Abstract
Many cervical cancer (CC) patients experience early cancer metastasis, resulting in poor therapeutic outcome after resection of primary cancer. Hence, there is a compelling requirement for understanding of the molecular mechanisms underlying the invasiveness control of CC. Pituitary tumor-transforming gene 1 (Pttg1) has been recently reported to promote cancer cell growth and metastasis in a number of various tumors. However, its regulation by microRNAs (miRNAs) as well as its role in CC have not been clarified. Here, we reported significantly higher levels of Pttg1 and significantly lower levels of miR-494 in the resected CC tissue, compared with the adjacent normal cervical tissue from the same patient. Interestingly, Pttg1 levels inversely correlated with miR-494 levels. In vitro, Pttg1 levels determined CC cell invasiveness and were inhibited by miR-494 levels. However, miR-494 levels were not affected by Pttg1 levels. Furthermore, miR-494 inhibited Pttg1 expression in CC cells, through directly binding and inhibition on 3′-UTR of Pttg1 mRNA. Together, our data suggest that Pttg1 may increase CC cell metastasis, which is negatively regulated by miR-494. Our work thus highlights a novel molecular regulatory machinery in metastasis of CC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many cervical cancer (CC) patients experience early cancer metastasis, resulting in poor therapeutic outcome after resection of primary cancer [1–4]. Hence, there is a compelling requirement for understanding of the molecular mechanisms underlying the invasiveness control of CC.
Pituitary tumor-transforming gene 1 (Pttg1, also known as securin) has been shown to promote a cell to enter an active cell cycle [5–8]. Recently, Pttg1 was reported to promote cancer cell growth and metastasis in a number of various tumors [9–28]. However, a role of Pttg1 in the tumorigenesis of CC is not determined. Moreover, the molecular mechanisms underlying the downstream signal transduction of Pttg1 as well as the regulation of Pttg1 expression in cancer cells remain largely unknown.
MicroRNA (miRNA) is a class of non-coding small RNA of comprised of about 22 nucleotides. MiRNA has been found to regulate the gene expression post-transcriptionally, through its base-pairing with the 3′-untranslated region (3′-UTR) of target mRNA [29, 30]. It has been shown that miRNA regulates many biological events including cancer development. Among all miRNAs, miR-494 has just been recently recognized as a tumor-suppressive microRNA in various types of cancer, including gastric cancer [31, 32], lung cancer [33, 34], oral cancer [35, 36], and brain cancer [37, 38]. However, a role of miR-494 in the pathogenesis of CC and its relationship with Pttg1 have not been acknowledged before.
Here, we reported significantly higher levels of Pttg1 and significantly lower levels of miR-494 in the resected CC tissue, compared with the adjacent normal cervical tissue from the same patient. Interestingly, Pttg1 levels inversely correlated with miR-494 levels. In vitro, Pttg1 levels determined CC cell invasiveness and were inhibited by miR-494 levels. However, miR-494 levels were not affected by Pttg1 levels. Furthermore, miR-494 inhibited Pttg1 expression in CC cells, through directly binding and inhibition on 3′-UTR of Pttg1 mRNA. Together, our data suggest that Pttg1 may increase CC cell metastasis, which is negatively regulated by miR-494.
Materials and methods
Specimens from patients
A total of 22 patients with CC were included in the study. The resected specimens (paired CC tissue and the adjacent normal cervical tissue (NCT)) from the patients were used for analyzing transcripts of Pttg1 and miR-494. All specimens had been histologically and clinically diagnosed at Department of Obstetrics and Gynecology of Air Force General Hospital of PLA from 2008 to 2013. For the use of these clinical materials for research purposes, prior patient’s consents and approval from the Institutional Research Ethics Committee were obtained.
