Abstract
Sugarcane is one of the most important commercial crops in India, supporting the second largest agro-based sugar industry. However for more than a century, the production is imperiled by the most devastating fungus Colletotrichum falcatum Went, which causes red rot in sugarcane. Complex polyploidy and limited understanding on the inheritance to red rot resistance makes breeding efforts more complicated in sugarcane. Transcription factors (TFs) play a key role in regulating defense response, which offers much promise for the manipulation of metabolic pathways, potential in regulating multiple signaling networks leading to plant defense against pathogens. In order to better manage red rot disease in sugarcane, we sought to screen the TFs possibly involving in pathogen defense by analyzing 5 major defense-related TF classes (WRKY, bZIP, MYB, NAC and TLP). In this study two parallel sets of experiments were carried out to compare the differential regulation of these classes of TFs upon pathogen challenge and SAR priming. Among the 41 TFs studied, differential regulation of 24 TFs upon pathogen challenge and 15 TFs upon SAR inducer priming were observed. Comparison of incompatible interaction and SAR inducer priming revealed that 8 TFs were highly induced in both the cases. Collectively, the results showed that TFs which are significantly induced early may involve actively in triggering or co-ordinating defense against pathogen invasion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugarcane is an important commercial crop cultivated for its stalks that account for nearly 60 % of the crystal sugar produced worldwide. The crop is cultivated in more than 90 countries all over the world, the largest area being in Brazil and India. Sugarcane is a renewable, agricultural bio-resource because it provides sugar, besides bio-fuel, fiber, and by-products with ecological sustainability. Among the various production constraints of sugarcane, red rot caused by the fungus Colletotrichum falcatum Went (Perfect stage: Physalospora tucumanensis Speg.) is one of the serious problems which reduces the yield by 30–40 % (Viswanathan 2010). Sugarcane production is severely affected by this disease in various parts of the world, including India, Java, Thailand, Fiji, and parts of USA (Singh and Singh 1989).
Most of the attempted fungicides have failed to control the disease under field conditions and hence releasing disease-resistant varieties has been the prime management strategy to contain the disease. Complex polyploidy and lack of information on inheritance of red rot resistance have limited the efforts on the development of disease resistant varieties using conventional breeding approaches. Further, the frequent breakdown of red rot resistance in varieties hitherto graded as resistant has been attributed to the change in pathotypes. Added to this, continuous cultivation of highly susceptible cultivars, otherwise agronomically superior has resulted in epiphytotic nature of the disease (Viswanathan et al. 2003). Obviously, there is a demand for novel strategy of management to supplement breeding for resistance and fungicidal control of sugarcane diseases. To a certain degree, susceptibility of the crop against pathogens could be altered with enhanced level of resistance without genetic manipulation by inducing Systemic acquired resistance (SAR). Salicylic acid (SA) dependent SAR pathway seems to be the most robust to be exploited for practical crop protection (Durrant and Dong 2004). In sugarcane, SAR could be induced by inducers of both biotic (Cf elicitor) and synthetic origin (BTH), which resulted with the expression of specific pathogenesis-related (PR) proteins and initial defense response (Ramesh sundar et al. 2006, 2008, 2009). Overlaps of gene expression in incompatible interaction and SAR induction were also reported by Schenk et al. 2003.
Transcription factors (TFs) are the proteins containing DNA binding domains that binds with DNA sequences and facilitates transcriptional activation or repression of a gene with the coordination of other cis/trans acting elements (Latchman 1997). Regulation of TFs is a complex process which ensures exact spatial and temporal expression of genes during various developmental process and cellular responses especially to stress (Rushton and Somssich 1998). Plant TFs are classified into various families based on the structure of DNA binding domain and sequence similarity. Research efforts in the recent past have been promising in identifying TFs, which are important for regulating plant responses to stress factors (Singh et al. 2002). Specific classes of TFs viz., WRKY, bZIP, MYB, NAC, TLP, etc., are reported to have a decisive role in plant defense (Ooka et al. 2003; Chen et al. 2005). The present scenario offers ample scope for modulating TFs which are potential targets in regulating multiple signaling networks for plant defense (Naoumkina et al. 2008).
