Abstract
Aim
Cervical cancer is the most common gynecological malignancy in the developing countries like India. In addition to human papillomavirus (HPV) infection, host genetic factors play an important role in viral persistence and neoplastic growth. IL-10, a multifunctional cytokine, plays an active role to promote tumor growth in the presence of HPV. The present study aims to find out the impact of IL-10 promoter polymorphisms at -1082A/G (rs1800896), -819C/T (rs1800872), and -592C/A (rs1800871) sites along with IL-10 production and HPV infection in the progression of cervical cancer.
Methods
We have genotyped a total of 506 subjects, 256 cases (208 cervical cancer + 48 precancer), and 250 healthy controls by using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) method followed by sequencing. IL-10 serum concentration was measured by enzyme-linked immunosorbent assay.
Results
The frequency of IL-10 -592 variant genotype (AA) was found significantly reduced in cases as compare to controls while -1082 variant genotype (GG) was found ~4-fold higher risk of cervical cancer (p = <0.0001, OR = 3.667, 95 % CI = 2.329–5.773). On construction of haplotypes, GTC haplotype was emerged as a major risk haplotype while ACA haplotype was seemed as a marker for precancerous lesions. IL-10 serum concentration was observed higher in HPV-infected precancer and cancer cases. GTC haplotype was found to be coupled with higher serum concentration of IL-10 and HPV infection.
Conclusion
IL-10 polymorphisms play a role in cervical cancer development and that GTC haplotype, which is closely related to its serum concentration, maybe a useful biomarker for HPV-mediated cervical cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer of uterine cervix is the fourth most common gynecological cancer worldwide and a leading cause of cancer-related deaths among women in developing countries like India. In India, more than 1, 34,420 new cases, roughly 1/5th of global burden, are reported every year with estimated 75,000 deaths annually [1]. Although for the development of cervical cancer, human papillomavirus (HPV) is known to be the most important etiologic agent but, unaided, it is not sufficient for disease manifestation. Other factors intrinsic to the host put their role in the progression of tumorigenesis [2, 3]. In this context, the general or local immune status plays a role in regulating the growth of HPV-transformed cells. Specifically, cell-mediated immune responses are important for controlling both HPV infections and HPV-associated neoplasm [4]. Cell-mediated immunity is regulated by cytokines that are secreted by T helper cells. On the basis of cytokines production pattern, the cells can be classified in two types, i.e., Th1 and Th2 [5]. A predominance of the Th2 cytokine profile, in association with a diminished Th1 profile, has been demonstrated in patients with neoplasm. This shift from Th1 to Th2 cytokines might be responsible for tumorigenesis [6].
Interleukin-10 (IL-10), a multifunctional Th2-cytokine, located on chromosome 1 (1q31-1q32) [7] acts as an immune response modulator [8]. In the presence of HPV oncoproteins, IL-10 was found to function as an anti-inflammatory agent [9]. Two viral oncoproteins of HPV-16, E6 and E7, play an active role in the malignant growth properties of cervical cancer cells [10]. IL-10 is found to promote tumor growth when E6 oncoprotein activity is high [11]. Moreover, HPV E2 protein binds to the regulatory region of the IL-10 gene and induces the expression of elevated levels of IL-10 mRNA in HPV-infected cells.
Three dimorphic polymorphisms in the promoter region, at positions -1082 A/G (rs1800896), -819 T/C (rs1800871) and -592 A/C (rs1800872) have been identified in many diseases [12–17]. According to several previous studies, altered genetic make-up appears to be responsible with modulated levels of IL-10 [18, 19]. Although many studies have investigated the roles of IL-10 promoter polymorphisms and the risk of cervical cancer, the findings remain inconclusive, which may be due to differences in study design and/or ethnic differences in the populations studied.
However, with the preeminent comprehension, no report is available to address the role of IL-10 promoter polymorphisms in the progression of cervical cancer. Therefore, the present study aimed to investigate the effects of IL-10 genetic variants, HPV infection, variation in serum concentration, and their interactions in the development of cervical carcinogenesis. We have also examined the effects of these genotypes/haplotypes with the invasiveness of cervical carcinoma.
Materials and methods
Sample collection
In the present study, a total of 506 consecutive subjects consisting of 256 cases (48 precancer + 208 invasive carcinoma) and 250 healthy controls were included. The patients were enlisted from Lok Nayak Jai Prakash Hospital and Safdarjung Hospital, New Delhi with histological confirmed precancer (cervical intraepithelial neoplasia (CIN) I, II, III)/invasive carcinoma of the uterine cervix. Classification and grading of the precancerous and cancerous lesions were done according to World Health Organization criteria and CIN classification system. The tumor stage and histological grade were determined according to the criteria laid down by International Federation of Gynecology and Obstetrics (FIGO). Age and ethnicity matched 250 control samples (cervical scrapes/tissue biopsy) with normal cervical cytology and having no history of cancer were obtained from women who visit the hospital for other gynecological reasons. Age at the time of diagnosis was recorded. Written consent from all the patients was obtained, and the study was carried out in accordance with the principles of Helsinki Declaration. The study was approved by the ethics committee of the institute.
