Abstract
Lung cancer is the leading cause of death worldwide. Non-small-cell lung cancer (NSCLC) accounts for most of these cases. T-cell immunoglobulin- and mucin-domain-containing molecule 3 (TIM-3) has been established as a negative regulatory molecule and plays a critical role in immune tolerance. Studies have shown that polymorphisms in TIM-3 gene can be associated with various diseases. The aim of this study was to investigate whether polymorphisms in the TIM-3 gene were associated with susceptibility to NSCLC. Three polymorphisms in TIM-3 gene (−1516G/T, −574G/T, and +4259T/G) were identified by polymerase chain reaction–restriction fragment length polymorphism in 432 NSCLC patients and 466 healthy controls. Results showed that frequencies of TIM-3 +4259TG genotype for cases and controls were 10.9 and 4.1 %, respectively; subjects carrying the +4259TG genotype had a 2.81-fold increased risk of NSCLC compared to the wild-type genotype (P < 0.0001). The TIM-3 −1516G/T and −574G/T polymorphisms did not show any correlation with NSCLC. In addition, when analyzing the survival time of NSCLC patients with TIM-3 +4259T/G polymorphism, cases with +4259TG genotype had significantly shorter survival time compared to the wild-type patients (15.2 months vs. 26.7 months, P = 0.007). These results suggested polymorphism in TIM-3 gene is associated with increased susceptibility to NSCLC and could be used as prognostic factor for this malignancy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer is the leading cause of death worldwide; more than a million people die from this disease each year [1]. Non-small-cell lung cancer (NSCLC) accounts for up to 80 % of all lung cancer cases. Despite the development of antitumor therapy, the prognosis for patients with lung cancer remains poor, with a 5-year survival rate of less than 20 % [2]. Recent studies have provided evidence that genetic factors may influence the development and the prognosis of NSCLC.
T-cell immunoglobulin- and mucin-domain-containing molecule 3 (TIM-3) is expressed on Th1, Th17 cells, and CD8 T cells, but not Th2 cells [3–5]. Interaction between TIM-3 and its ligand galectin-9 inhibits Th1 and Th17 responses [6] and induces peripheral tolerance [7, 8], suggesting an inhibitory role of TIM-3 in T cell responses. TIM-3 expression has also been identified in exhausted T cells during chronic infection [8]. TIM-3-expressing CD4+ and CD8+ T cells produce reduced amounts of cytokines or are less proliferative in response to antigen [9]. Blockade of the TIM-3 signaling pathway restores proliferation and enhances cytokine production in HIV-1-specific T cells [9]. Recent studies have shown an important role of TIM-3T cell exhaustion in cancer. Tim-3 and PD-1, another marker of T cell exhaustion, are co-expressed on CD8 tumor infiltrating lymphocytes (TILs) in mice-bearing transplanted tumors as well as on NY-ESO-1-specific CD8+ T cells in patients with advanced melanoma [10, 11]. TIM-3+PD-1+ T cells exhibit the most severe exhausted phenotype as defined by failure to proliferate and produce IL-2, TNF, and IFN-gamma. Blockade of both Tim-3 and PD-1 pathways is more effective in controlling tumor growth than targeting either pathway alone, suggesting these two pathways work synergistically in establishing T cell exhaustion [10, 11]. TIM-3 may play important roles in the development of NSCLC [12, 13]. It has been shown that TIM-3 is expressed on tumor cells and TILs in lung cancer tissues. The expression levels of TIM-3 may be correlated with patients’ survival [12]. In this study, we tested three single-nucleotide polymorphisms (SNPs; −1516G/T, −574G/T, and +4259T/G) of TIM-3 gene and analyzed whether the genetic variants would be involved in the susceptibility to NSCLC in the Chinese population.
