Abstract
In the late 1990s, a novel Botryosphaeria-like fungal pathogen was observed causing a disease on wheat in Queensland, characterised as white grain disorder (WGD). In recent years, this disease has sporadically appeared across the eastern states of Australia. In this study, internal transcribed spacer (ITS) region sequences were used to compare these fungi to other Botryosphaeriaceae spp. to show that they should be reclassified as members of the Eutiarosporella genus. Using a small population of WGD isolates, we built a three-loci maximum likelihood tree, using ITS, β-tubulin, and Elongation Factor1-α sequences to show that there are three separate Eutiarosporella spp. found in infected grain. This multigene tree, with the support of phenotypic differences between clades observed in vitro, show that that the causal agents of WGD should be delimited into three divergent species; Eutiarosporella tritici-australis, Eutiarosporella darliae, and Eutiarosporella pseudodarliae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In Australia, wheat is an economically significant industry that is worth over $4.5 billion dollars per year (Murray and Brennan 2009), a significant amount of which is exported to foreign markets. As an important export crop, understanding the abiotic and biotic factors that result in yield loss is critical. In the Australian wheat industry the major source of financial loss from disease is attributed to fungal pathogens (Murray and Brennan 2009). These losses are caused by reduction in overall yield due to disease or by reduction of overall grain quality and thus final sale price of the harvested grain.
In the late 1990s, a fungal pathogen of wheat was observed causing mummification of wheat grain in Queensland (Platz 2011). This fungus was initially identified as Botryosphaeria zeae (Platz 2011; Wildermuth et al. 2001), the pathogen responsible for grey ear rot in maize (De Leon et al. 2004). This disease, which shrivels and discolours wheat grain, became known as white grain disorder (WGD) (Platz 2011). Toxicity assays performed, showed this grain poses no detectable human health risk (Kopinski and Blaney 2010). Symptoms in wheat consistent with those of WGD were first recorded in Victoria (Vic) and South Australia (SA) in 2010 (Evans 2013; Penfold 2011). Since 2010, this disease has appeared sporadically across the Eastern Australian states.
To determine the causal agent(s) of WGD, the ITS region was sequenced from collected isolates and phylogenetic trees constructed. This analysis revealed two distinct clades (Stephen Neate, unpublished data). Most of the isolates from QLD and NSW were identified and labelled as B. zeae. B. zeae was also found in infected grain from VIC and SA, however the majority of isolates collected from these states were differentiable from B. zeae and were designated as Clade 1 (Stephen Neate, unpublished data). The geographic localisation of Clade 1 in the Southern states and presence of B. zeae across all Eastern states, suggested that there was not a single species causing mummification of wheat grains in Australia, but instead a complex of closely related species. To date, however, no comprehensive analysis has been conducted that would enable growers or pathologists to accurately identify this disease in their harvested grain.
Species of Botryosphaeriaceae are widespread across the world, and are observed as both pathogens and endophytes (or potential pathogens) on a wide range of plant hosts (Sakalidis et al. 2011a, 2013; Alvarez-Loayza et al. 2011; Slippers and Wingfield 2007). In Australasia, a selection of these includes conifers (Slippers et al. 2005), eucalypts (Barber et al. 2005; Slippers et al. 2004b), mangoes (Sakalidis et al. 2011a), grapes (Taylor et al. 2005) and baobabs (Sakalidis et al. 2011a). Closely related fungal species, or recently diverged species, can remain morphologically similar for a long period of time (Luque et al. 2005; Slippers et al. 2004c, 2009; Sakalidis et al. 2011b). Accordingly, it can be a flawed exercise to delimit species of Botryosphaeriaceae with the use of morphological observations alone (Phillips et al. 2013). Analyses within the fungal kingdom, and more specifically within the Botryosphaeriaceae family, have shown the power of using multiple gene loci comparisons, complemented with morphological and observations, to perform species delimitation (Aoki et al. 2014; Dettman et al. 2003; McDonald et al. 2012; Jami et al. 2014; Slippers et al. 2004a, c; Zhou et al. 2014).
Modern genetic data is vital for continued examination and differentiation of fungal pathogens (Hyde et al. 2010). Currently, there are no sequenced genes of B. zeae present on GenBank or similar publicly available gene databases. However, since the original observation of wheat-infecting B. zeae in the late 1990s, there has been a significant increase in the available gene sequence data for various Botryosphaeriaceae species. Many genera of this family have been described with the support of multiple genetic loci. This has lead to a significant refinement in the resolution of these genera (Phillips et al. 2013; Slippers et al. 2013). Previously, genera that were poorly resolved, such as the genus of Tiarosporella (now Tiarosporella and Eutiarosporella (Crous et al. 2015)), have now been characterised and their genetic relationship to other Botryosphaeriaceae described (Phillips et al. 2013; Slippers et al. 2013; Crous et al. 2006, 2015).
Using this increase in the available genetic data to perform phylogenetic analyses, we sought to verify the validity of the initial placement of these fungi within the Botryosphaeriaceae genera. In addition, we used phylogenetic analyses and morphological evidence displaying differences between isolates to assess whether a complex of separate but closely related species cause WGD.
Materials and methods
Isolate sampling and primary culturing
Isolates were provided by the Queensland Department of Primary Industries (QLD DPI) and the South Australian Research and Development Institute (SARDI). Additional isolates were isolated on tap water agar (12 g agar/L) (TWA) from WGD grain samples provided by the QLD DPI and SARDI.
