Abstract
Chrysanthemum has several hundreds of commercialized cultivars with similar morphological characters including flower color and type. Correct labelling of the cultivars and protecting the intellectual property rights of the breeder requires an accurate and reliable method to discriminate the cultivars. Simple sequence repeat (SSR) markers were used to identify and discriminate the standard type cultivars in chrysanthemum. 23 SSR markers from the chrysanthemum SSR database were used to identify eleven standard type chrysanthemum cultivars. Of the 23 SSRs, 7 markers showed polymorphic fragments. A total of 187 alleles were obtained using 7 SSR markers in 11 cultivars. The number of alleles per locus ranged from two to four with an average of 2.7 per locus. The polymorphism information content (PIC) of 7 SSRs ranged from 0.25 to 0.60 with an average of 0.41. Genetic relationship by cluster analysis based on unweighted pair group method with arithmetic mean (UPGMA) method revealed a genetic distance of 0.25–1.00 among the 11 cultivars. The 11 standard type cultivars were discriminated by the ChSSR_16 marker alone. However, two cultivars (Sei_no_issei and Makoto) showed complete genetic similarity and could not be distinguished. Hence, the results showed that these SSR markers may prove useful in identifying additional chrysanthemum cultivar types and to assess genetic relationships among cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Chrysanthemum (Dendranthema grandiflourm) is a species of perennial herbaceous plant from the family Asteraceae and is commercially important for the floral industry in various countries including Korea. Chrysanthemum is one of the most well-known ornamental plants and is the second most popular cut flower after rose, traded in the global flower market with higher economic value (Kumar et al. 2006). Some chrysanthemum species, such as Chrysanthemum morifolium and Chrysanthemum indicum, have also been widely used as a medicinal tea and for cosmetic purposes, respectively. About 200 species were originally contained within the genus Chrysanthemum, but the majority has been subdivided into 38 satellite genera of the chrysanthemum complex. This genus shows considerable variation in morphology and ploidy levels (from 2n = 2x = 18, to 2n = 36, 54, 72, up to 90) with complex evolutionary history (Liu et al. 2012). Chrysanthemum morifolium has a high ornamental value due to the abundant diversity in the shape and color of the flower. The high diversity in flower architecture is the result of greater genome complexity and the allohexaploid background of the cultivated chrysanthemum (Dowrick and El-Bayoumi 1966). The highly heterozygous nature of cultivated chrysanthemum, due to the self-incompatibility and out-breeding, has led to a delay in its genetic improvement (Drewlow et al. 1973).
Globally, several hundreds of chrysanthemum cultivars have been commercialized as it has the advantage of substantial inbuilt genetic variations which can easily be manipulated under cultivation methods leading to a wide array of phenotypic variations. Moreover, propagation of the cultivars is primarily achieved by vegetative cuttings and suckers (Shudo et al. 2011). Diverse morphological features of chrysanthemum cultivars have been evaluated, including flower size, color, and shape. However, the vast number of varieties has made it difficult to identify and discriminate them solely on morphological characters as they are highly unstable and can originate from distinct environmental conditions. In chrysanthemum, hundreds of cultivars are commercialized with similar flower morphological characters. This poses a major problem in cultivar assignment because of the existence of numerous synonyms among local varieties and incorrect labelling of the cultivar. Cultivar identification is therefore highly essential for breeding, registration, trade, and inspection. Identification and characterization of cultivars will provide valuable information for their introduction into market and genetic improvement. Hence, an effective method for accurate identification of cultivars is essential to ensure the correct labelling of the cultivars in the market and to protect the intellectual property rights of the breeder (Korir et al. 2013). Although phenotypic approaches are an option for cultivar identification, they are less informative and time consuming. Moreover, morphological characters are multigenic, are often visible only at a specific growth stage, and may be influenced by environmental factors, making it difficult to assess them. Hence, the use of molecular markers serves as a practical alternative for cultivar identification as it is more rapid and cost effective.
