Abstract
Background
Mitoxantrone (MTX) is a synthetic compound used as a second line chemotherapeutic drug for prostate cancer. It has been reported to trigger immunogenic cell death (ICD) in animal model studies, but the underlying mechanism is not fully understood yet, especially not in prostate cancer cells.
Methods
ICD was determined by assessing the release of damage-associated molecular patterns (DAMPs) in the prostate cancer-derived cell lines LNCaP, 22RV1 and PC-3. Short hairpin RNAs (shRNAs) were used to knock down target gene expression. Phagocytosis was assessed using a dual labeling technology in dendric cells co-cultured with cancer cells. The PERK gene promoter was cloned for dual luciferase assays. Chromatin immunoprecipitation (ChIP) was used to determine p53 protein-DNA binding activity. Immunocompetent mice and murine RM-1 prostate cancer cells were used for vaccination experiments.
Results
MTX treatment induced typical characteristics of DAMP release, including increased cell surface exposure of calreticulin (CALR), and extracellular release of ATP and high mobility group box-1 (HMGB1) protein. MTX also enhanced phagocytosis by dendritic cells. Moreover, MTX treatment increased eukaryotic initiation factor 2α (eIF2α) S51 phosphorylation, which was reduced when PERK and GCN2 were silenced using shRNAs. In addition, PERK or GCN2 silencing significantly reduced MTX-induced release of DAMPs in vitro and anti-tumor immunity in vivo. MTX treatment also resulted in dendritic cell activation in mice, which was attenuated when PERK or GCN2 were silenced in cancer cells used for vaccination. Further analysis revealed that PERK and GCN2 expression was enhanced by MTX treatment, of which PERK, but not GCN2, was enhanced via a p53-dependent mechanism.
Conclusion
MTX triggers ICD by activating eIF2α via PERK/GCN2 upregulation in prostate cancer cells. MTX-induced PERK expression upregulation depends on the p53 pathway, while that of GCN2 requires further investigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cell death classification has evolved in the last one and a half centuries from solely morphology-based grouping to the inclusion of multiple parameters, such as morphological manifestations, biochemical features and functional outcomes [1]. Stemming from the scientific advances of the past two decades [2], the conceptual distinction between apoptosis and necrosis has become indistinguishable in that they both occur in a regulated fashion and are involved in tissue homeostasis. Moreover, under certain circumstances apoptotic cells are detected by the immune system for clearance, similar to necrotic cells [3,4,5]. In the scenario of cancer chemotherapy, robust immunological memory is established against dying cancer cells exposed to chemo-drugs to evade new tumor formation, which is termed immunogenic cell death (ICD) [6, 7]. Recent studies have revealed that multiple insults can trigger ICD, including infectious pathogens, immunogenic chemotherapy, radiation, photodynamic therapy, high hydrostatic pressure and necroptosis [8, 9]. In current models of ICD induction, the process is triggered by a particular insult that induces a spatiotemporally coordinated emission of danger signaling traits, which are collectively termed damage-associated molecular patterns (DAMPs) [6, 10]. Several DAMPs, including cell surface exposure of CALR [11], extracellular release of ATP [12, 13] and nuclear protein HMGB1 [14], are considered universal contributors to virtually all ICD types. Therefore, these DAMPs are commonly used as hallmarks for in vitro evaluation of ICD inducers [7].
The endoplasmic reticulum (ER) is a key organelle involved in ICD induction [10, 15]. ICD inducers either selectively target the ER to induce both cell death and DAMPs, or trigger an ER-independent cell death but provoke DAMPs via ER stress [15, 16]. ER stress is a tightly orchestrated collection of activated sensors, including PERK, inositol-requiring enzyme 1α (IRE1α) and activating transcription factor 6 (ATF6) [17]. PERK protein responds to ER stress by inducing eIF2α phosphorylation, leading to increased C/EBP-homologous protein (CHOP) expression. IRE1α cleaves the transcription factor X-box binding protein (XBP-1) mRNA to produce a spliced XBP-1 variant (XBP-1 s) that triggers ER stress by up-regulating a large number of genes involved in the unfolded protein response (UPR) [18]. Under ER stress, ATF6 is cleaved by two proteases, serine protease site-1 protease (S1P) and metalloprotease site-2 protease (S2P), and cleaved ATF6 increases the expression of its target genes such as the ER resident chaperone binding immunoglobulin protein (BiP, also known as GRP78/HSPA5), glucose-response protein 94 (GRP94) and CHOP (reviewed in ref. [17, 18]). During ICD induction, eIF2α phosphorylation at residue serine-51 (S51) is essential for CALR translocation from the ER lumen to the cell surface, resulting in subsequent anti-cancer immune responses [19,20,21].
Prostate cancer is a leading cause of cancer deaths in men, especially in many western countries [22]. Although a recent study showed that oxaliplatin induced ICD in murine prostate cancer cells [23], the underlying molecular mechanism remains unknown. Mitoxantrone (MTX) is a type-2 DNA topoisomerase inhibitor that interferes with DNA synthesis and double-strand break repair in benign and malignant cells [24]. MTX has widely been used as the first-line chemotherapy treatment for metastatic prostate cancer before the Taxane era and has remained as the second line chemotherapy due to its low toxicity in the clinic [25, 26]. Previous reports have shown that MTX is a strong and bona fide ICD inducer in melanoma, osteosarcoma and mouse colon cancer cells, acting through an eIF2α phosphorylation-dependent mechanism [19, 21, 27, 28]. In this study, we investigated MTX-mediated ICD and the underlining mechanism in prostate cancer cells in vitro and in vivo. Our results reveal a novel mechanism of p53-dependent up-regulation of PERK expression, one of the four upstream kinases for the critical ICD mediator eIF2α, in prostate cancer cells upon MTX treatment.
2 Materials and methods
2.1 Cell lines, reagents, antibodies and shRNAs
Human prostate cancer LNCaP, 22RV1 and PC-3 cells, and mouse prostate cancer RM-1 cells were obtained from the ATCC (Manassas, VA, USA). Cells were cultured in RPMI-1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS, Gibco) plus 100 U/ml penicillin/streptomycin and 2 mmol/l L-glutamine.
Antibodies directed against CALR (ab2907), HMGB1 (ab79823), XBP-1 (ab220783), HRI (ab84980), phospho-GCN2 (ab75836) and GCN2 (ab134053) were purchased from Abcam (Cambridge, MA, USA). Antibodies directed against PARP (sc-8007) and p53 (sc-126) were obtained from Santa Cruz Biotech (Santa Cruz, CA, USA). Antibodies directed against eIF2α pS51 (#9721), CHOP (#2895), BiP (#3177), PERK (#5683), phospho-PERK (#3179) and eIF2α (#97225) were obtained from Cell Signaling (Danvers, MA, USA). PKR (18244–1-AP), HRP-conjugated goat anti-mouse IgG (SA00001–1) and goat anti-rabbit IgG (SA00001–2) were obtained from Proteintech (Chicago, IL, USA). CD14-FITC (130–110-576) and MicroBeads (130–050-201) were from Miltenyi Biotec (Bergisch Gladbach, Germany). PE/Cy7 anti-mouse CD11c, FITC anti-mouse CD8a, PerCP/Cy5.5 anti-mouse B220 and FITC anti-rabbit IgG were obtained from Biolengend (San Diego, CA, USA). PE-conjugated CD69, CD80 and CD86, as well as isotype control antibodies were obtained from BD Biosciences (Oxford, UK).
Cycloheximide (HY-12320), CFSE (HY-D0938) and Mitoxantrone (HY-13502A) were obtained from MCE (New Jersey, NJ, USA). RIPA Buffer (#9806) was obtained from Cell Signaling (Danvers, MA, USA). Cell Tracker Deep Red (C34565) was obtained from ThermoFisher Scientific (Shanghai, China). Plasmid constructs harboring the shRNA sequences of human HRI (TRCN0000010231), human PKR (TRCN0000197012), human PERK (TRCN0000262381), human GCN2 (TRCN0000304214), mouse PERK (TRCN0000028772) and GCN2 (TRCN0000361612) were purchased from Sigma (St Louis, MO, USA).
2.2 Western blotting and qRT-PCR
Western blotting was performed as previously described [29, 30]. Briefly, cells were lysed in RIPA buffer containing a complete protease inhibitor cocktail (Roche, Switzerland). Equal amounts of proteins from each cell lysate were subjected to SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto PVDF membranes. The membranes were incubated with primary antibodies overnight at 4 °C followed by a HRP-conjugated secondary antibody for 1 h at room temperature. Visualization was performed with ECL reagent (Santa Cruz Biotech).
Total cellular RNAs were extracted from cells using Trizol reagent (Invitrogen) according to the manufacturer’s instructions. RNA (1 μg) in a final volume of 20 μl was subjected to reverse transcription (RT) using a high-capacity RNA-to-cDNA master mix (Applied Biosystems, Foster City, CA, USA). The resulting RT products (0.5 μl) were subjected to real-time PCR analysis in a final volume of 20 μl using a SYBR Green-based protocol. 18S rRNA was included as an endogenous control. Real-time PCR was performed using a LightCycler® 480 machine (ROCHE Diagnostic Spa). Relative expression levels were calculated using the 2[−∆∆Ct] method and expressed as “fold change”. All data were normalized to 18S rRNA expression. The primer sequences used are listed on Table S1.
2.3 Lentivirus vector constructs, packaging and cell infection
Small hairpin interfering RNAs for the human p53 gene were cloned into a PLKO.1-puro vector using AgeI/EcoRI restriction enzyme sites [31, 32]. The pLKO.1-puro vector was a gift from Bob Weinberg (Addgene plasmid #8453). Lentiviruses were packaged using the psPAX2/pMD2.G system obtained from Dr. Didier Trono (Addgene plasmid #12259–12,260), as described before [33]. Briefly, psPAX2/pMD2.G and lentivirus vectors were co-transfected into 293 T cells and cultured for 24 h. Next, the cells were infected with supernatants containing the lentivirus. Stable expression clones were selected using puromycin (2 μg/ml).
2.4 Flow cytometry
For apoptosis analysis, cells were treated with MTX or vehicle control for 20 h. The treated cells were evaluated for apoptosis using a Dead Cell Apoptosis Kit (ThermoFisher Scientific, catalog #V13242). Briefly, 5 × 105 cells were resuspended in 100 μl binding buffer and next incubated with 5 μl FITC-conjugated Annexin-V antibody and 5 μl propidium iodide (PI) for 10 min according to manufacturer’s instructions.
CALR membrane translocation was analyzed using flow cytometry as reported before [34]. Briefly, 5 × 105 cells per treatment were incubated with an anti-CALR antibody for 40 min, followed by incubation with FITC-labeled anti-rabbit IgG (1:200) for 30 min. CALR-positive cells were gated on propidium iodide-negative cells.