Materials and methods
Patients and controls
The study group included 432 NSCLC cases recruited from the Affiliated Hospital of Academy of Military Medical Sciences and the East Hospital. The histological type of lung cancer was identified according to the World Health Organization classifications. The pathologic stage was determined according to the International System for Staging Lung Cancer [14]. Samples from the healthy control population were collected from individuals residing in the same geographic areas without histories of malignancy or other major diseases. The healthy control group included 302 males and 164 females. To exclude the possible effects of ethnicity, only Han Chinese were included in this study. Informed consent was obtained from all study participants according to the Helsinki Declaration. This study was approved by the institutional review boards of the Affiliated Hospital of Academy of Military Medical Sciences and the East Hospital.
DNA extraction and genotyping
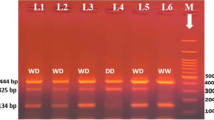
Genomic DNA was extracted from 5 ml frozen whole blood using the DNA Extraction Kit (Qiagen Inc., Hilden, Germany) according to the manufacturer’s protocol. The three polymorphisms within the TIM-3 gene promoter and encoding regions were identified by the polymerase chain reaction–restriction fragment length polymorphism (PCR–RFLP) assay. The primers and PCR conditions are listed in Table 1. PCR was performed in a total reaction volume of 20 ul containing 2 ul of 10X PCR buffer (Qiagen Inc., Hilden, Germany), 1.5 mM MgCl2, 0.5 uM of each primer (shown in Table 1), 0.2 mM dNTP, 1.2 U Taq polymerase (Qiagen Inc., Hilden, Germany), and 200 ng of genomic DNA. After an initial denaturation at 95 °C for 5 min, the DNA was amplified for 35 cycles at 94 °C for 30 s, 55–60 °C for 40 s (detailed annealing temperatures shown in Table 1), and 72 °C for 45 s, with a final elongation at 72 °C for 10 min on the Gene-Amp PCR System 9700 (PE Applied Biosystems, Foster City, CA, USA). PCR products containing the three polymorphic sites were then digested with the restriction enzymes Bsl I, Taq I, and Pst I (New England Biolabs, Beverly, MA, USA), respectively, by using the conditions recommended in the manufacturer’s instructions. The digested PCR products were fractionated on 2 % agarose Tris–borate–EDTA gel (Agarose 1000; Gibco BRL, Rockville, MD, USA) and stained with ethidium bromide (product size after digestion shown in Table 1). To confirm the genotyping results, more than 15 % of PCR-amplified DNA samples were examined by DNA sequencing. Results between PCR and DNA sequencing analysis were 100 % concordant.
Statistical analysis
The SPSS statistical software package ver.13.0 (SPSS Inc., Chicago, USA) and the Prism 5.0 were used for statistical analysis. Demographic data between the study groups were compared by the chi-square test and by the Student t test. Hardy–Weinberg equilibrium was analyzed using the chi-square test. For SNP analysis, genotype and allele frequencies of TIM-3 were compared between groups using the chi-square test, and odds ratios (OR) and 95 % confidence intervals (CIs) were calculated using unconditional logistic regression. P values less than 0.05 were considered significant. A survival curve was drawn with the Kaplan–Meier method for each of the different genotypes. Comparisons were made with the logrank test. Hazard ratios of death with 95 % CI were estimated using the multivariate Cox model.
Results
A total of 432 NSCLC cases and 466 controls were recruited for the present study. All subjects were ethnic Chinese. Demographic and other selected characteristics of the cases and controls are presented (Table 2). Cases and controls did not show statistically significant differences with regard to age (P > 0.05) and sex (P > 0.05), while those established risk factors such as smoking status and pack-year value showed significant differences (P < 0.001 and P < 0.001; Table 2). Most of the patients received chemotherapy (78.9 %), whereas others received radiotherapy (9.3 %), chemoradiotherapy (7.2 %), or other therapies.
