-122 is modulated by multiple transcription factors involved in hepatocyte development and regulation such as HNF1A, HNF3A, and HNF3B [43]. Within liver tissue, miR-122 has been found to be a key regulator of cholesterol and fatty-acid metabolism [44]. Mir-122 has also been found to be increased in multiple disease states including hepatitis B and C infections, drug-induced liver injury, and NAFLD [38, 44–49, 50•]. Due to its increased expression in a broad range of diseases with necro-inflammatory activity, miR-122 has been suggested as a serum or tissue marker of general hepatic inflammation. Unfortunately, the circulating level of mirR-122 does not correlate with fibrosis stage and functional capacity of the liver and cannot serve as a noninvasive means of staging chronic liver disease [46].
In the setting of acute inflammation, miR-122 shows more promise. It has been found that circulating levels of miR-122 correlated with the age of patients with HBV acute-on-chronic liver failure [50•, 51]. Two independent studies addressing acetaminophen (APAP) toxicity, a common toxin precipitating DILI, in experimental murine models identified miR-122 and miR-192 as potential circulating biomarkers in this setting of acute toxicity [52, 53].
Most recently, a study demonstrated that miR-122 serum levels were robustly increased early on in a diet-induced model of NAFLD and correlated with ALT and AST levels. Though most work has supported miR-122 as a broad marker for liver inflammation, this work supports the notion that miR-122 correlates to and can indicate inflammation in the setting of NASH [54•].
One function of miR-122 is post-transcriptional regulation of CYP7A1, the rate-limiting enzyme controlling bile acid synthesis in human hepatocytes [55]. Recent studies showed that miR-122 knockout mice accumulate triacylglycerol in the liver, due to an up regulation of enzymes responsible for triacylglycerol synthesis and storage [56••].
Within this same miR-122 knock out model, the microRNA was also found to regulate fibrogenic factors, such as Kruppel-like factor 6 that targets transforming growth factor β1. When miR-122 is increased, there is an activation of hepatic stellate cells and fibrogenic processes. Also coordinated role for miR-122 in liver health is supported the development of steatosis and abnormal levels of very-low density lipoproteins and high-density lipoproteins, in these miR-122 knockout mice [56••].
MiR-33a
MiR-33a is another microRNA that has been found to be involved in bile acid regulation, in addition to fatty-acid oxidation and cholesterol homeostasis [57, 58••]. When cellular cholesterol levels decrease, miR-33a expression is co-induced with its host gene sterol regulatory element-binding protein 2 (SREBP2) mRNA. MiR-33a silencing studies demonstrated regression of atherosclerosis in mice, which suggests that miR-33a acts in synergy with SREBP2 to regulate cholesterol homeostasis [59]. Unfortunately, this decrease in atherosclerosis comes with the price of increased intrahepatic cholesterol. It was found when SREBP2 and miR-33a are activated; hepatic efflux transporters and bile acid synthesis are down regulated [60•]. This was confirmed in work with hepatic stellate cells exposed to free cholesterol, which resulted in enhanced SREBP2 and miR-33a signaling with suppression of peroxisome proliferator-activated receptor γ signaling. This altered signaling results increased free cholesterol in the hepatic stellate cells, leading TGFβ-induced activation and exaggerated liver fibrosis in NASH [61••]. The picture is complex, however, miR-33a knockout mice develop obesity and liver steatosis, possibly due to expression of SREBP1 which was enhanced in the absence of miR-33a [62•]. MiR-33a and -33b have been also described to inhibit genes involved in fatty-acid metabolism and insulin signaling in hepatocytes [63], further implying their pathogenic role during NAFLD.
MiR-34a
As previously discussed, expression profiling studies have identified several differentially expressed microRNAs, including miR-21, miR-34a, miR-122, and miR-155, in human [37] and mouse NASH, [64–66] and that the altered expression of these microRNAs suggests a significant role in the pathogenesis of NASH. However, despite some recent success in the investigation and clinical management of NASH, major gaps remain in our understanding. This includes our limited understanding of the biochemical etiology, the broad spectrum of its presentation, and the basic process of the diseases progression from NAFLD to NASH [67, 68]. In particular, whether there is different susceptibility of certain patients or tissues to progress to NASH, and could this susceptibility be associated with an altered expression of microRNAs.