DNA extraction and HPV detection
Genomic DNA was extracted from fresh cervical cancer/precancer tissue biopsy samples as well as control (cervical scrapes/hysterectomy biopsy) samples by standard proteinase K digestion followed by phenol/chloroform/isopropanol treatment [20, 21]. HPV diagnosis was performed by PCR amplification using consensus primers MY09 and MY11 [22], and further typing was done by PCR using type specific primers for HPV 16 and HPV18 [20, 21].
Detection of IL-10 polymorphism by PCR-RFLP
We used polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) approach to genotype the three loci of the IL-10 gene promoter region. All three loci -1082 A/G, -819 C/T, and -592 A/C were genotyped by method employed by Roh et al. [23]. The RFLP analysis was performed on 10 % native polyacrylamide gel. In Brief, for -1082 loci, a PCR product of 139 bp was digested overnight at 37 °C with 2U of MnlI (Fermentas Life Sciences). The digested PCR products were separated in 10 % polyacrylamide gel electrophoresis. The wild type A allele produced a single 139 bp fragment, and the polymorphic G allele produced fragments of 106 and 33 bp. (Fig. 1). For the genotyping of -819 loci, a PCR product of 209 bp was amplified and digested with 2U of Mae III at 55 ° C for 1 hour. In the presence of wild type C allele, splitting gives rise to two fragments of 125 and 84 bp while in case of T allele, no splitting occurs (Fig. 2). RsaI enzyme was used for the genotyping of -592 loci. A PCR product of 412 bp was digested with 2U of the enzyme at 37 °C overnight and was resolved by 10 % PAGE. With the A allele, a cleavage of PCR product into two fragments 176 and 236 bp occurred. However, with the C allele, no cleavage occurred (Fig. 3).
RFLP IL-10 -(A1082G) on 15 % PAGE (after digestion with restriction enzyme Mnl1), M mol. wt. marker (Hae III digested φX174 DNA), lanes 1, 6, 8: AA genotype: 139 bp fragment, lanes 2, 5, 7, 8, 11, 12: GA genotype: 139, 106 bp fragment, lanes 3, 4, 9: GG genotype: 106, 33 bp fragments. Number sign indicates precancerous lesion. Asterisk indicates HPV-positive cases
RFLP IL-10 -(C819T) on 10 % PAGE (after digestion with restriction enzyme Mae III), M mol. wt. marker (Hae III digested φX174 DNA), lanes 3, 5: CC genotype: 125, 84 bp fragments, lanes 2, 4, 6, 7, 8, 9, 12: CT genotype: 209, 125, 84 bp fragments, lanes 1, 10: TT genotype: 209 bp. Number sign indicates precancerous lesion. Asterisk indicates HPV-positive cases
RFLP IL-10 -(C592A) on 10 % PAGE (after digestion with restriction enzyme Rsa I). M mol. wt. marker (Hae III digested φX174 DNA), Lanes 3, 4, 8, 9, 12: CC genotype: 412 bp. Lanes 1, 5, 6, 7, 10: CA genotype: 412, 236, 176 bp fragments, lanes 2, 11: AA genotype: 236, 176, bp fragments. Number sign indicates precancerous lesion. Asterisk indicates HPV-positive cases
Sequencing
We sequenced 20 % of the samples randomly to validate the data generated by PCR-RFLP method. Sequencing reactions were performed according to the conventional dideoxy chain termination method using ABI Prism™ 310 automated DNA sequencer (Applied Biosystem, USA). Same results were observed by both the techniques.
Enzyme-linked immunosorbent assay (ELISA) for IL-10
The concentration of IL-10 in human serum samples were measured by ELISA using Human IL-10 ELISA Kit as per the manufacturer protocol (GEN-PROBE, Diaclone (Cat. No.950.060.192)). ELISA kit contained IL-10 coated plate, standard, standard diluents, control, biotinylated anti-IL-10, streptavidin-HRP, and TMB substrate and H2SO4 stop reagent. Briefly, 100 μl of sample (human serum) was added in IL-10 coated wells along with diluted standard/controls which were followed by the 2-h incubation with 50-μl biotinylated anti-IL-10 at room temperature. The plate was washed thrice to remove any excess unbound analyte and secondary antibody. Bound antibodies were detected with conjugate streptavidin-HRP and adding a chromogen substrate TMB solution which result into the development of a blue-colored complex. The color reaction was stopped by adding H2SO4 as stop reagent which produced the yellow color product, and absorbance was evaluated and recorded at 450 nm. The concentration of IL-10 in samples, standards, and control was directly proportional to intensity of the color complex.