Cell line culture and transfection
Human CC cell line Hela S3 (H3) is a clonal derivative of the parent Hela line. H3 was cloned in 1955 by T.T. Puck, P.I. Marcus, and S.J. Cieciura [39] and was purchased from American Type Culture Collection (ATCC, Rockville, MD, USA). Cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM, Invitrogen, Carlsbad, CA, USA) supplemented with 15 % fetal bovine serum (FBS; Sigma-Aldrich, St Louis, MO, USA) in a humidified chamber with 5 % CO2 at 37 °C. The plasmids that contain transgene or shRNA for Pttg1 were kindly provided by Dr. Shengquan Huang (Xinqiao Hospital, Third Military Medical University, China) and have been described before [21]. The constructs for miR-494, miR-494 antisense (as-miR-494) were all purchased from GeneCopoeia (Rockville, MD, USA). Transfection was performed with 2 μg plasmids using the Lipofectamine 2000 according to the manufacturer’s instructions (Invitrogen).
Scratch wound healing assay
Scratch wound healing assay was performed as has been described previously [40]. Cells were seeded in 24-well plates at a density of 104 cells/well in complete DMEM and cultured to confluence. The cell monolayer was serum starved overnight in DMEM prior to initiating of the experiment. Confluent cell monolayer were then scraped with a yellow pipette tip to generate scratch wounds and washed twice with media to remove cell debris. Cells were incubated at 37 °C for 24 hours. Time lapse images were captured after 12 h. Images were captured from five randomly selected fields in each sample, and the wound areas were calculated by NIH ImageJ software.
Transwell cell migration assay
Cells (105) were plated into the top side of polycarbonate transwell filter coated with Matrigel in the upper chamber of the BioCoat™ Invasion Chambers (Becton-Dickinson Biosciences, Bedford, MA, USA) and incubated at 37 °C for 22 h. The cells inside the upper chamber with cotton swabs were then removed. Migratory and invasive cells on the lower membrane surface were fixed, stained with hematoxylin, and counted for 10 random 100× fields per well. Cell counts are expressed as the mean number of cells per field of view. Five independent experiments were performed and the data are presented as mean ± standard deviation (SD).
RNA extraction, reverse transcription, and quantitative RT-PCR
MiRNA and total RNA were extracted from resected specimen from the patients or from cultured cells with miRNeasy mini kit or RNeasy kit (Qiagen, Hilden, Germany), respectively, for complementary DNA (cDNA) synthesis. cDNA was randomly primed from 2 μg of total RNA using the Omniscript reverse transcription kit (Qiagen). RT-qPCR was subsequently performed in triplicate with a 1:4 dilution of cDNA using the Quantitect SyBr green PCR system (Qiagen) on a Rotorgene 6000 series PCR machine. Quantitative PCR (RT-qPCR) were performed in duplicates with QuantiTect SYBR Green PCR Kit (Qiagen). All primers were purchased from Qiagen. Data were collected and analyzed with the Rotorgene software accompanying the PCR machine, using 2–△△Ct method for quantification of the relative mRNA expression levels. Values of genes were first normalized against α-tubulin and then compared to controls.
Western blot
Total protein was extracted from the cultured cells by RIPA buffer (Sigma-Aldrich). Equal amount of proteins was loaded in the gel. Primary antibodies for Western blot are rabbit anti-Pttg1 and anti-α-tubulin (all purchased from Cell Signaling, St. Jose, LA, USA). Secondary antibody is HRP-conjugated anti-rabbit (Jackson ImmunoResearch Labs, West Grove, PA, USA). Images shown in the figure were representatives from five repetitions.
Luciferase-reporter activity assay
Luciferase-reporters were successfully constructed using molecular cloning technology. Target sequence for Pttg1 miRNA 3′UTR clone was purchased from Creative Biogene (Shirley, NY, USA). H3-miR-494, H3-scr, or H3-as-miR-494 cells were seeded in 24-well plates for 24 h, after which they were transfected with 1 μg of luciferase-reporter plasmids per well using PEI Transfection Reagent. Luciferase activities were measured using the dual luciferase-reporter gene assay kit (Promega, Beijing, China), according to the manufacturer’s instructions.
Statistical analysis
Statistical analyses were performed with SPSS 19.0 software (SSPS Inc., Chicago, IL, USA). All data were statistically analyzed using one-way ANOVA with a Bonferroni correction, followed by Fisher’s exact test for comparison between two groups. All values are depicted as mean ± standard deviation from ten individuals and are considered significant if p < 0.05. Bivariate correlations were calculated by Spearman’s rank correlation coefficients.