MiR-34a has been implicated as a central factor in the development of NASH. Past work has shown miR-34a involvement in cell maintenance through activity and activation of senescence, cell cycle arrest, and apoptosis [69]. MiR-34a induction of apoptosis has been theorized to involve interactions with target proteins in the proper biological setting. The best characterized target of miR-34a is Sirtuin 1 (SIRT1), a NAD-dependent deacetylase that modulates apoptosis in the context of oxidative stress and genomic damage [70]. Specifically, SIRT1 decreases p53 activity. When upregulated, miR-34a suppresses SIRT1, thereby increasing p53 acetylation and transcription and in turn leads to the induction of proapoptotic genes such as PUMA. This effect on apoptosis may be partially due to miR-34a suppression of KLF4 (Kruppel-like factor 4) [71••], which is transcriptionally activated by p53 following DNA damage [72] and causes cell cycle arrest at both the G1/S and G2/M boundaries [73, 74].
This miR-34a/SIRT1/p53 proapoptotic pathway in NAFLD represents a positive feedback loop whereby miR-34 and similar microRNAs are direct transcriptional targets upregulated by p53 [75]. Other work has observed that the inflammation in human liver that occurs with NAFLD and NASH is paralleled with increases in p53 [76]. This has also been supported by p53 activation of mitochondrial pathways of apoptosis in an animal model of NASH [77]. Together these data suggest a key role of miR-34a in hepatocyte apoptosis in the pathogenesis of NAFLD and NASH. Inhibition of miR-34a and p53 activity, or preserving SIRT1 function is logical therapeutic approaches that may benefit patients who have NAFLD and NASH.
One such advance has been made in modulating the miR-34a/SIRT1/p53 proapoptotic pathway through treatment with ursodeoxycholic acid (UDCA) [78••]. Rui et al. treated rat hepatocytes with UDCA resulting in increased SIRT1 expression and subsequent decreased acetylation of p53. As expected by the reduced p53 activity, apoptosis was also reduced. This protective activity of UDCA was effective even in a model of miR-34a overexpression. Their work indicates that the mechanism of UDCA and its inhibition of miR-34a are partly due to the prevention of p53 transactivation, and thus preventing the positive feedback loop. In turn, the decrease of miR-34a (by reducing p53 positive feedback) leads to increased SIRT1.
Recent results have also demonstrated that miR-34a expression in both human liver and human serum is increased in patients with NAFLD. Importantly a correlation was made of the liver miR-34a level and the severity of NAFLD [79]. Due to both of these factors, miR-34a may not only represent a therapeutic target in the treatment of NAFLD progression, but may also be useful as a noninvasive biomarker to assess disease progression. However, an association of increased circulating miR-34a with type 2 diabetes may complicate the use of this marker [80].
MiR-24
One group has started to investigate miR-24 in the development of NASH stemming from hepatocyte-specific microRNA profiling studies in mouse livers. They showed that miR-24 is highly and specifically expressed in hepatocytes [81••]. Ng et al. performed this study by microRNA profiling of livers from hepatocyte-specific Dicer1 knockout and wild-type (WT) mice. The microRNAs whose expression was at least 1.7-fold higher in WT than Dicer1 knockout mice were defined as hepatocyte-specific microRNAs. This experiment also confirmed the importance of proinflammatory miR-122 and miR-192, which were also enriched in this study.
They then confirmed the importance of miR-24 in NASH through high fat diet (HFD) treatment of C57Bl/6 mice, which demonstrated up regulation of miR-24. MiR-24 up regulation in the setting of fatty acid exposure was then confirmed with Oleic acid treatment of primary human hepatocyte and HepG2 cell cultures. Both cell cultures also demonstrated increased miR-24 expression.
Using the PubMed GEO Database, five normal and eight human NAFLD/NASH liver samples were compared to identify possible targets of miR-24. With this analysis, 411 down-regulated genes were identified in human fatty livers. The 3′ UTR of mRNA of these potential targets was then compared with the conserved seed region of miR-24. This led the group to identify 48 potential target genes that have binding motifs for miR-24. These 48 targets were cross-referenced with the 411 down-regulated genes found in the control fatty liver array comparison, identifying 3 genes Insig1, KFL6 (Kruppel-like factor 6), and CXADR (Coxsackie virus and adenovirus receptor).