Genotype and allele frequencies of the TIM-3 −1516G/T, −574G/T, and +4259T/G polymorphisms in NSCLC cases and controls are summarized in Table 3. The genotype distributions of these SNPs among the controls were in agreement with the Hardy–Weinberg equilibrium (P > 0.05). The homozygous variants of the three TIM-3 polymorphisms, −1516TT, −574TT, and +4259GG, were not detected in our study population. The −1516G/T and −574G/T SNPs did not show any association between NSCLC cases and controls (Table 3). As for the TIM-3 +4259T/G SNP, prevalence of TT genotype and TG genotype were, respectively, 89.1 and 10.9 % in patients and 95.9 and 4.1 % in controls. The +4259TG genotype and A allele revealed significantly increased frequencies in patients than in controls (OR = 2.81; 95 % CI, 1.59–4.91, P < 0.0001 and OR = 2.67; 95 % CI, 1.60–4.82, P = 0.0001, respectively). Also, we generated the haplotypes of the SNPs. The four most common haplotypes are shown in Table 3. There were no linkage disequilibrium observed and the frequency of GGG haplotype was higher in cases (P = 0.001). These data suggested that TIM-3 +4259T/G polymorphism is associated with increased susceptibility to NSCLC in the Chinese population.
We further analyzed the overall survival rates of NSCLC patients using Kaplan–Meier method and Log Rank Test (Fig. 1). Data from 381 cases were included for survival analysis whereas the remaining 61 cases were excluded due to loss of follow-up. The follow-up time ranged from 0 to 56 months. Data showed that the mean survival rates were statistically different based on the TIM-3 +4259T/G genotypes. Patients carrying the TG genotype presented a significantly lower survival rate than those with wild-type TT genotype (15.2 months vs. 26.7 months, P = 0.007). Thus, individuals with TG genotype indicated a worse prognosis of NSCLC. Results of Cox multivariate regression survival analysis are shown in Table 4. We found a decreased overall survival time for +4259TG genotype patients compared with wild-type, with age, sex, smoking status, tumor stage, and histological type as covariates (hazard ratio (HR), 1.43; 95 % CI, 1.15–1.81; P = 0.015). From analysis of other potential factors, data revealed that smoking status and tumor stage also contributed to survival rate (P = 0.018 and P = 0.012, respectively). These results indicated that TIM-3 +4259TG genotype is an adverse indicator for NSCLC prognosis.
Discussion
TIM-3 is a molecule expressed on terminally differentiated Th1 cells but not on Th2 cells, which negatively regulate Th1 immunity [15], and it is also a phosphatidylserine receptor to mediate phagocytosis of apoptotic cells [16]. The polymorphism studies have suggested that SNPs of TIM-3 are associated with rheumatoid arthritis [17, 18], gastric cancer [19], pancreatic cancer, atopic disease [18] and diabetes [20], etc. Our study indicated that +4259T/G SNP was associated with increased risk of NSCLC. This result was coincident with recently published papers about TIM-3 polymorphism with gastric cancer and pancreatic cancer, in which the +4259T/G SNP was associated with metastasis or vascular infiltration of these cancers [19]. These data indicated that +4259T/G SNP of TIM-3 might play important roles in affecting the metastasis of cancers.
The effect of TIM-3 on cancer is poorly understood. It is possible that TIM-3 plays a significant role in tumor progression by maintaining the tumor immunosuppressive environment via regulatory T cells (Tregs). Study has suggested that TIM-3+ Tregs in lung cancer tissues could be derived from natural Tregs upon chronic TCR stimulation by tumor antigens [13]. PD-1 ligand B7-H1 and TIM-3 ligands such as galectin-9 and apoptotic cells within the tumor tissue might be important for maintaining the number and function of TIM-3+PD-1+ Tregs. In the transplantation setting, Tim-3 has been shown to regulate allo-specific Treg activation [8]. It has been reported that Tim-3-Tim-3L-sensitive pathway is involved in the functional generation of donor-specific Tregs upon administration of tolerating treatments [8]. Recent study has revealed high levels of TIM-3 expression on Tregs within human lung tumor, which suggests that TIM-3 might play a direct role in the functional maturation of tumor infiltrating Tregs in the development of NSCLC. In addition, the higher TIM-3 level might result in an elevated CD80 expression of cell, and then the CD80 would preferentially interact with the inhibitory molecule CTLA-4 (cytotoxic T lymphocyte-associated antigen-4). This would eventually lead to a local immunosuppression [21, 22]. A recent research has demonstrated that NSCLC patients with TIM-3-positive tumor cells had a significantly shorter survival time than those with TIM-3-negative tumors [23]. Similarly, our data observed that patients carrying the TIM-3 +4259TG genotype presented a significantly lower survival rate than those with wild-type TT genotype (15.2 months vs. 26.7 months, P = 0.007; Fig. 1). All the results indicated that TIM-3 may play important roles in the prognosis of this cancer.
