Abstract
The role of the clock gene period (per) in photoperiodism was investigated in the jewel wasp Nasonia vitripennis (Walker) (Hymenoptera: Pteromalidae). Typically, females reared under long days lay eggs that develop to adulthood without developmental interruption, whereas those reared under short days lay eggs that enter larval diapause. We found that RNA interference (RNAi) of per disrupts the photoperiodic response; RNAi wasps failed to produce diapause-destined eggs in response to short days. This indicates that per is essential for the photoperiodic response in this species. Exposure to low temperature swiftly affects maternal physiology and prompts N. vitripennis females to lay diapause-destined eggs. Interestingly, per RNAi females also laid diapause-destined eggs in response to the low temperature treatment in the current study. This finding indicates that per RNAi did not disrupt the maternal physiology that was directly involved in production of diapause-destined eggs, but it did affect photoperiodic perception. We also investigated the expression patterns of several clock genes, per, mammalian-type cryptochrome (cry-m), cycle (cyc), PAR-domain protein 1 (Pdp1), and clockwork orange (cwo), to determine whether their expression patterns were affected by different photoperiods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Photoperiodism is a seasonal timing system that enables organisms to anticipate upcoming seasons and uses day length (photoperiod) as a cue to coordinate their development, reproduction, and physiology to seasonal changes in biotic and abiotic factors in the environment (Nelson et al. 2009). Using this system, organisms are able to concentrate their reproductive efforts into seasons that are favorable for the development and survival of their offspring. Physiological mechanisms underpinning photoperiodism are considered to comprise a sequence of several events, and one of them is the photoperiodic time measurement system, which assesses the length of day or night. Although many studies focused on the physiological nature of this system, its molecular mechanism remains largely unknown (Dolezel 2015; Saunders 2014).
Bünning (1936) first proposed the involvement of a circadian clock in photoperiodic time measurement (Bünning’s hypothesis). Later, several photoperiodic time measurement models were proposed (Vaz Nunes and Saunders 1999) based on the hypothesis, and one of the most influential of these is the internal coincidence model (Tyshchenko 1966). This model proposes two circadian oscillators that are entrained by dawn and dusk, respectively, and their internal phase relationships change with the length of the photophase (Tyshchenko 1966). These two oscillators independently activate particular downstream cascades, called ‘active phases’. Long-day responses are induced when they overlap (see Danilevskii et al. 1970). This model is still highly conceptual, but the photoperiodic induction of diapause in the jewel wasp Nasonia vitripennis (Walker) (Hymenoptera: Pteromalidae) fits well with this model (Saunders 1974).
Nasonia vitripennis is a cosmopolitan parasitic wasp that stings and lays eggs upon the pupae of various flies, including blow flies, flesh flies, and house flies (Saunders et al. 1970). When adult females of N. vitripennis are exposed to long-day conditions, they lay eggs that develop to adulthood without developmental interruption. In contrast, when adult females are exposed to short-day conditions, they lay eggs that enter diapause at the final (4th) instar larval stage (Saunders 1965, 1966). Larvae themselves cannot change their developmental destination in response to photoperiods (Saunders 1966). Thus, mothers photoperiodically regulate the developmental fate of their offspring; i.e., there is a maternal effect on photoperiodism.
Although the molecular mechanisms of the circadian clock involved in insect photoperiodism have long been undetermined, one possible theory is that the circadian clock is established by period (per), mammalian-type cryptochrome (cry-m, also known as cry2), Clock (Clk) and cycle (cyc), so-called circadian clock genes, like a circadian clock governing circadian locomotor activity rhythms (Goto 2013). The circadian clock governing locomotor activity rhythm has been extensively studied in many insect species and is considered to be mainly composed of three loops (reviewed by Tomioka and Matsumoto 2015). In the first loop, protein products of Clk and cyc genes form heterodimers and activate the transcription of per, timeless (tim), and cry-m during the late day or early night. In the middle of the night, PER and TIM proteins form heterodimers and CRY-m enters the nucleus, with the help of PER, to suppress its own transcription by inactivating the transcriptional activity of CLK/CYC. This negative feedback produces the rhythmic expression of per, tim, and cry-m. CLK/CYC also activates transcription of vrille (vri) and PAR-domain protein 1ε (Pdp1ε). The VRI protein accumulates and represses the transcription of Clk. PDP1ε accumulates later than VRI and activates Clk transcription during the day so that CLK accumulates during the day. CLK/CYC also activates the transcription of clockwork orange (cwo), which regulates the amplitudes of the oscillations of per and tim mRNA (reviewed by Hardin 2006; Tomioka and Matsumoto 2010, 2015).