Insig1 is a polytopic membrane protein of the ER that regulates lipid synthesis by retaining sterol regulatory element–binding proteins (SREBPs) in the ER and preventing their proteolytic activation in the Golgi apparatus [82–84]. The movement of SREBPs from ER to the Golgi complex is a central event in lipid homeostasis in animal cells [85–87]. Significantly, 18-20 SREBPs have been identified as membrane-bound transcription factors that activate genes encoding enzymes required for synthesis of cholesterol and triglycerides [86, 87].
The three SREBP isoforms, SREBP1a, SREBP1c, and SREBP2, have different roles in lipid synthesis. SREBP1c is involved in fatty acid and triglyceride synthesis, whereas SREBP2 is relatively specific to cholesterol synthesis [87]. Immediately after their synthesis, SREBPs bind to SCAP (SREBP cleavage-activating protein). When Insig1 protein levels are low, SCAPs escort SREBPs to the Golgi where they are processed and released into the cytosol, and can then enter the nucleus and activate transcription of lipogenic genes [86].
Next, they assessed the functional contribution of increased Insig1 expression to the development of fatty liver by reducing miR-24 expression in obese mice. They synthesized LNA (locked nucleic acid) anti-miR-24 anti-sense oligonucleotide (ASO) specifically targeting miR-24. To confirm that the phenotype observed in miR-24-ASO-injected mice was due to specific microRNA deficiency, and not toxicity caused by the ASO, they generated miR-24-mismatched-anti-sense oligo (miR-24-MM-ASO), a control ASO that differs from microRNAs in 4 mismatched base pairs. C57Bl/6 mice, which had been on a HFD for 8 weeks, were injected with either miR-24-ASO or miR-24-MM-ASO for 4 weeks. They observed an 88 % reduction of hepatic miR-24 expression in mice that received miR-24-ASO compared to miR-24-MM-ASO, and a threefold increase of Insig1, as revealed by qRT-PCR and immunostaining [81••].
MiR-24-ASO treatment had no effect on body and liver weight. However, antagonizing miR-24 significantly reduced high levels of triglycerides and cholesterol in liver and triglycerides in plasma in HFD animals treated with miR-24-ASO, in contrast to plasma total cholesterol levels. Nile-Red and H&E staining further confirmed that miR-24 knockdown reduced hepatic lipid accumulation in livers of HFD-treated mice. These findings suggest that the crosstalk between miR-24 and Insig1 plays an important role in NAFLD and hypertriglyceridemia, and miR-24-ASO may be a potential therapeutic target for these disorders. This is supported by past work where the overexpression of Insig1 in liver inhibits lipogenesis and knockout of Insig1 leads to increased total content of both liver and plasma triglycerides [88, 89], suggesting that the relationship between miR-24 and Insig1 may play an important role in the development of NAFLD and hyperlipidemia.
MiR-24 has been shown to be upregulated in NAFLD/NASH and hepatocyte cell culture models of fatty acid exposure. Up regulation of miR-24 leads to deceased Insig1, increased nuclear SREBP1c and SREBP2, and subsequently elevated lipid accumulation in primary human hepatocytes and HepG2 cells. In contrast, antagonizing miR-24 leads to the opposite and more potentially therapeutic effects. These data suggest that miR-24 may have a significant role in the development of NAFLD/NASH and so far shows promise as a therapeutic target.
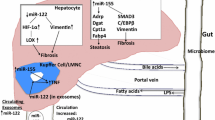
A summary of the discussed microRNAs can be found in Table 1. Figure 1 is a representation of the expression and activity of the discussed microRNAs.
