Abstract
Recent studies have provided new insights into the diagnostic value of circulating microRNAs (miRNAs) for breast cancer (BCa). However, the inconsistent results between studies have prevented the widespread usage of miRNAs in clinics. To systematically assess the potential diagnostic value of circulating miRNAs in BCa, we performed a comprehensive meta-analysis. Eligible studies were retrieved by searching electronic databases. The quality of the studies was assessed on the basis of quality assessment for studies of diagnostic accuracy (QUADAS) criteria. The bivariate meta-analysis model was employed to summarize the diagnostic indices and plot the summary receiver operator characteristic (SROC) curve. A total of 15 studies were included in this meta-analysis, involving 1368 BCa patients and 849 healthy controls. Our bivariate random effects meta-analysis yielded an area under curve (AUC) value of 0.9217, with a sensitivity of 0.82 (95 % confidence interval (CI) 0.80–0.83) and specificity of 0.82 (95 % CI 0.80–0.85) for the use of miRNAs in differentiating BCa patients from healthy controls. Notably, our subgroup analysis suggested that a combination of multiple miRNAs (AUC, sensitivity, and specificity of 0.9518, 0.87, and 0.88, respectively) seemed to harbor higher accuracy than single miRNA-based assays (AUC, sensitivity, and specificity of 0.8923, 0.79, and 0.77, respectively). Altogether, our data indicate that circulating miRNA profiling has a potential to be used as a screening test for BCa, among which, the detection of a combined multiple miRNAs may be a more comprehensive indicator than individual miRNA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BCa) is the leading cause of cancer-related deaths among women worldwide, accounting for about 30 % of all types of cancer cases [1]. Early detection is important for BCa treatment, as the overall 5-year survival can reach to 90 % when diagnosed at an early stage as opposed to 20 % when the cancer has already spread to distant organs [2]. Currently, mammography and ultrasound have been successful and standard screening tools for identifying early-stage BCa. Nevertheless, the diagnostic accuracies of these techniques are subject to various factors, including the breast density, professional experiences of the examiner, etc. [3, 4]. In addition, the high examination cost also hampers the wide application of these procedures. On the other hand, blood-based screening method characterized by the detection of circulating tumor biomarkers is easy, convenient, and low cost. Such biomarkers like carcinoembryonic antigen (CEA), carbohydrate antigen 15-3 (CA15-3), or tissue polypeptide-specific antigen (TPS) have gained a lot of recognitions in the diagnosis or prediction of BCa. Unfortunately, these serological tests also yielded low sensitivity and specificity, especially with respect to their applications in diagnosing BCa in the early phase [5, 6]. Therefore, there is a need to develop new and minimally invasive diagnostic approaches to supplement breast imaging and conventional blood-based tests, so as to better improve the early detection rates for BCa.
Recently, the discovery of microRNAs (miRNAs) has opened new opportunities in cancer biology. The miRNAs are a class of short, single-stranded, approximately 22-nucleotide noncoding RNAs that have proven to be essential and indispensable in the regulation of gene expression as well as other biological processes [7, 8]. Since their discovery, abnormal miRNA profiles have been reported to associate with the diagnosis, prognosis, and even survival in a variety of neoplastic diseases involving BCa [9–15]. Importantly, it has been demonstrated that some miRNAs are presented in body fluids such as plasma, serum, urine, and saliva in a stable and reproducible fashion, as they are protected from degradation by association with secreted membrane vesicles and/or RNA-binding proteins [16–18]. At present, the circulating miRNA signatures in BCa have been described by numerous studies, hinting a tremendous prospect to be developed as useful biomarkers [19–33]. Unfortunately, many of the published studies yielded conflicting results, which may be explained in part by the differences in study design, sample size, race, sample type, and miRNA profiling. Therefore, the present meta-analysis was conduced to estimate the pooled accuracy of circulating miRNA detection in diagnosing BCa.
Methods
Search strategy and inclusion criteria
For this meta-analysis, we followed the guidelines of the Preferred Reporting Items for Systematic Reviews and Meta-analysis (PRISMA) statement and methods [34]. We carefully searched online PubMed and EMBASE databases for all published articles from 31 December 1993 to 31 March 2014. The search terms were utilized as “breast cancer/carcinoma,” “serum/plasma microRNA/miRNA,” “circulating microRNA/miRNA,” and “diagnosis/sensitivity/specificity”. We manually searched the reference lists of eligible studies identified from the databases as well.