Statistical analysis
The data analysis was performed by using statistical software PLINK v.1.07 (http://pngu.mgh.harvard.edu/purcell/plink) and GraphPad InStat version 3.0. Chi-square test/Fisher’s exact test (for smaller numbers on subgroup analysis) was used to compare the distributions of IL-10 polymorphisms between cervical cancer/precancer patients and healthy controls. Risk estimates were calculated for dominant, codominant, and recessive genetic models using the most common homozygous genotype as reference. The odds ratio (OR) and its 95 % confidence intervals were also calculated as a measure of the association between these three polymorphisms and cervical cancer risk. Haplotypes were constructed from genotypes of these three polymorphic markers by using PLINK v.1.07 (http://pngu.mgh.harvard.edu/purcell/plink). LD estimates were determined by Haploview (http://www.broad.mit.edu/mpg/haploview/). The significance of statistical chi-square/Fisher’s exact test was considered as two-tailed. P value <0.05 was considered as significant. Genotypes were further checked for the conformance of Hardy Weinberg equilibrium. Statistical power of the study determined using Quanto software (http://hydra.usc.edu/gxe/) was >80 %.
Results
Population characteristics
The basic characteristics of the study population are listed in Table 1. No significant difference was observed in the distribution of ages among controls, precancer, and cancer group. In cancer patients, 96 % were squamous cell carcinoma (SCC) while 4 % were adenocarcinoma (ADC). In SCC cases, 26, 49, and 21 % of patients had well differentiated SCC (WDSCC), moderately differentiated SCC (MDSCC) and poorly differentiated SCC (PDSCC), respectively. Out of 208 cancer cases, 77 (37 %) were of stage I + II and 131(63 %) were of stage III + IV.
Prevalence of HPV in cervical cancer
Frequency of HPV in the studied population has been presented in Table 1. 89.4 % (229/256) (91 % (190/208) in cancerous cases and 81 % (39/48) in precancerous cases) of total cases were found to be positive for HPV DNA sequence, while only 5 % (12/250) healthy controls were found to be HPV positive. The difference in the prevalence of HPV among the three groups, i.e., healthy controls, precancer, and cancer was found statistically significant (p < 0.0001). Out of the HPV-positive cases, 94.8 % (217/229) (98.4 % (187/190) cancerous cases and 76.9 % (30/39) precancerous cases) were positive for HPV-16. HPV-18 was found positive in 1.57 % (3/190) of cancerous cases. But all of HPVs in controls were infected with HPV type-16. Prior to HPV specific PCR, quality of genomic DNA from all the samples was checked by β-globin gene amplification which was used as internal control.
Interleukin-10 polymorphisms in cervical cancer
The genotypic distribution and allelic frequencies of -1082 (A/G), -819 (C/T), and -592 (C/A) in the cases (cancer/precancer) and control population are presented in Table 2. We used three different types of genetic models (dominant, codominant, and recessive) to evaluate the association of IL-10 genotypes with cervical cancer.
Genotypic distribution of IL-10 -592 (C/A) polymorphism
The genotypic distribution of -592 locus CC, CA, and AA were 40, 46, and 14 % in cases and 24, 49, and 27 % in controls, respectively (Table 2). When we compared control group with the cancer group, higher frequency of carrier genotype (CA + AA) was observed in controls which was found to be statistically significant in dominant model (p = 0.0002, OR = 0.4768, 95 % CI = 0.30–0.68). But in recessive model, it was observed that AA genotype differ significantly in cases (includes cancerous and precancerous group) as compared to controls (p = 0.0009, OR = 0.4615, 95 % CI = 0.29–0.72). These findings illustrated the dominant effect of AA genotype in the development of cervical cancer. Simultaneously, it was interesting to note that there was gradual reduction in frequency of variant genotype AA from control group (27 %) to precancerous group (17 %) and further in cancerous group (14 %). Therefore, the frequency of the A allele was found significantly lesser in cases (p = 9.4 × 10−6) including cancerous group (p = 6.6 × 10−6) as compared to control group. Frequency of -592 CC genotype was found higher in HPV-infected women as compared to AA genotype which shows the possible impact of C allele in the persistent infection of HPV.