Recently, Bertossa et al. (2014) revealed differential expression patterns of per and cry-m in N. vitripennis under long-day and short-day conditions. mRNA levels of both genes showed clear oscillation, and reached a maximum at the end of the light phase in long-day conditions and the middle of the dark phase under short-day conditions. Thus, the phase is advanced in the long-day conditions over the short-day conditions. Based on the results, Bertossa et al. (2014) emphasized the potential roles of per and cry-m in photoperiodic time measurement. Paolucci (2014) performed reciprocal crosses between two iso-female lines from northern and southern locations, respectively, in Europe. The northern line produced diapause offspring much more quickly in response to short-day exposure (referred to as an early ‘switching point’ from nondiapause breeding to diapause breeding), than the southern line. Subsequent microsatellite genotyping and QTL analyses revealed two genomic loci involved in the variation of the switching point, and the highest QTL peak of the first chromosome corresponds to the loci of per and cyc. In addition, Paolucci (2014) identified three alleles of the per (per S , per N1 , and per N2 ) gene in the northern and southern lines. Crossing experiments revealed that per N1 is associated with a stronger diapause response than per N2 and per S . Furthermore, a clear latitudinal cline was observed in allele frequency for the two per alleles per N1 and per S , which corresponds to the latitudinal cline in the switching point. These results suggest that the per gene is involved in the adaptive variation in photoperiodic diapause in N. vitripennis. However, its significance in photoperiodism has not been physiologically validated.
In the present study, we first investigated expression patterns of the clock genes per, cry-m, Clk, cwo, and Pdp1 in the heads of N. vitripennis, under long-day and short-day conditions, to determine whether their expression patterns are affected by photoperiods. Next, we investigated the role of per in the photoperiodic response of N. vitripennis using the RNAi technique.
Materials and methods
Insects
Nasonia vitripennis individuals were collected from the pupae of the flesh fly Sarcophaga similis Meade in Osaka, Japan (34.6°N, 135.5°E) in 2011. The wasp stock culture was maintained under long-day conditions (light:dark [LD] 16:8 h) at 20 ± 1 °C. Approximately 50–80 newly emerged adult wasps were transferred to a glass vial (30 mm in diameter, 100 mm in height) and fed on 10 % (w/v) sucrose solution. On days 12–14 (day 0 was defined as the day of emergence of the adult), approximately 60 females were transferred into individual vials, and two nondiapause pupae of S. similis were supplied as a host for each female. Wasps emerged from the fly pupae 20–22 days after parasitism.
RNAi
Total RNA was extracted from the whole bodies of adult females using TRIzol Reagent (Thermo Fisher Scientific, Waltham, MA, USA). cDNA was synthesized using M-MLV reverse transcriptase and random hexamers (Thermo Fisher Scientific) and was used to make templates for double-stranded (ds) per RNA. Two types of dsRNA were designed from distinct regions of the per gene, to validate suppression of per expression itself. ‘Region A’ corresponds to nucleotide positions 1709–2177 of the per transcript (GenBank Accession Number: XM_008211025.1), and ‘region B’ corresponds to positions 332–834 of the transcript. As a control, dsRNA of β-lactamase (bla), which provides bacteria with ampicillin resistance, was synthesized using a pTA2 vector (Toyobo, Osaka, Japan). Template fragments of dsRNA were amplified by PCR with Pwo DNA Polymerase (Roche, Basel, Switzerland). The following primers were used in the reactions: per 1F (5′-CAC CTG GAG CAG AAG AAA GG-3′), per 1F-T7 (5′-TAA TAC GAC TCA CTA TAG GCA CCT GGA GCA GAA GAA AGG-3′), per 1R (5′-TTG CCA GAG ATG GAT TTT CC-3′), and per 1R-T7 (5′-TAA TAC GAC TCA CTA TAG GTT GCC AGA GAT GGA TTT TCC-3′) for per region A; per 2F (5′-TTC AAC GCT ACC TCA TGT GC-3′), per 2F-T7 (5′-TAA TAC GAC TCA CTA TAG GTT CAA CGC TAC CTC ATG TGC-3′), per 2R (5′-TTG ACG AAG GAG GAC CAA TC-3′), and per 2R-T7 (5′-TAA TAC GAC TCA CTA TAG GTT GAC GAA GGA GGA CCA ATC-3′) for per region B; and, bla F (5′-TCG CCG CAT ACA CTA TTC TC-3′), bla F-T7 (5′-TAA TAC GAC TCA CTA TAG GTC GCC GCA TAC ACT ATT CTC-3′), bla R (5′-TAC GAT ACG GGA GGG CTT AC-3′), and bla R-T7 (5′-TAA TAC GAC TCA CTA TAG GTA CGA TAC GGG AGG GCT TAC-3′) for bla (the underlined sequences indicate the T7 RNA polymerase promoter). A T7 Ribomax Express RNAi System (Promega, Madison, WI, USA) was used to synthesize dsRNAs according to the manufacturer’s instructions. The dsRNAs were dissolved in nuclease-free water and stored at −20 °C until they were used. Injection into N. vitripennis pupae was performed according to Lynch and Desplan (2006). In brief, yellow pupae (aged 14 days after oviposition) were fixed on a glass slide using double-stick tape, and dsRNA was injected into them using a glass capillary (4748, World Precision Instrument, Sarasota, FL). per region A dsRNA and per region B dsRNA were injected at 1 and 5 μg/μl, respectively. We were unable to regulate the volume of injection precisely, but we usually injected approximately 0.1 μl of each solution into a single pupa. Talcum powder was used to keep the adult wasps from adhering to the tape.