Murine dendritic cell activation was evaluated by subset classification based on surface markers (CD69, CD80 and CD86). Splenocytes were isolated from mice, after which immunocytofluorescent staining was conducted with PE-conjugated antibodies for the respective surface markers, PE/Cy7-conjugated CD11c or FITC-conjugated CD8a for 30 min. Next, the samples were gated on CD8a+ DCs (CD8a+/B220−/CD11c+). Isotype-matched IgG antibodies were used as negative controls. Mean fluorescence intensities (MFI) were determined by flow cytometric analysis (Beckman CytoFLEX, Germany).
2.5 Extracellular ATP and HMGB1 assessment
After treatment, cell culture media were collected and the supernatants were processed by centrifugation. ATP levels were determined using a luciferin-based ENLITEN ATP assay obtained from Promega (Madison, WI, USA). HMGB1 protein levels were assessed using an enzyme-linked immunosorbent assay (ELISA) kit from IBL International (Hamburg, Germany).
2.6 Phagocytosis assay
Monocyte-derived dendritic cell (MoDC) isolation and phagocytosis assays were performed as previously described [28, 34]. Briefly, human peripheral blood mononuclear cells (PBMCs) were isolated from buffy coat fractions obtained from human blood donated by healthy adults (DRK-Blutspendedienst, Hessen GmbH). Monocytes were enriched up to 95% using CD14 MicroBeads (Miltenyi Biotec, catalog #130–050-201) and next stimulated with 100 ng/ml granulocyte-macrophage colony-stimulating factor (GM-CSF) and 100 ng/ml IL-4 (Peprotech, Rocky Hill, NJ, USA) for 6 days to obtain a population of immature DCs. LNCaP prostate cancer cells were labeled with 0.5 μM CMFDA for 10 min followed by MTX treatment for 20 h. After treatment, CMFDA-labelled LNCaP cells were co-cultured with Cell Tracker Deep Red-labeled MoDCs for 2 h followed by flow cytometry analysis. The phagocytic index (%) was calculated using the following formula: number of phagocytes harboring cancer (double positive) cells/total number of phagocytes (Deep Red positive) × 100 [35].
2.7 Animal vaccination
C57BL/6 J male mice were acquired from Vital River Laboratory Animal Technology (Beijing, China). All animal procedures were performed in accordance with the procedure guidelines and the experimental protocol approved by the Institutional Animal Care and Use Committee of Jining Medical University. RM-1 prostate cancer cells were stably transfected with plasmid constructs harboring shRNA for PERK or GCN2, or a control shRNA with scramble sequence. After treatment with 1 μM MTX for 20 h, 1 × 105 cells were inoculated subcutaneously in the left flank of the mice. One week after RM-1 cell inoculation, the mice were re-challenged by subcutaneous injection of 5 × 105 RM-1 cells in the right flank. Tumor sizes were measured every 3 days with a caliper. Tumor volumes were defined as [length x width2]/2. Mice bearing a tumor with a volume exceeding 1500 mm3 were euthanized.
2.8 EIF2AK3 gene promoter cloning and luciferase reporter construction
A 2-kb fragment of the EIF2AK3 gene (encoding PERK protein) promoter sequence was cloned into the firefly luciferase reporter pGL4.20 (Promega) using XhoI/HindIII restriction sites. P53 binding sites were predicted using the JASPAR platform (http://jaspar.genereg.net/) [36]. A mutant promoter was generated by deleting the putative p53 binding site using a PCR-based three-step mutagenesis approach as described before [33]. The oligonucleotide primer sequences used are listed in Table S1. Plasmid transfections were performed by electroporation (Gene Pulser Xcell, Bio-Rad). Luciferase reporter assays were conducted using a Dual Luciferase Reporter Assay kit with the pGL4.73-Renilla luciferase reporter (Promega) as internal control. Reporter activity was measured using the luciferase assay system GLOMAX™ luminometer (Promega, Sweden).
2.9 Chromatin-immunoprecipitation (ChIP)
ChIP was performed as previously described [37]. Briefly, after treatment, cells were subjected to cross-linking in 1% formaldehyde solution (50 mM Hepes-KOH, 100 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 11% formaldehyde) for 10 min, after which the formaldehyde was quenched with 1/20 volume of 2.5 M glycine. Cytosolic proteins were removed with the buffer systems LB1 (50 mM Hepes-KOH, pH 7.5; 140 mM NaCl; 1 mM EDTA; 10% Glycerol; 0.5% NP-40; 0.25% Triton X-100) and LB2 (10 mM Tris-HCL, pH 8.0; 200 mM NaCl; 1 mM EDTA; 0.5 mM EGTA). Nuclei were sonicated after which the sonication efficiency was determined by DNA isolation and gel electrophoresis. Immunoprecipitation was performed using an anti-p53 antibody. A normal rabbit IgG was used as negative control. P53-bound genomic DNA was eluted and quantitatively assessed using real-time PCR. The oligonucleotide primer sequences used are listed in Table S1.
2.10 Statistical analysis
Data are presented as means ± SEM from at least three independent experiments. Protein band densities were quantified using Image J software and the ratio to the control was calculated after normalization to the β-actin band density in the corresponding lane. The band density of the control was set at 1.0 for each comparison. The final density data were plotted using GraphPad Prism software. Differences of multiple groups were examined by one-way ANOVA analysis. Statistical significance between two groups was analyzed by unpaired Student’s t test. Comparisons between growth curves were performed using two-way ANOVA analysis. Survival and tumor-free curves were analyzed using Kaplan-Meier analysis. Statistical analyses were performed using SPSS 20.0 software (Chicago, IL, USA).
3 Results
3.1 MTX induces apoptotic cell death in prostate cancer cells
Although MTX was previously used as first line chemotherapy for prostate cancer [38, 39], its mechanistic action on prostate cancer cell death has not been clearly defined. Therefore, we first set out to assess whether MTX-induced cell death is apoptotic or necrotic in prostate cancer cells. Apoptotic and necrotic cell death was evaluated using Annexin-V/PI staining coupled with flow cytometric analysis. We found that MTX treatment for 20 h induced a profound apoptotic cell death with a slight induction of necrosis (5–8%) in LNCaP and 22RV1 cells (Supplementary Fig. S1a, S1b). Consistently, PARP cleavage, a characteristic of apoptosis, was induced by MTX treatment in LNCaP and 22RV1 cells (Supplementary Fig. S1c). These results indicate that MTX mainly induces apoptosis in prostate cancer cells.
3.2 MTX provokes immunogenic cell death characteristics in prostate cancer cells
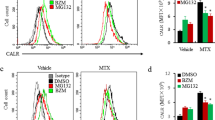
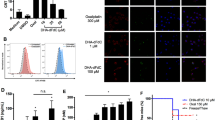
Next, we determined whether MTX triggers ICD induction in prostate cancer cells as reported in other cancer types [19]. ICD induction was evaluated in cell culture models by assessing DAMPs that are strictly required, including CALR display on the cell surface, and extracellular release of ATP and HMGB1 [40]. We found that MTX treatment in LNCaP and 22RV1 cells resulted in a significant increase in cell populations with positive CALR exposure (Fig. 1a, b). Concurrently, HMGB1 and ATP levels in cell culture media were significantly increased after MTX treatment (Fig. 1c, d). These data suggest that MTX treatment provoked DAMPs for ICD induction, as supported by previous studies [6].
MTX triggers the release of DAMPs. LNCaP and 22RV1 cells were treated with vehicle or MTX (1 μM) for 20 h and then harvested for CALR translocation analysis using anti-CALR antibody staining coupled with flow cytometry. An isotype IgG was used as internal control. Cell culture media were collected for HMGB1 and ATP measurements. a. Representative flow cytometry curves for CALR-positive cell populations. b. Quantitative data of CALR translocation assays. c. HMBG1 levels in cell culture media determined by ELISA. d. ATP levels in cell culture media determined by luciferase assay. The assays were repeated four times and the asterisks indicate significant differences compared to the vehicle controls (Student t-test, *** p < 0.001)
CALR cell surface exposure/translocation is considered to be a “eat-me” signal for the immune surveillance system through phagocytosis [11]. To assess whether MTX treatment triggers phagocytosis by dendritic cells (DCs), LNCaP and 22RV1 cells were treated with MTX for 20 h and subsequently labeled with the fluorescent dye CMFDA. Next, CMFDA-labeled cells were co-cultured with monocyte-derived DCs (MoDCs) pre-labeled with CellTracker™ Deep Red Dye. Flow cytometric analysis revealed that the cell populations exhibiting dual fluorescence were significantly higher in MTX-treated LNCaP and 22RV1 cells compared to the vehicle controls (Fig. 2a, b). These data indicate that MTX treatment induced a profound phagocytosis, a key event in ICD induction.
MTX stimulates phagocytosis by dendritic cells. LNCaP and 22RV1 cells were treated with vehicle or MTX (1 μM) for 20 h and then labeled with CMFDA before culture with CellTracker Deep Red-labeled MoDCs for 2 h. Cells were harvested for phagocytosis analysis using flow cytometry. a. Flow cytometry plots from a representative experiment. b. Phagocytosis index calculated as the percentage of double positive cells from the total population of the Deep Red positive cells. Quantitative data were deduced from triplicate experiments. The asterisks indicate significant differences compared to the vehicle controls (Student t-test, *** p < 0.001)
3.3 MTX induces eIF2α phosphorylation and DAMPs depending on PERK and GCN2
To understand the mechanism involved in MTX-induced DAMPs, eIF2α phosphorylation at residue S51, a critical event of ICD induction, was examined in LNCaP and 22RV1 cells. We found that MTX treatment largely increased the levels of eIF2α S51 phosphorylation (Fig. 3a, b). Concomitantly, CHOP protein, an eIF2α downstream effector, was also drastically induced. Notably, two other ER stress responding proteins, BiP and XBP1s, did not show any typical alterations. MTX treatment reduced XBP1s protein levels but had no effect on BiP protein expression. These data indicate that MTX treatment specifically induced eIF2α S51 phosphorylation, a critical indicator for ICD induction [19, 21].
