Abstract
Myxofibrosarcoma (MFS) is one of the most aggressive sarcomas with highly complex karyotypes and genomic profiles. Although a complete resection is required in the treatment of MFS, it is often not achieved due to its strong invasive nature. Additionally, MFS is refractory to conventional chemotherapy, leading to poor prognosis. Therefore, it is necessary to develop novel treatment modalities for MFS. Patient-derived cell lines are important tools in basic research and preclinical studies. However, only 10 MFS cell lines have been reported to date. Furthermore, among these cell lines, merely two MFS cell lines are publicly available. Hence, we established a novel MFS cell line named NCC-MFS3-C1, using a surgically resected tumor specimen from a patient with MFS. NCC-MFS3-C1 cells had copy number alterations corresponding to the original tumor. NCC-MFS3-C1 cells demonstrate constant proliferation, spheroid formation, and aggressive invasion. In drug screening tests, the proteasome inhibitor bortezomib and the histone deacetylase inhibitor romidepsin demonstrated significant antiproliferative effects on NCC-MFS3-C1 cells. Thus, the NCC-MFS3-C1 cell line is a useful tool in both basic and preclinical studies for MFS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Myxofibrosarcoma (MFS) is a malignant fibroblastic neoplasm with variably myxoid stroma and pleomorphism [1]. Although MFS is rare, it represents approximately 5% of soft tissue sarcoma diagnoses and is one of the most common sarcomas diagnosed in elderly patients [2, 3]. The typical site of origin is the lower extremities, and notably, more than half of MFS cases arise in dermal or subcutaneous tissues [4]. Genetically, MFS is characterized by highly complex karyotypes and genomic profiles [5]. Although there are no specific genetic aberrations for MFS, an integrated genomic study revealed frequent amplification of the chromosome 5p region, the biological significance of which has not been entirely clarified [6]. Regarding treatment for MFS, surgical resection is considered to be the optimal method. However, due to the strong infiltrative nature of MFS, complete resection is often difficult to achieve [7, 8]. In cases when complete resection is not achieved, it has been reported that local recurrence occurs at a high rate of 30–40% [2, 3, 9, 10]. Additionally, the distant metastasis rate is high, ranging from 20 to 35%, leading to an unfavorable clinical outcome with a 5-year mortality rate of 30–35% [2, 4, 9,10,11]. To improve clinical outcomes, systemic chemotherapy is essential. However, the efficacy of chemotherapy has not been clarified because randomized clinical trials assessing the efficacy of chemotherapy on MFS have not been conducted [3]. A small number of cohort studies revealed that the use of conventional chemotherapy was not clinically beneficial against distant metastases or for improving overall survival [2, 12, 13]. Thus, a novel treatment strategy has been urgently required.
Patient-derived cell lines are important tools in basic research and preclinical studies. Although long-term culturing causes genetic alterations in the cell lines, short-term cultured cell lines preserve the characteristics of the original tumor [14,15,16]. Therefore, efforts have been made for genomic profiling or determining the drug sensitivity of tumors using a number of cell lines in recent years [17,18,19,20]. However, these large-scale studies included only 28 sarcoma cell lines, which accounts for only 2.2% of all 1300 cell lines examined [21]. This indicates that latest technologies have not benefited research on sarcomas. This problem occurs due to the inadequate number of sarcoma cell lines owing to its rarity [22]. As for MFS, according to Cellosaurus [23], only 10 MFS cell lines have been reported worldwide (Table 1) [24,25,26,27,28,29,30,31]. Furthermore, among these cell lines, merely two MFS cell lines are publicly available. To promote research on MFS, it is necessary to establish novel MFS cell lines.
Here, we reported a novel MFS cell line, NCC-MFS3-C1, established from a surgically resected specimen from a patient with MFS. We characterized the NCC-MFS3-C1 cells and conducted a drug screening test.
Materials and methods
Patient history
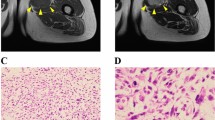
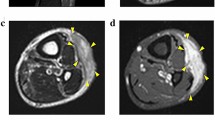
The patient donor was a 74-year-old woman with myxofibrosarcoma. The patient visited the previous hospital with the chief complaint of a mass on her posterior right thigh. Magnetic resonance imaging revealed a soft tissue tumor in the hamstrings, which was considered malignant (Fig. 1a, b). The patient was referred to the National Cancer Center Hospital (Tokyo, Japan). Subsequently, needle biopsy was performed, and the tumor was suspected to be a myxofibrosarcoma. Because there was no evident distant metastasis, wide resection was performed. Pathologically, the tumor showed pleomorphic atypical spindle cell proliferation with abundant myxoid stroma and curvilinear vessels (Fig. 1c, d). A part of the resected tumor at the operation was used to establish the cell line described in this study. The ethical committee of the National Cancer Center approved the use of clinical materials for this study, and written informed consent was obtained from the patient donor.