Genetic markers have wide potential applications in varietal identification and revealing the genetic diversity among germplasms. Earlier studies have used various markers including RAPD (Random Amplified Polymorphic DNA), AFLP (Amplified Fragment Length Polymorphism), SRAP (Sequence Related Amplified Polymorphism), ISSR (Inter Simple Sequence Repeats), and SCoT (Start Codon Targeted polymorphism) markers for genetic linkage map construction and quantitative trait loci (QTL) detection (Bhattacharya and da Silva 2006; Shao et al. 2010; Su et al. 2017; Zhang et al. 2010, 2011; Feng et al. 2016) in cultivated chrysanthemum. However, most of these markers have limitations in cultivar identification as they are dominantly scored, difficult to reproduce, and are laborious (Esselink et al. 2003). In contrast, SSR markers have various advantages over other markers as they have co-dominance and are multi-allelic, relatively abundant, and widely spread across the genome, which renders them easily scored (Powell et al. 1996). Earlier, SSR data bases were constructed and used for various purposes including cultivar identification, seed purity tests and identification of parent-offspring relationships in crops like citrus, pear and oriental melon (Hong et al. 2016; Kim and Nou 2016; Nguyen et al. 2019). In recent years, SSR markers have been used in the variety identification of chrysanthemum cultivars and a number of other crops (Zhang et al. 2014; Kwon et al. 2010; Caramante et al. 2011). Recently, next generation sequencing of the Chrysanthemum nakingense transcriptome was performed and the SSR markers derived from the expressed sequence tags (ESTs) were used for the identification of species (Wang et al. 2013). Jo et al. (2015) also addressed the identification of 50 chrysanthemum cultivars using EST-derived SSR markers developed using next generation sequencing. An SSR marker database was constructed and 147 chrysanthemum cultivars collected in Korea were distinguished using 14 SSR markers (Shim et al. 2015). Chrysanthemum is a commercially important crop, especially standard type cultivars with long, sturdy stems and bigger flowers, which are desired as cut flower and as part of flower arrangements, and therefore fetch high market prices. As the morphological features of standard type cultivars are largely similar, identification and fingerprinting of these cultivars at the genetic level is critical for the protection of breeder’s intellectual property rights. In this view, the present study was focused on developing a method using SSR markers to identify and distinguish standard type chrysanthemum cultivars that are routinely cultivated in Korea, Japan, and China.
2 Materials and methods
2.1 Plant material
A total of eleven standard type chrysanthemum cultivars were evaluated in this study. Iwa_no_hakusen, Zinba, Makoto, Yuka, Sei_no_issei, and Nami are commercial cultivars from Japan. Breeding line 11B1-609, Baekma, Baekgang, Youngma, and Gumo are standard cultivars that were bred at National Institute of Horticulture and Herbal Science (NIHHS), Korea. Rooted seedlings were planted, raised, and maintained in a greenhouse under natural light at NIHHS. For the first 40 days, artificial light with an intensity of 100 µmol photons m−2 s−1 was provided to the plants from 21:00 h to 3:00 h. The daily photoperiod varied between 14 h to 16 h. The temperature of the greenhouse was maintained at 22 ± 5 °C and a relative humidity of 70–75%. Eight plants from each cultivar were used both for phenotypic and SSR analyses. Morphological characters including plant height, flower diameter, nodal length, and days to flowering were monitored in the selected cultivars. To investigate white rust resistance in the tested cultivars, rooted cuttings were planted in the greenhouse in early April and the inoculation of white rust was performed by placing the diseased plants with sporulating lesions among the chrysanthemum cultivar plants. Resistance test was conducted under the minimum night temperature of 15 °C and a relative humidity of about 100%, by spraying water on the leaves and covering them with polyethylene film.