Results
High Pttg1 and low miR-494 was detected in CC specimen
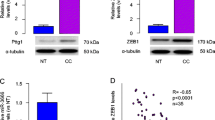
The recent findings highlighting a role of Pttg1 and miR-494 in the tumorigenesis inspired us to examine their involvement in CC. The resected specimens (paired CC and the adjacent normal cervical (NCT) tissue) from the 22 CC patients were analyzed for Pttg1 and miR-494 transcript levels by RT-qPCR. We found that compared to NCT, CC tissue had a significantly higher Pttg1 levels (Fig. 1a, p < 0.05) and a significantly lower miR-494 levels (Fig. 1b).
Pttg1 and miR-494 levels inversely correlated in CC specimen. The resected specimens (paired CC and the adjacent normal cervical (NCT) tissue) from the 22 CC patients were analyzed for Pttg1 and miR-494 transcript levels by RT-qPCR. a–b We found that compared to NCT, CC tissue had a significantly higher Pttg1 levels (a, p < 0.05) and a significantly lower miR-494 levels (b). c A strong and inverse correlation was detected between the levels of miR-494 and Pttg1 in the CC specimen (R = −0.67, p < 0.0001). *p < 0.05. n = 22. Statistics: one-way ANOVA with a Bonferroni correction. Bivariate correlations: Spearman’s rank correlation coefficients
Pttg1 and miR-494 levels inversely correlated in CC specimen
In order to find out whether there is a possible relationship between Pttg1 and miR-494, we checked the Pttg1 and miR-494 levels in each patient. We detected a strong and inverse correlation between the levels of miR-494 and Pttg1 in the CC specimen (Fig. 1c, R = −0.67, p < 0.0001). These data suggest presence of a relationship between miR-494 and Pttg1 in the development of CC.
Pttg1 increased CC invasiveness without affecting miR-494 levels
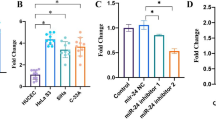
Then, we aimed to examine whether modification of Pttg1 levels in CC cells may alter cell metastasis and miR-494 levels. We used either a Pttg1-overexpressing plasmid or a plasmid carrying short hairpin small interfering RNA (shRNA) for Pttg1 (shPttg1) to transfect a human CC cell line, Hela S3 (H3) and to increase or decrease Pttg1 levels, respectively. H3 cells were also transfected with a plasmid carrying a scrambled sequence (scr) as controls. The modifications of Pttg1 levels in H3 cells were confirmed by RT-qPCR (Fig. 2a). We found that modifications of Pttg1 levels in H3 cells did not alter miR-494 levels (Fig. 2b), suggesting that miR-494 expression is not regulated by Pttg1. Moreover, overexpression of Pttg1 in H3 cells (H3-Pttg1) significantly increased cell invasiveness in either a scratch wound healing assay (Fig. 2c) or a transwell cell migration assay (Fig. 2d). On the other hand, depletion of Pttg1 in H3 cells (H3-shPttg1) significantly decreased cell invasiveness in either a scratch wound healing assay (Fig. 2c) or a transwell cell migration assay (Fig. 2d). These data suggest that Pttg1 increased CC invasiveness without affecting miR-494 levels, while miR-494 levels did not directly regulate cell invasion and metastasis.