Genotypic distribution of IL-10 -819 (C/T) polymorphism
The frequency of CC, CT, TT genotype of IL-10 -819 C/T locus was 28, 48, and 24 % in controls while 21, 50, and 29 % in cases, respectively. We observed that there was a difference in carrier genotype (CT + TT) distribution between cases and controls with 79 % (191/256) in cases and 72 % (181/250) in controls but the difference was not found statistically significant in either of the model. While comparing the genotype distribution between control group and precancer/cancer group, no significant p values were observed but it was interesting to note that there is increment of the frequency of variant TT genotype from control group (24 %) to precancerous group (27 %) and then precancerous (27 %) to cancerous (29 %) group. Difference in the distribution of allelic frequencies of C and T allele was observed in cancer patients, but the difference was not found significant. We could not find any association of HPV infection and -819 C/T polymorphism.
Genotypic distribution of IL-10 -1082 (A/G) polymorphism
The genotypic distributions of AA, GA, and GG of -1082 A/G locus were 40, 43, and 17 % for controls while 17, 38, and 45 %, respectively, in cases. When dominant model was taken into consideration, we observed a more than threefold increased risk of cervical cancer in cases having carrier genotype (GA + GG) as compared to controls (p = <0.0001, OR = 3.302, 95 % CI = 2.183–4.99). When we compared controls with cases (precancer/cancer group), we observed that the patients from precancer group who had GA + GG genotype were at ~2-fold higher risk of cervical cancer as compared to controls (p = 0.0376, OR = 2.242, 95 % CI = 1.09–4.60) but for cancer patients, the risk was ~4-fold higher than controls (p = <0.0001, OR = 3.667, 95 % CI = 2.329–5.773). The data from recessive model shows that the patients carrying GG genotype were at 4-fold higher risk of cervical cancer (p = <0.0001, OR = 3.989, 95 % CI = 2.646–6.013). In recessive model, we further stratified the data into precancer and cancer group; the risk was ~2.5-fold more for precancer (p = 0.0177, OR = 2.409, 95 % CI = 1.214–4.772) and ~4.5 times higher for cancer (p = <0.0001, OR = 4.457, 95 % CI = 2.909–6.829) as compared to controls. The risk was also shown to be significantly increased in both the groups, i.e., precancer and cancer with homozygous variant GG genotype in codominant model. Therefore, the heterozygous mutant GA genotype presented a 2.1-fold higher risk of cervical cancer, overall, while 1.1-fold for precancerous and 2.2-fold for cancerous. In consideration of allelic frequencies, our data indicates that as the frequency of G allele elevates in cases, the risk of cervical cancer also increases. On further analysis of -1082 genotypic data in association with HPV infection, we observed that GG genotype was more frequent in HPV-infected cases.
Whether all the three polymorphisms of IL-10 play some role in the progression of cervical cancer?
We further stratified our results in different clinical stages and histological grades of cervical cancer which has been presented in Table 3. We observed a decrease in the frequency of carrier genotype (CA + AA) of IL-10, -592 loci with the progression of cervical cancer (Table 3). As compared to controls, the frequency was significantly much lesser in advance clinical stages (p = 0.0029, OR = 0.386, 95 % CI = 0.2108–0.7068) as well as histological grades (p = 0.0046, OR = 0.4019, 95 % CI = 0.2194–0.7364) of the cancer. On the other hand, we could not found any trend in the frequency of carrier genotypes (CT + TT) of -819 loci with the progression of cervical cancer (Table 3). Interestingly, a constant increase in the frequency of carrier genotypes (AG + GG) of -1082 loci was observed from control group to the advance stages of cervical cancer (Table 3). As compared to controls, the frequency of carrier genotypes was ~2 times higher in precancerous group (p = 0.0149, OR = 2.232, 95 % CI = 1.208–4.125) which was found statistically significant. As the invasion of cancerous cells was increased, the frequency of AG + GG genotypes showed fold increment, i.e., 2-fold in initial stages (I + II) (p = 0.0093, OR = 2.364, 95 % CI = 1.27–4.39) while 5-fold in advance stages (III + IV) (p = <0.0001, OR = 5.394, 95 % CI = 2.565–11.344) as compared to controls. Interestingly, we observed the same trend of carrier genotype frequency in different histological grades of the cervical cancer where the risk factor increased by the degree of 2, i.e., 2.0 in WDSCC, 4.0 in MDSCC, and 6.0 in PDSCC.
What haplotype does in the succession of cervical carcinoma?
