Abstract
Honey bee abdominal pigmentation is one of the most recognisable traits and it is often used by beekeepers as an indicator of M-lineage subspecies purity. However, this approach may negatively impact population diversity and is futile if there is no association between tergite colour patterns and the genetic background. To assess whether this trait can be used as a proxy for introgression proportions in M-lineage subspecies, we genotyped, with highly informative SNP assays, A. m. mellifera and A. m. iberiensis individuals displaying four different colour phenotypes. The SNP data detected highly introgressed bees exhibiting a black phenotype and, at the same time, pure or marginally introgressed bees with yellow banding patterns, in both subspecies. Despite these observations, contrary to A. m. iberiensis, in A. m. mellifera, introgression proportions revealed to be a significant predictor of abdominal pigmentation. Therefore, abdominal pigmentation could be used by A. m. mellifera conservationists to guide colony selection when genetic tools are unavailable.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Colour was one of the first characters employed in early honey bee systematics (Alpatov 1929; Goetze 1940; Ruttner 1988; Woyke 1977). Yet, this character is not well suited for discriminant analysis used in morphometry because it is bimodally distributed among populations (Ruttner 1988). In spite of this difficulty, colour is an important discriminating factor easily classified in the field (Ruttner 1988). Furthermore, assessment of colour traits (e.g. pigmentation of tergites 2, 3 and 4), in combination with wing venation, pilosity and body size, is still recommended in honey bee identification, especially when unknown variation is under scrutiny (Meixner et al. 2013).
The consistency of distinct colour patterns in different honey bee subspecies was interpreted as indicative of high heritability, which stimulated research on colour inheritance in the past. Roberts and Mackensen (1951) performed reciprocal crosses between subspecies exhibiting yellow and black phenotypes and concluded that abdominal colour was governed by seven genes. In a similar experiment, Woyke (1977) found that colour inheritance was determined by only three major genes, although their expression was modified by six to seven polygenes. Later on, Szabo and Lefkovitch (1992) showed that colour heritability varied among castes with drones having higher values (0.39) than workers (0.32) or queens (0.21–0.23). While these heritability values suggest there is an important genetic component underlying colour phenotypes, environment can also influence this trait. Indeed, it has been shown that temperature during pupal development plays an important role in the phenotypic expression of colour in workers or queens of different honey bee species with the dark morphs being developed at lower temperatures than the yellow morphs (Spivak et al. 1992; Tsuruta et al. 1989).
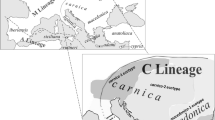
In the geographically large and environmentally diverse territory where the honey bee evolved, there are 31 currently recognized subspecies (Chen et al. 2016; Engel 1999; Meixner et al. 2011; Sheppard and Meixner 2003), which have been grouped into five lineages. Two of these lineages occur naturally in Europe: M in western and northern Europe and C in south-eastern Europe including Italy and the Balkans (Alburaki et al. 2013; Ruttner 1988). In Europe, M-lineage groups only two subspecies, Apis mellifera mellifera and Apis mellifera iberiensis, both being black in colour (Ruttner 1988). Lineage C groups eight subspecies, including one of the most widely spread and phenotypically distinct, the yellow Apis mellifera ligustica from Italy. Another breed widely used by beekeepers is Buckfast, a commercial hybrid with a significant C-lineage component commonly derived from A. m. ligustica (Borsuk and Olszewski 2010; Henriques et al. 2018a; Parejo et al. 2016).
Owing to recurrent introductions of C-lineage queens over the last decades, honey bee’s natural range has been changing dramatically in Europe, especially in the territories of A. m. mellifera in northern Europe and of A. m. iberiensis in the Canaries and Azores (De la Rúa et al. 2006; De la Rúa et al. 2001; De la Rúa et al. 2002; Ferreira et al. 2019; Jensen et al. 2005; Muñoz et al. 2013; Muñoz et al. 2014; Pinto et al. 2014; Soland-Reckeweg et al. 2009). To counteract the demise of the dark bee in northern Europe and of the Canarian black bee in some Canary Islands, conservation efforts have been implemented to protect these subspecies or local variants/ecotypes from further decline (De la Rúa et al. 2009; Muñoz and De la Rúa 2012). In order to do this, beekeepers need to base breeding decisions on a set of characters specific for the subspecies/breed of interest. In Ireland, one such character has been abdominal colour since it can be easily distinguished by eye. Irish beekeepers often re-queen colonies that contain variable yellow-banding phenotypes in order to protect and conserve the dark bee. In contrast, in the Azores, many beekeepers prefer and select for the yellow-banding phenotypes that were originally introduced with C-lineage queens in the 1980s (Ferreira et al. 2019). However, colour-based selection may negatively impact population diversity and is futile if there is no real association between the abdominal colour and the genetic background of the honey bees in a particular colony.