MTX induces eIF2α phosphorylation via PERK and GCN2. a-b. LNCaP and 22RV1 cells were treated with MTX (1 μM) or vehicle for 20 h. Whole cell lysates were subjected to Western blot analysis with antibodies as indicated. Actin served as endogenous protein loading control. Quantitative data (panel b) of protein band densities were normalized to vehicle-treated cells from three experiments. c-f. LNCaP (panel c-d) and 22RV1 (panel e-f) cells transfected with shRNAs as indicated were treated with MTX (1 μM) or vehicle for 20 h, followed by Western blot analysis with antibodies as indicated. Actin served as endogenous protein loading control. Relative densities of pS51 vs total eIF2α bands from LNCaP cells (panel d) and 22RV1 cells (panel f) were quantified using Inage J software. Data were deduced from three independent experiments. The asterisks indicate significant differences compared to the shScr group in the corresponding groups (Student t -test, * p < 0.05, ** p < 0.01, *** p < 0.001)
Four kinases are known to be responsible for eIF2α S51 phosphorylation, including heme-regulated inhibition factor-2α (HRI) encoded by EIF2AK1, protein kinase RNA-activated (PKR) encoded by EIF2AK2, protein kinase RNA (PKR)-like ER kinase (PERK) encoded by EIF2AK3 and general control non-derepressible-2 (GCN2) encoded by EIF2AK4 [41]. To determine which of these kinases contributed to eIF2α S51 phosphorylation during MTX-induced DAMPs, these genes were individually knocked down in LNCaP and 22RV1 cells using gene-specific small hairpin RNAs (shRNAs). A scrambled shRNA (shScr) was used as negative control. Compared to the control shRNA, PERK but not GCN2 shRNA largely reduced the basal eIF2α pS51 levels in LNCaP cells, and both PERK and GCN2 shRNAs largely reduced MTX-induced eIF2α S51 phosphorylation (Fig. 3c, d). The shRNAs for HRI or PKR had no significant effect. Interestingly, in 22RV1 cells both PERK and GCN2 knockdown not only reduced the basal levels of eIF2α pS51, but also abolished the MTX-induced effect (Fig. 3e, f). These results indicate that MTX induced eIF2α S51 phosphorylation depending on both PERK and GCN2 kinases and that the basal eIF2α S51 phosphorylation levels are differently regulated by PERK and GCN2 in a cell-specific manner.
To elucidate whether PERK or GCN2 is involved in MTX-triggered ICD induction, we analyzed CALR translocation, HMGB1 release and phagocytosis in LNCaP cells. Similar to the eIF2α S51 phosphorylation effect, PERK and GCN2 knockdown, but not HRI and PKR knockdown, significantly reduced MTX-triggered CALR exposure on the cell surfaces (Fig. 4a, b), extracellular release of HMBG1 (Fig. 4c) and phagocytosis by DCs (Fig. 4d, e). These data indicate that both PERK and GCN2 are involved in MTX-induced eIF2α S51 phosphorylation, the release of DAMPs and subsequent ICD induction. To verify whether gene silencing affects cell survival and, subsequently, interferes with MTX-induced ICD induction, we assessed apoptotic events after shRNA transfection. We found that HRI, PKR or PERK knockdown in LNCaP cells did not significantly attenuate basal or MTX-induced apoptosis, indicating that the attenuated ICD induction after PERK silencing was not due to increased cell death (Supplementary Fig. S2). On the other hand, GCN2 knockdown induced a moderate increase in apoptotic cell death with or without MTX treatment, indicating that GCN2 is an essential survival factor as has previously been reported in T lymphocytes [42], which may potentially interfere with MTX-induced eIF2α phosphorylation and ICD induction.
MTX induces DAMPs and phagocytosis depending on PERK and GCN2. a-c. LNCaP cells transfected with shRNAs as indicated were treated with MTX (1 μM) or vehicle for 20 h. Cells were harvested for CALR analysis by flow cytometry. Representative flow cytometry curves are shown in panel a and quantitative data are summarized in panel b from three independent experiments. Cell culture media were collected for HMGB1 measurement in triplicates (panel c). d-e. LNCaP cells transfected with shRNAs as indicated were treated with MTX (1 μM) or vehicle for 20 h followed by phagocytosis assay. Representative flow cytometry plots are shown in panel e and quantitative phagocytosis index data were deduced from three independent experiments (panel d). The asterisks indicate significant differences compared to MTX-treated shScr cells (Student t-test, *** p < 0.001)
3.4 PERK and GCN2 are required for MTX-induced anti-tumor immunity in vivo
We next determined the role of PERK and GCN2 kinases in MTX-triggered ICD and anti-cancer immunity in vivo using a vaccination approach, the gold standard method for ICD studies [7]. MTX-induced eIF2α pS51 was first verified in vitro in murine prostate cancer RM-1 cells. Similar to the human prostate cancer cells, RM-1 cells also exerted a significant increase in eIF2α S51 phosphorylation after MTX treatment (Fig. 5a, b), whereas PERK and GCN2 knockdown in RM-1 cells significantly reduced MTX-induced eIF2α S51 phosphorylation (Fig. 5c, d).
MTX stimulates eIF2a phosphorylation depending on PERK/GCN2 in RM-1 cells. a. RM-1 cells were treated with vehicle or MTX (1 μM) for 20 h. Whole cell proteins were used for Western blot analysis with the antibodies as indicated. Actin served as protein loading control. b. Relative band densities of p-PERK/total PERK, p-GCN2/total GCN2 or pS51/total eIF2a were calculated using Image-J software from three independent experiments. The asterisks indicate significant differences compared to the vehicle control (Student t-test, *** p < 0.001). c. RM-1 cells transfected with shRNAs as indicated were treated with MTX (1 μM) or vehicle for 20 h, followed by Western blot analysis with the indicated antibodies. d. Relative band densities of pS51/total eIF2α were determined using Image-J software from three independent experiments. The asterisks indicate significant differences compared to MTX-treated shScr cells (Student t-test, ** p < 0.01, *** p < 0.001)
For the in vivo vaccination assay, RM-1 subline cells stably transfected with gene-specific or scramble shRNAs were treated with MTX for 20 h before inoculation into immune-competent mice (Fig. 6a). Three additional control groups were included, i.e., untransfected RM-1 cells (RM-1), shScr-transfected RM-1 cells (shScr), and frozen-thawed shScr-transfected RM-1 cells (FT + shScr). One week after inoculation, the animals were re-challenged with exponentially grown RM-1 cells. We found that RM-1 xenograft tumors exhibited a steadily growth curve in all control groups (Fig. 6b), whereas animals vaccinated with MTX-treated shScr-transfected RM-1 cells (MTX + shScr group) showed a dramatic reduction in xenograft tumor growth (Fig. 6b), a longer tumor-free survival (Fig. 6c) and a better overall survival (Fig. 6d). These results indicate a successful ICD-mediated vaccination after MTX treatment. Interestingly, compared to the MTX + shScr group, mice vaccinated with PERK (MTX + shPERK) or GCN2 (MTX + shGCN2) knockdown cells exhibited a moderately less suppression in xenograft tumor growth (Fig. 6b), a reduced tumor-free survival (Fig. 6c) and a worse overall survival (Fig. 6d). These data indicate that PERK and GCN2 are involved in MTX-triggered anti-tumor immunity in vivo.
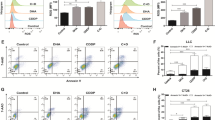
MTX promotes anti-tumor immunity and induces dendritic cell activation in vivo. a. Scheme of the vaccination experiments. RM-1 cells stably transfected with shScr, shPERK or shGCN2 constructs were left untreated or treated with MTX (1 μM) for 20 h, followed by subcutaneous injection (1 × 105 cells) into the left flank of C57BL/6 mice (n = 10). The three control groups were: untreated RM-1 cells (RM-1 cells), shScr-transfected (shScr) or frozen-thawed shScr-transfected RM-1 cells (FT + shScr). The experimental groups were vaccinated with MTX-treated shRNA-transfected RM-1 cells (MTX + shScr, MTX + shPERK or MTX + shGCN2). One week later, all mice were re-challenged by subcutaneous inoculation of 5 × 105 exponentially grown RM-1 cells in the right flank. Tumor growth was monitored twice a week using a caliper. b-d. Quantitative data for tumor growth curve (b), tumor-free progression (c) and animal overall survival (d) were summarized. Statistical analysis was performed using two-way ANOVA analysis. The colored asterisks indicate significant differences between the indicated groups (* p < 0.05). e-h. Splenocytes were isolated at the end of the experiments from mice vaccinated with shRNA-transfected RM-1 cells with or without MTX treatment as indicated (n = 3 per group). Surface markers CD69, CD80 and CD86 on dendritic cells (CD8a+/B220−/CD11c+) were analyzed using flow cytometry. Representative curves are shown in panel e and quantitative data are summarized in panel f-h. The asterisks indicate a significant difference compared to the MTX + shScr group, *** p < 0.001
To next determine whether vaccination with MTX-treated cells results in DC activation, we isolated splenocytes from mice vaccinated with MTX- or vehicle-treated RM-1 subline cells and analyzed the expression patterns of co-stimulatory molecules (CD80 and CD86) and the early activation marker CD69 molecule on CD8a+ DCs (CD8a+/B220−/CD11c+). We found that vaccination with MTX-treated cells resulted in significant increases in the surface markers CD69, CD80 and CD86 on DC cells, which is a clear sign of DC activation (Fig. 6e-h). This increased DC activation was blunted after PERK or GCN2 knockdown. These data indicate that PERK and GNC2 kinases are significant contributors of MTX-triggered DC activation in vivo.
3.5 MTX treatment enhances PERK and GCN2 gene expression
While analyzing the involvement of PERK and GCN2 kinases in MTX-induced eIF2α S51 phosphorylation, we noticed that MTX treatment increased PERK and GCN2 protein levels, but not PKR or HRI protein levels, in LNCaP and 22RV1 cells (Fig. 3c, e). To further investigate the mechanism underlying MTX-induced alterations in PERK and GCN2 expression, we evaluated whether MTX treatment increases the phosphorylation levels of PERK at T980 or GCN2 at T899, which are required for a full activation of these kinases [43, 44]. We found that MTX treatment in LNCaP and 22RV1 cells indeed resulted in elevation of the PERK and GCN2 phosphorylation levels, but their total protein levels were also largely increased (Fig. 7a). Consequently, the ratios of p-PERK/PERK or p-GCN2/GCN2 were not significant altered after MTX treatment (Fig. 7b). Similarly, we found that MTX treatment increased the levels of PERK and GCN2 proteins in RM-1 cells, in parallel with elevated phosphorylation of both proteins (Fig. 5a). These data indicate that the increased PERK and GCN2 phosphorylation levels are potentially due to total protein alterations. Further analysis using a real-time RT-PCR approach revealed that MTX treatment significantly increased the mRNA levels of the EIF2AK3 (PERK) and EIF2AK4 (GCN2) genes by approximately 3 to 6-fold compared to the vehicle controls (Fig. 7c). These data indicate that MTX treatment enhances EIF2AK3 (PERK) and EIF2AK4 (GCN2) gene expression, leading to increased PERK and GCN2 protein levels and subsequent eIF2α S51 phosphorylation levels.