Haplotype analysis using statistical software PLINK showed the presence of 8 haplotype which has been shown in Table 4. All the haplotypes were present in >1 % of population. Two haplotypes: GTC (32.7 %) and ACA (26.7 %) were the most frequent. The frequency of GTC was found almost twice in cases as compared to controls which was found statistically significant (p = <0.0001, OR = 2.99). Out of eight, five haplotypes were found under-presented in cases or higher risk group with respect to controls of which only two were found significantly higher in controls, i.e., ATA and ATC. ATA was found significant in controls vs. cancer (p = <0.0001, OR = 0.0164) as well as in control vs. precancer (p = 0.035, OR = 0.281) while another, i.e., ATC was found significant only in cancer cases (p = <0.0001, OR = 0.0764). Interestingly, the frequency of ACA haplotype was higher in precancerous group (35 %) as compared to controls (26.8 %) and cancer (26 %) which may be due to the less sample size. On the other hand, GTC haplotype was found to be positively associated with control vs. cases (p = <0.0001, OR = 2.99), control vs. cancer (p = <0.0001, OR = 3.16) and control vs. precancer (p = <0.0001, OR = 3.44). On further stratification of the haplotyping results with the invasiveness of cervical carcinoma, some interesting findings were exposed which have been shown in Table 5. The two haplotypes which were found most frequent in our population were further checked for their role during different stages of cervical carcinogenesis. We observed that the frequency of ACA haplotype was reduced in the advance stages of cancer clinically (22 %) and histologically (18 %) as compared to controls (27 %) while in contrast, the frequency of GTC haplotype was found higher and the difference in the frequencies of both the haplotypes was found statistically significant in both clinical (p = 0.0136, OR = 2.842) as well as histological (p = 0.0038, OR = 3.47) grades.
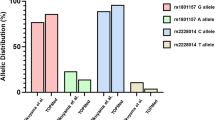
To complement our knowledge, we also tried to find out the association of HPV with GTC haplotype if any. On analysis, GTC haplotype was interestingly found more frequent (>95 %) in HPV-infected women as compared to any other haplotype which shows the possible role of IL-10 gene promoter polymorphisms in HPV-related cervical cancer (Fig. 4).
Serum concentration of IL-10
The total serum concentrations of IL-10 have been presented in Table 1. We observed a constant increase in serum concentration of IL-10 from controls (37.03 ± 0.4 pg/ml) to precancer (77.13 ± 0.35 pg/ml) and cancer (146.8 ± 0.4 pg/ml) group and found to be statistically significant by using ANOVA test (p < 0.0001). We could not found any correlation between serum level and genotypes of IL-10. But on construction of haplotypes, we found a significant association of GTC haplotype with higher serum concentration of IL-10 (Fig. 5) while ACA haplotype showed the lowest serum levels of IL-10. Interestingly, we noticed that all the samples having GTC haplotype along with high serum concentration were infected with HPV.
Discussion
Cervical cancer is the most common gynecological cancer in the developing countries including India. Though extensive screening program has been lowered down, the incidence in west but in resource poor countries like India cervical cancer shares the major health burden in females. Commercially, two prophylactic HPV vaccines are available and are very promising too but unfortunately have no therapeutic value. Therefore, it is important to evaluate the role of different biomarkers in cervical cancer susceptibility for the better understanding of the disease etiology, which may contribute towards early detection and preventive care for cervical cancer. Interleukin-10 (IL-10) polymorphisms are reported to be associated with the inflammatory and immunomodulatory diseases including cancer. IL-10 is a multifunctional cytokine with both immunosuppressive and anti-angiogenic functions, consequently resulting in both tumor-promoting and tumor-inhibiting properties [8]. Abnormal IL-10 production has already been reported in patients with cervical precancerous lesion and invasive cancer. -1082 G allele has been reported with higher serum concentration while in contrast, -819 T allele and -592 A allele have also been reported for lower serum concentration [18, 19]. In the present study, we also tried to find out some correlation between serum concentration and genotype/haplotype.
In the present case-control study, we analyzed the association of IL-10 promoter polymorphisms with the risk of cervical carcinoma. We observed a significant difference in the allele frequency and genotype distribution for IL-10, -592 loci in controls and cases. This finding is similar to a previous report from India [24], where AC + CC genotype was at higher risk of cervical cancer. On the analysis of data in codominant model, we observed a significant difference in the distribution of AC genotype among healthy controls, precancer, and cancer patients which is in a good agreement to the findings by Zoodsma et al. [25], where the frequency of AC genotype was found higher in precancer and cancer cases as compared to controls. In contrast to our findings, several studies failed to find any association of IL-10 -592 polymorphism with cervical cancer [23, 26]. Interestingly, the frequency of C allele was observed to be higher in HPV-positive samples. We observed an increase in the frequency to CT + TT genotype at -819 loci of IL-10 gene, but the difference was not found statistically significant. This observation is very similar with the previous finding from India [27]. Furthermore, we noticed a gradual increase in the frequency of IL-10 -1082 carrier genotype from healthy controls to precancer and then precancer to cancer patients. This finding is in good agreement to the previous finding in Japanese population [28]. On the other hand, several studies could not find any association of IL-10 -1082 polymorphism with cervical cancer [23, 25, 29–31]. In the present study, G allele was found more frequent in HPV-infected women; this finding is in contrast with a recent report from Brazil [32].