From an academic perspective, colour is no longer a key element in the systematic classification of honey bee diversity. Additionally, morphology-based characters have been complemented with molecular markers for identification of honey bee subspecies and ecotypes (Meixner et al. 2013). Among these, single nucleotide polymorphisms (SNPs) have revealed themselves to be particularly powerful for identifying individuals with variable admixture proportions originating from M- and C-lineage subspecies (Henriques et al. 2018a; Henriques et al. 2018b; Muñoz et al. 2017; Parejo et al. 2016). However, these molecular tools are more expensive and time-consuming than colony selection solely based on visual assessment of abdominal colour. On the other hand, colour assessment is subjective and is thought to be inadequate for subspecies determination (Ruttner 1988).
So far, there has been no direct comparison in determining subspecies status or levels of introgressive hybridisation between molecular data and abdominal colour patterns. In the absence of such comparisons, erroneous assumptions may be made by beekeepers in their breeding and conservation activities. For example, bees showing some yellow marks may be natural colour morphs of A. m. mellifera or A. m. iberiensis (Adam 1983), and consequently their elimination from the breeding pool may lead to the loss of unique combinations of genes. On the other hand, a queen heading a colony of black bees may have an admixed ancestry and maintenance of its descendants in the breeding population would continue the genetic pollution of native populations. In this context, the aim of this study was to verify if there is an association between the abdominal colour pattern and the introgression level inferred from SNPs in the two European M-lineage subspecies. For this purpose, we classified 98 individuals of A. m. iberiensis and 163 individuals of A. m. mellifera into four colour phenotypes and genotyped them using the best two ancestry-informative SNP assays recently designed for the two subspecies (Henriques et al. 2018a; Henriques et al. 2018b).
2 Material and methods
2.1 Sampling
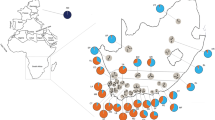
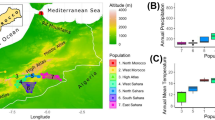
A total of 260 individuals were randomly sampled from the geographical range of A. m. iberiensis and A. m. mellifera. A. m. iberiensis is represented by 84 colonies (98 individuals) and A. m. mellifera by 48 colonies (162 individuals; Figure 1; Table S1). Colonies of A. m. iberiensis originated from Iberia (Portugal N = 58, Spain N = 2) and from the archipelagos of the Azores (N = 36) and Madeira (N = 2). Among the 84 A. m. iberiensis colonies, 77 were represented by a single individual, whereas seven were represented by three individuals exhibiting different colour phenotypes (Table S2). The A. m. mellifera colonies originated from Ireland (N = 71) and the United Kingdom (N = 91). Among the 48 A. m. mellifera colonies, 20 were represented by a single individual and 28 were represented by a variable number of individuals (between four and eight) displaying different phenotypes (Table S2).
Four phenotypes were identified in both subspecies (Figure S1): metasoma entirely black (BLK), presence of a yellow spot in opposite sides of tergite 2 (SPT), presence of a yellow band in tergite 2 (B1T) and presence of yellow bands in tergites 1, 2 and, sometimes, 3 (B2T). The sample size per phenotype was variable, being BLK = 154 (48 A. m. iberiensis and 106 A. m. mellifera), SPT = 24 (15 A. m. iberiensis and 9 A. m. mellifera), B1T = 36 (15 A. m. iberiensis and 21 A. m. mellifera), and B2T = 46 (20 A. m. iberiensis and 26 A. m. mellifera).
2.2 DNA extraction and genotyping
For A. m. iberiensis, total DNA was extracted from the mesosoma of adult workers using the Ron’s Tissue DNA Mini Kit (®Bioron), following the manufacturer instructions and including an additional step of RNA degradation (RNase A). For A. m. mellifera, total DNA was extracted from the legs of adult workers using the Omega bio-tek EZNA kit. All DNA extracts were diluted to 10–15 ng/μl and sent to Instituto Gulbenkian de Ciência (Portugal) for SNP genotyping using the Agena Bioscience iPLEX chemistry and the MassARRAY® MALDI-TOF platform (Gabriel et al. 2009).