With the availability of the sugarcane EST database (Vettore et al. 2001), it has become possible to specifically look into the differential expression of targeted candidate genes, which have been established to have a role in defense against pathogens (Viswanathan et al. 2009). In order to better understand the defense response triggered by pathogen challenge (compatible and incompatible interactions) and SAR inducers (BTH and Cf elicitor), this study primarily focuses on profiling differential expression of sugarcane TFs at various time intervals.
Materials and methods
Plant material
Two sugarcane varieties, namely CoC 671 (susceptible to red rot) and Co 93009 (resistant to red rot), were planted during 2009 – 2011 seasons in 20 ft rows in the institute farm, with three biologically independent replications. Nutritional supplements and proper irrigation were provided to ensure agronomically ideal growing conditions.
Inducer treatments
Foliar spray of inducers (BTH - 250 μM and Cf elicitor - 60 μg glucose equivalents) was done at 3rd and 5th month from the date of planting. The efficacy of SAR priming (BTH and Cf elicitor) in controlling the disease severity was evaluated (Srinivasan and Bhat 1961) and depicted in Fig. 1.
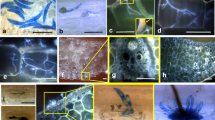
Efficacy evaluation of SAR priming (sugarcane cv. CoC 671) after 45 days of pathogen inoculation by nodal swabbing method. (a) Un-inoculated control (0 %) (b) BTH primed showing restricted lesion (6.3 % infection) (c) Cf elicitor primed (17.5 % infection) (d) Unprimed inoculated control showing extended progressive lesions (100 % infection)
Challenge inoculation
Pathogen was inoculated by nodal swabbing method at 30 days after foliar spray during September, 2010. In the nodal swabbing method, the leaf sheaths of the 6th and 7th leaf from the top were removed and a layer of absorbent cotton saturated with 2 ml of conidial suspension of C. falcatum (106 conidia / ml) was placed around the nodes. The inoculated nodes were then wrapped with polythene sheets in order to prevent drying of the inoculum (Ramesh Sundar et al. 2009). Three biologically independent replications were maintained.
Sample collection
Stalk tissue samples (CoC 671) primed with SAR inducers were collected at 0, 6, 12, 24, 48 and 72 hpi (hours post inoculation) with appropriate controls. Pathogen challenged stalk tissues of CoC 671 and Co 93009 were collected at 0, 6, 12, 24 and 48 hpi to study the pathogen responsive TFs.
RNA extraction
Tissue samples of 5 g were scooped around the pathogen inoculated region and ground to fine powder in liquid nitrogen. Total RNA was extracted using TRI reagent (Sigma, USA) following manufacturer’s instructions and resuspended in 80–100 μl of warm RNA resuspension solution (Ambion®, Canada). RNA samples were quantified using NanodropTM (USA). An equimolar pool of good quality RNA samples were used as template for RT-PCR analysis.
Primer designing
The sugarcane transcription factor databases - planttfdb.cbi.pku.edu.cn (v1.0) (Guo et al. 2008; Zhang et al. 2011) and grassius.org (Gray et al. 2009) were used for sequence retrieval. Five major defense related TF classes (WRKY, bZIP, MYB, NAC and TLP) as represented in PlantTFDB v1.0 were targeted and sequences of each targeted TF classes were aligned using ClustalW and trees were constructed based on the similarity. Representative sequences from each class of TF were selected randomly from each small cluster of the aligned tree and totally 60 sets of sequence specific forward and reverse primers targeting the five TF classes were designed using the FPCR ver.3.6.62 software.
RT-PCR
The RNA extracted at different time intervals (0, 6, 12, 24, 48 and 72 hpi) along with respective un-inoculated controls were reverse transcribed using RevertAid H-cDNA synthesis kit (MBI Fermentas, USA) and the cDNA was used for screening 60 sets of sequence specific primers. Initially the RNA concentration was normalized using the GAPDH (GenBank: HQ242713.1) primers. PCR amplification was performed using 2 μl of 10 mM dNTPs (2.5 mM each), 1 U Taq polymerase, 1.0 μM each of forward and reverse primers for 30 cycles. The PCR cycle comprised an initial denaturation for 5 min at 94 °C, annealing temperature depending on the primer for 30 s, extension at 72 °C for 30 s and a final extension for 10 min. The annealing temperature was initially standardized using genomic DNA extracted from CoC 671 by gradient PCR in a thermal cycler (Mastercycler gradient, Eppendorf, Germany). Electrophoresed amplicons were analyzed using Syngene: G Box - GENEsys 1.0.9.0. Each RT-PCR reaction was repeated with at least three independent biological replications for reproducibility.