Photoperiodic response
The photoperiodic response of N. vitripennis can be assessed using two methods (Saunders 1966, 1974). One is by determining the proportion of females that produce diapause offspring, and the other is by determining the number of days required to switch the diapause status of the progeny. In the present study, we adopted both methods; the photoperiodic response curve was obtained using the former method and the effect of per RNAi was evaluated using the latter method.
To determine the photoperiodic response of our wasp colony, newly emerged adults reared under LD 16:8 h were transferred to various photoperiodic conditions (LD 16:8, 15:9, 14:10, 13:11, and 12:12 h) and mass-cultured. On day 12, females were separated into individual vials and two pupae of S. similis were supplied to each wasp as a host. One day later, parasitized fly pupae were collected and maintained under constant darkness (DD). Each fly puparium was opened 14 days later and the numbers of wasp larvae (diapause) and wasp pupae (nondiapause) were counted. When a female laid a brood in which 50 % or more individuals entered diapause, the female was regarded as a ‘diapause producer’. In contrast, a female which laid a brood in which less than 50 % individuals entered diapause was regarded as a ‘nondiapause producer’. It should be noted that in most cases each female laid a brood comprising more than 90 % diapause or nondiapause offspring. It is also important to note that the proportions of diapause producers at the young adult stage tend to be varied among experiments. Therefore, we used a single batch for each experiment to minimize variation. Wasps were reared at 20 ± 1 °C, unless otherwise mentioned.
To determine the effects of per RNAi on N. vitripennis photoperiodism, yellow pupae reared under LD 16:8 h were injected with either per or bla dsRNA and either maintained under LD 16:8 h or transferred to 12:12 h. As an intact group, pupae were only fixed on a glass slide using double-stick tape. Newly emerged adults were mass-cultured for 2 days to allow them to mate, and females were then transferred into individual glass vials. Two flesh fly pupae were supplied as a host to each female every 2 days. Fly pupae were collected 1 day after parasitism, and the diapause status of the wasp offspring was determined as described above.
It has been reported that a few days of low temperature exposure under DD prompt females to lay diapause-destined eggs (Rivers et al. 2000). This treatment is effective and most females immediately became diapause producers. In the present study, we investigated the ability of per RNAi insects to become diapause producers in response to low-temperature treatment. In this experiment, yellow pupae reared under LD 16:8 h were injected with 1 μg/μl of dsbla or dsper of region A and transferred to LD 12:12 h. Newly emerged adults were then allowed to mate for 1 day. Thereafter, some females were exposed to low temperatures (10 ± 1 °C) under DD for 3 days. After this exposure, females were transferred into individual glass vials and returned to LD 12:12 h at 20 ± 1 °C. Each female was provided with two flesh fly pupae on days 4 and 6. One day later, parasitized fly pupae were collected and maintained under DD. A control group was continuously reared under LD 12:12 h without exposure to low temperatures.
Quantitative real-time PCR (qPCR)
We investigated the temporal expression patterns of per, cry-m, cyc, cwo, and Pdp1. Wasps were reared under LD 16:8 h until the adults emerged. Some wasps were continuously maintained under LD 16:8 h, whereas the others were transferred to LD 12:12 h on the day of adult emergence. Adult wasps were fed on 10 % sucrose, and were flash-frozen in liquid nitrogen at Zeitgeber time (ZT) 0 (light on) 4, 8, 12, 16, and 20 on day 12. Some females under long-day conditions were transferred to DD on day 12 at ZT16 (the time of light-off) and flash-frozen at 8, 12, 16, 20, 24, 28, and 32 h after light-off, which correspond to ZT 24, 28, 32, 36, 40, and 44, respectively. Total RNA was isolated from the heads of five N. vitripennis using a PureLink RNA Micro Kit with TRIzol reagent (Thermo Fisher Scientific). cDNA was synthesized using a High Capacity cDNA Reverse Transcription kit (Thermo Fisher Scientific). For qPCR analysis, 0.2 % of the cDNA was used at a final concentration of 1× GoTaq qPCR Master Mix (Promega) and 0.05 μM of each primer pair using a 7500 Real-Time PCR System (Thermo Fisher Scientific). Each reaction was performed in duplicate. The following primer pairs were used: perF (5′-CGC AAT CCA CCG AGG AAT AT-3′) and perR (5′-CAG CAC AAG GCC GTC ATG-3′) for per; cycF (5′-CCC GAA CGA GTC AAC ACT-3′) and cycR (5′-CGA AAC TCG CTC CCT TGC T-3′) for cyc; cry-mF (5′-CCG CTG ACG TAC CAT CAG TTC-3′) and cry-mR (5′-CTC GGG CGG GTC CAT T-3′) for cry-m; Pdp1F (5′-CCC AAT CTC TGG GAC AAG ACA-3′) and Pdp1R (5′-TCG TCC AGG TAG ACG TAC TTC-3′) for Pdp1; and cwoF (5′-CCA CCG GCC AAC TAC AAG TT-3′) and cwoR (5′-CTC TGG TCG GCG CTG AA-3′) for cwo. Ribosomal protein L32 (RpL32, also known as rp49) was used as a control gene for normalization, and primers for the gene were rp49F (5′-AGA AAT TGC CCA TGG AGT TAG C-3′) and rp49R (5′-CTG CTG GGC ACG TTC GA-3′). In all reactions, the generation of only a single expected amplicon was confirmed by melting analysis. Quantification of cDNAs was performed by a standard curve methodology.