All isolates were sub-cultured on petri dishes of potato dextrose agar (PDA) (Difco, USA) with mycelia-covered plugs, and incubated at 23 °C under 12 h light/dark cycles. When isolates showed growth fatigue after numerous sub-culturing events, mycelia from 20 % glycerol stocks, stored at −80 °C, were plated onto PDA. Morphological characterisation on PDA was performed by sub-culturing mycelia-covered plugs from single-conidia cultures. For agar media inoculation, pycnidia were crushed and diluted in 1 mL of sterile water. This spore solution was filtered through cotton wool in a 1 mL syringe to decrease the amount of superfluous mycelial fragments. The filtered spore solution was inoculated over the surface of an agar plate. Plates were left overnight for conidia to germinate. Germinating conidia, forming single colonies, were observed under a dissecting microscope and picked from the agar with a toothpick and sub-cultured to new PDA plates. Mycelial measurements were performed at 400× magnification.
Induction of asexual reproduction and microscopy of conidia
Asexual reproduction was induced on oat media agar (OMA) as described by Mead et al. (2013) and on recommendation from SARDI for WGD fungi (without wheat-extract) (Mead et al. 2013). OMA has been previously shown to be appropriate for the induction of pycnidia in Botryosphaeriaceae spp. (Vasconcelos et al. 2001). Plugs of mycelia-covered PDA were transferred to petri dishes of OMA and left to grow in the dark at 23 °C for 5 days. The surface mycelia were scraped from these plates with a scalpel and 2 mL of sterile water was scraped over the surface. The plates were incubated at 23 °C under constant near u.v. light for 5–10 days until conidia-bearing pycnidia developed. Individual pycnidia were cut out from the agar surface and crushed in a drop of water between a microscope slide and coverslip. The conidia were photographed under 400× magnification with an EOS Rebel T3i camera (Canon, Japan). Fifty conidia of each type specimen were analysed to obtain the conidia dimension measurements. To allow conidia to germinate, pycnidia were taken from the agar plate with a sterile toothpick, crushed with a pestle in a 1.5 ml tube (Eppendorf, Germany) with 20 μL of sterile water, and incubated at room temperature for 3 h.
Secondary culturing and growth analyses
Phenotypic variation was observed on six minimal media (MM) and four OMA media. The base minimal media (25 mMol of a carbon source, 10 mM of a nitrogen source, 10 mL/L of 100× trace element stock) was altered to incorporate varied carbon and nitrogen sources. Glucose, fructose, sucrose, and sorbitol were interchanged as the carbon source (with glutamate as the nitrogen source) and glutamate, glutamine and sodium nitrate were interchanged as the nitrogen source (with sucrose as the carbon source). In addition to standard OMA, OMA with 60 mL of V8 juice (V8-OMA) and OMA with 10 mMol gamma-aminobutyric acid (GABA) (GABA-OMA). All media contained 12 g Bacto agar/L (Difco, AUS).
Plugs of mycelia from single-conidia colonies (described above) were sub-cultured to appropriate media for growth analyses. Inoculated MM and OMA plates were grown under 12 h light/dark cycles at 23 °C. 150 mL of liquid MM was inoculated in a 250 mL flask, and shaken at 140 rpm, under 12 h light/dark cycles at 23 °C. Inoculated PDA plates were grown in full light for 15 h at 23 °C to ensure all isolates were growing healthily. Isolates were then rapped in metal-foil and grown in the dark at 16, 23, 28 and 30 °C, respectively, for effect of temperature on growth to be studied.
Genome sequencing and assembly
DNA was extracted from potato dextrose broth (PDB) (Edwards, USA) inoculated separately with isolate 153 (Clade 1, South Australia), isolate V4B6 (B. zeae, South Australia), and isolate 2G6 (B. zeae, Queensland). DNA was extracted using a QIAGEN DNeasy Plant Mini Kit (QIAGEN, Netherlands). Illumina paired-end sequencing was performed using a HiSeq™ 2000 (Illumina, USA) at the John Curtain School of Medical Research, Australian National University (ANU). The resulting sequenced reads were quality trimmed using Trimmomatic v0.27 (Lohse et al. 2012) and assembled de novo with SPAdes v2.5.0, with –k mer values 21, 33, 55, and 77 (Bankevich et al. 2012).
DNA extraction and PCR
DNA was extracted from mycelia scraped off previously inoculated PDA plates. A QIAGEN DNeasy Plant Mini Kit (QIAGEN, Netherlands) was used to extract DNA. Universal fungal ITS1 and ITS4 primers were used to amplify the ITS region (White et al. 1990). Primers for β-tubulin and elongation-factor 1-α (EF1-α) regions were designed using Vector NTI® (Life Technologies, USA) (Supplementary Table 1). A local BLAST+ tools search using homologous sequences from Botryosphaeria dothidea (Table 1) (Altschul et al. 1990) was performed to locate the homologous ITS, β-tubulin, and EF1-α regions on the genomes of the four sequenced isolates.
Sequence alignment and phylogenetic analyses
All sequences (ITS, β-tubulin, EF1-α) were aligned with MEGA 5.2.2 (Tamura et al. 2011) and Geneious 7.0.6 (Biomatters, New Zealand). Regions of poor alignment were trimmed manually where necessary. Congruence between the three individual gene trees was determined using Concaterpiller 1.8a (Leigh et al. 2008). Phylogenetic analyses were performed using RaxML version 7.2.8 (Stamatakis 2006) with the ‘-f a’ rapid bootstrap function and 10,000 bootstrap replicates. A complete list of all species and accession numbers are listed in (Table 2). The resulting trees were plotted with FigTree v1.4.0 (http://tree.bio.ed.ac.uk/software/figtree/).
Results
Isolate sampling and phylogenetic analyses
Eleven isolates of white grain disorder fungi sourced from Queensland and South Australia were used in this study. ITS phylogenies performed by the QLD DPI have previously shown that there were likely two clades of WGD fungi (data not shown). The genomes of three isolates, isolate 153 (Clade 1), isolate V4B6 (B. zeae), and isolate 2G6 (B. zeae), were sequenced using Illumina paired-end sequencing. ITS sequences in these genomes were used as templates to assign a further eight isolates of WGD fungi into Clade 1 or B. zeae (Supplementary Fig. 1). The ITS region used was 367 bp in length and a maximum likelihood tree was generated using RAXML. This analysis was rooted with ITS sequences of Botryosphaeria agaves. According to this phylogeny, a total of four isolates resolved in Clade 1, and seven into B. zeae. B. zeae isolates separated into two independent clades, and will be referred to throughout the text as B. zeae and B. zeae Clade 2.