Pttg1 increased CC invasiveness without affecting miR-494 levels. We used either a Pttg1-overexpressing plasmid or a plasmid carrying short hairpin small interfering RNA (shRNA) for Pttg1 (shPttg1) to transfect a human CC cell line, Hela S3 (H3), to increase or decrease Pttg1 levels, respectively. H3 cells were also transfected with a plasmid carrying a scrambled sequence (scr) as controls. a The modifications of Pttg1 levels in H3 cells were confirmed by RT-qPCR. b Modifications of Pttg1 levels in H3 cells did not alter miR-494 levels. c–d Cell invasiveness was examined in either a scratch wound healing assay (c) or a transwell cell migration assay (d). *p < 0.05. NS non-significant. n = 5. Statistics: one-way ANOVA with a Bonferroni correction
miR-494 inhibited CC invasiveness through Pttg1
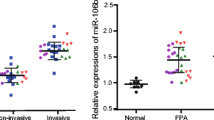
Then, we examined whether modification of miR-494 levels in CC cells may alter Pttg1 levels. We used either a miR-494-expressing plasmid or a plasmid carrying antisense (as) for miR-494 (as-miR-494) to transfect H3 cells. H3 cells were also transfected with a plasmid carrying a scrambled sequence (scr) as controls. The modifications of miR-494 levels in H3 cells were confirmed by RT-qPCR (Fig. 3a). We found that the increases in miR-494 levels (H3-miR-494) in H3 cells significantly decreased Pttg1 levels, while the decreases in miR-494 levels in H3 cells (H3-as-miR-494) significantly increased Pttg1 levels (Fig. 3b–c), suggesting that miR-494 inhibits Pttg1. Moreover, H3-miR-494 significantly decreased cell invasiveness in either a scratch wound healing assay (Fig. 3d) or a transwell cell migration assay (Fig. 3e). On the other hand, H3-as-miR-494 significantly increased cell invasiveness in either a scratch wound healing assay (Fig. 3d) or a transwell cell migration assay (Fig. 3e). Together, these data suggest that miR-494 inhibits CC invasiveness through Pttg1.
miR-494 inhibited CC invasiveness through Pttg1. We used either a miR-494-expressing plasmid or a plasmid carrying antisense (as) for miR-494 (as-miR-494) to transfect H3 cells. H3 cells were also transfected with a plasmid carrying a scrambled sequence (scr) as controls. a The modifications of miR-494 levels in H3 cells were confirmed by RT-qPCR. b–c The increases in miR-494 levels (H3-miR-494) in H3 cells significantly decreased Pttg1 levels, while the decreases in miR-494 levels in H3 cells (H3-as-miR-494) significantly increased Pttg1 levels, by RT-qPCR (b) and by Western blot (c). d–e Cell invasiveness was examined in either a scratch wound healing assay (d) or a transwell cell migration assay (e). *p < 0.05. NS non-significant. n = 5. Statistics: one-way ANOVA with a Bonferroni correction
miR-494 targets 3′UTR of Pttg1 mRNA to inhibit its expression
Since our data suggest that miR-494 inhibits Pttg1 expression in CC cells, we performed bioinformatics analysis of Pttg1 target sequence. Our data suggest that the miR-494 binding sites in the Pttg1 mRNA sequence 3′UTR ranged from 473th base site to 479th base site (Fig. 4a). H3-miR-494, H3-scr (control), and H3-as-miR-494 cells were then transfected with 1 μg of Pttg1-3′UTR luciferase-reporter plasmid. We found that the luciferase activities in H3-as-miR-494 cells were significantly higher than the control scr, while the luciferase activities in H3-miR-494 cells were significantly lower than the control scr (Fig. 4b). These data suggest that miR-494 targets 3′UTR of Pttg1 mRNA to inhibit its expression. Thus, Pttg1 may increase CC cell metastasis, which is negatively regulated by miR-494 (Fig. 5).
miR-494 targets 3′UTR of Pttg1 mRNA to inhibit its expression. a Bioinformatics analysis of Pttg1 target sequence. The miR-494 binding sites in the Pttg1 mRNA sequence 3′UTR ranged from 473th base site to 479th base site. b H3-miR-494, H3-scr (control) and H3-as-miR-494 cells were then transfected with 1 μg of Pttg1-3′UTR luciferase-reporter plasmid. The luciferase activities were determined
Discussion
It has been previously reported that Pttg1 levels significantly increase in some cancers, whereas its participation in the carcinogensis of CC is unknown [5–8]. Moreover, the molecular mechanisms that underlie the regulation of cancer growth and metastasis by Pttg1 are far from clarified [5–8].
So far, it has been known that Pttg1 may target securing to prevent sister chromatids from separation until ubiquitination by the anaphase-promoting complex (APC), resulting in proteolysis by the proteasome [5–8]. Moreover, Pttg1 may regulate some cellular processes, like DNA repair, apoptosis, and gene regulation [5–8]. Of note, the multipotent gene c-myc has been identified as a target for Pttg1, in that the phosphorylation and activation of c-myc may drive transactivation-mediated signal transduction [5–8]. In addition, Pttg1 promotes the transactivation of bFGF, which subsequently tumor-associated neo-angiogenesis [5–8]. Further, Pttg1 is capable of inhibiting p53-induced apoptosis through its direct binding to the p53 DNA-binding domain [5–8].