One-way ANOVA was used to test for significant differences in the temporal oscillations in mRNA abundance. Two-way ANOVA was used to test for significant differences in the expression patterns of the genes of interest in wasps under short- and long-day conditions. In addition, per and cry-m expression data were analyzed using R, version 3.1.2 (R core team 2014), according to the method used by Bertossa et al. (2014). Expression data were divided by the mean expression level within each treatment and a sine wave model (gene expression ~ sin (time × (2 × π/period) − π × acrophase/12) × amplitude + shift) was fitted to the data to determine circadian parameters. The nls2 function in R was used to determine the non-linear least-squares estimates of acrophases. Parameter intervals were set as follows: phase = 0°–360°, period = 24 h, and amplitude = 0–4.
Levels of per mRNA after dsRNA injection were also measured by qPCR. Six days after dsRNA injection, total RNA was isolated from the whole bodies of 4 pupae under short-day conditions at ZT 0, 6, 12, and 18. RNA isolation, cDNA synthesis, and qPCR were performed as described above.
Results
Photoperiodic response curve
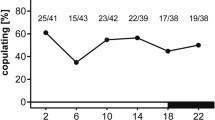
Our N. vitripennis colony showed a clear photoperiodic response as evidenced by the proportion of females laying diapause-destined eggs (diapause producers; Fig. 1). Proportions of diapause producers were higher under LD 12:12 and 13:11 h, than under LD 15:9 and 16:8 h. The critical day-length, which induces half of the population to become diapause producers, was approximately 13.5 h.
Temporal expression patterns of clock genes
Temporal variation in mRNA abundance of the clock genes per, cry-m, cyc, cwo, and Pdp1 in the heads of N. vitripennis was investigated under LD 16:8 h, LD 12:12 h, and DD (Fig. 2). per mRNA abundance increased from the end of the photophase or the early scotophase and decreased toward the middle of the photophase under LD 16:8 and LD 12:12 h, although significant diel oscillation was detected only in LD 16:8 h (one-way ANOVA, p < 0.05). A similar expression pattern was observed under DD, but this oscillation was not statistically significant (one-way ANOVA, p > 0.05). The expression of cry-m was similar to that of per, and diel oscillation was significant under LD 16:8 h and DD (one-way ANOVA, p < 0.05). The oscillation of Pdp1 expression was significant under DD (one-way ANOVA, p < 0.05), but no significance was detected under either LD condition. cyc and cwo mRNA abundance showed no oscillation under any condition (one-way ANOVA, p > 0.05).
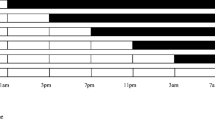
Temporal expression of mRNA of clock genes (per, cry-m, cyc, cwo and Pdp1) in the heads of Nasonia vitripennis (normalized by RpL32 mRNA abundance). The left column shows expression under short-day (light:dark [LD] 12:12 h; open circles and broken lines) and long-day (LD 16:8 h; closed circles and solid lines) conditions. The right column shows expression under constant darkness (DD). Each line is drawn through the mean of triplicate plots. Open and closed bars on the top indicate photophase and scotophase, respectively
Two-way ANOVA detected a significant difference in per and cry-m expression in LD 16:8 and 12:12 h (p < 0.05). For any other genes analyzed, no significant difference in expression was observed between photoperiods (two-way ANOVA, p > 0.05). We also calculated acrophases of per and cry-m expression using the sine curve fitting methodology, according to the methods used by Bertossa et al. (2014). However, we found only negligible differences (per had a an acrophase of 21.3 ± 0.6 h in LD 16:8 h and 20.3 ± 1.4 h in LD 12:12 h; cry-m had a an acrophase of 20.2 ± 1.3 h in LD 16:8 h and 19.2 ± 0.6 h in LD 12:12 h).
Effects of per RNAi on photoperiodic response
The levels of per mRNA were measured after dsRNA injection. Injecting the dsRNA of per regions A and B resulted in a significantly low level of per mRNA abundance than did injecting dsbla (two-way ANOVA, p < 0.01 for both, Fig. 3).