We further stratified our results according to different clinical and histological grade to evaluate the role of IL-10 promoter polymorphisms in the progression as well as advancement of cervical cancer. After which, we came out with some interesting findings that the frequency of carrier genotype of -592 loci (AC + AA) was significantly lowered down from healthy controls towards advance stages of cancer while in contrast, the frequency of carrier genotype at -1082 loci (AG + GG) was found in direct proportion with the development and advancement of cervical carcinoma.
To complement further, our understanding of the contribution of these genetic variants to cervical cancer three locus haplotypes were constructed and their distribution was compared in the cancer, precancer, and healthy control population. Haplotype GTC was found to increase the risk of cervical cancer with p value <0.0001 when compared to controls while ATA haplotype appeared to show its protective role. Amusingly, ACA haplotype was found significantly higher in precancer group as compared to controls as well as cancer patients. Overall, GTC haplotype emerged as a major risk haplotype which showed a statistically significant trend from controls to precancer and then to cervical cancer while ACA haplotype act as a marker for high risk group. Our study is in concordant with the previous studies in prostate, oral, and nasopharyngeal cancer in relation to -1082 and -592 polymorphism while in contrast with respect to -819 [33–35]. These discrepancies in the findings may be due to different ethnic groups, geographical regions, and environmental factors. It was interesting to analyze the association of GTC and ACA haplotype with the aggressiveness of cervical carcinoma. The analysis illustrates that the frequency of GTC haplotype increased while the frequency of ACA haplotype reduced with the aggressiveness of tumor. As far as our knowledge is concerned, this is the first study showing the role of IL-10 -1082, -819, and -592 genotype/haplotype in the susceptibility, development, and progression of cervical cancer.
IL-10 gene transcriptional control is of interest as IL-10 has pivotal role in the regulation of the immune response system. IL-10 production has been implicated in the tumorogenesis of various types of cancers. Abnormal IL-10 production at cervix level has been reported in cervical lesions as well as in cervical cancer patients [36–38]. Similar to this study, various previous studies have been reported in the elevated production of IL-10 in cervical carcinoma [38, 39]. We observed an increase of IL-10 serum concentration from healthy controls to precancerous lesion and from precancerous lesion to cervical cancer patients. We noticed that GTC haplotype is in direct proportion of HPV infection and IL-10 serum concentration. The substitution of *G for *A at position -1082 was found to result in significantly higher levels of IL-10 [18] which is in good agreement with our study.
In context to HPV, we observed higher frequency of -592 C and -1082 G allele in HPV-infected samples. Interestingly, we noticed that HPV-infected samples were the carrier of GTC haplotype. From previous studies, it is known that IL-10 may inhibit immune response against HPV in early cervical lesions and these polymorphisms are responsible to reduce down the HPV clearance in immuno-suppressed cells. In this way, from these findings, we can hypothesize that a person having GTC haplotype of -1082/-819/-592 loci of IL-10 gene is more prone to get infected with HPV and ultimately progress to cervical cancer.
In conclusion, GTC haplotype in the presence of HPV is responsible for higher expression of IL-10 which may help in HPV replication, persistence, and malignant transformation of HPV-infected cell. This genetic information might therefore be used to improve treatment, vaccine efficacy, and effectiveness and ultimately contribute to the discovery of personalized therapy and vaccination.
References
Ferlay J, Soerjomataram I, Ervik M, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray F: GLOBOCAN 2012 v1.0, Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 11; 2013.
Kohaar I, Hussain S, Thakur N, Tiwari P, Nasare V, Batra S, et al. Association between human leukocyte antigen class II alleles and human papillomavirus-mediated cervical cancer in Indian women. Hum Immunol. 2009;70:222–9.
Zur HH. Papillomavirus infections—a major cause of human cancers. Biochim Biophys Acta. 1996;1288:F55–78.
Wu TC. Immunology of the human papilloma virus in relation to cancer. Curr Opin Immunol. 1994;6:746–54.
Wu TC, Kurman RJ. Analysis of cytokine profiles in patients with human papillomavirus-associated neoplasms. J Natl Cancer Inst. 1997;89:185–7.