The 260 individuals were genotyped using the two best ancestry-informative SNP assays previously developed to estimate C-lineage introgression into A. m. iberiensis (Table S3; Henriques et al. 2018b) and A. m. mellifera (Table S4; Henriques et al. 2018a). To deal with poor or inconsistent amplification, SNP loci with missing data > 60% were discarded from the dataset.
2.3 Introgression estimation
Introgression proportions (Q-values) from C-lineage into both M-lineage subspecies were estimated for each individual using admixture (Alexander et al. 2009). Analyses of the Iberian samples were performed using reference populations of Henriques et al. (2018a), namely: A. m. iberiensis (N = 29), A. m. carnica (N = 5) and A. m. ligustica (N = 10). Analyses of the dark bee samples were performed using the reference populations of Henriques et al. (2018b), namely: A. m. mellifera (N = 28), A. m. carnica (N = 22) and A. m. ligustica (N = 17). Individuals from the same colony were analysed in separate admixture runs to avoid potential bias when estimating Q-values (Alexander et al. 2009; Rodríguez-Ramilo et al. 2014). Q-values were inferred for each individual for K = 2 using 10,000 iterations in 20 independent runs. The convergence between iterations was examined by comparing log-likelihood scores (LLS) using the default termination criteria set to stop when LLS increases by < 0.0001 between iterations. CLUMPAK was employed to summarize and visualize Q-plots (Kopelman et al. 2015). A threshold Q-value ≤ 0.05 was used to identify purebred A. m. iberiensis and A. m. mellifera individuals.
2.4 Statistical analysis
An intercolonial analysis was performed independently for each subspecies using a single individual from the 84 A. m. iberiensis and 48 A. m. mellifera colonies. Individuals were chosen to maximise the sample size per phenotypic class, as follows: BLK- A. m. iberiensis = 41 and A. m. mellifera = 24; SPT- A. m. iberiensis = 14 and A. m. mellifera = 6; B1T- A. m. iberiensis = 15 and A. m. mellifera = 10; B2T- A. m. iberiensis = 14; A. m. mellifera = 8 (Table S1). Variance analysis was performed using the Kruskal–Wallis test (Kruskal and Wallis 1952) as the data did not conform to the normal distribution (Shapiro–Wilk test; Shapiro and Wilk 1965) and the variance was not homogeneous across the four phenotypic groups (Levene test; Levene 1961). Pairwise multiple comparisons were implemented with the Dunn test (Dunn 1964), which is appropriate for groups with uneven sample sizes.
An intracolonial analysis was performed for A. m. mellifera using the 28 colonies with multiple genotyped individuals. While eight colonies were represented by a single phenotype (BLK), 20 were represented by two (11 colonies), three (eight colonies) and four (one colony) different phenotypes (Table S2). If introgression levels would differ among phenotypes, colonies containing individuals with multiple phenotypes would exhibit higher Q-value variability than colonies harbouring only the BLK phenotype. To test this hypothesis, the coefficient of variation (CV) was calculated for each colony by dividing the standard deviation by the mean. CV estimates the dispersion of the variable excluding the order of magnitude (Everitt 1998). Values of the CV were compared among the BLK and the phenotypically diverse colonies using the Kruskal–Wallis test.
To test whether C-lineage introgression was a predictor of abdominal pigmentation, a mixed-effect logistic regression analysis was performed for each subspecies, treating colony ID as a random effect. The presence/absence of yellow bands on the metasoma was treated as the dependent variable and the introgression level as the predictor variable. The phenotype was converted into a binary variable being BLK and SPT coded as ‘1’ and B1T and B2T as ‘0’. All statistical tests were performed in R (Team 2013).
3 Results
The 98 A. m. iberiensis individuals were genotyped using 79 highly informative SNPs (Henriques et al. 2018b). Of these, four SNPs did not pass the established threshold for missing data and were removed from the analysis (Table S3). The 162 A. m. mellifera individuals were genotyped using a different set of 66 highly informative SNPs (Henriques et al. 2018a). The missing data for A. m. mellifera was higher than for A. m. iberiensis, as eight SNPs had to be discarded (Table S4). After this quality control step, the missing data for the 260 individuals varied between 0 and 38.3%. For most individuals (76.9%), the missing data was lower than 10%, indicating a high genotyping success rate.
In the intercolonial analysis, the introgression proportions (Q-values) were inferred from the highly informative SNPs for 84 A. m. iberiensis and 48 A. m. mellifera individuals, each representing a single colony (Figure 2). Most A. m. iberiensis individuals (71, 84.5%) were purebred, with only 13 (15.5%), all from the Azores, showing introgression levels above the established threshold for purity (0.055 ≤ Q-value ≤ 0.193; Table I and S1). Admixture was higher in A. m. mellifera than in A. m. iberiensis as 44% of the individuals had Q-values ranging from 0.0621 to 0.654. This finding is not surprising given that 42% of the A. m. mellifera individuals were collected in the UK (Table I and S1), where recurrent large-scale importation of C-lineage queens is known.
