Abstract
The bread wheat TaSOS1 has been previously shown to be induced by salt stress treatment. To further investigate the regulation of the TaSOS1 gene, the two genomic fragments Pr SOS1-AB and Pr SOS1-D have been isolated and sequenced. Pr SOS1-AB and Pr SOS1-D are the promoter regions of SOS1 alleles, which are localised on genomes A and/or B, and on genome D, respectively. Sequence analysis of these two promoters revealed the presence of cis-regulatory elements which could be required for abiotic stress and abscisic acid (ABA) responsiveness. Histochemical assays of stably transformed Arabidopsis T3 plants showed that Pr SOS1-AB and Pr SOS1-D are active in this heterologous system, and their activities were almost the same at early developmental stages (4-, 8- and 12-day-old transgenic Arabidopsis seedlings). Nevertheless, β-glucuronidase (GUS) activity was detected only in plants carrying the Pr SOS1-AB –gusA construct grown for 20 or 30 days. Furthermore, in these plants, the application of abiotic stress produced an accumulation in gusA transcripts. Taken together, these results show that, in this heterologous dicot system and under normal growth conditions, Pr SOS1-AB and Pr SOS1-D are age-dependent and organ-specific promoters. However, in the presence of different stress conditions, the activities of these two promoters became different and only Pr SOS1-AB is an abiotic stress-inducible promoter at different developmental stages. Thus, Pr SOS1-AB can be used for the development of abiotic stress-tolerant transgenic plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salinity stress negatively impacts agricultural yield throughout the world, affecting production, whether it is for subsistence or economic gain. The plant response to salinity consists of numerous processes that must function in coordination to alleviate both cellular hyperosmolarity and ion disequilibrium (Tester and Davenport 2003). In addition, crop plants must be capable of satisfactory biomass production in a saline environment. Recent progress in the elucidation of salt stress signalling and effector output determinants that mediate ion homeostasis has uncovered some potential biotechnology tactics that may be used to obtain salt-tolerant crop plants, i.e. enhance the yield stability under salinity. However, it has been shown that high constitutive expression of the foreign gene may be detrimental to the host plant, with independent reports of increased sterility, retarded development, abnormal morphology, yield penalty, altered grain composition or transgene silencing (Sinha et al. 1993; Xu et al. 2006). The use of a strong, tissue-specific or inducible promoter to restrict gene expression to the required tissue, at a particular developmental stage and/or in response to a stress, may solve this type of problem (Karim et al. 2007; Pino et al. 2007). Until now, numerous inducible promoters have been isolated from a wide variety of organisms. Among these, the biotic and abiotic stress-inducible rd29A promoter has been widely used to minimise the otherwise negative effect on plant growth of transgene expression in plants such as tobacco, sugarcane and potato (Kasuga et al. 2004; Behnam et al. 2006; Wu et al. 2008).
Signal transduction networks for abiotic stress are divided into three major signalling types: osmotic/oxidative stress signalling involves the generation of reactive oxygen species (ROS)-scavenging enzymes and antioxidant compounds, as well as osmolytes; Ca2+-dependent signalling that leads to the activation of late embryogenesis abundant (LEA)-type genes; and Ca2+-dependent salt overly sensitive (SOS) signalling (Xiong et al. 2002). The SOS signalling pathway, which comprises SOS3, SOS2 and SOS1, is a pivotal regulator of, at least some, key transport systems required for ion homeostasis (Sanders 2000; Zhu 2000). Loss-of-function mutations in SOS3, SOS2 and SOS1 cause hypersensitivity to Na+ (Wu et al. 1996; Zhu et al. 1998). The direct downstream target of this pathway is SOS1, a plasma membrane Na+/H+ antiporter (Shi et al. 2000; Qiu et al. 2002; Quintero et al. 2002). SOS3 is a calcium-binding protein (Liu and Zhu 1998; Ishitani et al. 2000), while SOS2 is a Ser/Thr kinase. Both the catalytic and regulatory domains are essential for SOS2 function in salt tolerance. The C-terminal regulatory domain contains an auto-inhibitory FISL motif that binds to SOS3 or calcineurin B-like 10 (CBL10), thereby releasing SOS2 from auto-inhibition (Liu et al. 2000; Guo et al. 2001; Quan et al. 2007). SOS3–SOS2 and CBL10–SOS2 complexes regulate the plasma membrane Na+/H+ exchange activity of SOS1 through phosphorylation, which releases AtSOS1 from auto-inhibition (Quintero et al. 2002, 2011; Quan et al. 2007). In addition to Arabidopsis, the SOS1 gene has been identified in other plants like rice (Martínez-Atienza et al. 2007), wheat (Xu et al. 2008; Feki et al. 2011), tomato (Olías et al. 2009) and Thellungiella salsuginea (Oh et al. 2009). Despite the demonstrated role of some SOS1 genes in ion homeostasis and in the partitioning of the toxic ion Na+ between plant organs (Shi et al. 2002; Olías et al. 2009), only experimental functional analysis of Arabidopsis SOS1 and Salicornia brachiata SbSOS1 promoter was performed in transgenic Arabidopsis and tobacco plants, respectively (Shi et al. 2002; Goyal et al. 2013). Arabidopsis SOS1, SOS2 and SOS3 promoter regions present the same cis elements. However, the predicted cis elements present in SOS2 promoter are higher than SOS1 and SOS3, with the presence of many small RNA target sites. This in silico analysis indicates that these three SOS promoter regions could be involved in common features of transcriptional regulation, and the SOS2 gene is regulated by several upstream transcription factors and is involved in other outputs besides Na+ transport by SOS1 (Ji et al. 2013). Despite the extensive synteny of the open reading frames (ORFs) and the conservation of gene structures for SOS1 between Arabidopsis and its halophytic relative (Thellungiella parvula), the promoter region of this gene is conserved only between the Thellungiella species (Oh et al. 2010).