2.2 Genomic DNA isolation
Fresh leaf samples were collected from all standard type chrysanthemum cultivars, frozen in liquid nitrogen, and stored at – 80 °C. Genomic DNA was extracted using the DNeasy Plant Mini Kit (QIAGEN, Germany). DNA quality was checked by electrophoresis on 1% agarose gel and the DNA concentration was quantified using Quick Drop (Molecular Devices, USA). DNA was diluted to 25 ng µL−1 and used for SSR analysis.
2.3 SSR analysis
Fourteen EST-SSR primer pairs with high polymorphism and reproducibility were selected based on primary screening in a previous study (Shim et al. 2015). Nine SSR primer pairs with high polymorphism identified in another study were obtained (Han et al. 2019). Details of the primers are listed in Table 1. A total of 23 SSR markers were used for the identification and classification of 11 standard type chrysanthemum cultivars. The M13 tailing method was used to label the PCR products (Schuelke 2000). Each SSR marker was amplified independently in a 20 µL reaction volume containing 25 ng of genomic DNA, 8 pmol of each forward (5′ FAM labelled) and reverse primer, 0.4 mM dNTPs, 3 mM Mg2 + and 0.3 U of Taq DNA polymerase in 1X PCR supplied buffer. The PCR reaction was performed in a thermocycler (Veriti, Applied Biosystems, CA, USA) using the following cycling parameters: 94 °C (5 min); 35 cycles of 94 °C (1 min), 58 °C (30 s), 72 °C (45 s); and a final extension at 72 °C (30 min). 2 µL of PCR product was checked on 2.5% agarose gel electrophoresis and detected by DNA Loading STAR (Dyne Bio, South Korea). PCR products (1 mL each) were combined with 10 µL of Hi-Di formamide and 0.12 µL of GeneScan-500 ROX internal size standard per well and analyzed on an ABI PRISM 310 0XL Genetic Analyzer (Applied Biosystems). The PCR fragment sizes of the SSR loci were read using GeneMapper (version 3.7) software.
2.4 Data analysis
The total number of alleles, allele frequency, genetic diversity, heterozygosity, and PIC (polymorphism information content) of each SSR locus was calculated using Power Maker version 3.25 (Liu and Muse 2005). The SSR amplification bands were scored 1 for present and 0 for absent. SM (Simple Matching) similarity index was used to calculate the similarity of the qualitative data. Cluster analysis of genetic similarity data was performed using the unweighted pair group method of arithmetic averages (UPGMA) and a dendrogram was obtained using DendroUPGMA software. Principle Component Analysis (PCA) was performed using software PAST (version 3.26) with a bootstrap frequency of n = 500.
3 Results
3.1 Morphological evaluation
Growth and flowering parameters of eleven chrysanthemum cultivars were evaluated (Table 2). Baekma and Iwa_no_hakusen bear white colored flowers, Youngma and Gumo have yellow colored flowers, and the remaining cultivars bear ivory colored flowers (Fig. 1). Youngma exhibited maximum (88.3 cm) plant height, whereas Gumo showed the lowest (53.6 cm) plant height. Flower diameter ranged from 9.6 cm in Gumo to 14 cm in Baekma and 11-609 breeding line. Peduncle length was highest in Gumo (3 cm), followed by Baekma and Baekgang (2.7 cm), and shortest in Makoto (1.4 cm). The minimum number of days to flower ranged from 89 days in the majority of the cultivars to 102 days in Makoto. Yuka is a summer flowering cultivar, while all other tested cultivars flower in winter. Makoto, 11-609, and Zinba have higher number of side branches, whereas Nami, Baekang, and Gumo have medium number, and Sei_no_issei showed the lowest number of side branches. Makoto Youngma exhibited medium resistance to white rust disease, whereas Baekgang and Nami are highly resistant to white rust (Table 3). Baekma, Iwa_no_hakusen, Zinba, Sei_no_issei, Yuka, 11B1-609, and Gumo are susceptible to white rust.