Amplicon identification
Amplicon re-sequencing was performed to verify the identity of the amplified product, by comparison with the corresponding EST sequences from which the TFs were originally selected (Table 1).
Results and discussion
Genome-wide transcriptome analyses have identified hundreds of genes encoding TFs that are induced or repressed by a range of environmental stress factors (Chen and Zhu 2004); the next step is the identification of specific TFs involved in defense responses. Specific accumulation of defense gene transcripts is a key early component in the sequence of events leading to the activation of defense in infected tissues or during plant-pathogen interactions (Bell et al. 1986). The present study aims at screening of defense-related TFs by expression profiling of five major TF classes. Though, this study selected only representative TFs from each small cluster of an aligned tree, it covered all the internal nodes of closely related TFs of a class and so all types of TFs have been included. However, the possibility of missing out of potential defense-related TFs cannot be ruled out. Here, two sets of experiments were conducted to study the differentially regulated transcripts in both pathogen challenged (Co 93009 and CoC 671) and SAR-primed (CoC 671) sugarcane stalk tissues.
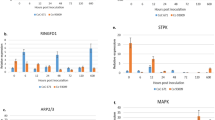
The expression levels of transcripts of all test samples were normalized using GAPDH as a reference gene. Among the 60 TFs subjected to RT-PCR, only 41 sets resulted in detectable amplification as depicted in representative profiles (Fig. 2a and b). The amplicons of expected size were confirmed by amplicon resequencing and respective similarities from NCBI as well as Transcript assemblies from TIGR and PlantGDB databases were represented in Table 1. Non-amplification of the remaining 19 sequences might be due to absence or insufficient expression at detectable levels in 30 PCR cycles. Total no. of PCR cycle was optimized by the Ct value (Cycle threshold) (33.6 ± 3.7) of representative TFs from each TF classes in response to different treatments and time intervals by relative quantification method using StepOne Plus Real-Time PCR system (Applied Biosystems, Foster City, CA, USA) (Results not shown).
Representative RT-PCR profiles of differentially regulated TFs. Transcript Expression levels were standardized to transcripts of GAPDH as a reference gene (a) Transcripts regulated differentially in compatible and incompatible interactions. S - Susceptible variety (CoC 671), R - Resistant variety (Co 93009) (b) Differentially regulated SAR - responsive transcripts. B - BTH, E - Cf elicitor, P - Pathogen challenge (Isolate Cf 671), c - Mock treatment. Arrows indicates up-regulation. 0, 6, 12, 24, 48 & 72 indicate hours post inoculation
For the sake of quick reference, these 41 TFs were named trivially, which are listed in Table 1, along with the respective PlantTFDB accession codes and respective matches in other databases. Of the 41 TFs, 24 TFs were found to be differentially regulated upon pathogen challenge (Table 2) and 15 TFs in response to SAR inducer priming (Table 3). Upon comparison of pathogen challenge (involving compatible and incompatible interactions) and SAR inducer priming, 13 TFs were found to be differentially regulated in both the cases (Fig. 3a) and on comparison of incompatible interaction (Co 93009) with SAR priming, 8 TFs were found to be differentially regulated in both the cases (Fig. 3b). In Tables 2 and 3, the up-regulation and down-regulation is represented as + and - respectively, which is an arbitrary indication of the intensity of expression arrived after pooling the data from at least three replications. The expression pattern of over 90 % of TFs remains unaltered in all the three replicates and also corroborates more or less with the qPCR data of few TFs, thus indicating the reproducibility of semi-quantitative PCR.