Phylogenetic analyses
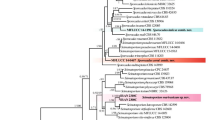
A type specimen was chosen to represent each separate clade of WGD fungi; Isolate 153 for Clade 1, Isolate 2E2 for B. zeae, and Isolate HR599118 for B. zeae Clade 2. The ITS sequences of these isolates of WGD fungi were compared to members of Botryosphaeriaceae. The resulting maximum likelihood tree was rooted to Endomelanconiopsis microspora, a close relative to the Botryosphaeriaeceae (Slippers et al. 2013). The partial ITS sequence, 381 bp, was used to construct the phylogenetic tree (Fig. 1). Of the sampled Botryosphaeriaceae spp., the WGD fungi form a well-supported clade with the Eutiarosporella genus. Based on this single locus, WGD fungi resolve closely with E. urbis-rosarum, E. tritici, T. madreeya. The genus of Tiarosporella was recently resolved into four genera (Tiarosporella, Marasasiomyces, Eutiarosporella, and Mucoharknessia) (Crous et al. 2015). Accordingly, a subsequent tree was performed comparing the ITS sequences (369 bp) from these new genera (and species) with all of the WGD isolates (Supplementary Fig. 2). All of the isolates still resolve between the Eutiarosporella spp.. Based on ITS sequence comparison, B. zeae Clade 2 isolates resolve closely with Mucoharknessia cortaderiae. However, the morphological descriptions of M. cartaderiae (Crous et al. 2015) are in stark contrast to those of the WGD isolates (see below), and so it is reasonable to conclude that these WGD isolates are still members of the Eutiarosporella and not Mucoharknessia.
Comparison of ITS sequences showed that WGD fungi resolved most closely with members of the Tiarosporella genus. A Maximum likelihood (ML) tree was performed comparing ITS sequences of the holotype isolates of WGD fungi to other Botryosphaeriaceae. The ML tree had 10,000 supporting bootstrap replicates, and was rooted to Endomelanconiopsis microspora. WGD fungi (black outlines) grouped within the Tiarosporella genus with high bootstrap support. Grey translucent boxes delineate the genera listed on the right
Multi-loci phylogenetic analyses
To further clarify the phylogenetic relationship both within clades of WGD fungi, and between WGD fungi and other Eutiarosporella spp., a phylogenetic analysis using ITS, β-tubulin, and EF1-α sequences was performed (Fig. 2). These loci were selected due to their prevalence in other studies that compared Botryosphaeriaceae spp. (Slippers et al. 2004a, b, c, 2005; Luque et al. 2005; Jami et al. 2012, 2013). Publically available sequence data for each of the three loci from Eutiarosporella spp., E. urbis-rosarum, T. graminis (now Marasasiomyces karoo), and E. tritici were used for comparison. Congruence between the three individual gene trees was determined using Concaterpiller 1.8a (Leigh et al. 2008), which produced a single concatenated set of the three loci, 1016 bp in length. The concatenated multi-gene phylogeny showed WGD fungi resolve most closely with E. tritici. Wheat-infecting B. zeae and B. zeae Clade 2 were separated both from each other and from Clade 1 with strong support (bootstrap values of 100). On the basis of these three loci, E. tritici, B. zeae and B. zeae Clade 2 branch separately to Clade 1, which all resolve on a monophyletic branch. A notable difference in the alignment of B. zeae, and B. zeae Clade 2 with the other species, is a 37–38 bp deletion in the EF1-α gene (Supplementary Fig. 3).
A comparison of ITS, β-tubulin, EF1-α from WGD fungi and Tiarosporella spp. distinctly separates the three clades of WGD fungi. A Maximum likelihood (ML) tree shows the genetic relationship between the clades of WGD fungi and with other Tiarosporella spp. The tree was rooted with Marasasiomyces karoo
Morphological analyses
In accordance with the phylogenetic results, the three clades displayed morphological characteristics reminiscent of the Eutiarosporella spp. described in the literature (Figs. 3, 4, 5, and 6). On nutrient rich media (such as PDA or V8-OMA), each clade displayed rapid white growth (Fig. 6, Supplementary Fig. 7), similar to that observed in other members of the Eutiarosporella genus (Jami et al. 2014; Phillips et al. 2013). Each of the clades also produced hyaline, thin-walled and aseptate conidia, some having mucosal appendages, which is a key feature of the Eutiarosporella spp. amongst the Botryosphaeriaceae (Phillips et al. 2013). The conidia from both Clade 1 and B. zeae Clade 2 had conidial dimensions similar to E. tritici, as opposed to the other Eutiarosporella spp. (Supplementary Fig. 5) (Jami et al. 2012, 2014; Phillips et al. 2013; Sutton and Marasas 1976). Surprisingly though, B. zeae had smaller conidia than the other two clades.