All eligible studies satisfied the following criteria were firstly included in our analysis: (1) circulating miRNAs were assessed in breast cancer diagnostic analysis and (2) studies mentioned the sample number, sensitivity, specificity, area under curve (AUC), and their 95 % confidence intervals (CIs) or other more detailed information. Studies were excluded based on the following criteria: (1) studies evaluating miRNAs in tissues instead of serum or plasma; (2) studies failed to explicitly state the control groups, or the controls enrolled were not healthy cases; and (3) non-English articles, review articles, letters, commentaries, abstracts presented in conferences, and studies without complete data.
Data extraction and quality assessment
Two reviewers independently assessed the final set of selected articles. The extracted data elements of this study for diagnosis included the author, year of publication, country of origin, number of patients, control sources, sample types, circulating miRNA profiles, test method, diagnostic parameters, and other substantial information. In studies containing both a training and a validation group, data of each group was regarded as a single study in the meta-analysis. Any disagreement was resolved by consensus.
According to an evidence-based and critical review checklist of quality assessment for studies of diagnostic accuracy (QUADAS) tool [35], a quality index was generated with a maximum score value of 14. All the 14-item checklists were evaluated in the included studies, each of which was defined as “Yes (low risk/high concern),” “No (high risk/low concern),” or “Unclear (unclear risk/unclear concern)”. An answer of “Yes” got a score of 1, while a “No” or “Unclear” answer gained a score of 0. The quality scores for each study were finally calculated on the basis of the evaluation standard, and any scoring discrepancies were resolved through discussion.
Statistical analysis
Statistical analysis was undertaken utilizing Meta-disc 1.4 (XI Cochrane Colloquium, Barcelona, Spain) and Stata 12.0 (Stata Corporation, College Station, TX, USA) software. The bivariate meta-analysis model was employed to summarize the sensitivity, specificity, diagnostic odds ratio (DOR), positive likelihood ratio (PLR), and negative likelihood ratio (NLR) and generate the bivariate summary receiver operator characteristic (SROC) curve with their corresponding 95 % CIs. For the heterogeneity analysis, Spearman correlation coefficient was performed to analyze the threshold effect, while Cochran’s Q test and inconsistency index (I 2) test were used to assess heterogeneity from a non-threshold effect. P < 0.05 for Spearman correlation coefficient, or P < 0.01 for Cochran’s Q test, or I 2 > 50 %, all indicated an existence of significant heterogeneity [36]. Moreover, influence analysis and meta-regression were employed to trace potential sources of study heterogeneity. Deeks’ funnel plot asymmetry test was applied to estimate the potential publication bias among studies, and P < 0.01 was considered to be representative of a significant statistical publication bias.
Results
Search results
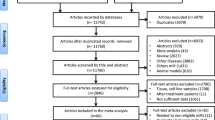
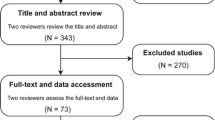
As shown in Fig. 1, a total of 195 trials were identified by our electronic database search (n = 186) and through other sources (n = 9). After the duplicates were removed, 110 unique abstracts remained. The titles, abstracts, and key words were then carefully evaluated, and 61 studies were excluded either because they were unrelated to miRNAs or BCa or the status of review articles, letters, or non-English papers. The retrieved 49 studies were conducted for more detailed evaluation, and 31 of them were further excluded due to the investigation of tissue miRNAs instead of circulating miRNAs in BCa or unrelated to diagnosis. In the 18 studies left, 2 of them contained benign BCa cases and 1 had unsufficient data for further analysis; they were all discarded. Therefore, only 15 publications [19–33] seemed to meet all of the inclusion criteria and none of the exclusion criteria.