Venn diagram depicting total no. of TFs differentially regulated with respect to pathogen challenge and SAR priming (a) Differentially regulated TFs with pathogen challenge (compatible and incompatible interaction) and SAR primed tissues (b) Comparison between the number of TFs expressed in incompatible interaction and SAR response
An overall observation of incompatible and compatible interactions, indicated induction of WRKY 40 and 44 between 6 hpi and 48 hpi in incompatible interaction (Co 93009), while in case of compatible interaction (CoC 671), the expression of WRKY 44 was up-regulated later from 12 hpi and the expression pattern of WRKY 40 was unstable (Table 2). SAR inducer treatments viz., BTH and C. falcatum elicitor induced WRKY 40 and 44 at earlier time interval (6 hpi). However, the expression of WRKY 44 and 40 was down-regulated between 12 and 48 hpi in primed samples, with an exceptional induction of WRKY 40 at 12hpi in BTH treatment (Table 3). Collectively, the higher level of earlier expression of WRKY 40 in SAR primed samples suggests its possible involvement in the activation of defense. TF-regulated gene expression involves multiple signals forming a network of gene activation cascade, which is exemplified by the earlier expression of WRKY 44 in primed samples at a very high level followed by down-regulation at later stages. This might be attributed to a distinct regulation of specific TFs by SAR inducers in a temporal pattern. Several WRKY genes have been reported from different crops and they all seem to behave differentially under varying stress conditions (Desveaux et al. 2004). Transient earlier up-regulation of OsWRKY1 observed in rice upon treatment with fungal elicitor and further inoculation with blast pathogen Magnaporthe grisea, indicates its regulatory role during early signaling events (Kim et al. 2000). Wang et al. (2006) demonstrated that over expression of WRKY genes at earlier stages of pathogen infection on SAR primed Arabidopsis. A recent work suggested SA-mediated smut disease resistance in sugarcane, wherein involvement of WRKY gene (class II) was speculated (Liu et al. 2012).
The next group of genes screened in this study is bZIP TFs. Most of the bZIP class of TFs viz., bZIP 4, 18, 23, 27 were found to be induced in the incompatible interaction between 6 and 48 hpi, which is not in case of compatible interaction (Table 2). BTH priming resulted in differential expression pattern of bZIP 4, however the profile of most of the other members of this family of TFs remained unaltered upon priming (Table 3). On the whole, an inconsistency in expression pattern was recorded at different time intervals for individual bZIPs. A total of 121 bZIPs were submitted in Sugarcane EST (SUCEST) database, which were classified into 13 groups based on the conserved regions (Vincentz et al. 2001). In tobacco, few of the bZIP TFs have the ability to induce basal defense during SAR induction (Thurow et al. 2005). bZIP class of TFs binds to the OCS-elements in plant promoters-responsive to SA-regulated signaling, which triggers a speculation for their up-regulation upon SAR priming and incompatible interaction.
The MYB factors are reported to be involved in plant secondary metabolism / cell morphogenesis and responsive to hormone and stress signaling (Chen et al. 2005). Among the three genes representing MYB class of TFs (MYB 78, 82, 83), expression of MYB 78 was up-regulated in all the intervals of incompatible interaction, whereas in compatible it was up-regulated only at an early interval. The expression of MYB 83 was un-stable in compatible interaction, while it was induced only at 24hpi in incompatible interaction (Table 2). In response to SAR priming, MYB 78 didn’t show significant difference in expression, but MYB 83 was up-regulated at earlier intervals of BTH treatment (Table 3). MYB 83 which possesses a putative R2R3 MYB domain, might play a key regulatory role in flavanoid synthesis, thus controlling the production of anthocyanin during pathogen invasion. In sugarcane, anthocyanin accumulation was reported to be higher in resistant cultivars than susceptible one, which implicates its mediation in disease resistance (Viswanathan 2000). Transient expression of AtMYB 30 was recorded in incompatible interaction of Arabidopsis to induce hypersensitive response (HR). Results of the present study corroborate with that of Arabidopsis (Vailleau et al. 2002) and elicitor-responsive expression in rice (Yanhui et al. 2006).
In NAC class of TFs, 11 members were screened (Table 1). In incompatible interaction, NAC J, L and O were up-regulated at earlier intervals between 6 and 24 hpi, but in compatible interaction, the induction was observed only at 48 hpi (Table 2). In BTH primed tissues NAC C, D & E were induced at earlier intervals between 6 and 12 hpi, but not in case of Cf elicitor treated samples (Table 3). The NAC family of TFs has been found to be associated with host response to pathogen infection (Ooka et al. 2003). One of the genes SsNAC23, a member of the NAC family was found to be involved in cold stress response in sugarcane and also in response to herbivore and water deficit conditions (Nogueira et al. 2003). In wheat against stripe rust fungal infection and hormone treatment (jasmonate, abscissic acid and ethylene), TaNAC4 was found to be up-regulated (Xia et al. 2010). Results of our study are drawing parallels to the established role of NAC genes in biotic and abiotic stress tolerance.