3.2 Genetic variation of SSR markers
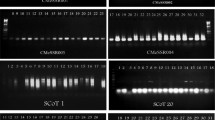
Polymorphism of SSR markers was analyzed in 11 standard type chrysanthemum cultivars. Among the 23 total markers, 7 were polymorphic among the 11 cultivars. Of the 7 SSR markers, 2 markers amplified 4 alleles each, followed by 4 markers that produced 3 alleles each, and 2 alleles were amplified by one marker. Analysis of genetic variation of 6 SSR markers resulted in a total of about 187 alleles. Allele size ranged from 136 to 274 bp. The average number of alleles was about 2.7 per locus, ranging from two (ChSSR_7) to four (ChSSR_16) alleles. Figure 2 shows the results obtained for loci ChSSR_51 and ChSSR_16. Major allele frequency (MAF) of 6 SSR markers ranged from 0.40 to 0.65 with a mean of 0.56 per locus. Observed heterozygosity (HO) ranged from 0.50 to 0.75 with an average of 0.66 (Table 4).
The gene diversity (Nei 1973) among the chrysanthemum cultivars ranged from 0.33 to 0.66 with an average of 0.50 for the 7 SSR markers. The PIC values, which reflect the frequency and allele diversity among the cultivars, were not uniformly high for all of the tested SSR markers. The PIC values ranged from a low of 0.25 (ChSSR_863) to a high of 0.60 (ChSSR_16) with an average PIC value of 0.41 (Table 4). Two markers, ChSSR_51 and ChSSR_16, were considered as informative markers for the identification of standard type chrysanthemum cultivars.
3.3 Genetic relationship analysis among the chrysanthemum cultivars
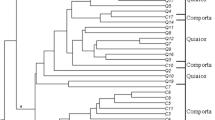
Genetic similarity was analyzed among all the 11 standard type chrysanthemum cultivars based on the Jaccard’s similarity coefficients for the 6 SSR markers. The similarity value for all SSR markers ranged from 0.25 to 1.00. UPGMA cluster analysis based on the genetic similarity values was utilized to construct a dendrogram. All 11 chrysanthemum cultivars could be clearly divided into three major groups with a similarity level of 0.74 (Fig. 3). Six cultivars including Yuka, Iwa_no_hakusen, Nami, Baekgang, Sei_no_Issei, and Makoto were grouped into cluster I. Cluster II consisted of 11-609 and Youngma, whereas Gumo and Zinba formed cluster III. Interestingly, cultivar Baekma failed to form a cluster and was individually separated. A principle component analysis (PCA) plot was derived from the analysis of the SSR data (Fig. 4). The grouping pattern of the PCA plot correlated with the dendrogram obtained by UPGMA-based cluster analysis. The cultivars were scattered throughout the plot. In the PCA plot, Baekma was placed farthest from the remaining cultivars. The grouping pattern of other cultivars also matched the UPGMA-based cluster analysis.
Dendrogram depicting the classification of 11 standard type chrysanthemum cultivars constructed using UPGMA analysis based on SSR markers. The clusters are marked on the right side of the dendrogram. The scale at the bottom is the Jaccard’s coefficient of similarity. Bootstrap values are given as nodal values along the branches
4 Discussion
Chrysanthemum species have complicated genetic backgrounds, often showing hexaploid-based aneuploidy, as well as highly similar morphological features making it difficult to discriminate among cultivars. To classify and identify chrysanthemum cultivars, different molecular markers including inter simple sequence repeats (ISSR) (Miao et al. 2007; Shao et al. 2010), sequence related amplified polymorphism (SRAP) (Fei et al. 2011), and simple sequence repeats (Zhang et al. 2014; Chang et al. 2018) were used. Among these, SSRs have the advantages of co-dominance, hypervariability, and high reproducibility. SSR markers have also been used in genetic diversity analysis, molecular map construction, and for the assessment of intellectual property rights for various crops (Esselink et al. 2003; Feng et al. 2016). SSRs were earlier used for the genetic analysis of chrysanthemum and related genera (Wang et al. 2013; Zhang et al. 2014; Feng et al. 2016; Chang et al. 2018). The present study reports an identification method for the discrimination of standard type cultivars in chrysanthemum.