Clinical and pathological data. Magnetic resonance imaging revealing a multilocular tumor in hamstrings with a low intensity on a T1-weighted image and b mixed intensity (high and low) on a T2-weighted image. c and d Hematoxylin and eosin staining showing the proliferation of pleomorphic atypical spindle cells with abundant myxoid stroma and curvilinear vessels
Histological analysis
Histological examination was performed on 4-μm-thick sections from a representative paraffin-embedded tumor sample. The sections were deparaffinized and stained with hematoxylin and eosin (H&E).
Cell culture
The surgically resected tumor tissue was used to establish the cell line, as previously described [32]. The cells were maintained in Dulbecco’s Modified Eagle Medium/Nutrient Mixture F-12 (Gibco, Grand Island, NY, USA), supplemented with 10% heat-inactivated fetal bovine serum (Gibco), 100 μg/mL penicillin, and 100 μg/mL streptomycin (Nacalai Tesque, Kyoto, Japan) at 37 °C in a humidified atmosphere with 5% carbon dioxide. The cells were maintained for more than 3 months under tissue culture conditions and passaged more than 20 times.
Authentication and quality control of the established cell line
The established cell line was authenticated by examining short tandem repeats (STRs) at 10 loci using the GenePrint 10 system (Promega, Madison, WI, USA) according to the manufacturer’s instructions and the procedure described in our previous study [32]. The STR pattern was analyzed using the GeneMapper software (Thermo Fisher Scientific) and matched to the data in the public cell banks using a function of Cellosaurus with a standard match threshold of 80% [33].
Mycoplasma contamination was examined using DNA in the tissue culture medium of the cell line as previously reported [32].
Genetic analysis
Single nucleotide polymorphism (SNP) array genotyping was conducted with the Infinium OmniExpressExome-8 version 1.4 BeadChip (Illumina, San Diego, CA, USA) following the manufacturer’s instructions and the procedure described in our previous study [32]. The SNP array data were analyzed using the R version 4.0.3 (R Foundation for Statistical Computing, http://www.R-project.org) and DNAcopy package version 1.64.0 (Bioconductor, https://bioconductor.org/). Chromosome regions with copy numbers > 3 and < 1 were defined as amplifications and deletions, respectively. Genes that showed copy number alterations (CNAs) were annotated using the biomaRt package version 2.46.0 (Bioconductor) and “Cancer Gene Census” in the Catalog Of Somatic Mutations In Cancer database (GRCh 37 version 91).
Cell proliferation assay
Cell proliferation assays were performed as described previously [34]. Briefly, the cells were seeded at a density of 2.5 × 104 cells/well in 24-well culture plates at day 0. The number of cells was counted at multiple time points. The doubling time was calculated based on the growth curve. All the experiments were performed in triplicates.
Spheroid formation assay
Spheroid formation was examined as described previously [32]. The obtained spherical colonies were prepared for paraffin sections using iPGell (Genostaff, Tokyo, Japan) according to the manufacturer’s instructions. Cell blocks were fixed with 10% formalin neutral buffer solution and embedded in paraffin. Four-micrometer-thick paraffin sections were prepared and stained with H&E.
Invasion assay using real-time cell analyzer
We examined the invasion potential using a real-time cell analyzer (xCELLigence, Agilent, Santa Clara, CA, USA) according to the manufacturer’s instructions and the procedure described in our previous study [32]. In this study, MG63 osteosarcoma cells (JCRB; Ibaraki Osaka, Japan) were used as controls due to their consistent growth and being a representative histological subtype of sarcoma [35]. This assay was monitored every 15 min for 72 h and plotted as a function of time after seeding.