Images of different aspects of Tiarosporella tritici-australis sp. nov.. Mycelial growth of E. tritici-australis on PDA. a–c. Mycelial growth 4DPI, bar = 4.5 cm (a). Underside growth of mycelia 4DPI (drawn lines indicate radial growth point at 2DPI), bar = 4.5 cm (b). Mycelial appearance after 4 WPI, bar = 4.5 cm (c). Growing hyphal tips, bar = 10 μm (d). Multi-branched pycnidium, bar = 1 mm (e). Erumpent pycnidium, bar = 1 mm (f). Conidia, bar = 10 μm (g) Conidiogenous cells, bar = 10 μm. Conidium, arrow indicates mucosal appendage, bar = 10 μm (H)
Images of different aspects of Tiarosporella darliae sp. nov.. Mycelial growth of E. darliae on PDA. a–c. Mycelial growth 4DPI, bar = 4.5 cm (a). Underside growth of mycelia 4DPI (drawn lines indicate radial growth point at 2DPI), bar = 4.5 cm (b). Mycelial appearance after 4 WPI, bar = 4.5 cm (c). Growing hyphal tips, bar = 10 μm (d). Pycnidia immersed in agar, bar = 0.5 mm (e). Pycnidia through mycelia, bar = 0.5 mm (f). Conidia, bar = 10 μm (g) Conidiogenous cells, bar = 10 μm. Conidium, arrow indicates mucosal appendage, bar = 10 μm (h)
Images of different aspects of Tiarosporella pseudodarliae sp. nov.. Mycelial growth of E. pseudodarliae on PDA. a–c. Mycelial growth 4DPI, bar = 4.5 cm (a). Underside growth of mycelia 4DPI (drawn lines indicate radial growth point at 2DPI), bar = 4.5 cm (b). Mycelial appearance after 4 WPI, bar = 4.5 cm (c). Growing hyphal tips, bar = 10 μm (d). Pycnidium, bar = 0.5 mm (e). Erumpent pycnidium, bar = 0.5 mm (f). Conidia, bar = 10 μm (g) Conidiogenous cells, bar = 10 μm. Conidium, arrow indicates mucosal appendage, bar = 10 μm (h)
On defined minimal media (MM), the extent to which growth was affected by supplied nutrients and the resulting growth morphology was different between the clades (Fig. 6). In general, growth of B. zeae was diminished on all defined media in comparison with the other clades. This is in contrast to growth on nutrient-rich media where the difference in healthy growth appearance was less distinct between the clades. As expected, sorbitol was the poorest carbon source for all of the clades. Each responded more positively to the remaining carbon sources. The responses of B. zeae to these were similar to each other, with wispy, white mycelial growth. B. zeae Clade 2 grew in a thick, striated pattern on glucose and fructose, but was more wispy and floccuous on sucrose. Clade 1 displayed thicker growth on glucose and sucrose, with the appearance of thinner growth on fructose. Growth for Clade 1 on all of the carbon sources was floccuous and white. For the nitrogen sources, glutamate appeared to provide the thickest growth, followed by glutamine. Interestingly, none of the clades responded well to nitrate as the carbon source, a commonly used nitrogen source in defined fungal media. On liquid MM (with sucrose as the carbon source and glutamine as the carbon source) (Supplementary Fig. 6) Clade 1 produced an iron-ore coloured pigment 2-3 dpi. B. zeae produced a similar pigment 5–7 dpi. No pigment formation was observed in this analysis for B. zeae Clade 2.
On PDA, all three clades responded similarly to each other under different growth temperatures (Supplementary Fig. 4). At 16 °C, growth was inhibited for all isolates. At 23 °C, radial growth was rapid (see below for more detailed descriptions of growth at this temperature). At 28 °C, radial growth was rapid, with thick mycelial growth for all isolates. At 3dpi at this temperature, Clade 1 mycelial growth was thick, white, and floccuous, becoming more appressed toward the site of inoculation; B. zeae mycelial growth was bright white with thick growth around the edge of the petri dish; B. zeae. Clade 2 growth was thick white at the border of the petri dish with a segregating band of thin-growth separating thick, white-to-olivaceous growth towards the site of inoculation. At 30 °C, radial growth was inhibited for each of the clades, however, mycelial growth was thick. According to this preliminary analysis, the optimum temperature for each of the clades is <23 °C but >30 °C.
Growth on OMA was observed as this is the base sporulation media used for these fungi (Supplementary Fig. 7). Each of the clades displayed thick, white mycelial growth. B. zeae and Clade 2 B. zeae were more floccuous in appearance than when observed on PDA. GABA-OMA was used as a media source in order to determine if GABA as a supplement may alter the growth of the fungi. GABA has previously been shown to alter sporulation in Parastagonospora nodorum and Clade 1 (previously acknowledged as “Botryosphaeria spp.”) (Mead et al. 2013) when added as a media supplement. Growth was not altered by the addition of GABA for any of the clades.
V8-OMA was used to observe growth on a more nutrient rich, complex media. Each of the clades responded well to the V8 juice with thicker mycelial growth, covering more of the plate’s surface.
We describe the type species of three novel species of WGD fungi (Eutiarosporella tritici-australis (HOLOTYPE: DAR 82485) (formerly Clade 1) (Fig. 3), Eutiarosporella darliae (HOLOTYPE: DAR 82491) (formerly B. zeae) (Fig. 4), and Eutiarosporella pseudodarliae (HOLOTYPE: DAR 82489)(formerly B. zeae Clade 2) (Fig. 5)) and continue to develop upon their specific morphology, in detail below.
Eutiarosporella tritici-australis E. Thynne, M.C. McDonald, M. Evans, H. Wallwork, S. Neate, P.S. Solomon sp. nov. (formerly Clade 1)
MycoBank no.: MB 811115
Etymology: Named after a close relative, Eutiarosporella tritici, (and because it infects wheat (Triticum aestivum)) in conjunction with the country in which it causes disease.