Admixture plots showing the membership partitioning (Q-value) into two clusters (K = 2). The vertical bars represent individuals, each corresponding to a single colony. Blue represents the M-lineage a A. m. iberiensis or b A. m. mellifera cluster and orange the C-lineage cluster. The black vertical lines separate individuals from different subspecies or phenotypes (BLK = black; SPT = Spot; B1T = yellow band in tergite 2; B2T = yellow bands in tergites 1, 2 and, sometimes, 3).
The statistics by phenotype are shown for both subspecies in Figure 3 and Table I. The four phenotypes contained individuals with a wide range of introgression proportions, comprising both pure and introgressed individuals. However, introgression was more variable in A. m. mellifera than in A. m. iberiensis. While in A. m. iberiensis pure individuals were nearly evenly distributed in each of the four phenotypes (BLK = 87.8%; SPT = 85.7%; B1T = 80.0%; B2T = 78.6%); in A. m. mellifera, there was a lower percentage of pure individuals in B1T (20.0%) and B2T (25.0%) than in BLK (75.0%) and SPT (83.3%). The minimum Q-values were 0.000 across the four phenotypes in A. m. iberiensis, whereas they were slightly higher in B1T (0.031) and B2T (0.017) than in BLK (0.000) in A. m. mellifera. The maximum Q-values showed a higher range in both subspecies. In A. m. iberiensis, the highest maximum Q-value was observed for BLK (0.193), whereas the lowest was observed for B2T (0.099). This unexpected finding contrasts with that obtained for A. m. mellifera in which the highest maximum Q-value (0.654) was obtained for B2T, although closely followed by BLK (0.547).
Boxplots showing the C-lineage introgression levels (Q-values) into A. m. iberiensis (N = 84) and A. m. mellifera (N = 48) by abdominal colour phenotype (BLK = black; SPT = Spot; B1T = yellow band in tergite 2; B2T = yellow bands in tergites 1, 2 and, sometimes, 3). The blue dots denote the Q-values obtained for each individual. The boxes indicate the first and third quartiles. The median accuracy is marked with a bold horizontal line within the box.
The mean Q-values estimated for A. m. iberiensis indicate a low (0.019 for BLK ≤ Q-value ≤ 0.028 for SPT; Table I) and similar level of introgression across the four phenotypes (P-value = 0.400, Kruskal-Wallis test), a result further supported by the mixed-effects logistic regression (P-value = 0.878). A different pattern was observed for A. m. mellifera as the average Q-values obtained for B1T (0.170) and B2T (0.299) were greater than those for BLK (0.085) and SPT (0.050; P- value = 0.005, Kruskal–Wallis test), being the phenotypes B1T and B2T significantly more introgressed than the BLK and SPT phenotypes (P-value ≤ 0.02, Dunn test; Table II). The mixed-effects logistic regression showed that the introgression level was a significant predictor of the presence of yellow bands (P-value = 0.003).
In the intracolonial analysis, the introgression proportions were inferred from seven A. m. iberiensis colonies, each with three different phenotypes and individuals, and 28 A. m. mellifera colonies, each with a variable number of phenotypes and individuals (Table S2). Of the seven A. m. iberiensis colonies, five carried only pure individuals, regardless of the phenotype. Two colonies showed signs of admixture, with the BLK phenotype exhibiting the highest Q-values (Figure S2). Of the 28 A. m. mellifera colonies, only five revealed to be purebred. Among the 20 phenotypically diverse colonies (eight only had the BLK phenotype), the BLK individuals frequently showed the lowest Q-values (Table S1), contrasting with the B2T individuals which frequently exhibited the highest Q-values (Table S1). Yet, there were also colonies where the BLK phenotype had the highest Q-values and the B2T phenotype had the lowest Q-values (Figure S2). Analysis of the intracolonial variability in A. m. mellifera indicates that the CV in the phenotypically diverse colonies and in colonies containing only BLK individuals (Figure 4) was not significantly different (P- value = 0.88, Kruskal–Wallis test).