Earlier studies have established the SSR marker database for the identification of cultivars in chrysanthemum (Han et al. 2019; Shim et al. 2015). Fourteen SSR markers from the chrysanthemum DNA profile database were used to analyze the genetic relationship among the 147 chrysanthemum varieties (Shim et al. 2015). However, only a few studies have explored the utilization of SSR markers for the discrimination of standard type chrysanthemum cultivars (Han et al. 2019). In the present investigation, 23 SSR markers were used to identify and discriminate 11 standard type chrysanthemum cultivars. Among the 23 SSRs, 7 SSR markers showed polymorphism in the 11 tested standard type cultivars. 7 SSRs generated a total of 154 alleles with 2 to 4 alleles per loci with an average of 2.7 alleles. The mean number of alleles per SSR locus is comparable with the results in 8 watermelon genotypes with 1 to 8 alleles (average of 3.6) per each locus tested using 18 SSR markers (Guerra-Sanz 2002). 2 to 6 alleles were reported in 13 watermelon varieties tested with 36 SSR markers (Joobeur et al. 2006). Similarly, Shim et al. (2015) reported 3 to 10 alleles (average of 5.6) per locus in 147 chrysanthemum cultivars tested with 14 SSR markers, which is similar to the results of Hong et al. 2013 for chrysanthemum cultivars. Further, the PIC values, indicating the degree of polymorphism for the 6 SSR markers for the 11 tested cultivars, ranged from 0.25 to 0.60 with an average PIC of 0.41. This result is close to the earlier result for 14 SSR markers in chrysanthemum cultivars with PICs value ranging from 0.28 to 0.78 and a mean PIC of 0.59 (Shim et al. 2015). A higher mean PIC value of 0.90 was reported for 32 chrysanthemum cultivars (Feng et al. 2016) and a mean PIC of 0.79 was reported for 88 chrysanthemum cultivars by Chang et al. (2018). Jo et al. (2015) reported PIC values of SSR markers ranging from 0.15 to 0.88 (mean PIC of 0.60) in 50 chrysanthemum cultivars. However, the mean PIC value was higher in Hong et al. (2013), which was 0.88 in chrysanthemum. This variation in PIC values is probably due to differences in the number of SSR markers used and the genetic diversity among the species. Two SSRs, ChSSR_51 and ChSSR_16, showed higher polymorphism among the chrysanthemum cultivars in this study.
The basis for selective breeding and cultivar identification is the evaluation of genetic relationships among populations. A UPGMA-based dendrogram, constructed based on the SSR data, showed that the tested chrysanthemum cultivars were grouped into three major groups with a similarity matrix ranging from 0.25 to 1, while Baekma alone separated individually. Based on the genotype of SSR markers used in the study, among the 6 cultivars including Yuka, Iwa_no_hakusen, Nami, Baekgang, Sei_no_issei, and Makoto that formed cluster I, Sei_no_issei and Makoto could not be distinguished and showed complete genetic similarity. This indicates that the genetic diversity is relatively low and that those cultivars might have been developed with a narrow genetic background. In an earlier study, standard type chrysanthemum cultivars White runner and Pingpong showed 100% genetic similarity and could not be distinguished (Shim et al. 2015). All cultivars in cluster I exhibit white rust disease resistance. However, Baekma and Zinba are genetically diverged in this study, which is similar to the results of Shim et al. (2015), who found that Baekma and Zinba showed a distant relationship.
In conclusion, this study demonstrates that our SSR marker method is a powerful tool to evaluate genetic relationships among closely related cultivars and to identify and discriminate among chrysanthemum cultivars. These microsatellites can be used to certify protected varieties and for pedigree analysis. Further studies are necessary to verify these results in other types of chrysanthemum cultivars.