It has been shown previously that Arabidopsis SOS1, SOS2 and SOS3 genes have different special expression profiles. The first two genes are expressed in both roots and shoots, while the latter is mainly expressed in root tissues (Liu et al. 2000; Shi et al. 2002; Quan et al. 2007). The expression of these genes is induced by salt stress in roots, but only SOS3 expression displays differential induction in various types of root cells located in different developmental root zones in response to salt stress (Ji et al. 2013). Arabidopsis AtSOS1 is preferentially expressed in epidermal cells at the root tip and in parenchyma cells at the xylem/symplast boundary of roots, stems and leaves (Shi et al. 2002). Concerning SOS1 gene expression, salt challenge induces a clear accumulation of SOS1 mRNA (Martínez-Atienza et al. 2007; Xu et al. 2008; Olías et al. 2009; Tang et al. 2010; Wang et al. 2010). Strikingly, in some cases, salinity produced little or even no alteration of SOS1 transcription (Taji et al. 2004; Kant et al. 2006; Mullan et al. 2007; Wu et al. 2007; Cosentino et al. 2010; Feki et al. 2011). To our knowledge, the expression profile of wheat SOS1 has been analysed under different abiotic stresses, but the wheat SOS1 promoter activity has not yet been experimentally analysed. In this study, we showed that the two isolated wheat SOS1 promoter regions (Pr SOS1-D and Pr SOS1-AB ) are active, age-dependent and organ-specific promoters in Arabidopsis plants. Moreover, we demonstrated that, contrary to Pr SOS1-D , Pr SOS1-AB is an abiotic stress-inducible promoter at different developmental stages (8, 20 and 30 days old). Thus, the Pr SOS1-AB promoter will be useful for specific spatiotemporal targeting and accumulation of proteins conferring tolerance to abiotic stresses in transgenic plants.
Materials and methods
Genomic library screening and isolation of wheat SOS1 promoter regions
The bacterial artificial chromosome (BAC) library from Triticum aestivum (cv. Chinese spring) was screened using polymerase chain reaction (PCR) amplifications as described by Isidore et al. (2005). Amplifications were done using wheat SOS1-specific primers, which are S1 as a sense primer and S2 as a reverse primer (Table 1). The 5′-flanking region of the wheat SOS1 gene was isolated using the inverse PCR method as described by Ochman et al. (1988). PCR reactions were carried out with the corresponding recombinant BAC clone as a template, and with the wheat SOS1-specific primers IS1 and IS2, designed close to the 5′UTR sequence (Table 1). Two promoter sequences, named Pr SOS1-D and Pr SOS1-AB of 2,660 and 2,745 bp, respectively, were obtained by sequencing.
In silico analysis of the wheat SOS1 promoters
The search for putative cis elements in the two promoter sequences was carried out using the signal scan search provided by the PLACE (http://www.dna.affrc.go.jp/PLACE/) and the PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) databases (Higo et al. 1999; Lescot et al. 2002).
Construction of the binary vectors and Arabidopsis transformation
Fragments of the two promoters Pr SOS1-D and Pr SOS1-AB were amplified using the two different couples of primers ES1–NS1 and ES2–NS2, respectively. The two forward primers ES1 and ES2 harbour the PstI restriction site, and the two reverse primers NS1 and NS2 harbour the NcoI restriction site. The two resulting fragments were cloned separately in front of the gusA gene into the pCAMBIA1391Z vector (Cambia, Canberra, Australia), using the PstI and NcoI restriction sites. The resulting recombinant binary vectors were named pCAMBIA1391Z–Pr SOS1-D –gusA and pCAMBIA1391Z–Pr SOS1-AB –gusA. Agrobacterium tumefaciens strain GV3101 (Konez and Schell 1986) was transformed separately by the freeze–thaw transformation method (Chen et al. 1994) with these two binary vectors and the binary vector pCAMBIA1301, which were subsequently used for Arabidopsis transformation. The Arabidopsis thaliana transformation was performed using the floral dipping technique (Clough and Bent 1998). Transgenic plants were selected on Murashige and Skoog (MS) agar medium (Murashige and Skoog 1962) containing 20 μg L−1 hygromycin. Seeds of the T2 generation were harvested and the seedlings from the homozygous T3 generation were used for histochemical β-glucuronidase (GUS) staining. The wild-type Arabidopsis and 35S-gusA transgenic plants were used as negative and positive controls, respectively.
Identification of the transgenic Arabidopsis plants
Genomic DNA was extracted as described by Michiels et al. (2003) from the leaves of Pr SOS1-AB –gusA and Pr SOS1-D –gusA transformed Arabidopsis, and was used in PCR amplifications. To detect positive lines, the couples of primers from the gusA gene (GR) and flanking the promoter Pr SOS1-AB region (ABF) or the promoter Pr SOS1-D region (DF) were used (Table 2). The amplified products were resolved on a 1 % agarose gel and visualised by ethidium bromide staining.
For each Arabidopsis transformation, four positive transgenic lines were randomly selected to analyse the expression of the gusA gene. From 20-day-old seedlings, total RNA was isolated using the Trizol method (Invitrogen), and to remove contaminating DNA, it was then treated with RNase-free DNaseI (Promega) at 37 °C for 15 min and further incubated at 65 °C for 10 min. DNase-treated RNA samples (0.5 μg) were reverse-transcribed using M-MLV reverse transcriptase (Invitrogen). The reverse transcription (RT) reactions were performed at 37 °C for 1 h and using the oligo-dT (18 mer) primer. One microlitre of each cDNA was used as the template for PCR amplification with 2 units of Taq DNA polymerase (Invitrogen), 200 μM dNTPs and 0.5 μM gusA-specific primers (GusF–GusR) (Table 2). An Arabidopsis thaliana β-tubulin gene fragment (GenBank accession no. XM_002863542.1), used as an internal control, was amplified with the forward TubF and the reverse TubR primers (Table 2). The PCR products (5 μl) were separated on 1.5 % agarose gel.
Growth conditions, abiotic stress treatment and gusA expression analysis
The effect of abiotic stress on gusA transcripts accumulation was monitored using seeds of homozygous transgenic lines and the non-transformed plants. Seeds of each T3 homozygous transgenic Arabidopsis lines were surface-sterilised and then grown on MS agar medium under light/dark cycle conditions of 16 h light/8 h dark cycles at 22 °C. Seedlings were grown in MS agar medium for 8, 20 or 30 days and then transferred to MS agar medium containing 100 mM NaCl, 100 mM mannitol or 20 μM abscisic acid (ABA), and kept for 2 days in each treatment. Then, the plants were harvested for analysis by histochemical GUS staining and for gusA expression analysis. Total RNA was extracted from 20- and 30-day-old seedlings, and treated with RNase-free DNaseI. The cDNA was synthesised by means of the M-MLV reverse transcriptase (Invitrogen). PCR amplifications were performed using the primers GusF and GusR (Table 2) and the PCR-amplified products were visualised on ethidium bromide-stained 1.5 % agarose gels and quantified using the Gel DocXR Gel Documentation System (Bio-Rad). This software was used to calculate the average band density, which was recorded and used in graphic analyses. The band density was determined by this software and was given in arbitrary units and graphed using Microsoft Excel. The error bars were determined from three separate biologic replicates. Each of the three biological replicates consisted of pooled plants subjected or not to different stress conditions.