MiR-122 increases in expression with steatosis which leads to Fibrosis, while being detected in the patient’s serum. MiR-24 increases with hepatic exposure to fatty acids, resulting in increased Insig1 expression which may increase steatosis. Steatosis may also increase miR-34a expression which then decreases SIRT1, leading to increased fibrosis
Conclusion
The incidence of NAFLD and NASH is increasing dramatically in the world population with the increase of metabolic syndrome across the globe. As the prevalence of more western HFDs spread, both NASH and metabolic syndrome will continue to become a major concern in human health. Furthermore, with the development of more effective HBV and HCV treatments, NASH and NAFLD will become the most dominant cause of liver disease throughout the world. These developments underscore the critical need to be able to detect and track the severity of NASH and NAFLD in patients, through noninvasive means. With the discovery and growing knowledge of microRNAs, we may be able to find better noninvasive markers of the disease to detect it earlier and to investigate therapeutic interventions. Furthermore, microRNAs show promise as possible targets for those interventions.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Alberti KG, Eckel RH, Grundy SM et al (2009) Harmonizing the metabolic syndrome: a joint interim statement of the International Diabetes Federation Task Force on Epidemiology and Prevention; National Heart, Lung, and Blood Institute; American Heart Association; World Heart Federation; International Atherosclerosis Society; and International Association for the Study of Obesity. Circulation 120:1640–1645
Lazo M, Clark JM (2008) The epidemiology of nonalcoholic fatty liver disease: a global perspective. Semin Liver Dis 28:339–350
• Lazo M, Hernandez R, Eberhardt MS, et al (2013) Prevalence of nonalcoholic fatty liver disease in the United States: the Third National Health and Nutritional Examination Survey, 1988–1994. Am J Epidemiol 178:38–45. Recent epidemiology study of the NASH epidemic
Finucane MM, Stevens GA, Cowan MJ et al (2011) National, regional, and global trends in body-mass index since 1980: systematic analysis of health examination surveys and epidemiological studies with 960 country-years and 9.1 million participants. Lancet 377(9765):557–567
Flegal KM, Carroll MD, Ogden CL et al (2010) Prevalence and trends in obesity among US adults, 1999–2008. JAMA 303:235–241
Liang TJ, Ghany MG (2014) Therapy of Hepatitis C—back to the future. NEJM. doi:10.1056/NEJMe1403619
Younossi ZM, Stepanova M, Afendy M et al (2011) Changes in the prevalence of the most common causes of chronic liver diseases in the United States from 1988 to 2008. Clin Gastroenterol Hepatol 9(6):524–530
Wanless IR, Lentz JS (1990) Fatty liver hepatitis (steatohepatitis) and obesity: an autopsy study with analysis of risk factors. Hepatology 12:1106–1110
Machado M, Marques-Vidal P, Cortez-Pinto H (2006) Hepatic histology in obese patients undergoing bariatric surgery. J Hepatol 45:600–606
Malik SM, deVera ME, Fontes P et al (2009) Outcome after liver transplantation for NASH cirrhosis. Am J Transplant 7:782–793
Siegel AB, Zhu AX (2009) Metabolic syndrome and hepatocellular carcinoma. Cancer 115:5651–5661
Ong JP, Pitts A, Younossi ZM (2008) Increased overall mortality and liver-related mortality in non-alcoholic fatty liver disease. J Hepatol 49:608–612
Blonski W, Kotlyar DS, Forde KA (2010) Non-viral causes of hepatocellular carcinoma. World J Gastroenterol 16(29):3603–3615
Clark JM (2006) The epidemiology of nonalcoholic fatty liver disease in adults. J Clin Gastroenterol 40(Suppl 1):S5–S10
Bosch FX, Ribes J, Diaz M, Cléries R (2004) Primary liver cancer: world-wide incidence and trends. Gastroenterology 127:S3–S16
El-Serag HB, Mason AC (1999) Rising incidence of hepatocellular carcinoma in the United States. N Engl J Med 340(10):745–750
Gomaa AI, Khan SA, Toledano MB, Waked I, Taylor-Robinson SD (2008) Hepatocellular carcinoma: epidemiology, risk factors and pathogenesis. World J Gastroenterol 14(27):4300–4308