On PDA media, growth is floccuous with thick aerial mycelia. Radial growth is rapid with 5.5–5.9 cm growth 2 days post inoculation (dpi) and the plate is covered by 3 dpi, with mycelia extending up the plate walls by 4 dpi. The mycelia, initially white, discolour with age to white-grey, dark grey, and olivaceous grey/black. Mycelia remain generally floccuous with age, however, are slightly more appressed close to the site of inoculation. Microscopically, healthy growing mycelia are hyaline and septate, with a granular surface appearance. Hyaline mycelia range from 3 to 8 μm in diameter. Pigmented mycelia range from 7 to 16 μm in diameter. Pycnidia form on PDA after an extended period of growth (beyond 4 weeks post inoculation (WPI)). However, pycnidia formation is more rapid under near u.v. lights and on OM. Pycnidia were observed both solitarily and/or grouped and were semi-immersed. Initially globular, a stem grows above surface mycelia (0.26(±0.06)mm × 0.93(±0.14)mm). Pycnidia can remain single stemmed or become highly branched. Pycnidia can become erumpent with maturity, exuding conidia bearing cirrus. Conidiogenous cells (7–12 μm × 3–5 μm) are hyaline, thin walled and holoblastic. Conidia are hyaline, thin-walled, aseptate, and range from clavate to fusiform in shape (12.25(±0.37) μm × 37.8(±0.65) μm). Some conidia bear a faint mucosal appendage reminiscent of a halo. Conidia were observed germinating from both the apical and basal ends.
Typification: AUSTRALIA, South Australia, isolated from Triticum aestivum (bread wheat), 2012, SARDI
(HOLOTYPE and EX-TYPE: DAR 82485)
Host and distribution: Individuals have been observed in infected wheat and wheat-stubble in South Australia and Victoria.
Optimum temperature range: >23 °C but <30 °C
Eutiarosporella darliae E. Thynne, M.C. McDonald, M. Evans, H. Wallwork, S. Neate, P.S. Solomon sp. nov. (Formerly B. zeae )
MycoBank no.: MB 811116
Etymology: Named after the location where it was first observed infecting wheat, Darling Downs, Queensland.
On PDA media, initially, mycelia are floccuous close to the site of inoculation with limited aerial growth. However, radially out, the mycelia become dense and close-knit. Microscopically, healthy growing mycelia are hyaline and septate, with a granular surface appearance. Viewed from beneath the media plate, the fungus darkens in a piebald pattern. On occasion, a secreted pigment stains the media a violaceous colour. Viewed from the surface, the mycelia appear to remain vividly white with age. These mycelia are highly appressed into a dense mat that can be pealed back from the agar surface in a single layer. However, light brown to olivaceous-gray mycelia can be present. Hyaline mycelia range from 1 to 5 μm in diameter. Pigmented mycelia range from 4 to 8 μm in diameter. Conidia bearing pycnidia form sporadically on OMA under near u.v. light. Conidia are semi-immersed to not immersed. They appear to remain small and predominantly globular, with some skewed to the cylindrical, ranging from 80 to 170 μm in diameter. Conidiogenous cells (7–16 μm × 3–7 μm) are hyaline, thin walled and holoblastic. Conidia are hyaline, thin-walled, aseptate, and range from clavate to fusiform in shape. The observed conidia were surprisingly small in comparison to E. tritici-australis and E. pseudodarliae (9.26(±0.33) μm × 27.37(±0.49) μm). Some conidia bear a faint mucosal appendage reminiscent of a halo. Conidia were observed germinating from both the apical and basal ends.
Typification: AUSTRALIA, Queensland, isolated from Triticum aestivum (bread wheat), 2012, QLD DPI
(HOLOTYPE and EX-TYPE: DAR 82491)
Host and Distribution: Individuals have been observed in infected wheat and wheat-stubble in Queensland and Northern New South Wales.
Optimum temperature range: >23 °C but <30 °C
Eutiarosporella pseudodarliae E. Thynne, M.C. McDonald, M. Evans, H. Wallwork, S. Neate, P.S. Solomon sp. nov. (Formerly B. zeae Clade 2)
MycoBank no.: MB 811117
Etymology : Named after its closest known relative Eutiarosporella darliae, with which it shares very close growth characteristics.
Individuals have been observed in infected wheat and wheat-stubble across the Eastern Australian states. On PDA media, mycelial growth is thick with a limited amount of dense aerial growth. Radial growth is rapid with 5.5–6 cm of growth, 2 days post inoculation (dpi) and the plate is covered by 3 dpi. Initial mycelial growth is white, but discolours. Viewed from the surface of the media plate, the mycelia darken to grey-white, olivaceous-grey/black, and light brown. Viewed from beneath the media plate, the fungus darkens rapidly from the site of inoculation. The agar beneath the surface mycelia becomes tar-like in appearance. Microscopically, healthy growing mycelia are hyaline and septate, with a granular surface appearance. Hyaline mycelia range from 1 to 7 μm in diameter. Pigmented mycelia range from 3 to 7 μm in diameter. On PDA and OMA, pycnidia form readily and profusely. Pycnidia were observed as semi-immersed, and single stemmed. Stems are smaller than observed on E. tritici-australis (0.11(±0.04)mm × 0.37(±0.15)mm). Pycnidia can become erumpent with maturity, exuding conidia bearing cirrus. Conidiogenous cells (7–16 μm × 3–7 μm) are hyaline, thin walled and holoblastic Conidia are hyaline, thin-walled, aseptate, and range from clavate to fusiform in shape (12.39(±0.47) μm × 35.84(±0.60) μm). Some conidia bear a faint mucosal appendage reminiscent of a halo. Conidia were observed germinating from both the apical and basal ends.
Typification: AUSTRALIA, Queensland, isolated from Triticum aestivum (bread wheat), 2011, QLD DPI
(HOLOTYPE and EX-TYPE: DAR 82489)
Host and distribution: Individuals have been observed in infected wheat and wheat-stubble across the Eastern Australian states.
Notes: Eutiarosporella pseudodarliae and Eutiarosporella darliae are closely related genetically, and can appear very similar in culture.
Optimum temperature range: >23 °C but <30 °C
Discussion
In this study fungal specimens isolated from diseased wheat-grain, previously described as Botryosphaeria zeae, were reclassified as members of the genus Eutiarosporella based on comparison of ITS sequences. Although originally grouped as a single species, our results revealed a species complex within the isolates analysed. With the support of a multigene phylogeny (ITS, β-tubulin and EF1-α), three novel species of Eutiarosporella are described; Eutiarosporella tritici-australis, Eutiarosporella darliae, and Eutiarosporella pseudodarliae.