Assessment of tumorigenicity in nude mice
The animal experiment in this study was performed in compliance with the guidelines of the Institute for Laboratory Animal Research, National Cancer Center Research Institute as previously described [34]. Briefly, we used female Balb/c nude mice purchased from CLEA Japan, Inc. (Tokyo, Japan). A 100-μL volume of cells in a 1:1 mixture of Matrigel (BD Biosciences) was subcutaneously injected into the mice (1 × 106 cells). Subsequently, the tumor size was measured weekly. The tumor volumes were calculated according to the following formula: volume = (length × width2)/2. After 2 months, the tumors were surgically resected, and the specimens were stained with H&E.
Screening for the antiproliferative effects of anticancer agents
Screening for the antiproliferative effects of 214 anticancer agents was performed according to the manufacturer’s instructions and the procedure described in our previous study [32]. A list of the anticancer agents is provided in Supplementary Table 1.
Dose–response experiments were also performed to validate the available hits in the pilot screening according to previously reported methods [32]. The IC50 (the sample concentration required to inhibit cell growth by 50% in comparison with the growth of the control cells) was determined from the dose–response curves.
Results
Authentication of the established cell line
We authenticated the NCC-MFS3-C1 cell line by analyzing the STR status of 10 microsatellites. We confirmed nearly identical STR allele patterns between the NCC-MFS3-C1 cells and the corresponding original tumor tissues (Table 2, Supplementary Fig. 1). The STR patterns of NCC-MFS3-C1 cells are unique as they did not match those of any other cell line available within public cell banks examined using the cell line database, Cellosaurus. Thus, we concluded that NCC-MFS3-C1 is a newly established cell line. The DNA sequence of mycoplasma was not detected in the tissue culture medium of NCC-MFS3-C1 cells (data not shown).
Characterization of the cell line
SNP array analysis revealed CNAs in NCC-MFS3-C1 cells, corresponding to the original tumor. Partial allelic deletions were identified in chromosomes 2q, 3p, 5p, 7q, 9p, 11q, 16q, 17q, 18q, 19q, and 22q. The deletion in chromosome 9p included the tumor suppressor genes CDKN2A and CDKN2B (Fig. 2, Supplementary Table 2).
NCC-MFS3-C1 cells comprised spindle cells in two-dimensional culture conditions (Fig. 3a–c). NCC-MFS3-C1 cells exhibited consistent proliferation. According to the growth curve time, it took approximately 36 h for the population to double (Fig. 3d). NCC-MFS3-C1 cells were also able to form spheroids when seeded on a low-attachment microplate. The H&E-stained spheroid section showed dense proliferation of pleomorphic atypical oval cells (Fig. 3e). We also found that NCC-MFS3-C1 cells exhibited more aggressive invasion than MG63 cells. The invasion ability of NCC-MFS3-C1 cells depended on the time after seeding (Fig. 3f).
Characterization of NCC-MFS3-C1 cells. a–c NCC-MFS3-C1 cells showing spindle cell morphology under two-dimensional culture conditions. d Growth curve of NCC-MFS3-C1 cells. The Y-axis indicates the relative cell proliferation of NCC-MFS3-C1 cells, and the X-axis represents the day after seeding. e The hematoxylin and eosin-stained spheroid section showing dense proliferation of pleomorphic atypical oval cells. f Real-time cell analyzer invasion assay showing the invasion ability of NCC-MFS3-C1 cells compared to that of MG63 osteosarcoma cells
Tumorigenesis in nude mice
NCC-MFS3-C1 cells transplanted into BALB/c nude mice formed small tumor masses under the described conditions (Supplementary Fig. 2a). One of the four masses showed areas of dense proliferation of atypical oval cells and myxoid components in H&E sections (Supplementary Fig. 2b and c). However, only sparse cell proliferation was observed in other three masses (data not shown). Overall, the tumor growth was not sufficient (Supplementary Fig. 2d).
Sensitivity to anticancer agents
The cell viability of NCC-MFS3-C1, after treatment with 214 anticancer agents at a fixed concentration of 10 µM, is shown in Supplementary Table 3.
Among the 214 anticancer agents, 24 agents that showed prominent antiproliferative effects on NCC-MFS3-C1 cells or are frequently used as standard chemotherapy for soft tissue sarcomas were further examined to calculate their IC50 values. The five agents with the lowest IC50 values are presented in Table 3, and the IC50 values of 24 agents are presented in Supplementary Table 4. We also showed the growth curves that served as the basis for the calculation of the IC50 values in Fig. 4 and Supplementary Fig. 3.