Study characteristics and quality assessments
In this meta-analysis, the final set of 15 diagnostic studies included a total of 1368 patients with BCa and 849 healthy control individuals. The sample size of BCa patients in each study varied from 20 to 185, with mean/median age varying from 50 to 62 years. All the BCa patients were diagnosed based on the histopathological examination, with a proportion in early stages (ductal carcinoma in situ (DCIS), I, and II) of 74.59 %. Among the 15 studies, 10 studies were conducted in Asia, 3 in Europe, and 2 in America. The published date was from 2010 to 2013. Besides, all the 15 studies used the method of reverse transcription quantitative PCR (RT-qPCR) for the miRNAs detection, and the specimen type includes serum ([19, 22, 23, 26–30, 32, 33]) and plasma ([20, 21, 24, 25, 31]). The main features of the included studies are described in Table 1.
We estimated the quality of the 15 included publications according to the 14-item QUADAS assessment tool [35]. All of the 15 studies had QUADAS scores more than 10 and revealed lower risks of bias, suggesting a high quality of the included investigations (Table 1 and Fig. 2).
Heterogeneity
In this meta-analysis, Spearman correlation coefficient was performed to analyze the heterogeneity from threshold effect, while Cochran’s Q and I 2 tests were conducted to assess the heterogeneity caused by non-threshold effect. As displayed in Table 2, the P values of Spearman correlation coefficient in the overall pooled analysis and subgroup analyses like single miRNA profiling, serum, and Asian population-based assays were all less than 0.05, indicating the existence of heterogeneity from threshold effect among these analyses. Moreover, heterogeneity generated by non-threshold effect appeared in all the pooled analyses, which the P values of Cochran’s Q test were all less than 0.1, accompanied by I 2 > 50 %.
Pooled diagnostic performance
As we found significant heterogeneity between studies, the random effects model was applied in our meta-analysis. Forest plots of the sensitivity and specificity for circulating miRNAs in diagnosing BCa are shown in Fig. 3. The overall pooled results for sensitivity, specificity, PLR, NLR, and DOR were 0.82 (95 % CI 0.80–0.83), 0.82 (95 % CI 0.80–0.85), 5.53 (95 % CI 3.74–8.20), 0.18 (95 % CI 0.13–0.25), and 34.82 (95 % CI 17.91–67.71), respectively. The SROC curve for the pooled studies is displayed in Fig. 4a, which the AUC value was 0.9217, accompanied by a Q value of 0.8553, suggesting an overall high accuracy for the diagnostic test.
The SROC curve of the subgroup analyses for the diagnosis of BCa. Sample size is indicated by the size of the square. The regression SROC curve indicates the overall diagnostic accuracy. a overall pooled study. b Single miRNA-based analysis. c Serum-based miRNA assay. d Ethnicity-based analysis of Asian population. AUC area under curve, Q* index, SE standard error, SROC curve summary receiver operator curve
Subgroup analyses and meta-regression
The results of our subgroup analyses are summarized in Table 3 and Fig. 4b–d. We first performed a subgroup analysis based on miRNA profiles. The result demonstrated that use of multiple miRNAs showed promising accuracy, with a sensitivity of 0.87, specificity of 0.88, DOR of 85.93, PLR of 9.98, NLR of 0.12, and AUC of 0.9518; by contrast, diagnosing based on single miRNA had an lower accuracy when compared with multiple miRNAs; the data reflected a higher potential diagnostic value of multiple miRNAs for BCa detection. Furthermore, a comparison of miRNA expression patterns in serum and plasma showed that the DOR (36.24 versus 32.08), NLR (0.17 versus 0.21), and AUC (0.9225 versus 0.8890) were higher in serum-based assay than in plasma, providing additional evidences for the use of serum miRNAs as relatively reliable diagnostic biomarkers. We also conducted an analysis based on ethnicity. Our data exhibited that Caucasian population group yielded a combined sensitivity of 0.84 and specificity of 0.81 under the SROC curve, accompanied by DOR of 49.14, PLR of 6.68, and NLR of 0.15; for the Asian population group, the pooled sensitivity, specificity, DOR, PLR, and NLR were 0.80, 0.83, 28.27, 5.21, and 0.21, respectively. Thus, a difference in diagnostic accuracy was displayed between these two ethnicities, with a better diagnosis accuracy found in Caucasian populations.