MTX treatment enhances PERK and GCN2 gene expression. a. LNCaP and 22RV1 cells were treated with vehicle or MTX (1 μM) for 20 h. Whole cell lysates were used for Western blot analysis with the antibodies indicated. Actin served as protein loading control. b. Relative band densities for PREK/Actin, GCN2/Actin, p-PERK/PERK or p-GCN2/GCN2 were determined using Image-J software from three independent experiments. c. LNCaP and 22RV1 cells were treated with vehicle or MTX (1 μM) for 20 h after which total RNAs were extracted for gene expression analysis by real-time PCR. Data are summarized from three independent experiments. The asterisks indicate significant differences compared to the vehicle controls (Student t-test, * p < 0.05, ** p < 0.01, *** p < 0.001)
3.6 p53 is involved in MTX-induced PERK upregulation
The tumor suppressor protein p53 functions primarily as a transcription factor to regulate numerous target genes, and recent data also suggest a functional role in ICD-related anti-tumor immunity [45]. In addition, MTX has been reported to enhance p53 expression in leukemic clam hemocytes [46]. To determine whether p53 is involved in the regulation of MTX-induced PERK/GCN2 expression, p53 expression was silenced in LNCaP cells expressing wild-type p53 [47]. We found that p53 silencing abolished MTX-induced PERK, but not GNC2 protein up-regulation in these cells (Fig. 8a, b). To verify these results, a TP53 gene-null PC-3 cell line [47] was tested for MTX-induced PERK expression. In PC-3 cells, the PERK protein level was very low at basal conditions and was not induced by MTX treatment. Notably, we found that MTX treatment markedly increased GCN2 protein and eIF2α phosphorylation levels in a dose-dependent manner in these cells (Fig. 8c, d). In addition, we found that MTX treatment induced significant increases in CALR translocation to the cell surfaces (Fig. 8e, f), extracellular release of HMGB1 (Fig. 8g) and ATP (Fig. 8h), as well as the phagocytosis index by dendritic cells (Fig. 8i, j) in PC-3 cells, consistent with the results from LNCaP and 22RV1 cells (Figs. 1 and 2). These data suggest that MTX-induced PERK up-regulation acts at the transcription level via a p53-dependent mechanism, while MTX-induced GCN2 up-regulation is p53 independent. In addition, GCN2 appears to be capable to compensate for the lack of p53-mediated PERK up-regulation by promoting eIF2α S51 phosphorylation and DAMP release in p53-negative cells. To confirm the key role of GCN2 in these events, GCN2 expression was silenced using the shRNA approach in PC-3 cells. By doing so, we found that GCN2 knockdown markedly reduced eIF2α phosphorylation at basal conditions and after MTX treatment (Fig. 8k, l). These data suggest that in p53-negative cells, while PERK expression is low, GCN2 is the major contributor in MTX-induced eIF2α phosphorylation.
MTX promotes p53-dependent PERK/EIF2AK3 gene up-regulation. a-b. LNCaP cells stably transfected with shRNAs as indicated were treated with vehicle or MTX (1 μM) for 20 h. Whole cell lysates were subjected to Western blot analysis with the antibodies as indicated. Representative blots are shown in panel a from three independent experiments and the relative band densities are summarized in panel b. c-d. PC-3 cells were treated with vehicle or MTX at the indicated concentrations for 20 h. Representative blots are shown in panel c from three independent experiments and the relative band densities of S51/total eIF2α are summarized in panel d. e-h. PC-3 cells were treated with vehicle or MTX (1 μM) for 20 h and next harvested for assessing CALR translocation. Cell culture media were collected for assessing HMGB1 and ATP levels. Representative curves of flow cytometry are shown in panel e and the quantitative data for CALR translocation, HMGB1 and ATP levels from three independent experiments are shown in panel f-h. i-j. PC-3 cells were treated with vehicle or MTX (1 μM) for 20 h and next harvested for CMFDA labeling. After co-culture with CellTracker Deep Red-labeled MoDCs for 2 h, the cells were subjected to flow cytometry to analyze phagocytosis events. Representative plots of flow cytometry are shown in panel i and the quantitative data for phagocytosis index were calculated as the percentage of double positive cells from the total population of the Deep Red positive cells (panel j). k-l. PC-3 cells stably transfected with shRNAs as indicated were treated with vehicle or MTX (1 μM) for 20 h. Representative blots from three independent experiments are shown in panel k and the relative band densities are summarized in panel l. The asterisks indicate a significant difference compared to the vehicle control (Student t-test, * p < 0.05, ** p < 0.01)
To elucidate the mechanism underlying p53-mediated PERK up-regulation, a 2-kb human PERK/EIF2AK3 gene promoter sequence was cloned into a firefly luciferase reporter (PERKwt-LUC). Within this promoter sequence a putative p53 binding site (−826 to −812) was predicted using the JASPAR platform [36]. A mutant PERKΔp53-LUC reporter was next created by deleting this potential p53-binding site (Fig. 9a and Table S2). We subsequently found that MTX treatment significantly increased PERKwt-LUC reporter activity in LNCaP cells, which was diminished when p53 was knocked down, consistent with the protein expression data (Fig. 9b). Deletion of the p53-binding site from the PERK promoter abolished MTX-induced PERK promoter activation (Fig. 9c). To further verify p53 binding to the PERK promoter, we performed an anti-p53 ChIP assay in LNCaP cells. We found that MTX treatment significantly enhanced p53 binding to the PERK promoter compared to the vehicle control (Fig. 9d). These data suggest that the p53 protein is involved in MTX-induced PERK gene up-regulation by binding to its promoter.
MTX enhances p53 transcriptional activity by increasing its protein stability. a. Scheme of PERK/EIF2AK3 gene promoter cloning and luciferase reporter construction. A putative p53 binding site (−826/−811) is illustrated within the 2-kb promoter region. A mutant reporter (Δp53-LUC) was established by deleting the putative p53 binding site. b. LNCaP cells stably transfected with control shRNA (shScr) or TP53 shRNA (shTP53) were used in the luciferase assay with the wt-LUC PERK gene reporter. The Renilla reporter was used as internal control. Cells were treated with MTX (1 μM) or vehicle overnight. Luciferase reporter activities were assessed using a dual-luciferase assay. Data are shown from three independent experiments. c. LNCaP cells were transfected with wt-LUC or Δp53-LUC reporters overnight and next treated with MTX (1 μM) or vehicle for 20 h, followed by luciferase assay. d. LNCaP cells were treated with MTX (1 μM) or vehicle for 20 h and next subjected to ChIP using anti-p53 antibodies. Normal IgG was used as negative control. The anti-p53-bound DNA eluates were quantitatively assessed using RT-PCR. e-f. LNCaP and 22RV1 cells were treated with MTX at the indicated concentrations for 20 h. Expression levels of the p53 protein were determined by Western blotting. Actin served as a protein loading control. Relative band densities of the p53 protein were calculated from three independent experiments. g. LNCaP and 22RV1 cells were treated with vehicle or MTX (1 μM) for 20 h after which total RNAs were extracted for assessing p53 expression by real-time RT-PCR in triplicate experiments. h-i. 22RV1 cells were treated with CHX (100 μM) alone or in combination of MTX (1 μM) for the indicated time periods. Relative band densities were acquired with Image-J software and the value at time-point 0 was set at 100% and plotted with Graphpad Prism 5.0 software. The asterisks indicate significant differences compared to the vehicle controls (Student t-test, * p < 0.05, ** p < 0.01, *** p < 0.001)
Finally, we investigated whether MTX treatment affects p53 expression in prostate cancer cells. We found that MTX treatment in LNCaP (wild-type p53) and 22RV1 (p53Q331R mutant) cells markedly increased p53 protein levels in a concentration-dependent manner (Fig. 9e, f), but had no effect on its mRNA levels (Fig. 9g), suggesting an increased stability of the p53 protein. Therefore, a cycloheximide (CHX)-based protein pulse-chase assay was conducted to evaluate p53 protein stability after MTX treatment. After LNCaP cells were treated with CHX plus or minus MTX for up to 8 h, we found that MTX treatment significantly slowed down p53 protein decay compared to the vehicle control, indicating that the p53 protein was stabilized after MTX treatment. (Fig. 9h, i).
4 Discussion
Cancer cells undergoing ICD are associated with the release of DAMPs and ER stress responses, which evoke anti-tumor immune responses. Recently, several ICD-related DAMPs have been reported, including cell surface translocation of the ER protein CALR [11], and extracellular release of HMGB1 [14] and ATP [12, 13]. Rapid translocation of CALR protein enhances the phagocytosis of dying cancer cells by DCs [11, 48]. HMGB1 exhibits multiple immunostimulatory functions upon binding to a wide range of pattern recognition receptors (PRRs) [49]. ATP that is released during ICD acts as a homing signal and activates the NLRP3 inflammasome [50, 51]. Our study showed that MTX treatment in human and murine prostate cancer cells resulted in the presentation of DAMPs, including CALR exposure, ATP and HMGB1 release, and DC phagocytosis.
ER stress is sensed by three transmembrane protein complexes, namely PERK/eIF2α, IRE1/XBP1 and ATF6 [17]. Previous studies have shown that eIF2α phosphorylation, but not of the other ER stress signals, is the key event in triggering ICD induction [19, 52]. CALR translocation to the plasma membrane occurs following eIF2α phosphorylation in dying cancer cells and triggers “find me” and “eat me” signals, thereby promoting phagocytic uptake of dying cancer cells by DCs [1, 6]. In this study, we have shown that MTX treatment increased eIF2α phosphorylation, but reduced IRE1a/XBP1 expression and had no effect on the ATF6/BiP pathway, indicating eIF2α as the sole ER factor that facilitates ICD induction in prostate cancer cells [19, 52].
Although it is well accepted that eIF2α phosphorylation is a pathogenomonic hallmark of ICD induction, the kinases responsible for ICD-related eIF2α S51 phosphorylation vary across different studies. Most studies have indicated that PERK is responsible for eIF2α S51 phosphorylation during ICD induction [19, 20, 53]. However, PKR and GCN2, but not PERK, were found to act as the key mediators of eIF2α S51 phosphorylation in human melanoma cells [21]. In this study, we showed that PERK and GCN2, but not HRI and PER, were responsible for MTX-induced eIF2α S51 phosphorylation in prostate cancer cells, which is new compared to previous studies [19,20,21, 53]. Moreover, we found that MTX treatment increased PERK and GCN2 protein levels and that PERK or GCN2 knockdown reduced MTX-induced ICD induction in vitro and anti-tumor immunity in vivo. These data indicate that MTX-induced PERK and GCN2 up-regulation is responsible for eIF2α phosphorylation and subsequently ICD induction.
The tumor suppressor protein p53 is a transcription factor that has recently been implicated in anti-tumor immunity [45]. In this study, we found that p53 is involved in MTX-induced PERK, but not GCN2, up-regulation. Interestingly, we found that in p53-negative prostate cancer cells GCN2 is capable of mediating MTX-induced eIF2α S51 phosphorylation and ICD induction, while PERK expression is extremely low, suggesting that the p53/PERK mechanism is dispensable in some cell types for ICD induction. Further analysis indicated that MTX enhanced p53 protein stability without affecting its mRNA transcription. Post-translational modifications of p53 are critical in modulating its stability [54, 55]. A series of modifications of the p53 protein are known to be involved in the regulation of p53 protein stability, including phosphorylation [55, 56], acetylation [54] and methylation [57, 58]. Additional studies are needed to determine how MTX enhances p53 protein stability.
Although immunotherapy by targeting immune checkpoints is a focus of current anti-tumor drug development, the success rate needs significant improvement for prostate cancer patients [59]. As a FDA-approved second-line chemotherapy after Taxane compounds for late-stage prostate cancer, MTX may be used together with anti-PD-1/PDL1 antibodies to improve immunotherapy in prostate cancer patients as recently discussed [60], since MTX is less toxic compared to Taxane compounds that do not act as bona fade ICD inducers [61].