Brunt E, Kleiner DE, Wilson LA, Belt P, Neuschwander Tetri BA (2011) Nonalcoholic fatty liver disease (NAFLD) activity score and the histopathologic diagnosis in NAFLD: distinct clinicopathologic meanings. Hepatology 53:810–820
Park EJ, Lee JH, Yu GY, He G, Ali SR, Holzer RG et al (2010) Dietary and genetic obesity promote liver inflammation and tumorigenesis by enhancing IL-6 and TNF expression. Cell 140:197–208
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294:853–858
Berezikov E (2011) Evolution of microRNA diversity and regulation in animals. Nat Rev Genet 12:846–860
Lagos-Quintana M, Rauhut R et al (2002) Identification of tissue-specific microRNAs from mouse. Curr Biol 12:735–739
Davidson BL, McCray PB Jr (2011) Current prospects for RNA interference-based therapies. Nat Rev Genet 12:329–340
Kaucsár T, Rácz Z, Hamar P (2010) Posttranscriptional gene-expression regulation by micro RNA (miRNA) network in renal disease. Adv Drug Deliv Rev 30:1390–1401
Murakami Y, Toyoda H, Tanaka M, Kuroda M, Harada Y et al (2011) The progression of liver fibrosis is related with overexpression of the miR-199 and 200 families. PLoS ONE 6:e16081
Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H et al (2006) Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene 25:2537–2545
Braconi C, Henry JC, Kogure T, Schmittgen T, Patel T (2011) The role of microRNAs in human liver cancers. Semin Oncol 38:752–763
He L, Hannon G (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5(7):522–531
Valadi H, Ekstrom K, Bossios A, Sjostrand M, Lee JJ et al (2007) Exosomemediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9:654–659
Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y et al (2010) Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem 285:17442–17452
Zhang Y, Liu D, Chen X, Li J, Li L et al (2010) Secreted monocytic miR-150 enhances targeted endothelial cell migration. Mol Cell 39:133–144
Pegtel DM, Cosmopoulos K, Thorley-Lawson DA, van Eijndhoven MA, Hopmans ES et al (2010) Functional delivery of viral miRNAs via exosomes. Proc Natl Acad Sci USA 107:6328–6333
Kogure T, Lin WL, Yan IK, Braconi C, Patel T (2011) Intercellular nanovesicle mediated microRNA transfer: a mechanism of environmental modulation of hepatocellular cancer cell growth. Hepatology 54:1237–1248
Kosaka N, Iguchi H, Ochiya T (2010) Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer Sci 101:2087–2092
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK et al (2008) Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA 105:10513–10518
Chen X, Ba Y, Ma L, Cai X, Yin Y et al (2008) Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 18:997–1006
Cheung O, Puri P, Eicken C et al (2008) Nonalcoholic steatohepatitis is associated with altered hepatic microRNA expression. Hepatology 48:1810–1820
Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T (2002) Identification of tissue-specific microRNAs from mouse. Curr Biol 12:735–739
Chang J, Nicolas E, Marks D, Sander C, Lerro A et al (2004) miR-122, a mammalian liver-specific microRNA, is processed from hcr mRNA and may downregulate the high affinity cationic amino acid transporter CAT-1. RNA Biol 1:106–113
Szabo G, Bala S (2013) MicroRNAs in liver disease. Nat Rev Gastroenterol Hepatol Sep 10(9):542–552
• Trebicka J, et al (2013) Hepatic and serum levels of miR-122 after chronic HCV induced fibrosis. J Hepatol 58(2):234–239. Additional support of miR-122 as secreted marker of HCV inflammation
Coulouarn C, Factor VM, Andersen JB, Durkin ME, Thorgeirsson SS (2009) Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic properties. Oncogene 28:3526–3536
Esau C, Davis S, Murray SF, Yu XX et al (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3:87–98
Pogribny IP, Starlard-Davenport A, Tryndyak VP (2010) Difference in expression of hepatic microRNAs miR-29c, miR-34a, miR-155, and miR-200b is associated with strain-specific susceptibility to dietary nonalcoholic steatohepatitis in mice. Lab Investig 90:1437–1446
Bihrer V, Friedrich-Rust M, Kronenberger B et al (2011) Serum miR-122 as a biomarker of necroinflammation in patients with chronic hepatitis C virus infection. Am J Gastroenterol 106:1663–1669