Furthermore, we added a total of eight pre-specified covariates (design type, study quality, ethnicity, sample type, miRNA profiling, reference method, number of cases, and number of controls) to the bivariate model to assess their impacts on sensitivity and specificity through meta-regression. In consideration of the study size of our analysis, we employed a permute meta-regression module with a check value of 10,000, and each time only two covariates were estimated by Stata 12.0 software. Our data exhibited that miRNA profiling (P = 0.022) and reference method (P = 0.039) might introduce significant heterogeneity in both sensitivity and specificity, while design type (P = 0.618), ethnicity (P = 0.564), sample type (P = 0.528), study quality (P = 0.462), number of cases (P = 0.071), and number of controls (P = 0.244) showed low likelihood of sources of inter-study heterogeneity (not shown).
Influence analysis and publication bias
Influence analysis was conducted via Stata 12.0 software, as shown in Fig. 5; the influence analysis and outlier detection identified one outlier study [23]. After excluding the outlier, the overall pooled sensitivity increased from 0.82 to 0.83, sensitivity increased from 0.82 to 0.83, DOR increased from 34.82 to 38.13, PLR increased from 5.53 to 5.77, NLR decreased from 0.18 to 0.17, and AUC increased from 0.9217 to 0.9261. Moreover, the P value of spearman correlation coefficient from threshold effect elevated from 0.002 to 0.005, and I 2 for sensitivity declined from 86.9 to 86.4 %, suggesting that the outlier is likely a source of heterogeneity; however, the P value of Cochran’s Q test as well as I 2 for the overall study altered inconspicuously, hinting substantial heterogeneity from non-threshold effect existing among studies.
Publication bias of the included studies was checked by Deeks’ funnel plot asymmetry test. The slope coefficient was associated with a P value of 0.119 and 0.142, respectively, for the overall and outlier excluded analysis, suggesting an existing low likelihood of publication bias (Fig. 6). For the subgroup analysis, the slope coefficient presented P values of 0.333, 0.163, and 0.730, respectively, for miRNA profile, matrix, and ethnicity-based assays, also showing that no obvious publication bias existed (not shown).
Funnel plot test for the assessment of potential bias for the overall study and outlier assays. a Deeks’ funnel plot asymmetry test for total circulating miRNAs assays, P = 0.119. b Deeks’ funnel plot asymmetry test for outlier excluded assays, P = 0.142. A P value less than 0.1 was considered to be representative of a significant statistical publication bias
Discussion
BCa remains one of the top threats to the health of women worldwide and early-time identification benefits the outcomes of treatment as well as the survival rate. However, early BCa detection remains a major challenge for clinicians. Currently, mammography is the major diagnostic tool in screening for early-stage BCa with a high specificity more than 0.90. Nevertheless, the sensitivity of mammography is low and subject to many restrictions, including the breast density and professional experience of the examiner [3, 4]. Alternative method such as ultrasound screening has a very operator-dependent sensitivity as well. On the other hand, blood-based biomarkers involve CEA, CA15-3, and TSP that are also nonspecific and have limited sensitivity and specificity [5, 6]. Recently, abnormal circulating miRNA signatures in BCa have been depicted by a number of investigations, indicating a potential diagnostic value for BCa detection [19–33, 37, 38].
Our data showed promising accuracy for circulating miRNA detection in diagnosing BCa, which the overall pooled sensitivity was 0.82, and specificity was 0.82, with an AUC of 0.9217, suggesting that circulating miRNAs achieved a relatively high overall accuracy for BCa detection. Diagnostic odds ratio (DOR), defined as the ratio of the odds of a true positive to the odds of a false-positive, is an important indicator of diagnostic accuracy [39]; a DOR value less than 1.0 indicates a low discriminating ability in the diagnostic test [40]. In our data, the pooled DOR was presented as 34.82, indicating a better discriminatory performance of circulating miRNAs for BCa. In addition, we determined the combined PLR and NLR to obtain a more comprehensive view of their diagnostic accuracy: the PLR of 5.53, which suggested that BCa patients had a five fold higher chance of being miRNAs test positive than other individuals without BCa. Meanwhile, the pooled NLR was found to be 0.18, implying that in a negative result from the miRNA test, only 18 % is likely to be false-negative.