Although many progresses have been obtained on the downstream signaling transduction of Pttg1, relatively less is known about its regulation. A recent study has detected Pttg1 mRNA and protein in H3 cells [41]. Moreover, a regulatory relationship between miR-494 and Pttg1 has been acknowledged in human cholangiocarcinoma [42]. These studies prompted us to examine a role of Pttg1 in CC, as well as its interaction with miR-494 in CC cells.
Here, we reported significantly higher levels of Pttg1 and significantly lower levels of miR-494 in the resected CC tissue, compared with the adjacent normal cervical tissue from the same patient. Interestingly, Pttg1 levels inversely correlated with miR-494 levels. These clinical findings are strong evidence for our hypothesis and important basis for our later metabolic study. In vitro, Pttg1 levels were found to positively affect CC cell invasiveness and were inhibited by miR-494 levels. However, miR-494 levels were not affected by Pttg1 levels. This is not surprising since miRNAs generally function and regulate the gene expression post-transcriptionally through their base-pairing with the 3′-UTR of target mRNA [29, 30]. To prove it, we performed luciferase-reporter assay using Pttg1-3′UTR plasmid. We found higher miR-494 levels resulted in lower bioluminescence levels, confirming our hypothesis that miR-494 inhibits Pttg1 expression in CC cells, through directly binding and inhibition on 3′-UTR of Pttg1 mRNA.
Together, our data suggest that Pttg1 may increase CC cell metastasis, which is negatively regulated by miR-494. Our work thus highlights a novel molecular regulatory machinery in metastasis of CC and sheds light on targeting interaction of Pttg1 and miR-494 as a therapeutic target for CC therapy.
References
Kanayama T, Mabuchi S, Fujita M, Kimura T. Calcaneal metastasis in uterine cervical cancer: a case report and a review of the literature. Eur J Gynaecol Oncol. 2012;33:524–5.
Setoodeh R, Hakam A, Shan Y. Cerebral metastasis of cervical cancer, report of two cases and review of the literature. Int J Clin Exp Pathol. 2012;5:710–4.
Peters P, Bandi H, Efendy J, Perez-Smith A, Olson S. Rapid growth of cervical cancer metastasis in the brain. J Clin Neurosci. 2010;17:1211–2.
Chen B, Zhang C, Dong P, Guo Y, Mu N. Molecular regulation of cervical cancer growth and invasion by VEGFA. Tumour Biol. 2014;35:11587–93.
Yu R, Ren SG, Horwitz GA, Wang Z, Melmed S. Pituitary tumor transforming gene (pttg) regulates placental JEG-3 cell division and survival: evidence from live cell imaging. Mol Endocrinol. 2000;14:1137–46.
Caporali S, Alvino E, Levati L, Esposito AI, Ciomei M, Brasca MG, et al. Down-regulation of the pttg1 proto-oncogene contributes to the melanoma suppressive effects of the cyclin-dependent kinase inhibitor PHA-848125. Biochem Pharmacol. 2012;84:598–611.
Vlotides G, Eigler T, Melmed S. Pituitary tumor-transforming gene: physiology and implications for tumorigenesis. Endocr Rev. 2007;28:165–86.
Tfelt-Hansen J, Kanuparthi D, Chattopadhyay N. The emerging role of pituitary tumor transforming gene in tumorigenesis. Clin Med Res. 2006;4:130–7.
Panguluri SK, Yeakel C, Kakar SS. Pttg: an important target gene for ovarian cancer therapy. J Ovarian Res. 2008;1:6.
El-Naggar SM, Malik MT, Kakar SS. Small interfering RNA against Pttg: a novel therapy for ovarian cancer. Int J Oncol. 2007;31:137–43.