Temporal expression of per mRNA (normalized by RpL32 mRNA abundance) in Nasonia vitripennis after injecting double-stranded RNA under short-day conditions (light:dark 12:12 h). a 1 μg/μl of dsbla (closed circles) or dsper region A (open circles) was injected. b 5 μg/μl of dsbla (closed circles) or dsper region B (open circles) was injected. Each line is drawn through the mean of quadruplicate plots. Two-way ANOVA detected significant differences between expression patterns in the dsbla- and dsper-injected wasps (p < 0.05)
We investigated the effects of per RNAi on the proportion of diapause producers (Fig. 4). In intact wasps, the proportion of diapause producers gradually increased in response to the number of short-day cycles received, while it remained low under long-day conditions. Thus, a clear photoperiodic response was observed. This also holds true for the dsbla-injected wasps. However, injection of dsRNA of both per region A and per region B suppressed the proportion of diapause producers under short-day conditions, though there were some differences in their efficacy; i.e., the proportion of diapause producers was lower until the end of the experiment in wasps injected with dsRNA of per region B, whereas in wasps injected with dsRNA of per region A, the number recovered on days 10 and 12 (Fig. 4).
Temporal variation in the proportion of Nasonia vitripennis diapause producers under short-day (left column) and long-day (right column) conditions. Upper panels: proportions of diapause producers in intact wasps and wasps injected with 1 μg/μl of dsbla or dsper region A. Lower panels: proportions of diapause producers in intact wasps and wasps injected with 5 μg/μl of dsbla or dsper region B. LD light:dark regime
To investigate whether per RNAi directly affected the diapause production process itself, we focused on the ability of per RNAi insects to become diapause producers in response to exposure to low temperature (Fig. 5). Proportions of diapause producers among the dsbla-injected wasps not exposed to low temperature increased in response to the number of short-day cycles received, although the difference was not statistically significant. In contrast, the dsper-injected wasps not exposed to low temperatures suppressed such increments. This is consistent with the results shown in Fig. 4 and confirms the suppressive effect of per RNAi on the incidence of diapause producers. Exposure to low temperature swiftly enhanced the proportion of diapause producers in the dsbla-injected group on day 4; this effect was also observed in the dsper-injected group. It should be noted that the proportion of diapause producers remained high on day 6 in the dsbla-injected wasps, but decreased in the dsper-injected wasps on the same day (Fig. 5; Tukey-type multiple comparisons for proportions, p < 0.05).
The effects of exposure to low temperature on the induction of diapause producers in Nasonia vitripennis injected with 1 μg/μl of dsbla or dsper region A. After dsRNA injection, wasps were reared under short-day conditions (LD 12:12 h) at 20 °C. One day after adult emergence, some wasps were exposed to 10 °C for 3 days in constant darkness (DD). After treatment, wasps were returned to the original conditions. The control wasps were continuously maintained under LD 12:12 h at 20 °C without exposure to low temperature. Flesh fly pupae were supplied on days 4 and 6
Discussion
Clock gene expression under different photoperiods
Bertossa et al. (2014) showed a clearly discernible oscillation in per and cry-m mRNA levels in the heads of N. vitripennis. Such discernible oscillation has also been observed in other insect species (Ingram et al. 2012; Zhu et al. 2008). In addition, changes in photoperiodic conditions cause the phase of their oscillations to shift, and the shift of the oscillation phase correlates to light-on (not light-off) in N. vitripennis, suggesting a possible role of per and cry-m oscillation in the dawn oscillator of the internal coincidence model (Bertossa et al. 2014). Although two-way ANOVA detected significant differences in per and cry-m expression under photoperiods, we found only negligible differences in their acrophases. Thus, phase shift in response to photoperiods was not observed in the present study. We further extended expression analyses to other clock genes, cyc, cwo, and Pdp1, but did not find significant differences in their expression patterns between photoperiods. Why there is an inconsistency between the previous study (Bertossa et al. 2014) and the present study is still unclear, but it may be due to a difference in experimental designs (photoperiods and sampling intervals), and from strain difference. Our N. vitripennis colony showed a clear photoperiodic response without significant difference in mRNA oscillation patterns between photoperiods, implying little role of transcriptional regulation of clock genes on photoperiodism. Extensive analyses of expression patterns of proteins as well as mRNA under various photoperiods and also analyses of the expression in the clock cells are necessary to assess the significance of their phase shifts in response to distinct photoperiods.