4 Discussion
In this study, A. m. mellifera showed higher levels of C-lineage introgression than A. m. iberiensis, as expected. A. m. mellifera genetic integrity has long been threatened by beekeeper-mediated C-lineage gene flow across most of its distributional range (Bertrand et al. 2015; Hassett et al. 2018; Jensen et al. 2005; Parejo et al. 2016; Pinto et al. 2014; Soland-Reckeweg et al. 2009). A. m. mellifera colonies from the UK (85% of the individuals had 0.055 ≤ Q-values ≤ 0.654) were more introgressed than those from Ireland (35% of the individuals had 0.055 ≤ Q-values ≤ 0.619). The same trend has been reported elsewhere (Henriques et al. 2018a; Jensen et al. 2005), supporting the findings of Hassett et al. (2018) who showed that Ireland is an important reservoir of A. m. mellifera diversity. The difference in introgression levels can be explained by the considerably higher number of imported queens in the UK (N.B.U 2018) than in Ireland (direct comm. from the Ireland’s Department of Agriculture Food and Marine; Table S5). In contrast with A. m. mellifera, the numerous genetic surveys carried out in Iberia have repeatedly documented a well-preserved A. m. iberiensis gene pool (Arias et al. 2006; Cánovas et al. 2008; Chávez-Galarza et al. 2015; Franck et al. 1998; Henriques et al. 2018c; Miguel et al. 2007; Pinto et al. 2013). In accordance with those surveys, all phenotypes collected in Iberia were revealed to be pure. Introgressed A. m. iberiensis individuals (0.055 ≤ Q-values ≤ 0.210) were all from the Azores, where a high frequency of C-lineage haplotypes still exists in several islands (De la Rúa et al. 2006; Ferreira et al. 2019; Muñoz et al. 2013). The ancestors of the extant C-derived colonies were mostly introduced in the 1980s in the framework of a breeding program using A. m. ligustica (Ferreira et al. 2019).
Interestingly, despite the differential introgression levels in A. m. mellifera and A. m. iberiensis, the same colour phenotypes were detected in the two M-lineage subspecies. Colour has been used differently by beekeepers in breeding activities. While in the Azores, many beekeepers prefer and have selected for the yellow phenotype (F. Aguiar, pers. comm.); in Ireland and in the UK, the black phenotype is used by many beekeepers to indicate A. m. mellifera purity. Particularly in areas where A. m. mellifera is threatened with hybridisation from introduced A. m. ligustica and Buckfast strains, colonies that show evidence of yellow banding are regularly re-queened. However, there is variability in the approach with some beekeepers being very intolerant of the existence of any colour variability, while others will accept a small level of banding as natural variation of the local strain (G. McCormack, pers. obs.).
The scientific literature has for some time indicated that honey bee colour by itself is unreliable for intra-species classification because (i) its assessment involves some degree of subjectivity, (ii) its variation is bimodal and not normally distributed (Ruttner 1988) and (iii) it is influenced by environmental factors such as brood nest temperature (Spivak et al. 1992). Assigning individual bees to a phenotype was carried out in this study by eye. Despite the fact that this approach may be perceived as inexact, it was remarkably easy to identify the four phenotypes in both subspecies. Furthermore, we wished to use an identification approach that is applied in practice by beekeepers so that it is highly relevant to beekeeping.
In A. m. iberiensis, introgression was similar among the four phenotypes and therefore a poor predictor of colour. In contrast, in A. m. mellifera, introgression revealed to be a significant predictor of abdominal pigmentation with the B1T and B2T exhibiting the highest introgression values. In the same way, while in A. m. iberiensis the frequency of introgressed individuals was evenly distributed across the four phenotypes, in A. m. mellifera there was a higher frequency of introgressed individuals in B1T and B2T than in BLK and SPT. However, the two most contrasting phenotypes, BLK and B2T, exhibited virtually overlapping introgression ranges 0.000 ≤ Q-values ≤ 0.547 and 0.017 ≤ Q-values ≤ 0.654, respectively. Even within colony, where individuals are genetically and environmentally related, there was no clear association between the introgression level and the colour pattern. Within most of the A. m. iberiensis colonies, the BLK individuals were more introgressed than their B1T and B2T counterparts. Within A. m. mellifera colonies, the trend was that BLK individuals had the lowest colony introgression and B2T the highest. Yet, there were also A. m. mellifera colonies in which the BLK individuals were more introgressed than the B2T individuals. A plausible explanation for this finding is C-lineage introgression originating from A. m. carnica, a non-yellow subspecies (Ruttner 1988).