In conclusion, we found that MTX treatment triggers ICD induction in prostate cancer cells in vitro and induces anti-tumor immunity in vivo. In addition, we found that MTX-induced ICD is associated with PERK/GCN2-dependent eIF2α phosphorylation and that MTX treatment enhances p53 protein stability, leading to EIF2AK3 (PERK) gene up-regulation through a p53-dependent transcriptional mechanism. MTX-induced EIF2AK2 (GCN2) gene expression occurs via a p53-independent pathway.
References
L. Galluzzi, I. Vitale, S.A. Aaronson, J.M. Abrams, D. Adam, P. Agostinis, E.S. Alnemri, L. Altucci, I. Amelio, D.W. Andrews, M. Annicchiarico-Petruzzelli, A.V. Antonov, E. Arama, E.H. Baehrecke, N.A. Barlev, N.G. Bazan, F. Bernassola, M.J.M. Bertrand, K. Bianchi, M.V. Blagosklonny, K. Blomgren, C. Borner, P. Boya, C. Brenner, M. Campanella, E. Candi, D. Carmona-Gutierrez, F. Cecconi, F.K. Chan, N.S. Chandel, E.H. Cheng, J.E. Chipuk, J.A. Cidlowski, A. Ciechanover, G.M. Cohen, M. Conrad, J.R. Cubillos-Ruiz, P.E. Czabotar, V. D'Angiolella, T.M. Dawson, V.L. Dawson, V. De Laurenzi, R. De Maria, K.M. Debatin, R.J. DeBerardinis, M. Deshmukh, N. Di Daniele, F. Di Virgilio, V.M. Dixit, S.J. Dixon, C.S. Duckett, B.D. Dynlacht, W.S. El-Deiry, J.W. Elrod, G.M. Fimia, S. Fulda, A.J. Garcia-Saez, A.D. Garg, C. Garrido, E. Gavathiotis, P. Golstein, E. Gottlieb, D.R. Green, L.A. Greene, H. Gronemeyer, A. Gross, G. Hajnoczky, J.M. Hardwick, I.S. Harris, M.O. Hengartner, C. Hetz, H. Ichijo, M. Jaattela, B. Joseph, P.J. Jost, P.P. Juin, W.J. Kaiser, M. Karin, T. Kaufmann, O. Kepp, A. Kimchi, R.N. Kitsis, D.J. Klionsky, R.A. Knight, S. Kumar, S.W. Lee, J.J. Lemasters, B. Levine, A. Linkermann, S.A. Lipton, R.A. Lockshin, C. Lopez-Otin, S.W. Lowe, T. Luedde, E. Lugli, M. MacFarlane, F. Madeo, M. Malewicz, W. Malorni, G. Manic, J.C. Marine, S.J. Martin, J.C. Martinou, J.P. Medema, P. Mehlen, P. Meier, S. Melino, E.A. Miao, J.D. Molkentin, U.M. Moll, C. Munoz-Pinedo, S. Nagata, G. Nunez, A. Oberst, M. Oren, M. Overholtzer, M. Pagano, T. Panaretakis, M. Pasparakis, J.M. Penninger, D.M. Pereira, S. Pervaiz, M.E. Peter, M. Piacentini, P. Pinton, J.H.M. Prehn, H. Puthalakath, G.A. Rabinovich, M. Rehm, R. Rizzuto, C.M.P. Rodrigues, D.C. Rubinsztein, T. Rudel, K.M. Ryan, E. Sayan, L. Scorrano, F. Shao, Y. Shi, J. Silke, H.U. Simon, A. Sistigu, B.R. Stockwell, A. Strasser, G. Szabadkai, S.W.G. Tait, D. Tang, N. Tavernarakis, A. Thorburn, Y. Tsujimoto, B. Turk, T. Vanden Berghe, P. Vandenabeele, M.G.V. Heiden, A. Villunger, H.W. Virgin, K.H. Vousden, D. Vucic, E.F. Wagner, H. Walczak, D. Wallach, Y. Wang, J.A. Wells, W. Wood, J. Yuan, Z. Zakeri, B. Zhivotovsky, L. Zitvogel, G. Melino, G. Kroemer, Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ 25, 486–541 (2018). https://doi.org/10.1038/s41418-017-0012-4
D.R. Green, The coming decade of cell death research: Five riddles. Cell 177, 1094–1107 (2019). https://doi.org/10.1016/j.cell.2019.04.024
T.L. Aaes, A. Kaczmarek, T. Delvaeye, B. De Craene, S. De Koker, L. Heyndrickx, I. Delrue, J. Taminau, B. Wiernicki, P. De Groote, A.D. Garg, L. Leybaert, J. Grooten, M.J. Bertrand, P. Agostinis, G. Berx, W. Declercq, P. Vandenabeele, D.V. Krysko, Vaccination with Necroptotic Cancer cells induces efficient anti-tumor immunity. Cell Rep 15, 274–287 (2016). https://doi.org/10.1016/j.celrep.2016.03.037
N. Casares, M.O. Pequignot, A. Tesniere, F. Ghiringhelli, S. Roux, N. Chaput, E. Schmitt, A. Hamai, S. Hervas-Stubbs, M. Obeid, F. Coutant, D. Metivier, E. Pichard, P. Aucouturier, G. Pierron, C. Garrido, L. Zitvogel, G. Kroemer, Caspase-dependent immunogenicity of doxorubicin-induced tumor cell death. J Exp Med 202, 1691–1701 (2005). https://doi.org/10.1084/jem.20050915
H. Yang, Y. Ma, G. Chen, H. Zhou, T. Yamazaki, C. Klein, F. Pietrocola, E. Vacchelli, S. Souquere, A. Sauvat, L. Zitvogel, O. Kepp, G. Kroemer, Contribution of RIP3 and MLKL to immunogenic cell death signaling in cancer chemotherapy. Oncoimmunology 5, e1149673 (2016). https://doi.org/10.1080/2162402X.2016.1149673
L. Galluzzi, A. Buque, O. Kepp, L. Zitvogel, G. Kroemer, Immunogenic cell death in cancer and infectious disease. Nat Rev Immunol 17, 97–111 (2017). https://doi.org/10.1038/nri.2016.107
O. Kepp, L. Senovilla, I. Vitale, E. Vacchelli, S. Adjemian, P. Agostinis, L. Apetoh, F. Aranda, V. Barnaba, N. Bloy, L. Bracci, K. Breckpot, D. Brough, A. Buque, M.G. Castro, M. Cirone, M.I. Colombo, I. Cremer, S. Demaria, L. Dini, A.G. Eliopoulos, A. Faggioni, S.C. Formenti, J. Fucikova, L. Gabriele, U.S. Gaipl, J. Galon, A. Garg, F. Ghiringhelli, N.A. Giese, Z.S. Guo, A. Hemminki, M. Herrmann, J.W. Hodge, S. Holdenrieder, J. Honeychurch, H.M. Hu, X. Huang, T.M. Illidge, K. Kono, M. Korbelik, D.V. Krysko, S. Loi, P.R. Lowenstein, E. Lugli, Y. Ma, F. Madeo, A.A. Manfredi, I. Martins, D. Mavilio, L. Menger, N. Merendino, M. Michaud, G. Mignot, K.L. Mossman, G. Multhoff, R. Oehler, F. Palombo, T. Panaretakis, J. Pol, E. Proietti, J.E. Ricci, C. Riganti, P. Rovere-Querini, A. Rubartelli, A. Sistigu, M.J. Smyth, J. Sonnemann, R. Spisek, J. Stagg, A.Q. Sukkurwala, E. Tartour, A. Thorburn, S.H. Thorne, P. Vandenabeele, F. Velotti, S.T. Workenhe, H. Yang, W.X. Zong, L. Zitvogel, G. Kroemer, L. Galluzzi, Consensus guidelines for the detection of immunogenic cell death. Oncoimmunology 3, e955691 (2014). https://doi.org/10.4161/21624011.2014.955691
M. Diederich, Natural compound inducers of immunogenic cell death. Arch Pharm Res 42, 629–645 (2019). https://doi.org/10.1007/s12272-019-01150-z
J. Zhou, G. Wang, Y. Chen, H. Wang, Y. Hua, Z. Cai, Immunogenic cell death in cancer therapy: Present and emerging inducers. J Cell Mol Med 23, 4854–4865 (2019). https://doi.org/10.1111/jcmm.14356
A.D. Garg, P. Agostinis, Cell death and immunity in cancer: From danger signals to mimicry of pathogen defense responses. Immunol Rev 280, 126–148 (2017). https://doi.org/10.1111/imr.12574
M. Obeid, A. Tesniere, F. Ghiringhelli, G.M. Fimia, L. Apetoh, J.L. Perfettini, M. Castedo, G. Mignot, T. Panaretakis, N. Casares, D. Metivier, N. Larochette, P. van Endert, F. Ciccosanti, M. Piacentini, L. Zitvogel, G. Kroemer, Calreticulin exposure dictates the immunogenicity of cancer cell death. Nat Med 13, 54–61 (2007). https://doi.org/10.1038/nm1523
I. Martins, Y. Wang, M. Michaud, Y. Ma, A.Q. Sukkurwala, S. Shen, O. Kepp, D. Metivier, L. Galluzzi, J.L. Perfettini, L. Zitvogel, G. Kroemer, Molecular mechanisms of ATP secretion during immunogenic cell death. Cell Death Differ 21, 79–91 (2014). https://doi.org/10.1038/cdd.2013.75
A.D. Garg, D.V. Krysko, P. Vandenabeele, P. Agostinis, Extracellular ATP and P(2)X(7) receptor exert context-specific immunogenic effects after immunogenic cancer cell death. Cell Death Dis 7, e2097 (2016). https://doi.org/10.1038/cddis.2015.411
P. Scaffidi, T. Misteli, M.E. Bianchi, Release of chromatin protein HMGB1 by necrotic cells triggers inflammation. Nature 418, 191–195 (2002). https://doi.org/10.1038/nature00858
A.D. Garg, L. Galluzzi, L. Apetoh, T. Baert, R.B. Birge, J.M.B.-S. Pedro, K. Breckpot, D. Brough, R. Chaurio, M. Cirone, A. Coosemans, P.G. Coulie, D. De Ruysscher, L. Dini, P. de Witte, A.M. Dudek-Peric, A. Faggioni, J. Fucikova, U.S. Gaipl, J. Golab, M.L. Gougeon, M.R. Hamblin, A. Hemminki, M. Herrmann, J.W. Hodge, O. Kepp, G. Kroemer, D.V. Krysko, W.G. Land, F. Madeo, A.A. Manfredi, S.R. Mattarollo, C. Maueroder, N. Merendino, G. Multhoff, T. Pabst, J.E. Ricci, C. Riganti, E. Romano, N. Rufo, M.J. Smyth, J. Sonnemann, R. Spisek, J. Stagg, E. Vacchelli, P. Vandenabeele, L. Vandenberk, B.J. Van den Eynde, S. Van Gool, F. Velotti, L. Zitvogel, P. Agostinis, Molecular and Translational Classifications of DAMPs in Immunogenic Cell Death. Front Immunol 6, 588 (2015). https://doi.org/10.3389/fimmu.2015.00588