Effects of per RNAi on photoperiodic response
In the present study, per dsRNA injection significantly reduced per mRNA abundance. Thus, RNAi is effective in this species, as reported previously (Lynch and Desplan 2006; Werren et al. 2009; Zwier et al. 2012).
per RNAi insects did not respond to short days with an increase in the proportion of diapause producers. dsRNAs of both per regions A and B affected the photoperiodic response, indicating that the loss of the response is not caused by an off-target effect, but by suppression of the expression of per itself. We conclude that the per gene is essential to the photoperiodic response in N. vitripennis. Causal involvement of the clock genes in photoperiodism has been reported in some insect species belonging to Heteroptera, Orthoptera, Lepidoptera, and Diptera (Ikeno et al. 2010, 2011, 2013; Meuti et al. 2015; Mohamed et al. 2014; Sakamoto et al. 2009). It has been verified that the circadian clock, composed of the clock genes, is involved in the photoperiodic response in the bean bug Riptortus pedestris (F.) (Ikeno et al. 2010). In this species, putative clock cells that are involved in the photoperiodic response were identified in the brain (Ikeno et al. 2014). However, it is still unknown which processes in the photoperiodic cascade are affected by the circadian clock. Conceptually, the circadian clock would be involved in photoperiodic time measurement (Bünning’s hypothesis). Alternatively, the circadian clock controls all the processes involved in the photoperiodic response, and dysfunction of the clock may affect almost all the processes (Veerman 2001). Indeed, the circadian clock regulates a wide array of physiological processes, such as behavior, learning, feeding, metabolism, chemosensation, and immunity (Allada and Chung 2010). Where the circadian clock is involved is still open to discussion, but a recent study in Drosophila melanogaster Meigen supports the involvement of the clock in the photoperiodic time measurement system. Pegoraro et al. (2014) focused on the photoperiodic modulation of chill-coma recovering time (CCRt) in D. melanogaster. Wild-type flies raised under short-day conditions exhibit significantly shorter CCRt than flies raised under long-day conditions. Interestingly, mutants with a long free-running period consistently showed shorter CCRt under both long and short photoperiods than mutants with a short free-running period. These results fit Bünning’s hypothesis (Bünning 1936). The hypothesis posited that the 24-h circadian clock consisted of two 12-h half-cycles: the photophil and scotophil, and that short-day effects are observed when light is restricted to occur within the photophil, while long-day effects are produced when the scotophil is illuminated. Under this concept, various day lengths are always shorter than the photophil phase and are interpreted as short days in Drosophila mutants with a long free-running period (Pegoraro et al. 2014). Although it is naive to apply the simplified Bünning’s hypothesis here, the different photoperiodic phenotypes of the slow and fast clock mutants suggest a causative role for the circadian clock in photoperiodic time measurement.
Schneiderman and Horwitz (1958) and Rivers et al. (2000) reported that exposure to low temperature immediately affects maternal physiology and prompts Nasonia females to be diapause producers. Photoperiodic induction of diapause producers (by rearing under short-day conditions) is less effective than induction by exposure to low temperatures, and requires more days to generate a high proportion of diapause producers (see Fig. 5 in the present study; Rivers et al. 2000). This indicates that exposure to low temperature directly affects the maternal physiology involved in the production of diapause-destined eggs, independent of the photoperiodic cascade. The per RNAi wasps, which failed to be diapause producers under short-day conditions, successfully laid diapause-destined eggs when exposed to low temperature. These results clearly indicate that per RNAi did not disrupt maternal physiology directly involved in production of diapause-destined eggs, but did affect photoperiodic perception. It is also worth noting that the proportion of diapause producers in the per RNAi wasps was lower than that in the control wasps on day 6 (Fig. 5). The dsbla RNA-injected wasps were able to perceive short days after the low temperature exposure and, therefore, retained a high proportion of diapause producers on day 6. On the other hand, the dsper RNA-injected wasps would have no or little ability to perceive short days, as shown in Fig. 4. Thus, they failed to retain a high proportion of diapause producers on day 6.
Although further studies are needed before making a conclusion, the present study, together with the previous studies (Bertossa et al. 2014; Paolucci 2014; Saunders 1974), supports a causative role for per in assessment of photoperiods, i.e., photoperiodic time measurement. The next point to be addressed is where the per-expressing cells governing photoperiodism reside. Because all the mechanisms regulating photoperiodism are assumed be in the brain (Bowen et al. 1984, Hasegawa and Shimizu 1987), the per-expressing cells would also reside in the Nasonia brain. In D. melanogaster, there are approximately 150 cells in the brain that express clock genes, and they are divided into eight subgroups, the ventral lateral neurons (s-LNv, l-LNv), the dorsal lateral neurons (LNd), the dorsal neurons (DN3, DN1p, DN1a, DN2), and the lateral posterior neurons (LPN) (Helfrich-Förster 2006; Tomioka and Yoshii 2006). Drosophila melanogaster shows a distinct bimodal rhythm in its locomotor activity, i.e., it exhibits morning and evening peaks, and these peaks are considered to be regulated by two oscillators (morning [M] oscillator and evening [E] oscillator). M and E oscillators track dawn and dusk, respectively, and therefore the phase angle between the two oscillators under different day lengths differs (reviewed by Yoshii et al. 2012). M oscillators are considered to reside in the four pigment-dispersing factor-positive s-LNvs, and E oscillators in the 3–4 LNds (Grima et al. 2004; Stoleru et al. 2004). These cells are the primary candidates for the dawn and dusk oscillators in the internal coincidence model in Nasonia. It is of interest to investigate oscillation patterns of per-expressing cells in the Nasonia brain under different photoperiods.