References
Bhattacharya A, da Silva JAT (2006) Molecular systematics in Chrysanthemum x grandiflorum (Ramat.) Kitamura. Sci Horticult 109:379–384
Caramante M, Corrado G, Monti LM, Rao R (2011) Simple sequence repeats are able to trace tomato cultivars in tomato food chains. Food Con 22:549–554
Chang L, Dongliang C, Xi C, Hua L, Yahui L, Conglin H (2018) SSR analysis of genetic relationship and classification in Chrysanthemum germplasm collection. Hort Plant J 4:73–82
Dowrick GJ, El-Bayoumi A (1966) The origin of new forms of the garden Chrysanthemum. Euphytica 15:32–38
Drewlow LW, Ascher PD, Widmer RE (1973) Genetic studies of self-incompatibility in the garden chrysanthemum, Chrysanthemum morifolium Ramat. Theor Appl Genet 43:1–5
Esselink GD, Smulders MJM, Vosman B (2003) Identification of cut rose (Rosa hybrida) and rootstock varieties using robust sequence tagged microsatellite site markers. Theor Appl Genet 106:277–286
Fei Z, Chen S, Chen F, Fang W, Yu C, Li F (2011) SRAP-based map ping and QTL detection for inflorescence-related traits in chrysanthemum (Dendranthema morifolium). Mol Breeding 27:11–23
Feng SG, Hea RF, Jianga MY, Lua JJ, Shenb XX, Liuc JJ, Wangb ZA, Wang HZ (2016) Genetic diversity and relationships of medicinal Chrysanthemum morifolium revealed by start codon targeted (SCoT) markers. Sci Hortic 201:118–123
Guerra-sanz JM (2002) Citrullus simple sequence repeats markers from sequence databases. Mol Ecol Not 2:223–225
Han JY, Kim JB, Lee JH, Hong CP, Pak HS, Kim TS (2019) Development of EST-SSR markers for cultivar determination and genetic diversity studies of commercial chrysanthemums in the Korean floral market. Kor J Breed Sci 51:201–208
Hong WJ, Khaing AA, Park YJ (2013) Cultivar identification of chrysanthemum (Dendranthema grandiflorum. Ramat.) using SSR markers. Korean J Intl Agri 25:385–394
Hong JH, Chae CW, Choi KJ, Kwon YS (2016) A database of Simple Sequence Repeat (SSR) marker-based DNA profiles of Citrus and related cultivars and germplasm. Korean J Hortic Sci Technol 34:142–153 (In Korean)
Jo KM, Jo YH, Chu HS, Lian S, Cho WK (2015) Development of EST-derived SSR markers using next-generation sequencing to reveal the genetic diversity of 50 chrysanthemum cultivars. Biochem Syst Ecol 60:37–45
Joobeur T, Gusmini G, Zhang X, Levi A, Xu Y, Wehner TC, Oliver M, Dean RA (2006) Construction of a watermelon BAC library and identification of SSRs anchored to melon or Arabidopsis genomes. Theor Appl Genet 112:1553–1562
Kim HT, Nou IS (2016) Confirmation of parentage of the Pear cultivar ‘Niitaka’ (Pyrus pyrifolia) based on self-incompatibility haplotypes and genotyping with SSR markers. Korean J Hortic Sci Technol 34:453–460
Korir NK, Han J, Shangguan L, Wang C, Kayesh E, Zhang Y, Fang J (2013) Plant variety and cultivar identification: advances and prospects. Crit Rev Biotechnol 33:111–125
Kumar S, Prasad KV, Choudhary ML (2006) Detection of genetic variability among chrysanthemum radio mutants using RAPD markers. Curr Sci 90:1108–1113
Kwon YS, Oh YH, Yi SI, Kim HY, An JM, Yang SG, Ok SH, Shin JS (2010) Informative SSR markers for commercial variety discrimination in watermelon (Citrullus lanatus). Genes Genom 32:115–122
Liu K, Muse SV (2005) Power Marker: an integrated analysis environment for genetic marker analysis. Bioinfor 21:2128–2129
Liu PL, Wan Q, Guo YP, Yang J, Rao GY (2012) Phylogeny of the genus Chrysanthemum L.: evidence from single-copy nuclear gene and chloroplast DNA sequences. PLoS ONE 7: e48970