Tubby-Like Proteins (TLPs) are the next family of TFs screened, in which the expression patterns of 10 TLP TFs were studied (Table 1). Comparison of CoC 671 and Co 93009 in response to pathogen challenge, showed slight variation in the level of expression pattern of TLB B, whereas the expression of other three TFs (TLP K, TLP L and TLP N) was inconsistent in both incompatible and compatible interactions (Fig. 2a). In BTH and Cf elicitor primed tissues, TLP B was up-regulated at 6 hpi followed by variable temporal expression. In case of TLP K, there wasn’t significant alteration except a feeble spike at 12hpi in BTH treatment (Table 3). In Arabidopsis, most of the TLP genes are involved in the regulation of molecular events during environmental stress and colonization of microbes (Reitz et al. 2012).
The up-regulation of TFs upon pathogen challenge in sugarcane varieties varying in red rot resistance showed a prominent difference in the induction patterns. Maximum number of TFs (20) was over expressed at 24 hpi in incompatible interaction, but in compatible interaction, only 13 TFs were up-regulated at a later interval of 48 hpi. Supporting evidence is available with the transcriptome analysis of legumes by Torregrosa et al. (2006) which showed differential expression of 3,900 genes in both susceptible and resistant varieties, with higher level of induction in genes involved in resistant interactions. A macroarray study in Cacao by Lopes et al. (2010) has indicated that the number of TFs induced in response to Moniliophthora perniciosa was higher (33 TFs) in incompatible interaction (Cacao-M. perniciosa), while only 18 TFs were induced upon compatible interaction, which strengthens the hypothesis that TFs play a pivotal role in regulating defense response against pathogen invasion.
Different types of TFs can display distinct temporal expression patterns in response to a specific stress, which could provide a basal platform and guidance to the use of other experimental approaches to test the hypothesis about a particular gene function (Chen and Zhu 2004). In SAR priming, the overall transcript abundance was observed to be moderately up-regulated at 6 hpi especially in BTH, but there was a drastic down-regulation of TF genes at 12 hpi in Cf elicitor primed tissues. However, in both the cases similar level of up-regulation was observed at 72 hpi, which was comparable with the 6 hpi pattern. This intermittent down-regulation may be only transient, which might get up-regulated even after 72 hpi, leading to induced resistance under field conditions (Ramesh Sundar et al. 2009). Rapid and early induction of TFs in response to BTH priming indicates the functional SAR response in sugarcane. bZIP 4, bZIP 15 and NAC H were found to be up-regulated only in Cf elicitor primed tissues at 6 hpi, but not in BTH treatment suggesting that these two SAR inducers may possess differential regulation of signaling cascades in activating induced defense. The similarity in the Cf elicitor priming and pathogen challenge responses could be attributed to the reason that the elicitor is capable of mimicking the pathogen response.
Collectively, 8 TFs were differentially regulated and activated early in both incompatible interaction and SAR priming, thus suggesting that these TFs might mediate defense against pathogen invasion. Even though, the expression pattern of few TFs represented in Tables 2 and 3 were found to be similar, there was a significant difference in the level of induction. However, the up-regulated TFs in both incompatible interaction and SAR response do not imply its exclusive role on pathogen defense, but also forms part of the housekeeping activity, thus might be involved in the overall co-ordination of defense signaling network (Schenk et al. 2003). Instability in the pace of transcriptional activation in response to pathogen challenge may indicate the dynamic interaction between the host and the pathogen.
The potential of TFs as a tool for the dissection of defense signaling and with the advent of high throughput genomic platforms, manipulation of multigenic traits has gained momentum. Development of new TF-based approaches thus would benefit gene discovery leading to crop improvement. This study is the first attempt in sugarcane to understand the TF-regulated defense response against C. falcatum using semi quantitative RT-PCR. In sequel of this preliminary work, further studies are underway using real time PCR validation of these TFs, which would result in delineating their regulatory roles in red rot defense. The expected outcome would open up new vistas in engineering durable red rot resistance in sugarcane.