References
Allada R, Chung BY (2010) Circadian organization of behavior and physiology in Drosophila. Annu Rev Physiol 72:605–624. doi:10.1146/annurev-physiol-021909-135815
Bertossa RC, van de Zande L, Beukeboom LW, Beersma DG (2014) Phylogeny and oscillating expression of period and cryptochrome in short and long photoperiods suggest a conserved function in Nasonia vitripennis. Chronobiol Int 31:749–760. doi:10.3109/07420528.2014.880451
Bowen MF, Saunders DS, Bollenbacher WE, Gilbert LI (1984) In vitro reprogramming of the photoperiodic clock in an insect brain-retrocerebral complex. Proc Natl Acad Sci USA 81:5881–5884. doi:10.1073/pnas.81.18.5881
Bünning E (1936) Die endonome Tagesrhythmik als Grundlage der photoperiodischen Reaktion. Ber Deutsch Bot Ges 54:590–607
Danilevskii AS, Goryshin NI, Tyshchenko VP (1970) Biological rhythms in terrestrial arthropods. Ann Rev Entomol 15:201–244. doi:10.1146/annurev.en.15.010170.001221
Dolezel D (2015) Photoperiodic time measurement in insects. Curr Opin Insect Sci 7:98–103. doi:10.1016/j.cois.2014.12.002
Goto SG (2013) Roles of circadian clock genes in insect photoperiodism. Entomol Sci 16:1–16. doi:10.1111/ens.12000
Grima B, Chélot E, Xia R, Rouyer F (2004) Morning and evening peaks of activity rely on different clock neurons of the Drosophila brain. Nature 431:869–873. doi:10.1038/nature02935
Hardin PE (2006) Essential and expendable features of the circadian timekeeping mechanism. Curr Opin Neurobiol 16:686–692. doi:10.1016/j.conb.2006.09.001
Hasegawa K, Shimizu I (1987) In vivo and in vitro photoperiodic induction of diapause using isolated brain-suboesophageal ganglion complexes of the silkworm, Bombyx mori. J Insect Physiol 33:959–966. doi:10.1016/0022-1910(87)90008-4
Helfrich-Förster C (2006) The neural basis of Drosophila’s circadian clock. Sleep Biol Rhythms 4:224–234. doi:10.1111/j.1479-8425.2006.00223.x
Ikeno T, Tanaka SI, Numata H, Goto SG (2010) Photoperiodic diapause under the control of circadian clock genes in an insect. BMC Biol 8:116. doi:10.1186/1741-7007-8-116
Ikeno T, Numata H, Goto SG (2011) Photoperiodic response requires mammalian-type cryptochrome in the bean bug Riptortus pedestris. Biochem Biophys Res Commun 410:394–397. doi:10.1016/j.bbrc.2011.05.142
Ikeno T, Ishikawa K, Numata H, Goto SG (2013) Circadian clock gene Clock is involved in the photoperiodic response of the bean bug Riptortus pedestris. Physiol Entomol 38:157–162. doi:10.1111/phen.12013
Ikeno T, Numata H, Goto SG, Shiga S (2014) Involvement of the brain region containing pigment-dispersing factor-immunoreactive neurons in the photoperiodic response of the bean bug, Riptortus pedestris. J Exp Biol 217:453–462. doi:10.1242/jeb.091801
Ingram KK, Kutowoi A, Wurm Y, Shoemaker D, Meier R, Bloch G (2012) The molecular clockwork of the fire ant Solenopsis invicta. PLoS One 7:e45715. doi:10.1371/journal.pone.0045715
Lynch JA, Desplan C (2006) A method for parental RNA interference in the wasp Nasonia vitripennis. Nat Protoc 1:486–494. doi:10.1038/nprot.2006.70
Meuti ME, Stone M, Ikeno T, Denlinger DL (2015) Functional circadian clock genes are essential for the overwintering diapause of the northern house mosquito, Culex pipiens. J Exp Biol 218:412–422. doi:10.1242/jeb.113233
Mohamed AA, Wang Q, Bembenek J, Ichihara N, Hiragaki S, Suzuki T, Takeda M (2014) N-acetyltransferase (nat) is a critical conjunct of photoperiodism between the circadian system and endocrine axis in Antheraea pernyi. PLoS One 9:e92680. doi:10.1371/journal.pone.0092680
Nelson RJ, Denlinger DL, Somers DE (2009) Photoperiodism: the biological calendar. Oxford University Press, Oxford
Paolucci S (2014) The genetics of adaptive photoperiodic response in Nasonia vitripennis. Ph.D. thesis, University of Groningen, Netherland
Pegoraro M, Gesto JS, Kyriacou CP, Tauber E (2014) Role for circadian clock genes in seasonal timing: testing the Bünning hypothesis. PLoS Genet 10:e1004603. doi:10.1371/journal.pgen.1004603