Miao HB, Chen FD, Zhao HB (2007) Genetic relationship of 85 chrysanthemum [Dendranthema × grandiflora (Ramat.) Kitamura] cultivars revealed by ISSR analysis. Acta Hort Sin 34:1243–1248 (in Chinese)
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nguyen NN, Kwon YS, Park JR, Sim SC (2019) Development of a core set of SSR markers for cultivar identification and seed purity tests in oriental melon (Cucumis melo L. var. makuwa). Korean J Hortic Sci Technol 37:119-129
Powell W, Machray GC, Provan J (1996) Polymorphism revealed by simple sequence repeats. Trends Plant Sci 1:215–222
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234
Shao QS, Guo QS, Deng YM, Guo HP (2010) A comparative analysis of genetic diversity in medicinal Chrysanthemum morifolium based on morphology, ISSR and SRAP markers. Biochem Syst Ecol 38:1160–1169
Shim EJ, Heo EJ, Yoon MK, Soh EH, Hong JH (2015) Construction of SSR marker database of Chrysanthemum varieties collected in Korea. Korean J Breed Sci 47:366–375 (in Korean)
Shudo A, Goeku S, Tarora K, Gima N, Urasaki N, Tokunaga T, Ureshino K, Miyagi E, Sekizuka S, Adaniya S (2011) Identification of chrysanthemum (Chrysanthemum morifolium) cultivars using flower pigment related genes. Trop Agr Develop 55:166–169
Su MH, Hsu TH, Wang CN, Lin KH, Chiang MC, Kang RD, Nguyen KM, Nguyen H (2017) Genetic diversity of a novel oil crop, Camellia brevistyla, revealed by ISSR DNA markers. Korean J Hortic Sci Technol 35:588–598
Wang H, Jiang J, Chen S, Qi X, Peng H, Li P, Song A, Guan Z, Fang W, Liao Y, Chen F (2013) Next-generation sequencing of the Chrysanthemum nankingense (Asteraceae) transcriptome permits large scale unigene assembly and SSR marker discovery. PLoSONE 8:62293
Zhang F, Chen SM, Chen FD, Fang WM, Li FT (2010) A preliminary genetic linkage map of chrysanthemum (Chrysanthemum morifolium) cultivars using RAPD, ISSR and AFLP markers. Sci Horticult 125:422–428
Zhang F, Chen S, Chen F, Fang W, Deng Y, Chang Q, Liu P (2011) Genetic analysis and associated SRAP markers for flowering traits of chrysanthemum (Chrysanthemummorifolium). Euphytica 177:15–24
Zhang Y, Dai SL, Hong Y, Song XB (2014) Application of genomic SSR locus polymorphisms on the identification and classification of Chrysanthemum cultivars in China. PLoSONE 9:104856
Acknowledgements
This study was financially supported by Rural Development Administration (Project No: PJ01098202) Republic of Korea.
Author information
Authors and Affiliations
Contributions
MM Performed the experiments and wrote the manuscript. OKK Analyzed the data and critically reviewed the manuscript. DYH and KJL Performed the fragment analysis by genetic analyzer. AMS and JTP Performed the morphological analysis of chrysanthemum cultivars. JAJ Conceived and designed the experiments and analyzed the data.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Data availability
All datasets generated and analyzed in the current study are available from the corresponding author on request.
Additional information
Communicated by Inhwa Yeam.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mekapogu, M., Kwon, O.K., Hyun, D.Y. et al. Identification of standard type cultivars in Chrysanthemum (Dendranthema grandiflorum) using SSR markers. Hortic. Environ. Biotechnol. 61, 153–161 (2020). https://doi.org/10.1007/s13580-019-00201-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-019-00201-0