Discussion
MFS is one of the most aggressive sarcomas with a poor prognosis. Although complete resection is required for the treatment of MFS, it is often challenging due to its strong invasive nature. When complete resection is not achieved in other tumors, postoperative chemotherapy is often considered. However, the effectiveness of conventional chemotherapy has not been proven in MFS patients because large-scale randomized clinical trials assessing the efficacy of conventional chemotherapy are insufficient. Therefore, novel treatment methods for MFS are required. In recent years, multiple large-scale genomic or drug sensitivity analyses have been performed in common cancers using a number of cell lines [17,18,19,20]. However, only 10 MFS cell lines have been reported to date, suggesting that MFS cannot benefit from these analyses. Hence, we established a novel MFS cell line, NCC-MFS3-C1.
The NCC-MFS3-C1 cell line was established from MFS observed in the deep thigh. Although MFS often arises in dermal or subcutaneous regions, deep-seated development is observed in approximately 30% of MFS cases [4]. Therefore, a NCC-MFS3-C1 cell line was established from a typical case of MFS.
Only a limited number of genetic studies about the CNAs of MFS have been reported. Although the frequent amplification of the chromosome 5p region was reported, the CNA is not specific to MFS [6]. Regarding the NCC-MFS3-C1 cell line, partial deletions in chromosomes 2q, 3p, 5p, 7q, 9p, 11q, 16q, 17q, 18q, 19q, and 22q were observed, and 5p amplification was not observed. These CNAs are different from those of the MFS cell lines previously reported (NCC-MFS1-C1 [30] and NCC-MFS2-C1 [31]), reflecting the highly complex karyotypes of MFS.
NCC-MFS3-C1 cells exhibited the typical morphology of spindle cell sarcoma. NCC-MFS3-C1 cells also demonstrated constant growth and spheroid formation. Although these characteristics are suitable for in vitro studies, NCC-MFS3-C1 cells were not sufficiently responsible for the growth of a mass in nude mice under the described conditions, indicating that NCC-MFS3-C1 cells may not be eligible for xenograft experiments.
In drug screening using 214 anticancer agents, the five agents with the lowest IC50 values were trabectedin, romidepsin, bortezomib, eribulin, and actinomycin D. Notably, NCC-MFS1-C1 [30] and NCC-MFS2-C1 [31] cell lines also showed high sensitivity to romidepsin and bortezomib. Bortezomib is one of the proteasome inhibitors that is often used for the treatment of multiple myeloma and mantle cell lymphoma [36]. Only one study has assessed the efficacy of bortezomib for MFS [37]. This study used MFS cell lines with amplification of the chromosome 5p region and suggested that the amplification of SKP2 on chromosome 5p was associated with the antiproliferative effects of bortezomib on MFS. However, NCC-MFS1-C1, NCC-MFS2-C1, and NCC-MFS3-C1 did not demonstrate amplification of the chromosome 5p region. Despite the lack of amplification, the growth of these MFS cell lines was commonly inhibited by bortezomib, indicating that other molecular mechanisms underlying MFS affected the sensitivity to bortezomib. Romidepsin is a histone deacetylase (HDAC) inhibitor, which is usually used for the treatment of peripheral T-cell lymphoma [38]. Only one study has reported the efficacy of HDAC inhibitor for MFS [39]. However, the molecular mechanisms of HDAC inhibitors for MFS remain unclear, warranting further studies.
In conclusion, we established a novel MFS cell line, NCC-MFS3-C1, which demonstrated constant proliferation, spheroid formation, and aggressive invasion. A genomic study revealed multiple allelic deletions, with no amplification of the chromosome 5p region as previously reported [6], reflecting the diversity of MFS. Our study also revealed the antiproliferative effect of bortezomib and romidepsin on the MFS cell line. However, considering the diversity and rarity of MFS, the number of MFS cell lines is still insufficient. Therefore, consistent efforts to further establish MFS cell lines are required for research.
References
WHO. WHO classification of tumours of soft tissue and bone, 5th edn. Lyon: IARC; 2020.
Sanfilippo R, Miceli R, Grosso F, et al. Myxofibrosarcoma: prognostic factors and survival in a series of patients treated at a single institution. Ann Surg Oncol. 2011;18:720–5.
Roland CL, Wang WL, Lazar AJ, Torres KE. Myxofibrosarcoma. Surg Oncol Clin N Am. 2016;25:775–88.
Mentzel T, Calonje E, Wadden C, et al. Myxofibrosarcoma. Clinicopathologic analysis of 75 cases with emphasis on the low-grade variant. Am J Surg Pathol. 1996;20:391–405.