Wheat is an economically significant industry in Australia that is under constant threat from fungal diseases (Murray and Brennan 2009). Accordingly, correct identification of the pathogen species that affect this industry is vital for growers and pathologists. Due to lack of sequence data, we were unable to compare these unknown wheat-infecting isolates to their original species classification, B. zeae. Fortunately, however, since the first observation of WGD fungi in 1999, there has been a significant expansion in the number of Botryosphaeriaceae spp. recognised. In various cases, the genera and species have been greatly refined (Crous et al. 2006, 2015; Slippers et al. 2013) and the species within described in detail (Phillips et al. 2013). This allows rapid identification of other Botryosphaeriaceae spp., as was shown in this study.
Herein, we re-designate WGD Botryosphaeriaceae species to now be contained within the genus Eutiarosporella. This is interesting because, to the best of our knowledge, the only species of the former Tiarosporella (now four generea) identified in Australia previously were saprophytic isolates, isolated from submerged trees in Queensland and described as T. paludosa (Hyde 1993). Unfortunately, gene sequence data is currently not available for these isolates. For many years, sequences confirming the identity of Tiarosporella and Eutiarosporella spp. have been limited to isolates collected from South Africa (Jami et al. 2014). The South African species T. madreeya, T. tritici (now Eutiarosporella tritici), and T. graminis (now Marasasiomyces karoo) (Crous et al. 2015)) were documented early in the literature (Sutton and Marasas 1976), but only had genes sequenced more recently (Crous et al. 2006; Phillips et al. 2013). Other species, E. urbis-rosarum and E. africana were only recently identified and characterised (Jami et al. 2012, 2014). Of all of these South African Eutiarosporella spp., the only species documented to have been interacting with wheat is E. tritici. Sutton and Marasas (1976) described E. tritici from wheat-straw in South Africa. The morphological descriptions of E. tritici from South Africa (Sutton and Marasas 1976) resemble those of the three, novel Australian wheat-infecting Eutiarosporella spp. described in this study. For example, the described conidia size for all species. We have also shown that E. tritici resolves closely with these three species, and is one of the roots for the etymology of E. tritici-australis.
This analysis does not yet shed light on the definitive origin of these pathogen species. Jami et al. discussed the overrepresentation of Tiarosporella spp. from South Africa, but note this is probably due to lack of more widespread sampling (Jami et al. 2014). From the perspective of WGD, a greater survey into potential alternative hosts for these pathogens may be required in order to determine the presence of other Eutiarosporella spp. in Australia.
Hyde et al. (2010) emphasised the need for up-to-date cataloguing of Australian fungal plant-pathogens (Hyde et al. 2010). They illustrate this need by highlighting discrepancies between gene-sequence and morphological data attributed to a single Botryosphaeriaceae isolate (Hyde et al. 2010). Our study supports Hyde’s assertion about the importance of performing a “re-inventory” of Australian fungal plant-pathogens (Hyde et al. 2010), as we too have highlighted nomenclature discrepancies that could only be resolved with genetic comparisons. To date, there are no published genetic studies analysing the maize pathogen, B. zeae, although morphologically they are believed to be similar to wheat-infecting Eutiarosporella spp. (Platz 2011). As a speculative side note, perhaps the initial morphology based comparison between these species was in fact accurate, but rather it is actually B. zeae that that is classified incorrectly, and should instead be of the Eutiarosporella. This further emphasises the need for continual gene and whole-genome sequencing to better understand the existing diversity of fungal species, particularly when they are found in close association with plants.
For the purposes of this study, we have merely utilised the genome sequences available to us to retrieve the ITS, β-tubulin and EF1-α from the three sequenced isolates. Our delimitation of the three novel species of wheat-infecting Eutiarosporella spp. was strongly supported by the multigene phylogeny of these three genes. These sequences have been used successfully in studies defining the boundaries of particular Botryosphaeriaceae spp. and so we felt that it was appropriate to utilise these sequences. We will now endeavour to use these genome sequences to assist in better understanding these recently emerged pathogens.
The ability to obtain nearly complete fungal genomes is becoming ever easier. This ease of access has made it feasible to rapidly compare multiple genomes and has led to major improvements in our understanding of the lifestyles, niche specialisation, and emergence of fungal phytopathogens (Gardiner et al. 2012; Klosterman et al. 2011; Thynne et al. 2015). Genomes of Botryosphaeriaceae family are now being exploited to identify basic biological processes such as homothalism vs. heterothallism, which once took years of careful experiments to correctly assign (Islam et al. (2012); Blanco-Ulate et al. (2013); Bihon et al. (2014)). For example, Bihon et al. (2014) used the genome sequences of Diplodia pinea to describe the mating type idiomorphs for this species, and in doing so described a novel putative MAT gene, characterised as MAT1-2-5 (Bihon et al. 2014).