A.D. Garg, A.M. Dudek-Peric, E. Romano, P. Agostinis, Immunogenic cell death. Int J Dev Biol 59, 131–140 (2015). https://doi.org/10.1387/ijdb.150061pa
A. Almanza, A. Carlesso, C. Chintha, S. Creedican, D. Doultsinos, B. Leuzzi, A. Luis, N. McCarthy, L. Montibeller, S. More, A. Papaioannou, F. Puschel, M.L. Sassano, J. Skoko, P. Agostinis, J. de Belleroche, L.A. Eriksson, S. Fulda, A.M. Gorman, S. Healy, A. Kozlov, C. Munoz-Pinedo, M. Rehm, E. Chevet, A. Samali, Endoplasmic reticulum stress signalling - from basic mechanisms to clinical applications. FEBS J 286, 241–278 (2019). https://doi.org/10.1111/febs.14608
C. Hetz, F.R. Papa, The unfolded protein response and cell fate control. Mol Cell 69, 169–181 (2018). https://doi.org/10.1016/j.molcel.2017.06.017
L. Bezu, A. Sauvat, J. Humeau, L.C. Gomes-da-Silva, K. Iribarren, S. Forveille, P. Garcia, L. Zhao, P. Liu, L. Zitvogel, L. Senovilla, O. Kepp, G. Kroemer, eIF2alpha phosphorylation is pathognomonic for immunogenic cell death. Cell Death Differ 25, 1375–1393 (2018). https://doi.org/10.1038/s41418-017-0044-9
T. Panaretakis, O. Kepp, U. Brockmeier, A. Tesniere, A.C. Bjorklund, D.C. Chapman, M. Durchschlag, N. Joza, G. Pierron, P. van Endert, J. Yuan, L. Zitvogel, F. Madeo, D.B. Williams, G. Kroemer, Mechanisms of pre-apoptotic calreticulin exposure in immunogenic cell death. EMBO J 28, 578–590 (2009). https://doi.org/10.1038/emboj.2009.1
P. Giglio, M. Gagliardi, N. Tumino, F. Antunes, S. Smaili, D. Cotella, C. Santoro, R. Bernardini, M. Mattei, M. Piacentini, M. Corazzari, PKR and GCN2 stress kinases promote an ER stress-independent eIF2alpha phosphorylation responsible for calreticulin exposure in melanoma cells. Oncoimmunology 7, e1466765 (2018). https://doi.org/10.1080/2162402X.2018.1466765
Global Burden of Disease Cancer Collaboration, C. Fitzmaurice, D. Abate, N. Abbasi, H. Abbastabar, F. Abd-Allah, O. Abdel-Rahman, A. Abdelalim, A. Abdoli, I. Abdollahpour, A.S.M. Abdulle, N.D. Abebe, H.N. Abraha, L.J. Abu-Raddad, A. Abualhasan, I.A. Adedeji, S.M. Advani, M. Afarideh, M. Afshari, M. Aghaali, D. Agius, S. Agrawal, A. Ahmadi, E. Ahmadian, E. Ahmadpour, M.B. Ahmed, M.E. Akbari, T. Akinyemiju, Z. Al-Aly, A.M. AlAbdulKader, F. Alahdab, T. Alam, G.M. Alamene, B.T.T. Alemnew, K.A. Alene, C. Alinia, V. Alipour, S.M. Aljunid, F.A. Bakeshei, M.A.H. Almadi, A. Almasi-Hashiani, U. Alsharif, S. Alsowaidi, N. Alvis-Guzman, E. Amini, S. Amini, Y.A. Amoako, Z. Anbari, N.H. Anber, C.L. Andrei, M. Anjomshoa, F. Ansari, A. Ansariadi, S.C.Y. Appiah, M. Arab-Zozani, J. Arabloo, Z. Arefi, O. Aremu, H.A. Areri, A. Artaman, H. Asayesh, E.T. Asfaw, A.F. Ashagre, R. Assadi, B. Ataeinia, H.T. Atalay, Z. Ataro, S. Atique, M. Ausloos, L. Avila-Burgos, E. Avokpaho, A. Awasthi, N. Awoke, B.P. Ayala Quintanilla, M.A. Ayanore, H.T. Ayele, E. Babaee, U. Bacha, A. Badawi, M. Bagherzadeh, E. Bagli, S. Balakrishnan, A. Balouchi, T.W. Barnighausen, R.J. Battista, M. Behzadifar, M. Behzadifar, B.B. Bekele, Y.B. Belay, Y.M. Belayneh, K.K.S. Berfield, A. Berhane, E. Bernabe, M. Beuran, N. Bhakta, K. Bhattacharyya, B. Biadgo, A. Bijani, M.S. Bin Sayeed, C. Birungi, C. Bisignano, H. Bitew, T. Bjorge, A. Bleyer, K.A. Bogale, H.A. Bojia, A.M. Borzi, C. Bosetti, I.R. Bou-Orm, H. Brenner, J.D. Brewer, A.N. Briko, N.I. Briko, M.T. Bustamante-Teixeira, Z.A. Butt, G. Carreras, J.J. Carrero, F. Carvalho, C. Castro, F. Castro, F. Catala-Lopez, E. Cerin, Y. Chaiah, W.F. Chanie, V.K. Chattu, P. Chaturvedi, N.S. Chauhan, M. Chehrazi, P.P. Chiang, T.Y. Chichiabellu, O.G. Chido-Amajuoyi, O. Chimed-Ochir, J.J. Choi, D.J. Christopher, D.T. Chu, M.M. Constantin, V.M. Costa, E. Crocetti, C.S. Crowe, M.P. Curado, S.M.A. Dahlawi, G. Damiani, A.H. Darwish, A. Daryani, J. das Neves, F.M. Demeke, A.B. Demis, B.W. Demissie, G.T. Demoz, E. Denova-Gutierrez, A. Derakhshani, K.S. Deribe, R. Desai, B.B. Desalegn, M. Desta, S. Dey, S.D. Dharmaratne, M. Dhimal, D. Diaz, M.T.T. Dinberu, S. Djalalinia, D.T. Doku, T.M. Drake, M. Dubey, E. Dubljanin, E.E. Duken, H. Ebrahimi, A. Effiong, A. Eftekhari, I. El Sayed, M.E.S. Zaki, S.I. El-Jaafary, Z. El-Khatib, D.A. Elemineh, H. Elkout, R.G. Ellenbogen, A. Elsharkawy, M.H. Emamian, D.A. Endalew, A.Y. Endries, B. Eshrati, I. Fadhil, V. Fallah, M. Faramarzi, M.A. Farhangi, A. Farioli, F. Farzadfar, N. Fentahun, E. Fernandes, G.T. Feyissa, I. Filip, F. Fischer, J.L. Fisher, L.M. Force, M. Foroutan, M. Freitas, T. Fukumoto, N.D. Futran, S. Gallus, F.G. Gankpe, R.T. Gayesa, T.T. Gebrehiwot, G.G. Gebremeskel, G.A. Gedefaw, B.K. Gelaw, B. Geta, S. Getachew, K.E. Gezae, M. Ghafourifard, A. Ghajar, A. Ghashghaee, A. Gholamian, P.S. Gill, T.T.G. Ginindza, A. Girmay, M. Gizaw, R.S. Gomez, S.V. Gopalani, G. Gorini, B.N.G. Goulart, A. Grada, M. Ribeiro Guerra, A.L.S. Guimaraes, P.C. Gupta, R. Gupta, K. Hadkhale, A. Haj-Mirzaian, A. Haj-Mirzaian, R.R. Hamadeh, S. Hamidi, L.K. Hanfore, J.M. Haro, M. Hasankhani, A. Hasanzadeh, H.Y. Hassen, R.J. Hay, S.I. Hay, A. Henok, N.J. Henry, C. Herteliu, H.D. Hidru, C.L. Hoang, M.K. Hole, P. Hoogar, N. Horita, H.D. Hosgood, M. Hosseini, M. Hosseinzadeh, M. Hostiuc, S. Hostiuc, M. Househ, M.M. Hussen, B. Ileanu, M.D. Ilic, K. Innos, S.S.N. Irvani, K.R. Iseh, S.M.S. Islam, F. Islami, N. Jafari Balalami, M. Jafarinia, L. Jahangiry, M.A. Jahani, N. Jahanmehr, M. Jakovljevic, S.L. James, M. Javanbakht, S. Jayaraman, S.H. Jee, E. Jenabi, R.P. Jha, J.B. Jonas, J. Jonnagaddala, T. Joo, S.B. Jungari, M. Jurisson, A. Kabir, F. Kamangar, A. Karch, N. Karimi, A. Karimian, A. Kasaeian, G.G. Kasahun, B. Kassa, T.D. Kassa, M.W. Kassaw, A. Kaul, P.N. Keiyoro, A.G. Kelbore, A.A. Kerbo, Y.S. Khader, M. Khalilarjmandi, E.A. Khan, G. Khan, Y.H. Khang, K. Khatab, A. Khater, M. Khayamzadeh, M. Khazaee-Pool, S. Khazaei, A.T. Khoja, M.H. Khosravi, J. Khubchandani, N. Kianipour, D. Kim, Y.J. Kim, A. Kisa, S. Kisa, K. Kissimova-Skarbek, H. Komaki, A. Koyanagi, K.J. Krohn, B.K. Bicer, N. Kugbey, V. Kumar, D. Kuupiel, C. La Vecchia, D.P. Lad, E.A. Lake, A.M. Lakew, D.K. Lal, F.H. Lami, Q. Lan, S. Lasrado, P. Lauriola, J.V. Lazarus, J. Leigh, C.T. Leshargie, Y. Liao, M.A. Limenih, S. Listl, A.D. Lopez, P.D. Lopukhov, R. Lunevicius, M. Madadin, S. Magdeldin, H.M.A. El Razek, A. Majeed, A. Maleki, R. Malekzadeh, A. Manafi, N. Manafi, W.A. Manamo, M. Mansourian, M.A. Mansournia, L.G. Mantovani, S. Maroufizadeh, S.M.S. Martini, T.P. Mashamba-Thompson, B.B. Massenburg, M.T. Maswabi, M.R. Mathur, C. McAlinden, M. McKee, H.A.A. Meheretu, R. Mehrotra, V. Mehta, T. Meier, Y.A. Melaku, G.G. Meles, H.G. Meles, A. Melese, M. Melku, P.T.N. Memiah, W. Mendoza, R.G. Menezes, S. Merat, T.J. Meretoja, T. Mestrovic, B. Miazgowski, T. Miazgowski, K.M.M. Mihretie, T.R. Miller, E.J. Mills, S.M. Mir, H. Mirzaei, H.R. Mirzaei, R. Mishra, B. Moazen, D.K. Mohammad, K.A. Mohammad, Y. Mohammad, A.M. Darwesh, A. Mohammadbeigi, H. Mohammadi, M. Mohammadi, M. Mohammadian, A. Mohammadian-Hafshejani, M. Mohammadoo-Khorasani, R. Mohammadpourhodki, A.S. Mohammed, J.A. Mohammed, S. Mohammed, F. Mohebi, A.H. Mokdad, L. Monasta, Y. Moodley, M. Moosazadeh, M. Moossavi, G. Moradi, M. Moradi-Joo, M. Moradi-Lakeh, F. Moradpour, L. Morawska, J. Morgado-da-Costa, N. Morisaki, S.D. Morrison, A. Mosapour, S.M. Mousavi, A.A. Muche, O.S.S. Muhammed, J. Musa, A.R. Nabhan, M. Naderi, A.J. Nagarajan, G. Nagel, A. Nahvijou, G. Naik, F. Najafi, L. Naldi, H.S. Nam, N. Nasiri, J. Nazari, I. Negoi, S. Neupane, P.A. Newcomb, H.A. Nggada, J.W. Ngunjiri, C.T. Nguyen, L. Nikniaz, D.N.A. Ningrum, Y.L. Nirayo, M.R. Nixon, C.A. Nnaji, M. Nojomi, S. Nosratnejad, M.N. Shiadeh, M.S. Obsa, R. Ofori-Asenso, F.A. Ogbo, I.H. Oh, A.T. Olagunju, T.O. Olagunju, M.M. Oluwasanu, A.E. Omonisi, O.E. Onwujekwe, A.M. Oommen, E. Oren, D.D.V. Ortega-Altamirano, E. Ota, S.S. Otstavnov, M.O. Owolabi, A. M. P, J.R. Padubidri, S. Pakhale, A.H. Pakpour, A. Pana, E.K. Park, H. Parsian, T. Pashaei, S. Patel, S.T. Patil, A. Pennini, D.M. Pereira, C. Piccinelli, J.D. Pillay, M. Pirestani, F. Pishgar, M.J. Postma, H. Pourjafar, F. Pourmalek, A. Pourshams, S. Prakash, N. Prasad, M. Qorbani, M. Rabiee, N. Rabiee, A. Radfar, A. Rafiei, F. Rahim, M. Rahimi, M.A. Rahman, F. Rajati, S.M. Rana, S. Raoofi, G.K. Rath, D.L. Rawaf, S. Rawaf, R.C. Reiner, A.M.N. Renzaho, N. Rezaei, A. Rezapour, A.I. Ribeiro, D. Ribeiro, L. Ronfani, E.M. Roro, G. Roshandel, A. Rostami, R.S. Saad, P. Sabbagh, S. Sabour, B. Saddik, S. Safiri, A. Sahebkar, M.R. Salahshoor, F. Salehi, H. Salem, M.R. Salem, H. Salimzadeh, J.A. Salomon, A.M. Samy, J. Sanabria, M.M. Santric