Chen G, Li J, Li F, Li X, Zhou J, Lu Y, et al. Inhibitory effects of anti-sense Pttg on malignant phenotype of human ovarian carcinoma cell line SK-OV-3. J Huazhong Univ Sci Technol Med Sci = Hua zhong ke ji da xue xue bao Yi xue Ying De wen ban = Huazhong keji daxue xuebao Yixue Yingdewen ban. 2004;24:369–72.
Yan S, Zhou C, Lou X, Xiao Z, Zhu H, Wang Q, et al. Pttg overexpression promotes lymph node metastasis in human esophageal squamous cell carcinoma. Cancer Res. 2009;69:3283–90.
Zhou C, Liu S, Zhou X, Xue L, Quan L, Lu N, et al. Overexpression of human pituitary tumor transforming gene (hpttg), is regulated by beta-catenin/TCF pathway in human esophageal squamous cell carcinoma. Int J Cancer. 2005;113:891–8.
Shibata Y, Haruki N, Kuwabara Y, Nishiwaki T, Kato J, Shinoda N, et al. Expression of Pttg (pituitary tumor transforming gene) in esophageal cancer. Jpn J Clin Oncol. 2002;32:233–7.
Zhang ML, Lu S, Zheng SS. Epigenetic changes of pituitary tumor-derived transforming gene 1 in pancreatic cancer. Hepatobil Pancreat Dis Int. 2008;7:313–7.
Ai J, Zhang Z, Xin D, Zhu H, Yan Q, Xin Z, et al. Identification of over-expressed genes in human renal cell carcinoma by combining suppression subtractive hybridization and cDNA library array. Sci China C Life Sci. 2004;47:148–57.
Dominguez A, Ramos-Morales F, Romero F, Rios RM, Dreyfus F, Tortolero M, et al. Hpttg, a human homologue of rat Pttg, is overexpressed in hematopoietic neoplasms. Evidence for a transcriptional activation function of hpttg. Oncogene. 1998;17:2187–93.
Zhou C, Tong Y, Wawrowsky K, Melmed S. Pttg acts as a stat3 target gene for colorectal cancer cell growth and motility. Oncogene. 2014;33:851–61.
Kim DS, Franklyn JA, Smith VE, Stratford AL, Pemberton HN, Warfield A, et al. Securin induces genetic instability in colorectal cancer by inhibiting double-stranded DNA repair activity. Carcinogenesis. 2007;28:749–59.
Huang SQ, Liao QJ, Wang XW, Xin DQ, Chen SX, Wu QJ, et al. RNAi-mediated knockdown of pituitary tumor-transforming gene-1 (Pttg1) suppresses the proliferation and invasive potential of PC3 human prostate cancer cells. Braz J Med Biol Res. 2012;45:995–1001.
Huang S, Liao Q, Li L, Xin D. Pttg1 inhibits smad3 in prostate cancer cells to promote their proliferation. Tumour Biol. 2014;35:6265–70.
Castilla C, Flores ML, Medina R, Perez-Valderrama B, Romero F, Tortolero M, et al. Prostate cancer cell response to paclitaxel is affected by abnormally expressed securin Pttg1. Mol Cancer Ther. 2014;13:2372–83.
Cao XL, Gao JP, Wang W, Xu Y, Shi HY, Zhang X. Expression of pituitary tumor transforming gene 1 is an independent factor of poor prognosis in localized or locally advanced prostate cancer cases receiving hormone therapy. Asian Pac J Cancer Prev. 2012;13:3083–8.
Castilla C, Flores ML, Conde JM, Medina R, Torrubia FJ, Japon MA, et al. Downregulation of protein tyrosine phosphatase ptpl1 alters cell cycle and upregulates invasion-related genes in prostate cancer cells. Clin Exp Metastasis. 2012;29:349–58.
Singh SV, Powolny AA, Stan SD, Xiao D, Arlotti JA, Warin R, et al. Garlic constituent diallyl trisulfide prevents development of poorly differentiated prostate cancer and pulmonary metastasis multiplicity in tramp mice. Cancer Res. 2008;68:9503–11.
Bacac M, Provero P, Mayran N, Stehle JC, Fusco C, Stamenkovic I. A mouse stromal response to tumor invasion predicts prostate and breast cancer patient survival. PLoS One. 2006;1:e32.