Widemann BC, Italiano A. Biology and management of undifferentiated pleomorphic sarcoma, myxofibrosarcoma, and malignant peripheral nerve sheath tumors: state of the art and perspectives. J Clin Oncol. 2018;36:160–7.
Barretina J, Taylor BS, Banerji S, et al. Subtype-specific genomic alterations define new targets for soft-tissue sarcoma therapy. Nat Genet. 2010;42:715–21.
Iwata S, Yonemoto T, Araki A, et al. Impact of infiltrative growth on the outcome of patients with undifferentiated pleomorphic sarcoma and myxofibrosarcoma. J Surg Oncol. 2014;110:707–11.
Waters B, Panicek DM, Lefkowitz RA, et al. Low-grade myxofibrosarcoma: CT and MRI patterns in recurrent disease. Am J Roentgenol. 2007;188:W193–8.
Dewan V, Darbyshire A, Sumathi V, Jeys L, Grimer R. Prognostic and survival factors in myxofibrosarcomas. Sarcoma. 2012;2012:830879.
Lee AY, Agaram NP, Qin LX, et al. Optimal percent myxoid component to predict outcome in high-grade myxofibrosarcoma and undifferentiated pleomorphic sarcoma. Ann Surg Oncol. 2016;23:818–25.
Huang HY, Lal P, Qin J, Brennan MF, Antonescu CR. Low-grade myxofibrosarcoma: a clinicopathologic analysis of 49 cases treated at a single institution with simultaneous assessment of the efficacy of 3-tier and 4-tier grading systems. Hum Pathol. 2004;35:612–21.
Mutter RW, Singer S, Zhang Z, Brennan MF, Alektiar KM. The enigma of myxofibrosarcoma of the extremity. Cancer. 2012;118:518–27.
Look Hong NJ, Hornicek FJ, Raskin KA, et al. Prognostic factors and outcomes of patients with myxofibrosarcoma. Ann Surg Oncol. 2013;20:80–6.
Saito S, Morita K, Kohara A, et al. Use of BAC array CGH for evaluation of chromosomal stability of clinically used human mesenchymal stem cells and of cancer cell lines. Hum Cell. 2011;24:2–8.
Tseng YY, Boehm JS. From cell lines to living biosensors: new opportunities to prioritize cancer dependencies using ex vivo tumor cultures. Curr Opin Genet Dev. 2019;54:33–40.
Ben-David U, Beroukhim R, Golub TR. Genomic evolution of cancer models: perils and opportunities. Nat Rev Cancer. 2019;19:97–109.
Basu A, Bodycombe NE, Cheah JH, et al. An interactive resource to identify cancer genetic and lineage dependencies targeted by small molecules. Cell. 2013;154:1151–61.
Barretina J, Caponigro G, Stransky N, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012;483:603–7.
Yang W, Soares J, Greninger P, et al. Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013;41:D955–61.
Shoemaker RH. The NCI60 human tumour cell line anticancer drug screen. Nat Rev Cancer. 2006;6:813–23.
Goodspeed A, Heiser LM, Gray JW, Costello JC. Tumor-derived cell lines as molecular models of cancer pharmacogenomics. Mol Cancer Res. 2016;14:3–13.
Hattori E, Oyama R, Kondo T. Systematic review of the current status of human sarcoma cell lines. Cells. 2019;8:157.
Bairoch A. The cellosaurus, a cell-line knowledge resource. J Biomol Tech. 2018;29:25–38.
Kawashima H, Ogose A, Gu W, et al. Establishment and characterization of a novel myxofibrosarcoma cell line. Cancer Genet Cytogenet. 2005;161:28–35.
Ariizumi T, Ogose A, Kawashima H, Hotta T, Umezu H, Endo N. Multinucleation followed by an acytokinetic cell division in myxofibrosarcoma with giant cell proliferation. J Exp Clin Cancer Res. 2009;28:44.
Moneo V, Serelde BG, Fominaya J, et al. Extreme sensitivity to Yondelis (Trabectedin, ET-743) in low passaged sarcoma cell lines correlates with mutated p53. J Cell Biochem. 2007;100:339–48.
Miserocchi G, De Vita A, Mercatali L, et al. Characterization and drug sensitivity of a new high-grade myxofibrosarcoma cell line. Cells. 2018;7:186.