Milicevic, B. Sartorius, A. Sarveazad, B. Sathian, M. Satpathy, M. Savic, M. Sawhney, M. Sayyah, I.J.C. Schneider, B. Schottker, M. Sekerija, S.G. Sepanlou, M. Sepehrimanesh, S. Seyedmousavi, F. Shaahmadi, H. Shabaninejad, M. Shahbaz, M.A. Shaikh, A. Shamshirian, M. Shamsizadeh, H. Sharafi, Z. Sharafi, M. Sharif, A. Sharifi, H. Sharifi, R. Sharma, A. Sheikh, R. Shirkoohi, S.R. Shukla, S. Si, S. Siabani, D.A.S. Silva, D.G.A. Silveira, A. Singh, J.A. Singh, S. Sisay, F. Sitas, E. Sobngwi, M. Soofi, J.B. Soriano, V. Stathopoulou, M.B. Sufiyan, R. Tabares-Seisdedos, T. Tabuchi, K. Takahashi, O.R. Tamtaji, M.R. Tarawneh, S.G. Tassew, P. Taymoori, A. Tehrani-Banihashemi, M.H. Temsah, O. Temsah, B.E. Tesfay, F.H. Tesfay, M.Y. Teshale, G.A. Tessema, S. Thapa, K.G. Tlaye, R. Topor-Madry, M.R. Tovani-Palone, E. Traini, B.X. Tran, K.B. Tran, A.G. Tsadik, I. Ullah, O.A. Uthman, M. Vacante, M. Vaezi, P. Varona Perez, Y. Veisani, S. Vidale, F.S. Violante, V. Vlassov, S.E. Vollset, T. Vos, K. Vosoughi, G.T. Vu, I.S. Vujcic, H. Wabinga, T.M. Wachamo, F.S. Wagnew, Y. Waheed, F. Weldegebreal, G.T. Weldesamuel, T. Wijeratne, D.Z. Wondafrash, T.E. Wonde, A.B. Wondmieneh, H.M. Workie, R. Yadav, A. Yadegar, A. Yadollahpour, M. Yaseri, V. Yazdi-Feyzabadi, A. Yeshaneh, M.A. Yimam, E.M. Yimer, E. Yisma, N. Yonemoto, M.Z. Younis, B. Yousefi, M. Yousefifard, C. Yu, E. Zabeh, V. Zadnik, T.Z. Moghadam, Z. Zaidi, M. Zamani, H. Zandian, A. Zangeneh, L. Zaki, K. Zendehdel, Z.M. Zenebe, T.A. Zewale, A. Ziapour, S. Zodpey and C.J.L. Murray, Global, Regional, and National Cancer Incidence, Mortality, Years of Life Lost, Years Lived With Disability, and Disability-Adjusted Life-Years for 29 Cancer Groups, 1990 to 2017: A Systematic Analysis for the Global Burden of Disease Study. JAMA Oncol 5, 1749–1768 (2019). doi: https://doi.org/10.1001/jamaoncol.2019.2996
J. Zhou, T. Yang, L. Liu, B. Lu, Chemotherapy oxaliplatin sensitizes prostate cancer to immune checkpoint blockade therapies via stimulating tumor immunogenicity. Mol Med Rep 16, 2868–2874 (2017). https://doi.org/10.3892/mmr.2017.6908
J. Kapuscinski, Z. Darzynkiewicz, Interactions of antitumor agents Ametantrone and Mitoxantrone (Novatrone) with double-stranded DNA. Biochem Pharmacol 34, 4203–4213 (1985). https://doi.org/10.1016/0006-2952(85)90275-8
J.S. de Bono, S. Oudard, M. Ozguroglu, S. Hansen, J.P. Machiels, I. Kocak, G. Gravis, I. Bodrogi, M.J. Mackenzie, L. Shen, M. Roessner, S. Gupta, A.O. Sartor, T. Investigators, Prednisone plus cabazitaxel or mitoxantrone for metastatic castration-resistant prostate cancer progressing after docetaxel treatment: A randomised open-label trial. Lancet 376, 1147–1154 (2010). https://doi.org/10.1016/S0140-6736(10)61389-X
G. Gravis, Systemic treatment for metastatic prostate cancer. Asian J Urol 6, 162–168 (2019). https://doi.org/10.1016/j.ajur.2019.02.002
J. Qin, N.M. Kunda, G. Qiao, K. Tulla, B.S. Prabhakar, A.V. Maker, Vaccination with Mitoxantrone-treated primary Colon Cancer cells enhances tumor-infiltrating lymphocytes and clinical responses in colorectal liver metastases. J Surg Res 233, 57–64 (2019). https://doi.org/10.1016/j.jss.2018.07.068
T. Colangelo, G. Polcaro, P. Ziccardi, L. Muccillo, M. Galgani, B. Pucci, M.R. Milone, A. Budillon, M. Santopaolo, G. Mazzoccoli, G. Matarese, L. Sabatino, V. Colantuoni, The miR-27a-calreticulin axis affects drug-induced immunogenic cell death in human colorectal cancer cells. Cell Death Dis 7, e2108 (2016). https://doi.org/10.1038/cddis.2016.29
C. Li, C. He, Y. Xu, H. Xu, Y. Tang, H. Chavan, S. Duan, A. Artigues, M.L. Forrest, P. Krishnamurthy, S. Han, J.M. Holzbeierlein, B. Li, Alternol eliminates excessive ATP production by disturbing Krebs cycle in prostate cancer. Prostate 79, 628–639 (2019). https://doi.org/10.1002/pros.23767
C. Li, W. Jiang, Q. Hu, L.C. Li, L. Dong, R. Chen, Y. Zhang, Y. Tang, J.B. Thrasher, C.B. Liu, B. Li, Enhancing DPYSL3 gene expression via a promoter-targeted small activating RNA approach suppresses cancer cell motility and metastasis, Oncotarget 7, 22893–22910 (2016). https://doi.org/10.18632/oncotarget.8290
K. Berns, E.M. Hijmans, J. Mullenders, T.R. Brummelkamp, A. Velds, M. Heimerikx, R.M. Kerkhoven, M. Madiredjo, W. Nijkamp, B. Weigelt, R. Agami, W. Ge, G. Cavet, P.S. Linsley, R.L. Beijersbergen, R. Bernards, A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428, 431–437 (2004). https://doi.org/10.1038/nature02371
M. Soto, J.A. Raaijmakers, B. Bakker, D.C.J. Spierings, P.M. Lansdorp, F. Foijer, R.H. Medema, p53 prohibits propagation of chromosome segregation errors that produce structural aneuploidies. Cell Rep 19, 2423–2431 (2017). https://doi.org/10.1016/j.celrep.2017.05.055
C. Li, H. Xu, L. Xiao, H. Zhu, G. Zhang, W. Wei, K. Li, X. Cao, D. Shen, J. Holzbeierlein, B. Li, CRMP4a suppresses cell motility by sequestering RhoA activity in prostate cancer cells. Cancer Biol Ther 19, 1193–1203 (2018). https://doi.org/10.1080/15384047.2018.1491507
C. Pozzi, A. Cuomo, I. Spadoni, E. Magni, A. Silvola, A. Conte, S. Sigismund, P.S. Ravenda, T. Bonaldi, M.G. Zampino, C. Cancelliere, P.P. Di Fiore, A. Bardelli, G. Penna, M. Rescigno, The EGFR-specific antibody cetuximab combined with chemotherapy triggers immunogenic cell death. Nat Med 22, 624–631 (2016). https://doi.org/10.1038/nm.4078
G.H. Nam, E.J. Lee, Y.K. Kim, Y. Hong, Y. Choi, M.J. Ryu, J. Woo, Y. Cho, D.J. Ahn, Y. Yang, I.C. Kwon, S.Y. Park, I.S. Kim, Combined rho-kinase inhibition and immunogenic cell death triggers and propagates immunity against cancer. Nat Commun 9, 2165 (2018). https://doi.org/10.1038/s41467-018-04607-9
A. Khan, O. Fornes, A. Stigliani, M. Gheorghe, J.A. Castro-Mondragon, R. van der Lee, A. Bessy, J. Cheneby, S.R. Kulkarni, G. Tan, D. Baranasic, D.J. Arenillas, A. Sandelin, K. Vandepoele, B. Lenhard, B. Ballester, W.W. Wasserman, F. Parcy, A. Mathelier, JASPAR 2018: Update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res 46, D1284 (2018). https://doi.org/10.1093/nar/gkx1188
D. Schmidt, M.D. Wilson, C. Spyrou, G.D. Brown, J. Hadfield, D.T. Odom, ChIP-seq: Using high-throughput sequencing to discover protein-DNA interactions. Methods 48, 240–248 (2009). https://doi.org/10.1016/j.ymeth.2009.03.001
C.P. Bergstrom, B. Ruffell, C.M. Ho, C.S. Higano, W.J. Ellis, M. Garzotto, T.M. Beer, J.N. Graff, Docetaxel and mitoxantrone before radical prostatectomy in men with high-risk prostate cancer: 10-year follow-up and immune correlates. Anti-Cancer Drugs 28, 120–126 (2017). https://doi.org/10.1097/CAD.0000000000000438
C. Sanchez, P. Mendoza, H.R. Contreras, J. Vergara, J.A. McCubrey, C. Huidobro, E.A. Castellon, Expression of multidrug resistance proteins in prostate cancer is related with cell sensitivity to chemotherapeutic drugs. Prostate 69, 1448–1459 (2009). https://doi.org/10.1002/pros.20991
L. Bezu, L.C. Gomes-de-Silva, H. Dewitte, K. Breckpot, J. Fucikova, R. Spisek, L. Galluzzi, O. Kepp, G. Kroemer, Combinatorial strategies for the induction of immunogenic cell death. Front Immunol 6, 187 (2015). https://doi.org/10.3389/fimmu.2015.00187