Histochemical GUS staining
GUS activity was assayed histochemically by incubating tissue under vacuum infiltration with GUS staining solution [50 mM Na2HPO4 buffer (pH 7.0), 0.5 mM potassium ferricyanide, 0.5 mM potassium ferrocyanide, 0.1 % Triton X-100 and 1 mg/l X-Gluc (5-bromo-4-chloro-3-indolyl β-D-glucuronide)] for several minutes and then incubated overnight at 37 °C (Jefferson et al. 1987). The pigments and chlorophyll were removed by soaking the Arabidopsis tissues for several hours in 70 % (v/v) ethanol. Three to six stained plants from three independent experiments were observed under binocular loupe and photographed using an Olympus W120 digital still camera, and most of the photos showed similar results.
Results
Isolation and in silico analysis of the two wheat SOS1 promoter regions
A genomic library from bread wheat (Triticum aestivum) was screened by PCR with specific primers of the wheat SOS1 gene, which resulted in the isolation of two different BAC clones, BAC-AB and BAC-D, containing the wheat SOS1 alleles that are localised on genomes A and/or B, and on genome D, respectively. In order to obtain the promoter region of the bread wheat TaSOS1 gene, we performed an inverse PCR using SOS1-specific primers and the two BAC clones obtained as a template. 2,661-bp (Pr SOS1-D ) and 2,745-bp (Pr SOS1-AB ) genomic DNA fragments were isolated from BAC-D and BAC-AB clones, respectively, sequenced and deposited into GenBank NCBI (Pr SOS1-D : accession no. KF169800, Pr SOS1-AB : accession no. KF169799).
During the inspection of the two promoter sequences, we found that 140 nucleotides at the 3′ end of the Pr SOS1-D promoter region exhibited similarity with the sequence upstream of the ATG of the TaSOS1 gene (accession no. AY326952), confirming that this cloned sequence was the upstream region of the wheat SOS1 allele, which is localised on genome D. Moreover, the Pr SOS1-AB promoter region was similar to Pr SOS1-D and contained five supplementary nucleotide sequences (Fig. 1a). Thus, this second SOS1 promoter region was the upstream region of the wheat SOS1 allele, which is localised on genomes A and/or B. This finding allowed us to determine the putative transcription start site (+1), which was located 140 and 133 bp upstream of the ATG codon of the Pr SOS1-D and Pr SOS1-AB promoters, respectively. Two potential TATA boxes (TAAATAA) were identified at −27 and −40 nucleotides upstream of the transcription start site of the Pr SOS1-D and Pr SOS1-AB promoters, respectively (Fig. 1b, c), which was consistent with the regular features of eukaryotic promoters (Ke et al. 1997).
a Schematic presentation of the five gaps in the Pr SOS1-D compared to the Pr SOS1-AB promoter regions. The numbers represent their lengths (positive numbers) and their positions (negative numbers). The ATG codon is indicated by the rectangle. Presentation of the putative transcription start site (designated as +1) and the putative TATA box in the Pr SOS1-D (b) and Pr SOS1-AB (c) promoter regions
In silico analysis of the two upstream regions from the transcription start site of the SOS1 allele was performed using the plant promoter databases PLACE and PlantCARE to obtain additional indications about the transcriptional regulation of this gene. In silico analysis revealed similarity in the type but differences in the number and the position of the cis elements among the two wheat SOS1 promoter regions. Many regulatory cis elements were found and some of them are related to abiotic (dehydration and salt), biotic (fungal elicitor) and hormone (ABA) stress responses. Like Arabidopsis and Salicornia brachiata SOS1 promoters (Goyal et al. 2013; Ji et al. 2013), in silico analysis showed also the presence of several potential binding sites for transcription factors such as MYB, DOF and WRKY (Table 3).
Generation of the transgenic Arabidopsis plants
In order to study the expression of the gusA gene under different abiotic stresses, the two T-DNA of the recombinant binary vectors pCAMBIA1391Z–Pr SOS1-D –gusA and pCAMBIA1391Z–Pr SOS1-AB –gusA were introduced separately by A. tumefaciens-mediated transformation in several independent transgenic Arabidopsis lines. These vectors contain the hptII gene conferring resistance to hygromycin as a selectable marker for plant transformation. After Agrobacterium-mediated transformation of Arabidopsis plants and selection with hygromycin, several transformants were produced. From each transformation event, three transgenic lines were propagated to the T3 generation, from which homozygous plants were isolated for further analyses. The two T-DNA regions of these two binary vectors are schematically presented in Fig. 2a.
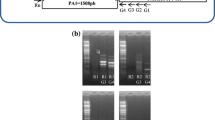
Molecular analysis of the different transgenic Arabidopsis plants. a The T-DNA schematic maps of the two binary vectors pCAMBIA1391Z–Pr SOS1-AB –gusA and pCAMBIA1391Z–Pr SOS1-D –gusA used for Arabidopsis transformation. The two wheat SOS1 promoters were inserted separately in this binary vector at the restriction sites PstI and NcoI, upstream of the gusA gene. The HPTII marker is flanked by the CaMV35S promoter (P35S) and terminator region (T35S). GR, ABF and DF indicate the primers used for the detection of the positive Arabidopsis lines. b PCR analysis of the seven and nine T2 transgenic Arabidopsis lines carrying pCAMBIA1391Z–Pr SOS1-AB –gusA (AB lines) and pCAMBIA1391Z–Pr SOS1-D –gusA (D lines), respectively, and the non-transformed Arabidopsis plants (Wt). M: molecular weight marker, (−): negative control without DNA. c RT-PCR analysis of the four AB lines (AB6, 7, 8 and 9) and the four D lines (D1, 4, 7 and 9) using specific primers for the gusA gene. No amplification was detected in the case of the non-transformed plant (Wt). A 0.5-kb β-tubulin gene fragment was amplified by RT-PCR as an internal control
T-DNA integration and gusA gene transcription were confirmed in the putative transgenic events by PCR and semi-quantitative RT-PCR, respectively. As expected, PCR products of 600- or 800-bp fragments were detected in the seven putative transgenic Arabidopsis lines carrying the Pr SOS1-AB –gusA construct and the nine putative transgenic plants carrying the Pr SOS1-D –gusA construct, respectively (Fig. 2b). The expression level of the gusA gene was analysed using RT-PCR, performed on young leaves of four AB lines (Arabidopsis carrying the Pr SOS1-AB –gusA construct), four D lines (Arabidopsis carrying the Pr SOS1-D –gusA construct) and non-transformed plants. As a control for cDNA amplification, the constitutively expressed β-tubulin gene was amplified. Semi-quantitative RT-PCR showed that the gusA transcript was expressed in these eight selected transgenic lines, and the expression level was almost similar for the AB and D lines, except the D4 line (Fig. 2c). Two representative transgenic lines AB6 and D9 were selected to further investigate the functional properties of Pr SOS1-AB and Pr SOS1-D , respectively, in T3 homozygous seedlings. Genetic segregation data performed on the selected lines using the hptII gene gave rise to a 3:1 ratio, confirming that this marker segregates as a single copy gene. GUS activity and gusA transcript accumulation were monitored by histochemical staining and RT-PCR, respectively.