Crosby ME, Jacobberger J, Gupta D, Macklis RM, Almasan A. E2f4 regulates a stable G2 arrest response to genotoxic stress in prostate carcinoma. Oncogene. 2007;26:1897–909.
Zhu X, Mao Z, Na Y, Guo Y, Wang X, Xin D. Significance of pituitary tumor transforming gene 1 (Pttg1) in prostate cancer. Anticancer Res. 2006;26:1253–9.
Di Leva G, Croce CM. MiRNA profiling of cancer. Curr Opin Genet Dev. 2013;23:3–11.
Pereira DM, Rodrigues PM, Borralho PM, Rodrigues CM. Delivering the promise of miRNA cancer therapeutics. Drug Discov Today. 2013;18:282–9.
He W, Li Y, Chen X, Lu L, Tang B, Wang Z, et al. Mir-494 acts as an anti-oncogene in gastric carcinoma by targeting c-myc. J Gastroenterol Hepatol. 2014;29:1427–34.
Zhou RP, Chen G, Shen ZL, Pan LQ. Cinobufacin suppresses cell proliferation via mir-494 in BGC-823 gastric cancer cells. Asian Pac J Cancer Prev. 2014;15:1241–5.
Bai Y, Sun Y, Peng J, Liao H, Gao H, Guo Y, et al. Overexpression of secretagogin inhibits cell apoptosis and induces chemoresistance in small cell lung cancer under the regulation of mir-494. Oncotarget. 2014;5:7760–75.
Romano G, Acunzo M, Garofalo M, Di Leva G, Cascione L, Zanca C, et al. Mir-494 is regulated by ERK1/2 and modulates trail-induced apoptosis in non-small-cell lung cancer through bim down-regulation. Proc Natl Acad Sci U S A. 2012;109:16570–5.
Liborio-Kimura TN, Jung HM, Chan EK. Mir-494 represses hoxa10 expression and inhibits cell proliferation in oral cancer. Oral Oncol. 2015;51:151–7.
Ries J, Vairaktaris E, Agaimy A, Kintopp R, Baran C, Neukam FW, et al. Mir-186, mir-3651 and mir-494: potential biomarkers for oral squamous cell carcinoma extracted from whole blood. Oncol Rep. 2014;31:1429–36.
Kwak SY, Yang JS, Kim BY, Bae IH, Han YH. Ionizing radiation-inducible mir-494 promotes glioma cell invasion through EGFR stabilization by targeting p190b RhoGap. Biochim Biophys Acta. 1843;2014:508–16.
Asuthkar S, Velpula KK, Nalla AK, Gogineni VR, Gondi CS, Rao JS. Irradiation-induced angiogenesis is associated with an mmp-9-mir-494-syndecan-1 regulatory loop in medulloblastoma cells. Oncogene. 2014;33:1922–33.
Puck TT, Marcus PI, Cieciura SJ. Clonal growth of mammalian cells in vitro; growth characteristics of colonies from single Hela cells with and without a feeder layer. J Exp Med. 1956;103:273–83.
Liang CC, Park AY, Guan JL. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc. 2007;2:329–33.
Solbach C, Roller M, Peters S, Nicoletti M, Kaufmann M, Knecht R. Pituitary tumor-transforming gene (Pttg): a novel target for anti-tumor therapy. Anticancer Res. 2005;25:121–5.
Yamanaka S, Campbell NR, An F, Kuo SC, Potter JJ, Mezey E, et al. Coordinated effects of microRNA-494 induce G(2)/M arrest in human cholangiocarcinoma. Cell Cycle. 2012;11:2729–38.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
The Publisher and Editor retract this article in accordance with the recommendations of the Committee on Publication Ethics (COPE). After a thorough investigation we have strong reason to believe that the peer review process was compromised.
An erratum to this article is available at http://dx.doi.org/10.1007/s13277-017-5487-6.
About this article
Cite this article
Chen, B., Hou, Z., Li, C. et al. RETRACTED ARTICLE: MiRNA-494 inhibits metastasis of cervical cancer through Pttg1. Tumor Biol. 36, 7143–7149 (2015). https://doi.org/10.1007/s13277-015-3440-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3440-0