Currently, very little is known about wheat-infecting Eutiarosporella spp. and their emergence in the Australian grain industry have yet to cause significant enough damage to warrant large-scale studies. We expect, however, that the sequenced genomes of wheat-infecting Eutiarosporella spp. will greatly assist in answering remaining questions about their lifecycle, especially related towards host specificity. Until this point, it is our hope that the three loci sequences provided in this study will contribute valuable data for the community, adding to the knowledge about the distribution and host-range of members of the Botryosphaeriaceae family.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alvarez-Loayza P, White JF Jr, Torres MS, Balsev H, Kristiansen T, Svenning JC, Gil N (2011) Light converts endosymbiotic fungus to pathogen, influencing seedling survival and niche-space filling of a common tropical tree, Iriartea deltoidea. PLoS One 6, e16386
Aoki T, Vaughan MM, McCormick SP, Busman M, Ward T, Kelly A, O’Donnell K, Johnston P, Geiser DM (2014) Fusarium dactylidis sp. nov., a novel nivalenol toxin-producing species sister to F. pseudograminearum isolated from orchard grass (Dactylis glomerata) in Oregon and New Zealand. Mycologia: doi:10.3852/14-213
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Barber PA, Burgess TJ, Hardy GE, Slippers B, Keane PJ, Wingfield MJ (2005) Botryosphaeria species from Eucalyptus in Australia are pleoanamorphic, producing dichomera synamorphs in culture. Mycol Res 109:1347–1363
Bihon W, Wingfield MJ, Slippers B, Duong TA, Wingfield BD (2014) MAT gene idiomorphs suggest a heterothallic sexual cycle in a predominantly asexual and important pine pathogen. Fungal Genet Biol 62:55–61
Blanco-Ulate B, Rolshausen P, Cantu D (2013) Draft genome sequence of Neofusicoccum parvum isolate UCR-NP2, a fungal vascular pathogen associated with grapevine cankers. Genome Announcements 1:e00339–13
Crous PW, Slippers B, Wingfield MJ, Rheeder J, Marasas WF, Philips AJ, Alves A, Burgess T, Barber P, Groenewald JZ (2006) Phylogenetic lineages in the Botryosphaeriaceae. Stud Mycol 55:235–253
Crous PW, Muller MM, Sanchez R, Giordano L, Bianchinotti MV, Anderson FE, Groenewald JZ (2015) Resolving Tiarosporella spp. allied to Botryosphaeriaceae and Phacidiaceae. Phytotaxa 202:73–93
De Leon C, Jeffers D, Maize I, Center WI (2004) Maize diseases: a guide for field identification, 4th edn. International Maize and Wheat Improvement Center (CIMMYT), Mexico, 119 p
Dettman JR, Jacobson DJ, Taylor JW (2003) A multilocus genealogical approach to phylogenetic species recognition in the model eukaryote Neurospora. Evolution 57:2703–2720
Evans M (2013) White grain rejection spreading. Ground Cover Supplement 102:4
Gardiner DM, McDonald MC, Covarelli L, Solomon PS, Rusu AG, Marshall M, Kazan K, Chakraborty S, McDonald BA, Manners JM (2012) Comparative pathogenomics reveals horizontally acquired novel virulence genes in fungi infecting cereal hosts. PLoS Pathog 8, e1002952
Hyde KD (1993) Tropical Australian freshwater fungi VI. Tiarosporella paludosa and Clohesyomyces aquaticus gen. et sp. Nov. (Coelomycetes). Aust Syst Bot 6:169–173
Hyde KD, Chomnunti P, Crous PW, Groenewald JZ, Damm U, Ko TWK, Shivas RG, Summerell BA, Tan YP (2010) A case for re-inventory of Australia’s plant pathogens. Persoonia: Mol Phylogeny Evol Fungi 25:50–60
Islam MS, Haque M, Emdad EM, Halim A, Hossen QMM, Hossain MZ, Ahmed B, Rahim S, Rahman MS (2012) Tools to kill: genomes of one of the most destructive plant pathogenic fungi Macrophomina phaseolina. BMC Genomics 13:493
Jami F, Slippers B, Wingfield MJ, Gryzenhout M (2012) Five new species of the Botryosphaeriaceae from Acacia Karroo in South Africa. Cryptogam Mycol 33:245–266
Jami F, Slippers B, Wingfield MJ, Gryzenhout M (2013) Greater Botryosphaeriaceae diversity in healthy than associated diseased Acacia karroo tree tissues. Australas Plant Pathol 42:421–430
Jami F, Slippers B, Wingfield MJ, Gryzenhout M (2014) Botryosphaeriaceae species overlap on four unrelated, native South African hosts. Fungal Biol 118:168–179
Klosterman SJ, Subbarao KV, Kang S, Veronese P, Gold SE, Thomma BP, Chen Z, Henrissat B, Lee YH, Pk J (2011) Comparative genomics yields insights into niche adaptation of plant vascular wilt pathogens. PLoS Pathog 7, e1002137
Kopinski JS, Blaney BJ (2010) Nutritive value and non-toxicity of Botryosphaeria zeae-infected wheat for weaner pigs. J Anim Physiol Anim Nutr 94:44–54
Leigh JW, Susko E, Baumgartner M, Roger AJ (2008) Testing congruence in phylogenomic analysis. Syst Biol 57:104–115
Lohse M, Bolger AM, Nagel A, Fernie AR, Lunn JE, Stitt M, Usadel B (2012) RobiNA: a user-friendly, integrated software solution for RNA-Seq-based transcriptomics. Nucleic Acids Res 40:W622–W627
Luque J, Martos S, Phillips AJL (2005) Botryosphaeria viticola sp. nov. on grapevines: a new species with a Dothiorella anamorph. Mycologia 97:1111–1121
McDonald MC, Razavi M, Friesen TL, Brunner PC, McDonald BA (2012) Phylogenetic and population genetic analyses of Phaeosphaeria nodorum and its close relatives indicate cryptic species and an origin in the Fertile Crescent. Fungal Genet Biol 49:882–895
Mead O, Thynne E, Winterberg B, Solomon PS (2013) Characterising the Role of GABA and Its Metabolism in the Wheat Pathogen Stagonospora nodorum. PLoS One 8, e78368
Murray GM, Brennan JP (2009) Estimating disease losses to the Australian wheat industry. Australas Plant Pathol 38:558–570
Penfold K (2011) White grains may be blight disease. GRDC Ground Cover 92