Activity of Pr SOS1-AB and Pr SOS1-D promoters in the transgenic Arabidopsis plants
GUS activity was analysed in the transgenic Arabidopsis seedlings carrying the Pr SOS1-AB –gusA (AB6 line) or Pr SOS1-AB –gusA constructs (D9 line) grown in normal MS medium and at different developmental stages. At an early developmental stage (4 days old), clear GUS activity was detected only in the young leaves of these two lines. Histochemical staining of 8- and 12-day-old AB6 and D9 seedlings did not enable the detection of any GUS activity. However, the GUS activity was different between AB6 and D9 lines grown for 20 or 30 days in normal MS medium. Indeed, at these two seedling developmental stages, no GUS activity was observed in the D9 line. Concerning the AB6 line, GUS activity was detected slightly in the roots and leaves of 20-day-old seedlings. In mature plants (30 days old), blue staining was observed in leaves and not in the stems, roots or flowers (Fig. 3). Taken together, these results showed that the Pr SOS1-AB and Pr SOS1-D promoters are active in Arabidopsis plants; their activities were almost the same only at early developmental stages, showing them to be age-dependent and organ-specific promoters.
a GUS activity in transgenic Arabidopsis plants carrying the pCAMBIA1391Z–Pr SOS1-AB –gusA construct (AB6 line) or the pCAMBIA1391Z–Pr SOS1-D –gusA construct (D9 line) at different developmental stages (4, 8, 12, 20 and 30 days old). Wt: non-transformed plants, 35S: transgenic Arabidopsis plants carrying pCAMBIA1301 (positive control). b Binocular observation of GUS staining in flowers of transgenic Arabidopsis plants (AB6 and D9) and of the control plants (335S and Wt)
Pr SOS1-AB is an abiotic stress-inducible promoter at different development stages
In a previous work, it has been demonstrated that the expression of the bread wheat TaSOS1 gene is induced by salt stress (Xu et al. 2008). Here, the activities of the two wheat SOS1 promoters were analysed in Arabidopsis under different abiotic stresses and at different developmental stages. For this, GUS activity was examined in the two transgenic AB6 and D9 lines grown for 8, 20 and 30 days in MS agar medium and then transferred to the same medium containing NaCl, mannitol or ABA for 2 days. At an early developmental stage (8 days old), the application of NaCl, mannitol or ABA produced a blue staining in the leaves of the two transgenic Arabidopsis AB6 and D9 lines. Contrary to the D9 line, GUS activity was observed also in the roots of the AB6 line challenged with NaCl, mannitol or ABA (Fig. 4a). Unlike the D9 line, abiotic and hormonal stresses produced a blue staining in a different part of 20-day-old seedlings of the AB6 line. Indeed, NaCl induced deeper blue staining in leaves and roots compared to ABA and mannitol treatment (Fig. 4b). Moreover, NaCl and ABA led to a strong GUS staining in the root tips of the AB6 line compared to mannitol treatment (Fig. 4c). In mature plants (30 days old), blue staining was observed in the stem and root of the AB6 line challenged with NaCl, mannitol or ABA stresses. In the case of the D9 line, no GUS activity was observed after the application of these different abiotic stresses (Fig. 4d). These data show that, contrary to Pr SOS1-D , the activity of Pr SOS1-AB is induced by different abiotic stresses independently of the vegetative stage of the transgenic Arabidopsis plants.
Histochemical GUS staining of (a) 8-, (b) 20- and (d) 30-day-old transgenic seedlings placed for 2 days in MS agar medium containing 100 mM NaCl, 100 mM mannitol or 20 μM ABA. The arrows indicate the blue colouration in 8-day-old seedling roots. c Binocular observation of GUS staining in the root tips of 20-day-old transgenic seedlings subjected or not to different stress conditions. AB6 and D9: Arabidopsis carrying the pCAMBIA1391Z–Pr SOS1-AB –gusA or pCAMBIA1391Z–Pr SOS1-D –gusA constructs, respectively
Relation between stress treatment and gusA expression in transgenic Arabidopsis carrying the Pr SOS1-AB –gusA construct
To further ascertain the induced activity of the Pr SOS1-AB promoter in 20- and 30-day-old seedlings of the AB6 and AB7 lines, RT-PCR was used to detect the presence of gusA transcripts under abiotic (salt and osmotic) and hormonal stress (ABA) conditions. Compared to the non-treated plants, an accumulation of gusA transcripts was detected and found to be up-regulated by NaCl, ABA and mannitol treatments in whole 20-day-old seedlings and in the shoots and roots of 30-day-old AB6 seedlings (Fig. 5). These results are in complete agreement with those of the histochemical staining assays.