Cermelli S, Ruggieri A, Marrero JA et al (2011) Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS One 6:e23937
Starkey Lewis PJ, Merz M et al (2012) Serum microRNA biomarkers for drug-induced liver injury. Clin Pharmacol Ther 92:291–293
Tryndyak VP, Latendresse JR, Montgomery B et al (2012) Plasma microRNAs are sensitive indicators of interstrain differences in the severity of liver injury induced in mice by a choline- and folate-deficient diet. Toxicol Appl Pharmacol 262:52–59
• Waidmann O, Bihrer V, Pleli T, et al (2012) Serum microRNA-122 levels in different groups of patients with chronic hepatitis B virus infection. J Viral Hepat 19:e58–e65. MiR-122 also as a marker of HCV infection and inflammation
Ji F, Yang B, Peng X, Ding H, You H et al (2011) Circulating microRNAs in hepatitis B virus-infected patients. J Viral Hepat 18:e242–e251
Wang K et al (2009) Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc Natl Acad Sci USA 106(11):4402–4407
Su YW et al (2012) A panel of serum microRNAs as specific biomarkers for diagnosis of compound- and herb-induced liver injury in rats. PLoS One 7(5):e37395
• Clarke JD, Sharapova T, Lake AD et al (2013) Circulating microRNA 122 in the methionine- and choline-deficient mouse model of non-alcoholic steatohepatitis. Appl Toxicol. doi:10.1002/jat.2960. Recent mouse diet model of NASH which correlates with miR-122 expression in serum
Song KH, Li T, Owsley E, Chiang JY (2010) A putative role of micro RNA in regulation of cholesterol 7alpha-hydroxylase expression in human hepatocytes. J Lipid Res 51:2223–2233
•• Tsai WC, Hsu SD, Hsu CS, et al (2012) MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Investig 122:2884–2897. The use of knock out 122 mice to investigate its role in liver disease. This knock out had increased fibrosis and tumor formation- partially attributed to the action of a miR-122a target, the Klf6 transcript
Najafi-Shoushtari SH, Kristo F, Li Y, Shioda T et al (2010) MicroRNA-33 and the SREBP host genes cooperate to control cholesterol homeostasis. Science 328:1566–1569
•• Allen RM, Marquart TJ, Albert CJ, Suchy FJ, et al (2012) miR-33 controls the expression of biliary transporters, and mediates statin- and diet-induced hepatotoxicity. EMBO Mol Med 4:882–895. The first work connected miR-33 with bile and cholesterol metabolism in the liver and the connection to NASH
Rayner KJ, Sheedy FJ, Esau CC et al (2011) Antagonism of miR-33 in mice promotes reverse cholesterol transport and regression of atherosclerosis. J Clin Investig 121:2921–2931
• Li T, Francl JM, Boehme S, Chiang JY (2013) Regulation of cholesterol and bile acid homeostasis by the CYP7A1/SREBP2/miR-33a axis. Hepatology 58:1111–1121. Additional support and detail into the mechanism of mir-33a and bile acid metabolism
•• Tomita K, Teratani T, Suzuki T et al (2014) Free cholesterol accumulation in hepatic stellate cells: mechanism of liver fibrosis aggravation in nonalcoholic steatohepatitis in mice. Hepatology 59:154–169. Evidence of how miR-33 is involved in inflammation that is part of fibrosis in NASH
• Horie T, Nishino T, Baba O, et al (2013) MicroRNA-33 regulates sterol regulatory element-binding protein 1 expression in mice. Nat Commun 4:2883. Communication on miR-33 and an additional SREBP it modulates
Davalos A, Goedeke L, Smibert P et al (2011) miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proc Natl Acad Sci USA 108:9232–9237
Wang B, Majumder S, Nuovo G et al (2009) Role of microRNA-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino-defined diet in C57BL/6 mice. Hepatology 50:1152–1161
Dolganiuc A, Petrasek J, Kodys K et al (2009) MicroRNA expression profile in Lieber–DeCarli diet-induced alcoholic and methionine choline deficient diet-induced nonalcoholic steatohepatitis models in mice. Alcohol Clin Exp Res 33:1704–1710
Hill-Baskin AE, Markiewski MM, Buchner DA et al (2009) Diet-induced hepatocellular carcinoma in genetically predisposed mice. Hum Mol Genet 18:2975–2988
Marra F, Gastaldelli A, Svegliati Baroni G et al (2008) Molecular basis and mechanisms of progression of non-alcoholic steatohepatitis. Trends Mol Med 14:72–81