Phillips A, Alves A, Abdollahzadeh J, Slippers B, Wingfield MJ, Groenewald J, Crous P (2013) The Botryosphaeriaceae: genera and species known from culture. Stud Mycol 76:51–167
Platz G (2011) Wheat and barley disease management in 2011. Yellow spot and head diseases in wheat. Strategies and products for barley leaf rust. GRDC Update Papers
Sakalidis M, Hardy GE, Burgess TI (2011a) Endophytes as potential pathogens of the baobab species Adansonia gregorii: a focus on the Botryosphaeriaceae. Fungal Ecol 4:1–14
Sakalidis M, Hardy GE, Burgess TI (2011b) Use of the Genealogical Sorting Index (GSI) to delineate species boundaries in the Neofusicoccum parvum–Neofusiccocum ribis species complex. Mol Phylogenet Evol 60:333–344
Sakalidis M, Slippers B, Wingfield B, Hardy GSJ, BURGESS T (2013) The challenge of understanding the origin, pathways and extent of fungal invasions: global populations of the Neofusicoccum parvum–N. ribis species complex. Divers Distrib 19:873–883
Slippers B, Wingfield MJ (2007) Botryosphaeriaceae as endophytes and latent pathogens of woody plants: diversity, ecology and impact. Fungal Biol Rev 21:90–106
Slippers B, Crous PW, Denman S, Coutinho TA, Wingfield BD, Wingfield MJ (2004a) Combined multiple gene genealogies and phenotypic characters differentiate several species previously identified as Botryosphaeria dothidea. Mycologia 96:83–101
Slippers B, Fourie G, Crous PW, Coutinho TA, Wingfield BD, Carnegie AJ, Wingfield MJ (2004b) Speciation and distribution of Botryosphaeria spp. on native and introduced Eucalyptus trees in Australia and South Africa. Stud Mycol 50:343–358
Slippers B, Fourie G, Crous PW, Coutinho TA, Wingfield BD, Wingfield MJ (2004c) Multiple gene sequences delimit Botryosphaeria australis sp. nov. from B. lutea. Mycologia 96:1030–1041
Slippers B, Summerell BA, Crous PW, Coutinho TA, Wingfield BD, Wingfield MJ (2005) Preliminary studies on Botryosphaeria species from Southern Hemisphere conifers in Australasia and South Africa. Australas Plant Pathol 34:213–220
Slippers B, Burgess T, Pavlic D, Ahumada R, Maleme S, Rodas C, Wingfield MJ (2009) A diverse assemblage of Botryosphaeriaceae infect Eucalyptus in native and non-native environments. South For 71:101–110
Slippers B, Boissin E, Phillips A, Groenewald J, Lombard L, Wingfield M, Postma A, Burgess T, Crous P (2013) Phylogenetic lineages in the Botryosphaeriales: a systematic and evolutionary framework. Stud Mycol 76:31–49
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Sutton BC, Marasas WFO (1976) Observations on Neottiosporina and Tiarosporella. Trans Br Mycol Soc 67:69–76
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Taylor A, Hardy GESJ, Wood P, Burgess T (2005) Identification and pathogenicity of Botryosphaeria species associated with grapevine decline in Western Australia. Australas Plant Pathol 34:187–195
Thynne E, McDonald MC, Solomon PS (2015) Phytopathogen emergence in the genomics era. Trends Plant Sci. doi:10.1016/j.tplants.2015.01.009
Vasconcelos AF, Dekker RH, Barbosa A, Paccola-Meirelles L (2001) Comparison of the laccases, molecular marker proteins, and induction of pycnidia by three species of Botryosphaeriaceous fungi. Mycoscience 42:543–548
White TJ, Bruns T, Lee SJWT, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc: Guide Methods Appl 18:315–322
Wildermuth G, Williamson P, Shivas R, McNamara R (2001) Premature head blight and white grain – a new disease of wheat. Proceedings of 13th Biennial Australasian Plant Pathology Conference, Queensland Department of Primary Industries
Zhou YJ, Zhang J, Wang XD, Yang L, Jang DH, Li GQ, Hsiang T, Zhuang WY (2014) Morphological and phylogenetic identification of Botryotis sinoviticola, a novel cryptic species causing grey mold disease of table grapes (Vitis vinifera) in China. Mycologia 106:43–56
Acknowledgments
This study was supported by the Grains Research and Development Corporation (GRDC) (ANU00019). Peter Solomon is an Australian Research Council Future Fellow and Elisha Thynne is a recipient of an Australian Postgraduate Award and a GRDC Grains Industry Research Scholarship.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplementary Table 1
(DOCX 37 kb)
Supplementary Fig 1
Phylogenetic analyses separate isolates of WGD fungi into three distinct clades. Tree is rooted to Botryosphaeria agaves. (JPEG 553 kb)
Supplementary Fig 2
Phylogenetic analyses comparing isolates of WGD fungi with a range of species from the former genus of Tiarosporella spp.. Tree is rooted to Marasasiomyces karoo. (JPEG 1739 kb)
Supplementary Fig 3
Alignment of EF1-α sequences of the isolates of wheat-infecting Tiarosporella spp. focusing on the region of base-deletion in B. zeae and Clade 2 B. zeae. (JPEG 473 kb)
Supplementary Fig 4
Growth of each of the clades of wheat-infecting Tiarosporella spp. at different temperatures. (JPEG 2495 kb)
Supplementary Fig 5
Histogram displaying the comparative conidial dimensions of Tiarosporella spp. in relation to WGD fungi. (JPEG 7644 kb)
Supplementary Fig 6
Image of Clade 1 pigment production on liquid minimal media (with sucrose and glutamine). (JPEG 1334 kb)
Supplementary Fig 7
Growth of each of the three clades on OMA, GABA-OMA,V8-OMA. (JPEG 979 kb)
Rights and permissions
About this article
Cite this article
Thynne, E., McDonald, M.C., Evans, M. et al. Re-classification of the causal agent of white grain disorder on wheat as three separate species of Eutiarosporella . Australasian Plant Pathol. 44, 527–539 (2015). https://doi.org/10.1007/s13313-015-0367-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-015-0367-2