Abbreviations
- TFs:
-
Transcription factors
- SAR:
-
Systemic acquired resistance
- BTH:
-
Benzothiadiazole
- SA:
-
Salicylic acid
- Cf :
-
Colletotrichum falcatum
- hpi:
-
hours post inoculation
References
Bell JN, Ryder TB, Vincent PM, Lamb CJ et al (1986) Differential accumulation of plant defense gene transcripts in a compatible and incompatible plant-pathogen interaction. Mol Cell Biol 1615–1623
Chen R, Zhongfu N, Nie X, Qin Y, Dong G, Sun Q (2005) Isolation and characterization of genes encoding MYB transcription factor in wheat (Triticum aestivem L.). Plant Sci 169(6):1146–1154
Chen WJ, Zhu T (2004) Networks of transcription factors with roles in environmental stress response. Trends Plant Sci 9(12):591–596
Desveaux D, Subramaniam R, Despres C, Mess JN, Lévesque C, Fobert PR, Dangl JL, Brisson N (2004) A “whirly” plant transcription factor is required for salicylic acid-dependent disease resistance in Arabidopsis. Dev Cell 6:229–240
Durrant WE, Dong X (2004) Systemic acquired resistance. Annu Rev Phytopathol 42:185–209
Gray J, Bevan M, Brutnell T, Buell R, Cone K, Hake S, Jackson D, Kellogg E, Lawrence C, McCouch S, Mockler T, Moose S, Paterson A, Peterson T, Rokshar D, Souza GM, Springer N, Stein N, Timmermans M, Wang GL, Grotewold E (2009) Naming transcription factors. Plant Physiol 149:4–6
Guo AY, Chen X, Gao G, Zhang H, Zhu QH, Liu XC, Zhong YF, Gu XC, He K, Luo JC (2008) PlantTFDB: a comprehensive plant transcription factor database. Nucleic Acids Res 36:D966–D969
Kim CY, Lee SH, Park HC, Bae CG, Cheong YH, Choi YJ, Han CD, Lee SY, Lim CO, Cho MJ (2000) Identification of rice blast fungal elicitor-responsive genes by differential display analysis. Mol Plant Microbe Interact 13:470–474
Latchman DS (1997) Transcription factors: an overview. Int J Biochem Cell Biol 29(12):1305–1312
Liu J, Que Y, Guo J, Xu L, Wu J, Chen R (2012) Molecular cloning and expression analysis of a WRKY transcription factor in sugarcane. Afr J Biotechnol 11(24):6434–6444
Lopes MA, Hora Junior BT, Dias CV, Santos GC, Gramacho KP, Cascardo JCM, Gesteira AS, Micheli F (2010) Expression analysis of transcription factors from the interaction between cacao and Moniliophthora perniciosa (Tricholomataceae). Genet Mol Res 9(3):1279–1297
Naoumkina M, He X, Dixon R (2008) Elicitor-induced transcription factors for metabolic reprogramming of secondary metabolism in Medicago truncatula. BMC Plant Biol 8:132
Nogueira FTS, De Rosa VE, Menossi M, Ulian EC, Arruda P (2003) RNA expression profiles and data mining of sugarcane response to low temperature. Plant Physiol 132(4):1811–1824
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10(6):239–247
Ramesh Sundar A, Viswanathan R, Malathi P, Padmanaban P (2006) Mechanism of resistance induced by plant activators against Colletotrichum falcatum in sugarcane. Arch Phytopathol PFL 39(4):259–272
Ramesh Sundar A, Velazhahan R, Nagarathinam S, Vidhyasekaran P (2008) Induction of pathogenesis-related proteins in sugarcane leaves and cell-cultures by a glycoprotein elicitor isolated from Colletotrichum falcatum. Biol Plant 52(2):321–328
Ramesh Sundar A, Viswanathan R, Nagarathinam S (2009) Induction of systemic acquired resistance (SAR) using synthetic signal molecules against Colletotrichum falcatum in sugarcane. Sugar tech 11(3):274–281
Reitz MU, Bissue JK, Zocher K, Attard A, Huckelhoven R, Becker K, Imani J et al (2012) The subcellular localization of tubby-like proteins and participation in stress signaling and root colonization by the mutualist Piriformospora indica. Plant Physiol 160(1):349–364
Rushton PJ, Somssich IE (1998) Transcriptional control of plant genes responsive to pathogens. Curr Opin Plant Biol 1:311–315