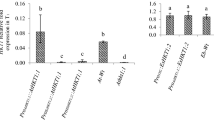
Analysis of gusA expression in the whole plants of 20-day-old seedlings (a) and in the shoots and roots of 30-day-old seedlings (b), challenged with various abiotic stresses [100 mM NaCl (Na), 100 mM mannitol (M) or 20 μM ABA (a)] for 2 days. NT: non-treated plants. 35S: positive control plant. Wt: non-transformed plant. AB6: the transgenic Arabidopsis plant carrying the pCAMBIA1391Z–Pr SOS1-AB –gusA construct. A 0.5-Kb β-tubulin gene fragment was amplified by RT-PCR as an internal control. (−): negative control without cDNA. The histograms correspond to the band densities in the gels, which are expressed in arbitrary units calculated by the Gel DocXR software. The standard errors were determined from three independent biologic replicates
Discussion
The SOS1 protein has been associated to salt stress response, and loss of function of the SOS1 gene results in the hypersensitivity of Arabidopsis to NaCl (Wu et al. 1996). After the description of the first SOS1 gene in Arabidopsis (Shi et al. 2000), other SOS1 genes have been identified in different plants, such as rice, wheat and tomato (Martínez-Atienza et al. 2007; Olías et al. 2009; Xu et al. 2008; Feki et al. 2011). So far, only the expression pattern of Arabidopsis thaliana AtSOS1 and Salicornia brachiata SbSOS1 have been examined using the SOS1 promoter GUS (Shi et al. 2002; Goyal et al. 2013). In this work, we report the expression of Triticum aestivum TaSOS1 in Arabidopsis plants under various abiotic stresses (salt, drought and hormonal). It is worth noting that there is a single SOS1 locus in the genome of the tetraploid durum wheat (Triticum durum L. subsp. durum) (AA BB) (Feki et al. 2011). Thus, it seems that there are at least two SOS1 copies in the hexaploid bread wheat (Triticum aestivum) (AA BB DD), one localised on genomes A or B, and the other one on genome D. Here, we report the isolation of two novel promoter regions Pr SOS1-AB and Pr SOS1-D of the TaSOS1 gene by screening the genomic library of bread wheat (Triticum aestivum). The cloned Pr SOS1-AB and Pr SOS1-D sequences exhibited homology with the 5′ end sequence of SOS1 alleles, which are localised on genomes A or B, and on genome D, respectively. Thus, the isolated sequences were in the upstream region of the two wheat SOS1 alleles.
Sequence analysis of these two promoter regions revealed the presence of the same potential abiotic stress responsive cis elements, transcription factor-binding sites such as DOF and WRKY, ABA responsive element (ABRE) and GT elements (Table 3). DOF factors play an important role in the genes induced by plant hormones and stress signals (Yanagisawa and Sheen 1998). WRKY is required for positive and negative regulatory behaviours of ABA signalling (Zhang et al. 2004). An ABA-responsive cis-acting element named ABRE (C/GACGTGGC) is present in the promoter regions of many ABA-inducible genes. ABA plays a central role as a signalling molecule in stress regulatory networks in plants. Several trans- and cis-acting regulatory elements have been characterised that function in ABA-dependent and/or ABA-independent manners, leading to stress-inducible gene expression. An ABRE-like sequence (ACGTG) is required for the aetiolation-induced expression of the erd1 (early responsive to dehydration) gene in Arabidopsis (Simpson et al. 2003). In addition, ABRE is the most conserved of the dehydration-inducible promoters in Arabidopsis thaliana, rice and soybean, suggesting that the transcriptional regulation of dehydration-inducible genes is similar among these species, with an ABRE-dependent transcriptional pathway (Maruyama et al. 2012). The GT-1 motif plays a role in pathogen- and salt-induced expression of the SCaM-4 promoter (Park et al. 2004). In the same way, in silico analysis of the SOS1 upstream sequence from A. thaliana, Vitis vinifera and Oryza sativa revealed the presence of almost the same cis-regulatory elements (Goyal et al. 2013; Ji et al. 2013).
Many SOS1 genes were isolated and then characterised in mutant yeast (Shi et al. 2002; Martínez-Atienza et al. 2007; Olías et al. 2009; Oh et al. 2009; Feki et al. 2011). So far, only the regulatory mechanism of the Arabidopsis SOS1 gene has been analysed. In Arabidopsis, SOS1 mRNA is unstable under normal growth conditions, and its stability is substantially increased upon salt stress treatment (Shi et al. 2003). Chung et al. (2008) identified the cis element in SOS1 mRNA responsible for the stability-regulation and demonstrated that stress-induced SOS1 mRNA stability is mediated by ROS. As an initial step towards understanding regulatory mechanisms controlling wheat SOS1 gene expression, the expression pattern of the Pr SOS1-AB and Pr SOS1-D promoters was investigated using a gusA reporter gene system in transgenic Arabidopsis seedlings grown under normal or stressed conditions. In a previous work, it has been demonstrated that monocot gene promoters are functional in a dicots plant (Liu et al. 2003; Iwamoto et al. 2004; Tittarelli et al. 2007). In this study, histochemical staining revealed that the monocotyledonous Pr SOS1-AB and Pr SOS1-D promoters are active in this heterologous transgenic system. Moreover, their activities were similar only at early developmental stages (4, 8 and 12 days old). To our knowledge, this is the first isolation and characterisation of two wheat SOS1 promoters. Histochemical staining revealed the ability of these promoters to direct GUS expression with age-dependent and organ-specific patterns. Indeed, GUS activity was detected only in young leaves of transgenic Arabidopsis carrying the Pr SOS1-D -gusA construct (D lines). Despite the presence of gusA mRNA in D lines, GUS activity was never detected in roots and leaves. In an attempt to explain these results, one may speculate that GUS activity is undetectable because of the absence of GUS or an inactive form of the enzyme. Concerning the transgenic Arabidopsis carrying the Pr SOS1-AB -gusA construct (AB lines), GUS activity was absent only in 8- and 12-day-old seedlings.
The over-accumulation of stress regulators by using strong constitutive promoters has, in many cases, improved stress tolerance (Hsieh et al. 2002; Ito et al. 2006). Nevertheless, this enhanced stress tolerance is sometimes conferred at the expense of plant development and growth (Xu et al. 2006). The use of stress-inducible promoters is expected to be optimised for driving candidate abiotic stress tolerance genes (Rai et al. 2009; Zhu et al. 2010; Ben Saad et al. 2011). In this study, we showed that the two wheat TaSOS1 promoters are induced by salt, drought and hormonal stresses. This is consistent with the presence in these promoter regions of the different cis-regulatory elements related to various stresses (Table 3). Contrary to our result, it has been demonstrated that the AtSOS1 and SbSOS1 promoters are induced by salt stress and not by ABA or cold stress (Shi et al. 2000; Goyal et al. 2013). Our data showed that the induction by these different abiotic stresses was different between these two TaSOS1 promoters. Indeed, contrary to the Pr SOS1-D promoter, Pr SOS1-AB is induced by NaCl, ABA and mannitol at different developmental stages (8, 20 and 30 days old). This finding was supported by RT-PCR analysis of steady-state gusA mRNA levels in transgenic Arabidopsis, which revealed that the reporter gene transcription under the control of Pr SOS1-AB was highly induced by the different abiotic stresses. Moreover, GUS activity was significant at root tips under NaCl and ABA stresses. These data are in good agreement with the findings of Shi et al. (2002), which showed that Arabidopsis AtSOS1 is expressed in root epidermal cells, particularly at the root tip and in cells bordering the vascular tissue. Despite the presence of the same cis-regulatory elements in the two wheat SOS1 promoter regions, their activities were different under various abiotic stress conditions. This suggestion could be explained by the difference in the number of cis-regulatory elements within the two isolated promoter regions or by the presence of some cis-regulatory elements in the nucleotide sequences which are absent in the Pr SOS1-D promoter region.