R Core Team (2014) R: a language and environment for statistical computing. R foundation for statistical computing, Vienna, Austria. http://www.R-project.org/
Rivers DB, Lee RE, Denlinger DL (2000) Cold hardiness of the fly pupal parasitoid Nasonia vitripennis is enhanced by its host Sarcophaga crassipalpis. J Insect Physiol 46:99–106. doi:10.1016/S0022-1910(99)00106-7
Sakamoto T, Uryu O, Tomioka K (2009) The clock gene period plays an essential role in photoperiodic control of nymphal development in the cricket Modicogryllus siamensis. J Biol Rhythms 24:379–390. doi:10.1177/0748730409341523
Saunders DS (1965) Larval diapause of maternal origin: induction of diapause in Nasonia vitripennis (Walker) (Hymenoptera: Pteromalidae). J Exp Biol 42:495–508
Saunders DS (1966) Larval diapause of maternal origin—II. The effect of photoperiod and temperature on Nasonia vitripennis. J Insect Physiol 12:569–581. doi:10.1016/0022-1910(66)90095-3
Saunders DS (1974) Evidence for ‘dawn’ and ‘dusk’ oscillators in the Nasonia photoperiodic clock. J Insect Physiol 20:77–88. doi:10.1016/0022-1910(74)90125-5
Saunders DS (2014) Insect photoperiodism: effects of temperature on the induction of insect diapause and diverse roles for the circadian system in the photoperiodic response. Entomol Sci 17:25–40. doi:10.1111/ens.12059
Saunders DS, Sutton D, Jarvis RA (1970) The effect of host species on diapause induction in Nasonia vitripennis. J Insect Physiol 16:405–416. doi:10.1016/0022-1910(70)90181-2
Schneiderman HA, Horwitz J (1958) The induction and termination of facultative diapause in the chalcid wasps Mormoniella vitripennis (Walker) and Tritneptis klugii (Ratzeburg). J Exp Biol 35:520–551
Stoleru D, Peng Y, Agosto J, Rosbash M (2004) Coupled oscillators control morning and evening locomotor behaviour of Drosophila. Nature 431:862–868. doi:10.1038/nature02926
Tomioka K, Matsumoto A (2010) A comparative view of insect circadian clock systems. Cell Mol Life Sci 67:1397–1406. doi:10.1007/s00018-009-0232-y
Tomioka K, Matsumoto A (2015) Circadian molecular clockworks in non-model insects. Curr Opin Insect Sci 7:58–64. doi:10.1016/j.cois.2014.12.006
Tomioka K, Yoshii T (2006) Entrainment of Drosophila circadian rhythms by temperature cycles. Sleep Biol Rhythms 4:240–247. doi:10.1111/j.1479-8425.2006.00227.x
Tyshchenko VP (1966) Two-oscillatory model of the physiological mechanism of the photoperiodic reaction of insects. Zh Obshch Biol 27:209–222
Vaz Nunes M, Saunders DS (1999) Photoperiodic time measurement in insects: a review of clock models. J Biol Rhythms 14:84–104. doi:10.1177/074873049901400202
Veerman A (2001) Photoperiodic time measurement in insects and mites: a critical evaluation of the oscillator-clock hypothesis. J Insect Physiol 47:1097–1109. doi:10.1016/S0022-1910(01)00106-8
Werren JH, Loehlin DW, Giebel JD (2009) Larval RNAi in Nasonia (parasitoid wasp). Cold Spring Harb Protoc. doi:10.1101/pdb.prot5311
Yoshii T, Rieger D, Helfrich-Förster C (2012) Two clocks in the brain: an update of the morning and evening oscillator model in Drosophila. Prog Brain Res 199:59–81. doi:10.1016/B978-0-444-59427-3.00027-7
Zhu H, Sauman I, Yuan Q, Casselman A, Emery-Le M, Emery P, Reppert SM (2008) Cryptochromes define a novel circadian clock mechanism in monarch butterflies that may underlie sun compass navigation. PLoS Biol 6:e4. doi:10.1371/journal.pbio.0060004
Zwier MV, Verhulst EC, Zwahlen RD, Beukeboom LW, Van de Zande L (2012) DNA methylation plays a crucial role during early Nasonia development. Insect Mol Biol 21:129–138. doi:10.1111/j.1365-2583.2011.01121.x
Acknowledgments
This work was supported, in part, by KAKENHI (Grant No. 25450488).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mukai, A., Goto, S.G. The clock gene period is essential for the photoperiodic response in the jewel wasp Nasonia vitripennis (Hymenoptera: Pteromalidae). Appl Entomol Zool 51, 185–194 (2016). https://doi.org/10.1007/s13355-015-0384-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13355-015-0384-1