As reported, the accuracies of conventional CA15-3 and CEA in the diagnosis of BCa revealed high volatilities, which the discrepancy may due to ethnic group, sample size, or cancer subtype. A study from US researchers reported that the sensitivities of CA15-3 and CEA for BCa ranged from 0.54 to 0.90 and 0.29 to 0.53, while the specificities ranged from 0.86 to 0.99 and 0.70 to 0.99, respectively [41]. Yet, two studies conducted in Asia showed that the sensitivities of CA15-3 were 0.11 versus 0.16, and CEA were 0.09 versus 0.22 for BCa detection, showing a high volatility among studies [22, 32]. Besides, Zeng et al. presented the similar results that the overall sensitivities of CA15-3 and CEA for the diagnosis of BCa were 0.14 and 0.12 (range 0.07–0.33; range 0.0–0.30) in every TNM stage, with a highest sensitivity of 0.33 presented in DCIS and 0.30 in TNM III, implying that CA15-3 or CEA detection may be better for advanced BCa [24]. By comparison, circulating miRNAs seemed to harbor overall high diagnostic accuracy in diagnosing BCa, which the pooled sensitivity ranged from 0.80 to 0.83 and specificity from 0.80 to 0.84 in our study. However, for the diagnosis of early-stage cases, our BCa patients in early stages (DCIS, I, and II) only had a proportion of 74.59 %; hence, it is solid to say that circulating miRNAs is better than conventional biomarkers in screening BCa in the early stage.
Furthermore, we conducted subgroup analyses based on the following variables like miRNA profiling, sample type, and ethnicity. Interestingly, when according to miRNAs profiling, the result suggested that combinations of multiple miRNAs are more promising as biomarkers for BCa detection than single miRNA. Supporting evidence from Mar-Aguilar et al. demonstrated that combining miR-145, -155, and -382 could be a more powerful diagnostic tool in distinguishing BCa patients from healthy controls, with a sensitivity of 0.97 and specificity of 1.0 higher than individuals. Meanwhile, another proof also showed that the efficiency for BCa detection was enhanced when the circulating miRNAs (miR-148b, -409-3p, and -801) were combined into diagnostic panels.
However, other researches that validated the diagnostic value of single miRNA for BCa also presented robust results. To date, circulating miRNA-21 and -155 have been the most heavily reported miRNAs for BCa research. Lv et al. conducted a meta-analysis to distinguish the candidates for BCa detection from spurious circulating miRNAs and suggested that miR-21 and -155 are reliable candidate biomarkers for detection of BCa [42]. Gao et al. have proved that serum miR-21 has a higher sensitivity in the diagnosis of BCa, when compared with CEA and CA15-3 [32]. Another investigation also held the point that circulating miR-21 could serve as a potential broad-spectrum serum-based biomarker for the detection of some solid cancers involving BCa [33]. On the other hand, a newly published meta-analysis containing three studies for miRNA-155 showed that circulating miR-155 has the potential diagnostic value with a high pooled sensitivity (0.79) and specificity (0.85) for BCa [43]. However, more studies are required to confirm the actual diagnostic value of circulating miRNA-21 or -155 for BCa, and other biomarkers are also expected.
In addition, we traced the pooled diagnostic accuracy based on different sample types. As the data indicated, serum-based assay seemed to undergo a higher combined DOR, NLR, and AUC than plasma-based test. Our results provided evidence that serum may be a better matrix for diagnostic profiling of miRNAs in BCa. Shen et al. similarly observed a diagnostic difference between serum and plasma miRNA assays in lung cancer [39]. Research from Wang et al. demonstrated that the coagulation process may affect the spectrum of extracellular miRNAs in the blood, implying that different matrices may harvest different diagnostic accuracies for miRNA detection [44]. Notwithstanding, the plasma specimen group only included five studies, and large-scale prospective studies are needed to provide more accurate results.
It has been reported that different racial expression profiles are associated with circulating miRNA concentrations, hinting that circulating miRNA signature varies among ethnicity [45, 46]. Our findings also reached similar results that Caucasian population-based miRNA test yielded an overall higher accuracy against that of Asian: the pooled DOR was 49.14 versus 28.27, and PLR was 6.68 versus 5.21, accompanied by a lower NLR of 0.15 in Caucasians, suggesting that circulating miRNA assay may be more precise in Caucasian populations. However, our findings do not agree with the results from Li et al. in which, miRNAs assay seemed to be more precise in Asian populations when used in the diagnosis of hematologic malignancies [47].