This is the first report on the isolation and characterisation of wheat TaSOS1 promoters (Pr SOS1-AB and Pr SOS1-D ) from Triticum aestivum. Histochemical GUS transient analysis validated that Pr SOS1-AB and Pr SOS1-D are functional, age-dependent and organ-specific promoters. Interestingly, in this heterologous system, only the Pr SOS1-AB promoter is induced by NaCl, mannitol and ABA at different developmental stages. These results will lead to more interest in the Pr SOS1-AB promoter, because it could be an attractive candidate promoter for the development of transgenic crop plants. Moreover, Pr SOS1-AB may be an effective and desirable promoter for controlling stress tolerance candidate genes in terms of driving low constitutive transgene expression under normal conditions and high induction in response to salt (NaCl), hormonal (ABA) and osmotic (mannitol) stresses. For these reasons, using the Pr SOS1-AB promoter could avoid potential harmful effects related to an over-expression of the target gene under the control of constitutive promoters in transgenic plants.
References
Behnam B, Kikuchi A, Celebi-Toprak F, Yamanaka S, Kasuga M, Yamaguchi-Shinozaki K, Watanabe KN (2006) The Arabidopsis DREB1A gene driven by the stress-inducible rd29A promoter increases salt-stress tolerance in proportion to its copy number in tetrasomic tetraploid potato (Solanum tuberosum). Plant Biotechnol 23:169–177
Ben Saad R, Ben Romdhan W, Zouari N, Azaza J, Mieulet D, Verdeil JL, Guiderdoni E, Hassairi A (2011) Promoter of the AlSAP gene from the halophyte grass Aeluropus littoralis directs developmental-regulated, stress-inducible, and organ-specific gene expression in transgenic tobacco. Transgenic Res 20:1003–1018. doi:10.1007/s11248-010-9474-6
Chen H, Nelson RS, Sherwood JL (1994) Enhanced recovery of transformants of Agrobacterium tumefaciens after freeze–thaw transformation and drug selection. Biotechniques 16:664–670
Chung JS, Zhu JK, Bressan RA, Hasegawa PM, Shi H (2008) Reactive oxygen species mediate Na+-induced SOS1 mRNA stability in Arabidopsis. Plant J 53:554–565. doi:10.1111/j.1365-313X.2007.03364.x
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
Cosentino C, Fischer-Schliebs E, Bertl A, Thiel G, Homann U (2010) Na+/H+ antiporters are differentially regulated in response to NaCl stress in leaves and roots of Mesembryanthemum crystallinum. New Phytol 186:669–680. doi:10.1111/j.1469-8137.2010.03208.x
Feki K, Quintero FJ, Pardo JM, Masmoudi K (2011) Regulation of durum wheat Na+/H+ exchanger TdSOS1 by phosphorylation. Plant Mol Biol 76:545–556. doi:10.1007/s11103-011-9787-8
Goyal E, Singh RS, Kanika K (2013) Isolation and functional characterization of Salt overly sensitive 1 (SOS1) gene promoter from Salicornia brachiata. Biol Plant 57:465–473. doi:10.1007/s10535-013-0309-1
Guo Y, Halfter U, Ishitani M, Zhu JK (2001) Molecular characterization of functional domains in the protein kinase SOS2 that is required for plant salt tolerance. Plant Cell 13:1383–1400. doi:10.1105/TPC.010021
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res 27:297–300. doi:10.1093/nar/27.1.297
Hsieh T-H, Lee J-T, Charng Y-Y, Chan M-T (2002) Tomato plants ectopically expressing Arabidopsis CBF1 show enhanced resistance to water deficit stress. Plant Physiol 130:618–626. doi:10.1104/pp.006783
Ishitani M, Liu J, Halfter U, Kim CS, Shi W, Zhu JK (2000) SOS3 function in plant salt tolerance requires N-myristoylation and calcium binding. Plant Cell 12:1667–1678. doi:10.1105/tpc.12.9.1667
Isidore E, Scherrer B, Bellec A, Budin K, Faivre-Rampant P, Waugh R, Keller B, Caboche M, Feuillet C, Chalhoub B (2005) Direct targeting and rapid isolation of BAC clones spanning a defined chromosome region. Funct Integr Genom 5:97–103. doi:10.1007/s10142-004-0127-9
Ito Y, Katsura K, Maruyama K, Taji T, Kobayashi M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of rice DREB1/CBF-type transcription factors involved in cold-responsive gene expression in transgenic rice. Plant Cell Physiol 47:141–153. doi:10.1093/pcp/pci230
Iwamoto M, Higo H, Higo K (2004) Strong expression of the rice catalase gene CatB promoter in protoplasts and roots of both a monocot and dicots. Plant Physiol Biochem 42:241–249. doi:10.1016/j.plaphy.2004.01.008
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Ji H, Pardo JM, Batelli G, Van Oosten MJ, Bressan RA, Li X (2013) The Salt Overly Sensitive (SOS) pathway: established and emerging roles. Mol Plant 6:275–286. doi:10.1093/mp/sst017
Kant S, Kant P, Raveh E, Barak S (2006) Evidence that differential gene expression between the halophyte, Thellungiella halophila, and Arabidopsis thaliana is responsible for higher levels of the compatible osmolyte proline and tight control of Na+ uptake in T. halophila. Plant Cell Environ 29:1220–1234. doi:10.1111/j.1365-3040.2006.01502.x
Kaplan B, Davydov O, Knight H, Galon Y, Knight MR, Fluhr R, Fromm H (2006) Rapid transcriptome changes induced by cytosolic Ca2+ transients reveal ABRE-related sequences as Ca2+-responsive cis elements in Arabidopsis. Plant Cell 18:2733–2748. doi:10.1105/tpc.106.042713
Karim S, Aronsson H, Ericson H, Pirhonen M, Leyman B, Welin B, Mäntylä E, Palva ET, Van Dijck P, Holmström KO (2007) Improved drought tolerance without undesired side effects in transgenic plants producing trehalose. Plant Mol Biol 64:371–386. doi:10.1007/s11103-007-9159-6
Kasuga M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2004) A combination of the Arabidopsis DREB1A gene and stress-inducible rd29A promoter improved drought- and low-temperature stress tolerance in tobacco by gene transfer. Plant Cell Physiol 45:346–350. doi:10.1093/pcp/pch037