Clerici M, Shearer GM, Clerici E. Cytokine dysregulation in invasive cervical carcinoma and other human neoplasias: time to consider the TH1/TH2 paradigm. J Natl Cancer Inst. 1998;90:261–3.
Eskdale J, Kube D, Tesch H, Gallagher G. Mapping of the human IL10 gene and further characterization of the 5' flanking sequence. Immunogenetics. 1997;46:120–8.
Mocellin S, Panelli MC, Wang E, Nagorsen D, Marincola FM. The dual role of IL-10. Trends Immunol. 2003;24:36–43.
Woodworth CD, Lichti U, Simpson S, Evans CH, DiPaolo JA. Leukoregulin and gamma-interferon inhibit human papillomavirus type 16 gene transcription in human papillomavirus-immortalized human cervical cells. Cancer Res. 1992;52:456–63.
Madrigal M, Janicek MF, Sevin BU, Perras J, Estape R, Penalver M, et al. In vitro antigene therapy targeting HPV-16 E6 and E7 in cervical carcinoma. Gynecol Oncol. 1997;64:18–25.
Vairaktaris E, Yapijakis C, Serefoglou Z, Derka S, Vassiliou S, Nkenke E, et al. The interleukin-10 (-1082A/G) polymorphism is strongly associated with increased risk for oral squamous cell carcinoma. Anticancer Res. 2008;28:309–14.
Burada F, Dumitrescu T, Nicoli R, Ciurea ME, Rogoveanu I, Ioana M. Cytokine promoter polymorphisms and risk of colorectal cancer. Clin Lab. 2013;59:773–9.
Liu P, Song J, Su H, Li L, Lu N, Yang R, et al. IL-10 gene polymorphisms and susceptibility to systemic lupus erythematosus: a meta-analysis. PLoS One. 2013;8:e69547.
Tsai CW, Tsai MH, Shih LC, Chang WS, Lin CC, Bau DT. Association of interleukin-10 (IL10) promoter genotypes with nasopharyngeal carcinoma risk in Taiwan. Anticancer Res. 2013;33:3391–6.
Wu Z, Zheng W, Xu J, Sun F, Chen H, Li P, Chen S, Shen M, Zhang W, You X, Wu Q, Zhang F, Li Y: IL10 polymorphisms associated with Behcet's disease in Chinese Han. Hum Immunol 2013.
Zheng XY, Guan WJ, Mao C, Chen HF, Ding H, Zheng JP, et al. Interleukin-10 promoter 1082/-819/-592 polymorphisms are associated with asthma susceptibility in Asians and atopic asthma: a meta-analysis. Lung. 2014;192:65–73.
Zou L, Wang L, Gong X, Zhao H, Jiang A, Zheng S: The association between three promoter polymorphisms of IL-10 and inflammatory bowel diseases (IBD): A meta-analysis. Autoimmunity 2013.
Turner DM, Williams DM, Sankaran D, Lazarus M, Sinnott PJ, Hutchinson IV. An investigation of polymorphism in the interleukin-10 gene promoter. Eur J Immunogenet. 1997;24:1–8.
Westendorp RG, Langermans JA, Huizinga TW, Elouali AH, Verweij CL, Boomsma DI, et al. Genetic influence on cytokine production and fatal meningococcal disease. Lancet. 1997;349:170–3.
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, et al. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988;239:487–91.
Sambrook Ra: Molecular Cloninig: A laboratory manual.; Cold Spring Harbor Laboratory Press 3, 2001.
Manos MM, Ting Y, Wright DK, Lewis AJ, Broker TR, Wolinski SM. Use of polymerase chain reaction amplification for the detection of genital human papillomaviruses. Cancer Cells. 1989;7:pp 209–14.
Roh JW, Kim MH, Seo SS, Kim SH, Kim JW, Park NH, et al. Interleukin-10 promoter polymorphisms and cervical cancer risk in Korean women. Cancer Lett. 2002;184:57–63.
Shekari M, Kordi-Tamandani DM, MalekZadeh K, Sobti RC, Karimi S, Suri V. Effect of anti-inflammatory (IL-4, IL-10) cytokine genes in relation to risk of cervical carcinoma. Am J Clin Oncol. 2012;35:514–9.
Zoodsma M, Nolte IM, Schipper M, Oosterom E, van der Steege G, de Vries EG, et al. Interleukin-10 and Fas polymorphisms and susceptibility for (pre)neoplastic cervical disease. Int J Gynecol Cancer. 2005;15 Suppl 3:282–90.
Ivansson EL, Gustavsson IM, Magnusson JJ, Steiner LL, Magnusson PK, Erlich HA, et al. Variants of chemokine receptor 2 and interleukin 4 receptor, but not interleukin 10 or Fas ligand, increase risk of cervical cancer. Int J Cancer. 2007;121:2451–7.