Heterogeneity is a latent problem when interpreting the results of any meta-analysis. In our study, heterogeneity from threshold effect existed in the overall and subgroup analyses. The threshold effect is mainly generated by the different cut-off value settings or thresholds used in different studies. In our study, the cut-off values for miRNAs were not uniformed among studies, which may further contribute to the heterogeneity from the threshold effect. On the other hand, the pooled DOR is often used to discuss the heterogeneity caused by non-threshold effects [40]. We found that the DOR of each study did not distribute along a straight line with the pooled DOR in the forest plots, and the P values in Cochran’s Q test were all less than 0.01, accompanied by an I 2 of more than 50 %, also indicating substantial heterogeneity from non-threshold effect in studies. For its causes, different ethnicities, measurement methods, or sample types may contribute to heterogeneity sources. In our study, although the detecting methods for circulating miRNAs were all based on RT-qPCR, the ethnicities, sample types, and reference methods were different among studies. Additionally, the participants enrolled in the tests were not unified for their BCa stages, disease conditions, or other concomitant diseases (unclear information). Therefore, we further conducted influence and meta-regression analyses to assess the contribution of the factors above and found that the outlier studies, miRNA profiles, as well as different reference methods used among studies were the sources of heterogeneity.
Taken together,our results identified the potential value of circulating miRNAs as biomarkers for BCa detection, in particular finding that miRNA panels could be more powerful tools than a single one. Therefore, we suggest that further investigations into circulating miRNAs might open avenues for prospective trials of a noninvasive diagnostic test for BCa. However, we found substantial heterogeneity among the pooled studies as well as in the subgroup analyses. In consequence, the combined diagnostic indices of circulating miRNAs in this study are unable to completely mirror their actual diagnostic value for BCa, and further, large cohort studies are still warranted.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11–30. doi:10.3322/caac.21166.
Cuk K, Zucknick M, Madhavan D, Schott S, Golatta M, Heil J, et al. Plasma microRNA panel for minimally invasive detection of breast cancer. PLoS One. 2013;8:e76729. doi:10.1371/journal.pone.0076729.
Checka CM, Chun JE, Schnabel FR, Lee J, Toth H. The relationship of mammographic density and age: implications for breast cancer screening. AJR Am J Roentgenol. 2012;198:W292-5. doi:10.2214/AJR.10.6049.
Guadagni F, Ferroni P, Carlini S, Mariotti S, Spila A, Aloe S, et al. A reevaluation of carcinoembryonic antigen (CEA) as a serum marker for breast cancer: a prospective longitudinal study. Clin Cancer Res. 2001;7:2357–62.
Duffy MJ, Evoy D, McDermott EW. CA 15-3: uses and limitation as a biomarker for breast cancer. Clin Chim Acta. 2010;411:1869–74.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Pritchard CC, Cheng HH, Tewari M. MicroRNA profiling: approaches and considerations. Nat Rev Genet. 2012;13:358–69. doi:10.1038/nrg3198.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834–8.
Hamamoto J, Soejima K, Yoda S, Naoki K, Nakayama S, Satomi R, et al. Identification of microRNAs differentially expressed between lung squamous cell carcinoma and lung adenocarcinoma. Mol Med Rep. 2013;8:456–62. doi:10.3892/mmr.2013.1517.
Yong FL, Law CW, Wang CW. Potentiality of a triple microRNA classifier: miR-193a-3p, miR-23a and miR-338-5p for early detection of colorectal cancer. BMC Cancer. 2013;13:280. doi:10.1186/1471-2407-13-280.
Li W, Xie L, He X, Li J, Tu K, Wei L, et al. Diagnostic and prognostic implications of microRNAs in human hepatocellular carcinoma. Int J Cancer. 2008;123:1616–22. doi:10.1002/ijc.23693.
Bovell LC, Putcha BD, Samuel T, Manne U. Clinical implications of microRNAs in cancer. Biotech Histochem. 2013;88:388–96. doi:10.3109/10520295.2013.788735.