Genetic variation underlying the yellow abdominal patterns in A. m. mellifera and A. m. iberiensis samples has most likely originated from A. m. ligustica gene flow, as both M-lineage subspecies have been described as velvet or jet black (Ruttner 1988; Adam 1983). Nonetheless, the hypothesis that some level of yellow pigmentation has arisen from standing variation in M-lineage populations cannot be completely rejected. In a legendary trip across Iberia in the 1959, Br. Adam encountered black bees but rarely he also observed bees with “yellow markings confined to the area where the first three dorsal segments join the ventral plate” (Adam 1983). Colour variation in Iberian populations was also described by Luis Mendez de Torres in his beekeeping treatise published in 1586 (Adam 1983), long before large-scale C-lineage trading. These reports suggest that alleles coding for yellow pigmentation are present in A. m. iberiensis gene pool.
Our results suggest that while in A. m. iberiensis identification of pure bees from colour-based phenotypes is unreliable, in A. m. mellifera it is more likely that bees exhibiting yellow bands are introgressed. This finding has differential implications for managing the breeding populations of both subspecies. In the absence of molecular data, A. m. mellifera conservationists should maintain BLK and SPT phenotypes in the breeding population and eradicate B1T and B2T, as opposed to keeping the four phenotypes in A. m. iberiensis. When molecular tools are affordable, identification of introgressed colonies from genotype data should be used to avoid purging of valuable genetic diversity.
References
Adam, B. (1983) In search of the best strains of honey bee Northern Bee Books, 206p, UK 36:335–344
Alburaki M., Bertrand B., Legout H., Moulin S., Alburaki A., Sheppard W.S., Garnery L. (2013) A fifth major genetic group among honeybees revealed in Syria. BMC genetics 14, 117
Alexander D.H., Novembre J., Lange K. (2009) Fast model-based estimation of ancestry in unrelated individuals. Genome Research 19(9), 1655–1664
Alpatov W. (1929) Biometrical studies on variation and races of the honey bee (Apis mellifera L.). The Quarterly Review of Biology 4, 1–58
Arias M.C., Rinderer T.E., Sheppard W.S. (2006) Further characterization of honey bees from the Iberian Peninsula by allozyme, morphometric and mtDNA haplotype analyses. Journal of Apicultural Research 45, 188–196
Bertrand B., Alburaki M., Legout H., Moulin S., Mougel F., Garnery L. (2015) MtDNA COI-COII marker and drone congregation area: An efficient method to establish and monitor honeybee (Apis mellifera L.) conservation centres. Molecular Ecology Resources 15, 673–683
Cánovas F., De la Rúa P., Serrano J., Galián J. (2008) Geographical patterns of mitochondrial DNA variation in Apis mellifera iberiensis (Hymenoptera: Apidae). Journal of Zoological Systematics and Evolutionary Research 46, 24–30
Chávez-Galarza J., Henriques D., Johnston J.S., Carneiro M., Rufino J., Patton J.C., Pinto M.A. (2015) Revisiting the Iberian honey bee (Apis mellifera iberiensis) contact zone: maternal and genome-wide nuclear variations provide support for secondary contact from historical refugia. Molecular Ecology 24, 2973–2992
Chen C., Liu Z., Pan Q., Chen X., Wang H. et al (2016) Genomic analyses reveal demographic history and temperate adaptation of the newly discovered honey bee subspecies Apis mellifera sinisxinyuan n. ssp. Molecular Biology and Evolution 33, 1337–1348
De la Rúa P., Galián J., Pedersen B.V., Serrano J. (2006) Molecular characterization and population structure of Apis mellifera from Madeira and the Azores. Apidologie 37, 699–708
De la Rúa P., Galián J., Serrano J., Moritz R. (2001) Genetic structure and distinctness of Apis mellifera L. populations from the Canary Islands. Molecular Ecology 10, 1733–1742
De la Rúa P., Jaffé R., Dall'Olio R., Muñoz I., Serrano J. (2009) Biodiversity, conservation and current threats to European honeybees. Apidologie 40, 263–284
De la Rúa P., Serrano J., Galián J. (2002) Biodiversity of Apis mellifera populations from Tenerife (Canary Islands) and hybridisation with East European races. Biodiversity & Conservation 11, 59–67
Dunn O.J. (1964) Multiple comparisons using rank sums. Technometrics 6, 241–252
Engel M.S. (1999) The taxonomy of recent and fossil honey bees (Hymenoptera: Apidae; Apis). Journal of Hymenoptera Research 8, 165–196