Ke J, Choi JK, Smith M, Horner HT, Nikolau BJ, Wurtele ES (1997) Structure of the CAC1 gene and in situ characterization of its expression. The Arabidopsis thaliana gene coding for the biotin-containing subunit of the plastidic acetyl-coenzyme A carboxylase. Plant Physiol 113:357–365. doi:10.1104/pp.113.2.357
Konez C, Schell J (1986) The promoter of TL-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Gen Genet 204:383–396. doi:10.1007/BF00331014
Lacombe E, Van Doorsselaere J, Boerjan W, Boudet AM, Grima-Pettenati J (2000) Characterization of cis-elements required for vascular expression of the Cinnamoyl CoA Reductase gene and for protein–DNA complex formation. Plant J 23:663–676. doi:10.1046/j.1365-313x.2000.00838.x
Lam E, Chua NH (1989) ASF-2: a factor that binds to the cauliflower mosaic virus 35S promoter and a conserved GATA motif in cab promoters. Plant Cell 1:1147–1156. doi:10.1105/tpc.1.12.1147
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. doi:10.1093/nar/30.1.325
Liu J, Zhu JK (1998) A calcium sensor homolog required for plant salt tolerance. Science 280:1943–1945. doi:10.1126/science.280.5371.1943
Liu J, Ishitani M, Halfter U, Kim CS, Zhu JK (2000) The Arabidopsis thaliana SOS2 gene encodes a protein kinase that is required for salt tolerance. Proc Natl Acad Sci U S A 97:3730–3734. doi:10.1073/pnas.97.7.3730
Liu ZZ, Wang JL, Huang X, Xu WH, Liu ZM, Fang RX (2003) The promoter of a rice glycine-rich protein gene, Osgrp-2, confers vascular-specific expression in transgenic plants. Planta 216:824–833. doi:10.1007/s00425-002-0934-y
Martínez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu JK, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012. doi:10.1104/pp.106.092635
Maruyama K, Todaka D, Mizoi J, Yoshida T, Kidokoro S, Matsukura S, Takasaki H, Sakurai T, Yamamoto YY, Yoshiwara K, Kojima M, Sakakibara H, Shinozaki K, Yamaguchi-Shinozaki K (2012) Identification of cis-acting promoter elements in cold- and dehydration-induced transcriptional pathways in Arabidopsis, rice, and soybean. DNA Res 19:37–49. doi:10.1093/dnares/dsr040
Michiels A, Van den Ende W, Tucker M, Van Riet L, Van Laere A (2003) Extraction of high-quality genomic DNA from latex-containing plants. Anal Biochem 315:85–89. doi:10.1016/S0003-2697(02)00665-6
Mullan DJ, Colmer TD, Francki MG (2007) Arabidopsis–rice–wheat gene orthologues for Na+ transport and transcript analysis in wheat–L. elongatum aneuploids under salt stress. Mol Genet Genomics 277:199–212. doi:10.1007/s00438-006-0184-y
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. doi:10.1111/j.1399-3054.1962.tb08052.x
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120:621–623
Oh DH, Leidi E, Zhang Q, Hwang SM, Li Y, Quintero FJ, Jiang X, D’Urzo MP, Lee SY, Zhao Y, Bahk JD, Bressan RA, Yun DJ, Pardo JM, Bohnert HJ (2009) Loss of halophytism by interference with SOS1 expression. Plant Physiol 151:210–222. doi:10.1104/pp. 109.137802
Oh DH, Dassanayake M, Haas JS, Kropornika A, Wright C, d’Urzo MP, Hong H, Ali S, Hernandez A, Lambert GM, Inan G, Galbraith DW, Bressan RA, Yun DJ, Zhu JK, Cheeseman JM, Bohnert HJ (2010) Genome structures and halophyte-specific gene expression of the extremophile Thellungiella parvula in comparison with Thellungiella salsuginea (Thellungiella halophila) and Arabidopsis. Plant Physiol 154:1040–1052. doi:10.1104/pp.110.163923
Olías R, Eljakaoui Z, Li J, De Morales PA, Marín-Manzano MC, Pardo JM, Belver A (2009) The plasma membrane Na+/H+ antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of Na+ between plant organs. Plant Cell Environ 32:904–916. doi:10.1111/j.1365-3040.2009.01971.x
Park HC, Kim ML, Kang YH, Jeon JM, Yoo JH, Kim MC, Park CY, Jeong JC, Moon BC, Lee JH, Yoon HW, Lee SH, Chung WS, Lim CO, Lee SY, Hong JC, Cho MJ (2004) Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol 135:2150–2161. doi:10.1104/pp.104.041442
Pino MT, Skinner JS, Park EJ, Jeknić Z, Hayes PM, Thomashow MF, Chen TH (2007) Use of a stress inducible promoter to drive ectopic AtCBF expression improves potato freezing tolerance while minimizing negative effects on tuber yield. Plant Biotechnol J 5:591–604. doi:10.1111/j.1467-7652.2007.00269.x
Planchais S, Perennes C, Glab N, Mironov V, Inzé D, Bergounioux C (2002) Characterization of cis-acting element involved in cell cycle phase-independent activation of Arath;CycB1;1 transcription and identification of putative regulatory proteins. Plant Mol Biol 50:111–127
Qiu QS, Guo Y, Dietrich MA, Schumaker KS, Zhu JK (2002) Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci U S A 99:8436–8441. doi:10.1073/pnas.122224699
Quan R, Lin H, Mendoza I, Zhang Y, Cao W, Yang Y, Shang M, Chen S, Pardo JM, Guo Y (2007) SCABP8/CBL10, a putative calcium sensor, interacts with the protein kinase SOS2 to protect Arabidopsis shoots from salt stress. Plant Cell 19:1415–1431. doi:10.1105/tpc.106.042291
Quintero FJ, Ohta M, Shi H, Zhu JK, Pardo JM (2002) Reconstitution in yeast of the Arabidopsis SOS signaling pathway for Na+ homeostasis. Proc Natl Acad Sci U S A 99:9061–9066. doi:10.1073/pnas.132092099
Quintero FJ, Martinez-Atienza J, Villalta I, Jiang X, Kim WY, Ali Z, Fujii H, Mendoza I, Yun DJ, Zhu JK, Pardo JM (2011) Activation of the plasma membrane Na/H antiporter Salt-Overly-Sensitive 1 (SOS1) by phosphorylation of an auto-inhibitory C-terminal domain. Proc Natl Acad Sci U S A 108:2611–2616. doi:10.1073/pnas.1018921108