Chen WX, Hu Q, Qiu MT, Zhong SL, Xu JJ, Tang JH, et al. miR-221/222: promising biomarkers for breast cancer. Tumor Biol. 2013;34:1361–70. doi:10.1007/s13277-013-0750-y.
Chen L, Li Y, Fu Y, Peng J, Mo MH, Stamatakos M, et al. Role of deregulated microRNAs in breast cancer progression using FFPE tissue. PLoS One. 2013;8:e54213. doi:10.1371/journal.pone.0054213.
Chen X, Ba Y, Ma L, Cai X, Yin Y, Wang K, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997–1006. doi:10.1038/cr.2008.282.
Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58:1375–81. doi:10.1136/gut.2008.167817.
Madhavan D, Cuk K, Burwinkel B, Yang R. Cancer diagnosis and prognosis decoded by blood-based circulating microRNA signatures. Front Genet. 2013;4:116. doi:10.3389/fgene.2013.00116.
Kubiczkova L, Kryukov F, Slaby O, Dementyeva E, Jarkovsky J, Nekvindova J, et al. Circulating serum microRNAs as novel diagnostic and prognostic biomarkers for multiple myeloma and monoclonal gammopathy of undetermined significance. Haematologica. 2014;99:511–8. doi:10.3324/haematol.2013.093500.
Wu Q, Wang C, Lu Z, Guo L, Ge Q. Analysis of serum genome-wide microRNAs for breast cancer detection. Clin Chim Acta. 2012;413:1058–65. doi:10.1016/j.cca.2012.02.016.
Schrauder MG, Strick R, Schulz-Wendtland R, Strissel PL, Kahmann L, Loehberg CR, et al. Circulating micro-RNAs as potential blood-based markers for early stage breast cancer detection. PLoS One. 2012;7:e29770. doi:10.1371/journal.pone.0029770.
Cuk K, Zucknick M, Heil J, Madhavan D, Schott S, Turchinovich A, et al. Circulating microRNAs in plasma as early detection markers for breast cancer. Int J Cancer. 2013;132:1602–12. doi:10.1002/ijc.27799.
Guo LJ, Zhang QY. Decreased serum miR-181a is a potential new tool for breast cancer screening. Int J Mol Med. 2012;30:680–6. doi:10.3892/ijmm.2012.1021.
Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011;57:84–91. doi:10.1373/clinchem.2010.151845.
Zeng RC, Zhang W, Yan XQ, Ye ZQ, Chen ED, Huang DP, et al. Down-regulation of miRNA-30a in human plasma is a novel marker for breast cancer. Med Oncol. 2013;30:477. doi:10.1007/s12032-013-0477-z.
Heneghan HM, Miller N, Kelly R, Newell J, Kerin MJ. Systemic miRNA-195 differentiates breast cancer from other malignancies and is a potential biomarker for detecting noninvasive and early stage disease. Oncologist. 2010;15:673–82. doi:10.1634/theoncologist.2010-0103.
Mar-Aguilar F, Mendoza-Ramírez JA, Malagón-Santiago I, Espino-Silva PK, Santuario-Facio SK, Ruiz-Flores P, et al. Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis Markers. 2013;34:163–9. doi:10.3233/DMA-120957.
Zhao SY, Wu Q, Gao F, Zhang CB, Yang XW. Serum microRNA-155 as a potential biomarker for breast cancer screening. Chin Sci Bull. 2012;57:3466–8.
Sun Y, Wang M, Lin G, Sun S, Li X, Qi J, et al. Serum microRNA-155 as a potential biomarker to track disease in breast cancer. PLoS One. 2012;7:e47003. doi:10.1371/journal.pone.0047003.
Hu Z, Dong J, Wang LE, Ma H, Liu J, Zhao Y, et al. Serum microRNA profiling and breast cancer risk: the use of miR-484/191 as endogenous controls. Carcinogenesis. 2012;33:828–34. doi:10.1093/carcin/bgs030.
Zhao FL, Hu GD, Wang XF, Zhang XH, Zhang YK, Yu ZS. Serum overexpression of microRNA-10b in patients with bone metastatic primary breast cancer. J Int Med Res. 2012;40:859–66.
Ng EK, Li R, Shin VY, Jin HC, Leung CP, Ma ES, et al. Circulating microRNAs as specific biomarkers for breast cancer detection. PLoS One. 2013;8:e53141. doi:10.1371/journal.pone.0053141.