Everitt, B. (1998) The cambridge dictionary of statistics cambridge university press Cambridge
Ferreira, F., Henriques, D., Neves, J. C., Machado, C. A. S., Azevedo, J. C., Francoy, T. M., Pinto, M. A. (2019) Historical and contemporaneous human-mediated processes left a strong genetic signature on honey bee populations from the Macaronesian archipelago of the Azores. Apidologie (online) 1–13. https://doi.org/10.1007/s13592-019-00720-w
Franck P., Garnery L., Solignac M., Cornuet J.M. (1998) The origin of west European subspecies of honeybees (Apis mellifera): new insights from microsatellite and mitochondrial data. Evolution 52, 1119–1134
Gabriel S., Ziaugra L., Tabbaa D. (2009) SNP genotyping using the Sequenom MassARRAY iPLEX platform. Current Protocols in Human Genetics 60, 2.12. 11–12.12. 18
Goetze G. (1940) The best bee: methods for selecting bees for (great) length of tongue. Insectes Sociaux 3, 335–346
Borsuk G., Olszewski K. (2010) Morphometric traits of Buckfast and Caucasian bees. Journal of Apicultural Science 54, 43–48
Hassett J., Browne K.A., McCormack G.P., Moore E., Society N.I.H.B., Soland G., Geary M. (2018) A significant pure population of the dark European honey bee (Apis mellifera mellifera) remains in Ireland. Journal of Apicultural Research 57, 337–350
Henriques D., Browne K.A., Barnett M.W., Parejo M., Kryger P. et al (2018a) High sample throughput genotyping for estimating C-lineage introgression in the dark honeybee: an accurate and cost-effective SNP-based tool. Scientific Reports 8, 8552
Henriques D., Parejo M., Vignal A., Wragg D., Wallberg A., Webster M.T., Pinto M.A. (2018b) Developing reduced SNP assays from whole-genome sequence data to estimate introgression in an organism with complex genetic patterns, the Iberian honeybee (Apis mellifera iberiensis). Evolutionary Applications 11, 1270–1282
Henriques D., Wallberg A., Chávez-Galarza J., Johnston J.S., Webster M.T., Pinto M.A. (2018c) Whole genome SNP-associated signatures of local adaptation in honeybees of the Iberian Peninsula. Scientific Reports 8, 11145
Jensen A.B., Palmer K.A., Boomsma J.J., Pedersen B.V. (2005) Varying degrees of Apis mellifera ligustica introgression in protected populations of the black honeybee, Apis mellifera mellifera, in northwest Europe. Molecular Ecology 14, 93–106
Kopelman N.M., Mayzel J., Jakobsson M., Rosenberg N.A., Mayrose I. (2015) Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Molecular Ecology Resources 15, 1179–1191
Kruskal W.H., Wallis W.A. (1952) Use of ranks in one-criterion variance analysis. Journal of the American Statistical Association 47, 583–621
Levene, H. (1961) Robust tests for equality of variances. Contributions to probability and statistics Essays in honor of Harold Hotelling 279–292
Meixner M.D., Leta M.A., Koeniger N., Fuchs S. (2011) The honey bees of Ethiopia represent a new subspecies of Apis mellifera-Apis mellifera simensis n. ssp. Apidologie 42, 425–437
Meixner M.D., Pinto M.A., Bouga M., Kryger P., Ivanova E., Fuchs S. (2013) Standard methods for characterising subspecies and ecotypes of Apis mellifera. Journal of Apicultural Research 52, 1–28
Miguel I., Iriondo M., Garnery L., Sheppard W.S., Estonba A. (2007) Gene flow within the M evolutionary lineage of Apis mellifera: role of the Pyrenees, isolation by distance and post-glacial re-colonization routes in the western Europe. Apidologie 38, 141–155
Muñoz I., De la Rúa P. (2012) Temporal analysis of the genetic diversity in a honey bee mating area of an island population (La Palma, Canary Islands, Spain). Journal of Apicultural Science 56, 41–49
Muñoz I., Henriques D., Jara L., Johnston J.S., Chávez-Galarza J., De la Rúa P., Pinto M.A. (2017) SNPs selected by information content outperform randomly selected microsatellite loci for delineating genetic identification and introgression in the endangered dark European honeybee (Apis mellifera mellifera). Molecular Ecology Resources 17, 783–795
Muñoz I., Pinto M.A., De la Rúa P. (2013) Temporal changes in mitochondrial diversity highlights contrasting population events in Macaronesian honey bees. Apidologie 44, 295–305
Muñoz I., Pinto M.A., De la Rúa P. (2014) Effects of queen importation on the genetic diversity of Macaronesian island honey bee populations (Apis mellifera Linneaus 1758). Journal of Apicultural Research 53, 296–302
N.B.U (2018) National Bee Unit [online] http://www.nationalbeeunitcom/. Accessed 22 July 2019