Rai M, He C, Wu R (2009) Comparative functional analysis of three abiotic stress-inducible promoters in transgenic rice. Transgenic Res 18:787–799. doi:10.1007/s11248-009-9263-2
Sanders D (2000) Plant biology: the salty tale of Arabidopsis. Curr Biol 10:R486–R488. doi:10.1016/S0960-9822(00)00554-6
Shi H, Ishitani M, Kim C, Zhu JK (2000) The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter. Proc Natl Acad Sci U S A 97:6896–6901. doi:10.1073/pnas.120170197
Shi H, Quintero FJ, Pardo JM, Zhu JK (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477. doi:10.1105/tpc.010371
Shi H, Lee BH, Wu SJ, Zhu JK (2003) Overexpression of a plasma membrane Na+/H+ antiporter gene improves salt tolerance in Arabidopsis thaliana. Nat Biotechnol 21:81–85. doi:10.1038/nbt766
Simpson SD, Nakashima K, Narusaka Y, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Two different novel cis-acting elements of erd1, a clpA homologous Arabidopsis gene function in induction by dehydration stress and dark-induced senescence. Plant J 33:259–270. doi:10.1046/j.1365-313X.2003.01624.x
Sinha NR, Williams RE, Hake S (1993) Overexpression of the maize homeo box gene, KNOTTED-1, causes a switch from determinate to indeterminate cell fates. Genes Dev 7:787–795. doi:10.1101/gad.7.5.787
Taji T, Seki M, Satou M, Sakurai T, Kobayashi M, Ishiyama K, Narusaka Y, Narusaka M, Zhu JK, Shinozaki K (2004) Comparative genomics in salt tolerance between Arabidopsis and Arabidopsis-related halophyte salt cress using Arabidopsis microarray. Plant Physiol 135:1697–1709. doi:10.1104/pp.104.039909
Tang RJ, Liu H, Bao Y, Lv QD, Yang L, Zhang HX (2010) The woody plant poplar has a functionally conserved salt overly sensitive pathway in response to salinity stress. Plant Mol Biol 74:367–380. doi:10.1007/s11103-010-9680-x
Tester M, Davenport R (2003) Na+ tolerance and Na+ transport in higher plants. Ann Bot 91:503–527. doi:10.1093/aob/mcg058
Tittarelli A, Milla L, Vargas F, Morales A, Neupert C, Meisel L, Salvo-G H, Peñaloza E, Muñoz G, Corcuera L, Silva H (2007) Isolation and comparative analysis of the wheat TaPT2 promoter: identification in silico of new putative regulatory motifs conserved between monocots and dicots. J Exp Bot 58:2573–2582. doi:10.1093/jxb/erm123
Urao T, Yamaguchi-Shinozaki K, Urao S, Shinozaki K (1993) An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5:1529–1539. doi:10.1105/tpc.5.11.1529
Wang X, Yang R, Wang B, Liu G, Yang C, Cheng Y (2010) Functional characterization of a plasma membrane Na+/H+ antiporter from alkali grass (Puccinellia tenuiflora). Mol Biol Rep 38:4813–4822. doi:10.1007/s11033-010-0624-y
Wu SJ, Ding L, Zhu JK (1996) SOS1, a genetic locus essential for salt tolerance and potassium acquisition. Plant Cell 8:617–627. doi:10.1105/tpc.8.4.617
Wu Y, Ding N, Zhao X, Zhao M, Chang Z, Liu J, Zhang L (2007) Molecular characterization of PeSOS1: the putative Na+/H+ antiporter of Populus euphratica. Plant Mol Biol 65:1–11. doi:10.1007/s11103-007-9170-y
Wu Y, Zhou H, Que Y-X, Chen R-K, Zhang M-Q (2008) Cloning and identification of promoter Prd29A and its application in sugarcane drought resistance. Sugar Technol 10:36–41. doi:10.1007/s12355-008-0006-0
Xiong L, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14:S165–S183. doi:10.1105/tpc.000596
Xu R, Zhao H, Dinkins RD, Cheng X, Carberry G, Li QQ (2006) The 73 kD subunit of the cleavage and polyadenylation specificity factor (CPSF) complex affects reproductive development in Arabidopsis. Plant Mol Biol 61:799–815. doi:10.1007/s11103-006-0051-6
Xu H, Jiang X, Zhan K, Cheng X, Chen X, Pardo JM, Cui D (2008) Functional characterization of a wheat plasma membrane Na+/H+ antiporter in yeast. Arch Biochem Biophys 473:8–15. doi:10.1016/j.abb.2008.02.018
Yanagisawa S, Schmidt RJ (1999) Diversity and similarity among recognition sequences of Dof transcription factors. Plant J 17:209–214. doi:10.1046/j.1365-313X.1999.00363
Yanagisawa S, Sheen J (1998) Involvement of maize DOF zinc finger proteins in tissue-specific and light-regulated gene expression. Plant Cell 10:75–89. doi:10.1105/tpc.10.1.75
Zhang ZL, Xie Z, Zou X, Casaretto J, Ho TH, Shen QJ (2004) A rice WRKY gene encodes a transcriptional repressor of the gibberellin signaling pathway in aleurone cells. Plant Physiol 134:1500–1513. doi:10.1104/pp.103.034967
Zhu J-K (2000) Genetic analysis of plant salt tolerance using Arabidopsis. Plant Physiol 124:941–948. doi:10.1104/pp.124.3.941
Zhu J-K, Liu J, Xiong L (1998) Genetic analysis of salt tolerance in Arabidopsis: evidence for a critical role of potassium nutrition. Plant Cell 10:1181–1191. doi:10.1105/tpc.10.7.1181
Zhu LP, Yu Z, Zou CX, Li QL (2010) Plant stress-inducible promoters and their function. Yi Chuan 32:229–234
Acknowledgements
This study was supported by a grant from the Ministry of Higher Education, Scientific Research and Information and Communication Technology of Tunisia. The authors thank Dr. Boulos Chalhoub from the Laboratory of Genome Organization (URGV-INRA, France) for providing the genomic library, Cecile Huneau for her technical assistance and Mohammed Najib Saidi for his help in the quantitative analysis.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Feki, K., Brini, F., Ben Amar, S. et al. Comparative functional analysis of two wheat Na+/H+ antiporter SOS1 promoters in Arabidopsis thaliana under various stress conditions. J Appl Genetics 56, 15–26 (2015). https://doi.org/10.1007/s13353-014-0228-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-014-0228-7