Gao J, Zhang Q, Xu J, Guo L, Li X. Clinical significance of serum miR-21 in breast cancer compared with CA153 and CEA. Chin J Cancer Res. 2013;25:743–8. doi:10.3978/j.issn.1000-9604.2013.12.04.
Wang B, Zhang Q. The expression and clinical significance of circulating microRNA-21 in serum of five solid tumors. J Cancer Res Clin Oncol. 2012;138:1659–66.
Moher D, Liberati A, Tetzlaff J, Altman DG, PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. J Clin Epidemiol. 2009;62:1006–12. doi:10.1016/j.jclinepi.2009.06.005.
Whiting P, Rutjes AW, Reitsma JB, Bossuyt PM, Kleijnen J. The development of QUADAS: a tool for the quality assessment of studies of diagnostic accuracy included in systematic reviews. BMC Med Res Methodol. 2003;3:25.
Zamora J, Abraira V, Muriel A, Khan K, Coomarasamy A. Meta-DiSc: a software for meta-analysis of test accuracy data. BMC Med Res Methodol. 2006;6:31.
Liu J, Mao Q, Liu Y, Hao X, Zhang S, Zhang J. Analysis of miR-205 and miR-155 expression in the blood of breast cancer patients. Chin J Cancer Res. 2013;25:46–54. doi:10.3978/j.issn.1000-9604.2012.11.04.
Roth C, Rack B, Müller V, Janni W, Pantel K, Schwarzenbach H. Circulating microRNAs as blood-based markers for patients with primary and metastatic breast cancer. Breast Cancer Res. 2010;12:R90. doi:10.1186/bcr2766.
Shen Y, Wang T, Yang T, Hu Q, Wan C, Chen L, et al. Diagnostic value of circulating microRNAs for lung cancer: a meta-analysis. Genet Test Mol Biomarkers. 2013;17:359–66. doi:10.1089/gtmb.2012.0370.
Glas AS, Lijmer JG, Prins MH, Bonsel GJ, Bossuyt PM. The diagnostic odds ratio: a single indicator of test performance. J Clin Epidemiol. 2003;56:1129–35.
Harris L, Fritsche H, Mennel R, Norton L, Ravdin P, Taube S, et al. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol. 2007;25:5287–312.
Lv M, Zhu X, Chen W, Zhao J, Tang J. Searching for candidate microRNA biomarkers in detection of breast cancer: a meta-analysis. Cancer Biomark. 2013;13:395–401. doi:10.3233/CBM-130379.
Wang F, Hou J, Jin W, Li J, Yue Y, Jin H, et al. Increased circulating microRNA-155 as a potential biomarker for breast cancer screening: a meta-analysis. Molecules. 2014;19:6282–93. doi:10.3390/molecules19056282.
Wang K, Yuan Y, Cho JH, McClarty S, Baxter D, Galas DJ. Comparing the MicroRNA spectrum between serum and plasma. PLoS One. 2012;7:e41561. doi:10.1371/journal.pone.0041561.
Zhao H, Shen J, Medico L, Wang D, Ambrosone CB, Liu S. A pilot study of circulating miRNAs as potential biomarkers of early stage breast cancer. PLoS One. 2010;5:e13735. doi:10.1371/journal.pone.0013735.
Hu Z, Chen X, Zhao Y, Tian T, Jin G, Shu Y, et al. Serum microRNA signatures identified in a genome-wide serum microRNA expression profiling predict survival of non-small-cell lung cancer. J Clin Oncol. 2010;28:1721–6. doi:10.1200/JCO.2009.24.9342.
Li Q, Liu L, Li W. Identification of circulating microRNAs as biomarkers in diagnosis of hematologic cancers: a meta-analysis. Tumour Biol. 2014. doi:10.1007/s13277-014-2364-4.
Funding
This work was supported by the National Natural Science Foundation of China (No. 81371879).
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cui, Z., Lin, D., Song, W. et al. Diagnostic value of circulating microRNAs as biomarkers for breast cancer: a meta-analysis study. Tumor Biol. 36, 829–839 (2015). https://doi.org/10.1007/s13277-014-2700-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2700-8