Siegel AB, Zhu AX (2009) Metabolic syndrome and hepatocellular carcinoma: two growing epidemics with a potential link. Cancer 115:5651–5661
Hermeking H (2010) The miR-34 family in cancer and apoptosis. Cell Death Differ 17:193–199
Yamakuchi M, Ferlito M, Lowenstein CJ (2008) MiR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci USA 105:13421–13426
•• Chen, Q. Li Lei, et al (2014) MiR-34a regulates apoptosis in liver cells by targeting the KLF4 gene. Cell Mol Biol Lett 19:52–64. The research linking miR-34a to apoptosis through KLF4
Zhang W, Shields JM, Sogawa K et al (1998) The gut-enriched Kruppel-like factor suppresses the activity of the CYP1A1 promoter in a Spl-dependent fashion. J Biol Chem 273:17917–17925
Yoon HS, Chen X, Yang VW (2003) Kruppel-like factor 4 mediates p53-dependent G1/S cell cycle arrest in response to DNA damage. J Biol Chem 278:2101–2105
Yoon HS, Yang VW (2004) Requirement of Kruppel-like factor 4 in preventing entry into mitosis following DNA damage. J Biol Chem 279:5035–5041
Chang TC, Wentzel EA, Kent OA et al (2007) Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell 26:745–752
Panasiuk A, Dzieciol J, Panasiuk B, Prokopowicz D (2006) Expression of p53, Bax and Bcl-2 proteins in hepatocytes in non-alcoholic fatty liver disease. World J Gastroenterol 12:6198–6202
Farrell GC, Larter CZ, Hou JY et al (2009) Apoptosis in experimental NASH is associated with p53 activation and TRAIL receptor expression. J Gastroenterol Hepatol 24:443–452
•• Rui E, et al (2013) miR-34a/SIRT1/p53 is suppressed by ursodeoxycholic acid in the rat liver and activated by disease severity in human non-alcoholic fatty liver disease. J Hepatol 58:119–125. The use of urosodeoxycholic as a way to modulate the miR-34a/SIRT1/p53 apoptotic pathway in NASH
Cermelli S, Ruggieri A, Marrero JA et al (2011) Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS ONE 6:e23937
Kong L, Zhu J et al (2011) Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2 diabetes: a clinical study. Acta Diabetol Mar 48(1):61–69
•• Ng R, Wu H, Xiao H, et al (2014) Inhibition of miR-24 expression in liver prevents hepatic lipid accumulation and hyperlipidemia. Hepatology. doi:10.1002/hep.27153. The primary study of miR-24 demonstrating its connection to NASH but also possible therapeutic benefit when inhibited
Yabe D, Brown MS, Goldstein JL (2002) Insig-2, a second endoplasmic reticulum protein that binds SCAP and blocks export of sterol regulatory element-binding proteins. Proc Nat Acad Sci USA 99:12753–12758
Gong Y, Lee JN, Lee PC et al (2006) Sterol-regulated ubiquitination and degradation of Insig-1 creates a convergent mechanism for feedback control of cholesterol synthesis and uptake. Cell Metab 3:15–24
Yang T, Espenshade PJ, Wright ME et al (2002) Crucial step in cholesterol homeostasis: sterols promote binding of SCAP to INSIG-1, a membrane protein that facilitates retention of SREBPs in ER. Cell 110:489–500
Brown MS, Goldstein JL (1999) A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood. Proc Nat Acad Sci USA 96:11041–11048
Edwards PA, Tabor D, Kast HR, Venkateswaran A (2000) Regulation of gene expression by SREBP and SCAP. Biochim Biophys Acta 1529:103–113
Horton JD, Goldstein JL, Brown MS (2002) SREBPs: activators of the complete program of cholesterol and fatty acid synthesis in the liver. J Clin Investig 109:1125–1131
Engelking LJ, Kuriyama H, Hammer RE et al (2004) Overexpression of Insig-1 in the livers of transgenic mice inhibits SREBP processing and reduces insulin-stimulated lipogenesis. J Clin Investig 113:1168–1175
Engelking LJ, Liang G, Hammer RE et al (2005) Schoenheimer effect explained–feedback regulation of cholesterol synthesis in mice mediated by Insig proteins. J Clin Investig 115:2489–2498
Acknowledgments
This work was supported, in whole or in part, by NIH Grants R01 DK081450-03 and T32 07150-37 to AJS.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vincent, R., Sanyal, A. Recent Advances in Understanding of NASH: MicroRNAs as Both Biochemical Markers and Players. Curr Pathobiol Rep 2, 109–116 (2014). https://doi.org/10.1007/s40139-014-0049-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40139-014-0049-8