Singh H, Jain M, Sachan R, Mittal B. Association of TNFA (-308G > A) and IL-10 (-819C > T) promoter polymorphisms with risk of cervical cancer. Int J Gynecol Cancer. 2009;19:1190–4.
Matsumoto K, Oki A, Satoh T, Okada S, Minaguchi T, Onuki M, et al. Interleukin-10–1082 gene polymorphism and susceptibility to cervical cancer among Japanese women. Jpn J Clin Oncol. 2010;40:1113–6.
Barbisan G, Perez LO, Contreras A, Golijow CD. TNF-alpha and IL-10 promoter polymorphisms, HPV infection, and cervical cancer risk. Tumour Biol. 2012;33:1549–56.
Ni J, Ye Y, Teng F, Wu Q. Interleukin 10 polymorphisms and cervical cancer risk: a meta-analysis. Int J Gynecol Cancer. 2013;23:126–33.
Wang Q, Zhang C, Walayat S, Chen HW, Wang Y. Association between cytokine gene polymorphisms and cervical cancer in a Chinese population. Eur J Obstet Gynecol Reprod Biol. 2011;158:330–3.
Chagas BS, Gurgel AP, da Cruz HL, Amaral CM, Cardoso MV, Silva Neto JC, et al. An interleukin-10 gene polymorphism associated with the development of cervical lesions in women infected with Human Papillomavirus and using oral contraceptives. Infect Genet Evol. 2013;19:32–7.
Liu J, Song B, Bai X, Liu W, Li Z, Wang J, et al. Association of genetic polymorphisms in the interleukin-10 promoter with risk of prostate cancer in Chinese. BMC Cancer. 2010;10:456.
Wei YS, Kuang XH, Zhu YH, Liang WB, Yang ZH, Tai SH, et al. Interleukin-10 gene promoter polymorphisms and the risk of nasopharyngeal carcinoma. Tissue Antigens. 2007;70:12–7.
Yao JG, Gao LB, Liu YG, Li J, Pang GF. Genetic variation in interleukin-10 gene and risk of oral cancer. Clin Chim Acta. 2008;388:84–8.
de Gruijl TD, Bontkes HJ, van den Muysenberg AJ, van Oostveen JW, Stukart MJ, van Verheijen RH, et al. Differences in cytokine mRNA profiles between premalignant and malignant lesions of the uterine cervix. Eur J Cancer. 1999;35:490–7.
Giannini SL, Al-Saleh W, Piron H, Jacobs N, Doyen J, Boniver J, et al. Cytokine expression in squamous intraepithelial lesions of the uterine cervix: implications for the generation of local immunosuppression. Clin Exp Immunol. 1998;113:183–9.
Torres-Poveda K, Burguete-Garcia AI, Cruz M, Martinez-Nava GA, Bahena-Roman M, Ortiz-Flores E, et al. The SNP at -592 of human IL-10 gene is associated with serum IL-10 levels and increased risk for human papillomavirus cervical lesion development. Infect Agent Cancer. 2012;7:32.
Stanczuk GA, Sibanda EN, Perrey C, Chirara M, Pravica V, Hutchinson IV, et al. Cancer of the uterine cervix may be significantly associated with a gene polymorphism coding for increased IL-10 production. Int J Cancer. 2001;94:792–4.
Acknowledgment
We would like to thank Dr. Sudha Salhan, Department of Obstetrics and Gynaecology, Safdarjung Hospital, New Delhi and Dr. Swaraj Batra, Department of Obstetrics and Gynaecology, LNJP Hospital, New Delhi, India for providing clinical samples and valuable feedback for the manuscript. We also would like to thank Dr. Suresh Bhambhani, Division of Cytopathology for pathological analysis of the clinical sample. We would like to thank the patients and their family members too. We would like to thank Dr. Dwaipayan Bharadwaj, CSIR-IGIB, New Delhi for critical evaluation of the manuscript. This work was supported by the core funding of ICPO (ICMR) to MB. PS is grateful to the Indian Council of Medical Research (ICMR), and AK is grateful to the University Grant commission (UGC) for their Senior Research Fellowships.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Pallavi Singhal, Anoop Kumar, and Soham Bharadwaj contributed equally.
Rights and permissions
About this article
Cite this article
Singhal, P., Kumar, A., Bharadwaj, S. et al. Association of IL-10 GTC haplotype with serum level and HPV infection in the development of cervical carcinoma. Tumor Biol. 36, 2287–2298 (2015). https://doi.org/10.1007/s13277-014-2836-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2836-6