Schenk PM, Kazan K, Manners JM, Anderson JP, Simpson RS, Wilson IW, Somerville SC et al (2003) Systemic gene expression in Arabidopsis during an incompatible interaction with Alternaria brassicicola. Plant Physiol 132:999–1010
Singh K, Foley RC, Onate-Sanchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol (5):430–436
Singh K, Singh RP (1989) Red rot. In: Ricaud CG, Egan BT, Gillaspie AG, Hughes CG (eds) Sugarcane diseases of the world: major diseases. Elsevier, Amsterdam, pp 169–188
Srinivasan KV, Bhat NR (1961) Red rot of sugarcane criteria of grading resistance. J Indian Bot Soc 40:566–577
Thurow C, Schiermeyer A, Krawczyk S, Butterbrodt T et al (2005) Tobacco bZIP transcription factor TGA2.2 and related factor TGA2.1 have distinct roles in plant defense responses and plant development. Plant J 44:100–113
Torregrosa CA, Dumas B, Krajinski F, Esquerre-Tugaye MT, Jacquet C (2006) Transcriptomic approaches to unravel plant–pathogen interactions in legumes. Euphytica 147:25–36
Vailleau F, Daniel X et al (2002) A R2R3-MYB gene, AtMYB30, acts as a positive regulator of the hypersensitive cell death program in plants in response to pathogen attack. Proc Natl Acad Sci U S A 99(15):10179–10184
Vettore A, da Silva FR, Kemper EL, Arruda P (2001) The libraries that made SUCEST. Genet Mol Biol 24(1–4):1–7
Vincentz M, Schlogl PS, Correa LGG, Kuhne F, Leite A (2001) Phylogenetic relationship between Arabidopsis and bZIP transcriptional regulatory factors. Genet Mol Biol 24:55–60
Viswanathan R (2000) Possible involvement of anthocyanin compounds in resistance of sugarcane against red rot. Indian phytopath 53(3):311–313
Viswanathan R, Malathi P, Padmanaban P (2003) Variation in sugarcane red rot pathogen Colletotrichum falcatum Went. In: Rao GP, Manoharachari C, Bhat DJ, Rajak RC, Lakhanpal TN (eds) Frontiers of fungal diversity in India. International Book Distributing Co, Lucknow, pp 639–667
Viswanathan R, Ramesh sundar A, Malathi P, Rahul PR, Ganesh kumar V, Prathima PT, Raveendran M, Kumar KK, Balasubramanian P (2009) Interaction between sugarcane and Colletotrichum falcatum causing red rot: understanding disease resistance at transcription level. Sugar tech 11(1):44–50
Viswanathan R (2010) Plant disease: Red rot of sugarcane. Anmol pubilication pvt. ISBN: 978-81-261-4214-9
Wang D, Amornsiripanitch N, Dong X (2006) A genomic approach to identify regulatory nodes in the transcriptional network of systemic acquired resistance in plants. PLoS pathogens 2(11):e123. doi:10.1371/journal.ppat.0020123
Xia N, Zhang G, Liu XY, Deng L, Cai GL, Zhang Y, Wang XJ, Zhao J, Huang LL, Kang ZS (2010) Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses. Mol Biol Rep 37(8):3703–3712
Yanhui C, Xiaoyuan Y, Kun H, Meihua L, Jigang L, Zhaofeng G, Zhiqiang L, Yunfei Z, Xiaoxiao W, Xiaoming Q, Yunping S, Li Z, Xiaohui D, Jingchu L, Xing-Wang D, Zhangliang C, Hongya G, Li-Jia Q (2006) The MYB transcription factor superfamily of Arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60(1):107–124
Zhang H, Jin JP, Tang L, Zhao Y, Gu XC, Gao G, Luo JC (2011) PlantTFDB 2.0: update and improvement of the comprehensive plant transcription factor database. Nucleic Acids Res 39:D1114–D1117
Acknowledgments
The authors are grateful to Dr.N. Vijayan Nair, Director of the institute for providing facilities and encouragement. The financial support received from Indian Council of Agricultural Research, Department of Biotechnology and the Department of Science of Technology, New Delhi is greatly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Muthiah, M., Ramadass, A., Amalraj, R.S. et al. Expression profiling of transcription factors (TFs) in sugarcane X Colletotrichum falcatum interaction. J. Plant Biochem. Biotechnol. 22, 286–294 (2013). https://doi.org/10.1007/s13562-012-0157-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13562-012-0157-7