R.C. Wek, Role of eIF2alpha kinases in translational control and adaptation to cellular stress. Cold Spring Harb Perspect Biol 10 (2018). https://doi.org/10.1101/cshperspect.a032870
A. Rashidi, J. Miska, C. Lee-Chang, D. Kanojia, W.K. Panek, A. Lopez-Rosas, P. Zhang, Y. Han, T. Xiao, K.C. Pituch, J.W. Kim, M. Talebian, J. Fares, M.S. Lesniak, GCN2 is essential for CD8(+) T cell survival and function in murine models of malignant glioma. Cancer Immunol Immunother 69, 81–94 (2020). https://doi.org/10.1007/s00262-019-02441-6
P.R. Romano, M.T. Garcia-Barrio, X. Zhang, Q. Wang, D.R. Taylor, F. Zhang, C. Herring, M.B. Mathews, J. Qin, A.G. Hinnebusch, Autophosphorylation in the activation loop is required for full kinase activity in vivo of human and yeast eukaryotic initiation factor 2alpha kinases PKR and GCN2. Mol Cell Biol 18, 2282–2297 (1998). https://doi.org/10.1128/mcb.18.4.2282
H.P. Harding, Y. Zhang, A. Bertolotti, H. Zeng, D. Ron, Perk is essential for translational regulation and cell survival during the unfolded protein response. Mol Cell 5, 897–904 (2000). https://doi.org/10.1016/s1097-2765(00)80330-5
G. Guo, M. Yu, W. Xiao, E. Celis, Y. Cui, Local activation of p53 in the tumor microenvironment overcomes immune suppression and enhances antitumor immunity. Cancer Res 77, 2292–2305 (2017). https://doi.org/10.1158/0008-5472.CAN-16-2832
S. Bottger, E. Jerszyk, B. Low, C. Walker, Genotoxic stress-induced expression of p53 and apoptosis in leukemic clam hemocytes with cytoplasmically sequestered p53. Cancer Res 68, 777–782 (2008). https://doi.org/10.1158/0008-5472.CAN-06-0968
A.G. Carroll, H.J. Voeller, L. Sugars, E.P. Gelmann, p53 oncogene mutations in three human prostate cancer cell lines. Prostate 23, 123–134 (1993). https://doi.org/10.1002/pros.2990230206
M. Michaud, I. Martins, A.Q. Sukkurwala, S. Adjemian, Y. Ma, P. Pellegatti, S. Shen, O. Kepp, M. Scoazec, G. Mignot, S. Rello-Varona, M. Tailler, L. Menger, E. Vacchelli, L. Galluzzi, F. Ghiringhelli, F. di Virgilio, L. Zitvogel, G. Kroemer, Autophagy-dependent anticancer immune responses induced by chemotherapeutic agents in mice. Science 334, 1573–1577 (2011). https://doi.org/10.1126/science.1208347
J.S. Park, F. Gamboni-Robertson, Q. He, D. Svetkauskaite, J.Y. Kim, D. Strassheim, J.W. Sohn, S. Yamada, I. Maruyama, A. Banerjee, A. Ishizaka, E. Abraham, High mobility group box 1 protein interacts with multiple toll-like receptors. Am J Physiol Cell Physiol 290, C917–C924 (2006). https://doi.org/10.1152/ajpcell.00401.2005
I. Martins, A. Tesniere, O. Kepp, M. Michaud, F. Schlemmer, L. Senovilla, C. Seror, D. Metivier, J.L. Perfettini, L. Zitvogel, G. Kroemer, Chemotherapy induces ATP release from tumor cells. Cell Cycle 8, 3723–3728 (2009). https://doi.org/10.4161/cc.8.22.10026
A.D. Garg, D.V. Krysko, T. Verfaillie, A. Kaczmarek, G.B. Ferreira, T. Marysael, N. Rubio, M. Firczuk, C. Mathieu, A.J. Roebroek, W. Annaert, J. Golab, P. de Witte, P. Vandenabeele, P. Agostinis, A novel pathway combining calreticulin exposure and ATP secretion in immunogenic cancer cell death. EMBO J 31, 1062–1079 (2012). https://doi.org/10.1038/emboj.2011.497
L. Bezu, A. Sauvat, J. Humeau, M. Leduc, O. Kepp, G. Kroemer, eIF2alpha phosphorylation: A hallmark of immunogenic cell death. Oncoimmunology 7, e1431089 (2018). https://doi.org/10.1080/2162402X.2018.1431089
J. Han, S.H. Back, J. Hur, Y.H. Lin, R. Gildersleeve, J. Shan, C.L. Yuan, D. Krokowski, S. Wang, M. Hatzoglou, M.S. Kilberg, M.A. Sartor, R.J. Kaufman, ER-stress-induced transcriptional regulation increases protein synthesis leading to cell death. Nat Cell Biol 15, 481–490 (2013). https://doi.org/10.1038/ncb2738
S.M. Reed, D.E. Quelle, p53 acetylation: Regulation and consequences. Cancers (Basel) 7, 30–69 (2014). https://doi.org/10.3390/cancers7010030
M.F. Lavin, N. Gueven, The complexity of p53 stabilization and activation. Cell Death Differ 13, 941–950 (2006). https://doi.org/10.1038/sj.cdd.4401925
Y. Xia, R.C. Padre, T.H. De Mendoza, V. Bottero, V.B. Tergaonkar, I.M. Verma, Phosphorylation of p53 by IkappaB kinase 2 promotes its degradation by beta-TrCP. Proc Natl Acad Sci U S A 106, 2629–2634 (2009). https://doi.org/10.1073/pnas.0812256106
G.S. Ivanov, T. Ivanova, J. Kurash, A. Ivanov, S. Chuikov, F. Gizatullin, E.M. Herrera-Medina, F. Rauscher 3rd, D. Reinberg, N.A. Barlev, Methylation-acetylation interplay activates p53 in response to DNA damage. Mol Cell Biol 27, 6756–6769 (2007). https://doi.org/10.1128/MCB.00460-07
J. Huang, L. Perez-Burgos, B.J. Placek, R. Sengupta, M. Richter, J.A. Dorsey, S. Kubicek, S. Opravil, T. Jenuwein, S.L. Berger, Repression of p53 activity by Smyd2-mediated methylation. Nature 444, 629–632 (2006). https://doi.org/10.1038/nature05287
S.F. Slovin, Immunotherapy for castration-resistant prostate cancer: Has its time arrived? Expert Opin Biol Ther 20, 481–487 (2020). https://doi.org/10.1080/14712598.2020.1735345
I. Vanmeerbeek, J. Sprooten, D. De Ruysscher, S. Tejpar, P. Vandenberghe, J. Fucikova, R. Spisek, L. Zitvogel, G. Kroemer, L. Galluzzi, A.D. Garg, Trial watch: Chemotherapy-induced immunogenic cell death in immuno-oncology. Oncoimmunology 9, 1703449 (2020). https://doi.org/10.1080/2162402X.2019.1703449
J.W. Hodge, C.T. Garnett, B. Farsaci, C. Palena, K.Y. Tsang, S. Ferrone, S.R. Gameiro, Chemotherapy-induced immunogenic modulation of tumor cells enhances killing by cytotoxic T lymphocytes and is distinct from immunogenic cell death. Int J Cancer 133, 624–636 (2013). https://doi.org/10.1002/ijc.28070
Acknowledgments
This work was partially supported by the Shandong Provincial Natural Science Foundation of China (ZR2016HL25 and ZR2016CQ01), NSFC grant 81501770 and an internal grant from Jining Medical University (JYHL2018ZD02).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declare no conflicts of interest.
Ethical approval
All protocols and procedures in this study were approved by the institutional review committee of Jining Medical University for animal welfare.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 34 kb)
ESM 2
MTX induces apoptosis in prostate cancer cells. a. LNCaP and 22RV1 cells were treated with the vehicle or MTX (1 μM) for 20 h and then harvested for apoptosis analysis using Annexin-V/PI staining approach coupled with flow cytometry. b. Quantitative data of apoptosis were presented from three independent experiments. Early apoptotic was defined as Annexin positive population, later apoptotic as Annexin/PI positive and necrotic death as PI positive only. The asterisks indicate a significant difference compared to the vehicle control (Student t-test, *p < 0.05, **p < 0.01, ***p < 0.001). c. LNCaP and 22RV1 cells were treated with the vehicle or MTX (1 μM) for 20 h and then whole cell proteins were extracted for western blot assay with PARP antibodies. Actin blot served as protein loading control. Data were representative of two independent experiments. (PNG 70 kb)
ESM 3
Effect of PERK or GCN2 Knockdown on MTX-induced apoptosis. LNCaP cells transfected with the shRNAs as indicated were treated with MTX (1 μM) or the vehicle for 20 h, followed by Annexin-V/PI staining, as described earlier. Representative plots of flow cytometry from three independent experiments were shown in panel a. Quantitative data of apoptotic population (%) were summarized in panel b. The asterisks indicate a significant difference compared to the shScr control (Student t-test, *p < 0.05, ***p < 0.001). (PNG 64 kb)
Rights and permissions
About this article
Cite this article
Li, C., Sun, H., Wei, W. et al. Mitoxantrone triggers immunogenic prostate cancer cell death via p53-dependent PERK expression. Cell Oncol. 43, 1099–1116 (2020). https://doi.org/10.1007/s13402-020-00544-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-020-00544-2