Lohberger B, Stuendl N, Wolf E, Liegl-Atzwanger B, Leithner A, Rinner B. The novel myxofibrosarcoma cell line MUG-Myx1 expresses a tumourigenic stem-like cell population with high aldehyde dehydrogenase 1 activity. BMC Cancer. 2013;13:563.
Lohberger B, Stuendl N, Leithner A, et al. Establishment of a novel cellular model for myxofibrosarcoma heterogeneity. Sci Rep. 2017;7:44700.
Kito F, Oyama R, Sakumoto M, et al. Establishment and characterization of a novel cell line, NCC-MFS1-C1, derived from a patient with myxofibrosarcoma. Hum Cell. 2019;32:214–22.
Noguchi R, Yoshimatsu Y, Ono T, et al. Establishment and characterization of NCC-MFS2-C1: a novel patient-derived cancer cell line of myxofibrosarcoma. Hum Cell. 2021;34:246–53.
Sin Y, Yoshimatsu Y, Noguchi R, et al. Establishment and characterization of a novel alveolar rhabdomyosarcoma cell line, NCC-aRMS1-C1. Hum Cell. 2020;33:1311–20.
Capes-Davis A, Reid YA, Kline MC, et al. Match criteria for human cell line authentication: where do we draw the line? Int J Cancer. 2013;132:2510–9.
Tsuchiya R, Yoshimatsu Y, Noguchi R, et al. Establishment and characterization of NCC-DDLPS3-C1: a novel patient-derived cell line of dedifferentiated liposarcoma. Hum Cell. 2021;34:1008–18.
Billiau A, Edy VG, Heremans H, et al. Human interferon: mass production in a newly established cell line, MG-63. Antimicrob Agents Chemother. 1977;12:11–5.
Gandolfi S, Laubach JP, Hideshima T, Chauhan D, Anderson KC, Richardson PG. The proteasome and proteasome inhibitors in multiple myeloma. Cancer Metastasis Rev. 2017;36:561–84.
Li CF, Wang JM, Kang HY, et al. Characterization of gene amplification-driven SKP2 overexpression in myxofibrosarcoma: potential implications in tumor progression and therapeutics. Clin Cancer Res. 2012;18:1598–610.
Iyer SP, Foss FF. Romidepsin for the treatment of peripheral T-cell lymphoma. Oncologist. 2015;20:1084–91.
Kawarazaki A, Horinaka M, Yasuda S, Kawashima H, Numajiri T, Sakai T. The HDAC inhibitor OBP-801 suppresses the growth of myxofibrosarcoma cells. J BUON. 2020;25:464–71.
Acknowledgements
We thank Drs. F. Nakatani, E. Kobayashi, S. Fukushima, M. Nakagawa, T. Komatsubara, M Saito, C. Sato (Department of Musculoskeletal Oncology), Drs. T Shibayama, and H. Tanaka (Department of Diagnostic Pathology), National Cancer Center Hospital, for sampling tumor tissue specimens from surgically resected materials. We also appreciate the technical assistance provided by Ms. Y. Kuwata (Division of Rare Cancer Research). We appreciate the technical support provided by Ms. Y. Shiotani, Mr. N. Uchiya, and Dr. T. Imai (Central Animal Division, National Cancer Center Research Institute). We would like to thank Editage (www.editage.jp) for help with English language editing and constructive comments on the manuscript. This research was technically assisted by the Fundamental Innovative Oncology Core at the National Cancer Center.
Funding
This research was supported by the Japan Agency for Medical Research and Development (Grant number 20ck0106537h0001).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
The ethical committee of the National Cancer Center approved the use of clinical materials for this study (approval number 2004–050).
Informed consent
Written informed consent for publication was provided by the patient.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2021_548_MOESM2_ESM.tif
Supplementary file2 Tumorigenesis in nude mice. (A) NCC-MFS3-C1 cells transplanted into BALB/c nude mice formed a small tumor mass under the described condition. One of the four masses showed areas of (B) dense proliferation of atypical oval cells and (C) myxoid components in Hematoxylin and eosin staining sections. (D) The graph showing estimated tumor volume. Bars represent the mean standard error (TIF 1984 kb)
Rights and permissions
About this article
Cite this article
Tsuchiya, R., Yoshimatsu, Y., Noguchi, R. et al. Establishment and characterization of NCC-MFS3-C1: a novel patient-derived cell line of myxofibrosarcoma. Human Cell 34, 1266–1273 (2021). https://doi.org/10.1007/s13577-021-00548-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-021-00548-6