Parejo M., Wragg D., Gauthier L., Vignal A., Neumann P., Neuditschko M. (2016) Using whole-genome sequence information to foster conservation efforts for the European dark honey bee, Apis mellifera mellifera. Frontiers in Ecology and Evolution 4, 140
Pinto M.A., Henriques D., Chávez-Galarza J., Kryger P., Garnery L. et al (2014) Genetic integrity of the Dark European honey bee (Apis mellifera mellifera) from protected populations: a genome-wide assessment using SNPs and mtDNA sequence data. Journal of Apicultural Research 53, 269–278
Pinto M.A., Henriques D., Neto M., Guedes H., Muñoz I., Azevedo J.C., De la Rúa P. (2013) Maternal diversity patterns of Ibero-Atlantic populations reveal further complexity of Iberian honeybees. Apidologie 44, 430–439
Roberts W.C., Mackensen O. (1951) Breeding improved honey bees. American Bee Journal 91, 292–294
Rodríguez-Ramilo S.T., Toro M.A., Wang J., Fernández J. (2014) Improving the inference of population genetic structure in the presence of related individuals. Genetics Research 96, e003
Ruttner, F. (1988) Biogeography and taxonomy of honeybees. Springer Science & Business Media,
Shapiro S.S., Wilk M.B. (1965) An analysis of variance test for normality (complete samples). Biometrika 52, 591–611
Sheppard W.S., Meixner M.D. (2003) Apis mellifera pomonella, a new honey bee subspecies from Central Asia. Apidologie 34, 367–375
Soland-Reckeweg G., Heckel G., Neumann P., Fluri P., Excoffier L. (2009) Gene flow in admixed populations and implications for the conservation of the Western honeybee, Apis mellifera. Journal of Insect Conservation 13, 317
Spivak M., Zeltzer A., Degrandi-Hoffman G., Martin J.H. (1992) Influence of temperature on rate of development and color patterns of queen honey bees (Hymenoptera: Apidae). Environmental Entomology 21, 364–370
Szabo, T., Lefkovitch, P. (1992) Heritability of colour patterns in a closed population of honeybees. Apidologie 23:151–159
Team, R. C (2013) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna URL http://www.R-project.org
Tsuruta T., Matsuka M., Sasaki M. (1989) Temperature as a causative factor in the seasonal colour dimorphism of Apis cerana japonica workers. Apidologie 20, 149–155
Woyke, J. (1977) The heredity of colour patterns in the honeybee. Genetic, Selection and Reproduction of the Honeybee Bucharest: Bee Biology in Moscow
Acknowledgements
We would like to thank João Costa (Instituto Gulbenkian Ciência, Oeiras, Portugal) for genotyping the SNP panels, and to the beekeepers who supplied bees and insight into the abdominal coloration. Analyses were performed using computational resources at the Instituto Politécnico de Bragança.
Funding
KAB was funded by Dr. Tony Ryan research scholarship and an Irish Research Council scholarship. This work was financed by the Native Irish Honeybee Society (NIHBS), Federation of Irish Beekeeping Associations (FIBKA) and FEDER (Fundo Europeu de Desenvolvimento Regional) through the program COMPETE 2020–POCI (Programa Operacional para a Competividade e Internacionalização) and by the Portuguese funds through FCT (Fundação para a Ciência e a Tecnologia) in the framework of the project BeeHappy (POCI-01-0145-FEDER-029871).
Author information
Authors and Affiliations
Contributions
MAP, GPM and DH conceived the ideas and designed the methodology. CJN, ARL, KAB, RF and GPM identified the phenotypes. CJN, ARL, AQ, KAB and RF did the DNA extractions. DH performed most of the analyses with the assistance of ARL. DH and MAP wrote the manuscript with input from GPM. All the authors critically reviewed the manuscript for important intellectual content.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Manuscript editor: Marina Meixner
L'introgression chez les abeilles de la lignée M peut-elle être détectée par des patrons de couleur abdominale?
Apis mellifera mellifera / Apis mellifera iberiensis / SNPs / baguage jaune / conservation.
Kann Introgression in der M-Linie der Honigbienen durch das Farbmuster des Abdomens festgestellt werden?
Apis mellifera mellifera / Apis mellifera iberiensis / SNPs / gelbe Bienen / Naturschutz.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Henriques, D., Lopes, A.R., Ferrari, R. et al. Can introgression in M-lineage honey bees be detected by abdominal colour patterns?. Apidologie 51, 583–593 (2020). https://doi.org/10.1007/s13592-020-00744-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13592-020-00744-7