TIM-3 has emerged as a promising target for cancer immunotherapy [20]. Recent studies have focused on the role of TIM-3 expression on CD8+ T cells in peripheral blood as well as within tumors [10, 11]. This case–control study demonstrates for the first time that the TIM-3 SNP is associated with increased risk of NSCLC and could be a prognostic factor of the malignancy in the Chinese population. Our results provide important insights for understanding the genetics of NSCLC and would be helpful for the development of TIM-3 as a possible therapeutic approach to this disease.
References
Ishikawa M, Kitayama J, Yamauchi T, Kadowaki T, Maki T, Miyato H. Adiponectin inhibits the growth and peritoneal metastasis of gastric cancer through its specific membrane receptors AdipoR1 and AdipoR2. Cancer Sci. 2007;98:1120–7.
Lawrence RE, Salgia R. MET molecular mechanisms and therapies in lung cancer. Cell Adh Migr. 2010;4:146–52.
Anderson AC, Anderson DE, Bregoli L, Hastings WD, Kassam N, et al. Promotion of tissue inflammation by the immune receptor Tim-3 expressed on innate immune cells. Science. 2007;318:1141–3.
Hastings WD, Anderson DE, Kassam N, Koguchi K, Greenfield EA, et al. TIM-3 is expressed on activated human CD4+ T cells and regulates Th1 and Th17 cytokines. Eur J Immunol. 2009;39:2492–501.
Monney L, Sabatos CA, Gaglia JL, Ryu A, Waldner H, et al. Th1-specific cell surface protein Tim-3 regulates macrophage activation and severity of an autoimmune disease. Nature. 2002;415:536–41.
Zhu C, Anderson AC, Schubart A, Xiong H, Imitola J, et al. The Tim-3 ligand galectin-9 negatively regulates T helper type 1 immunity. Nat Immunol. 2005;6:1245–52.
Sabatos CA, Chakravarti S, Cha E, Schubart A, Sanchez-Fueyo A, et al. Interaction of Tim-3 and Tim-3 ligand regulates T helper type 1 responses and induction of peripheral tolerance. Nat Immunol. 2003;4:1102–10.
Sanchez-Fueyo A, Tian J, Picarella D, Domenig C, Zheng XX, et al. Tim-3 inhibits T helper type 1-mediated auto- and alloimmune responses and promotes immunological tolerance. Nat Immunol. 2003;4:1093–101.
Jones RB, Ndhlovu LC, Barbour JD, Sheth PM, Jha AR, et al. Tim-3 expression defines a novel population of dysfunctional T cells with highly elevated frequencies in progressive HIV-1 infection. J Exp Med. 2008;205:2763–79.
Fourcade J, Sun Z, Benallaoua M, Guillaume P, Luescher IF, et al. Upregulation of Tim-3 and PD-1 expression is associated with tumor antigen-specific CD8+ T cell dysfunction inmelanoma patients. J Exp Med. 2010;207:2175–86.
Sakuishi K, Apetoh L, Sullivan JM, Blazar BR, Kuchroo VK, et al. Targeting Tim-3 and PD-1 pathways to reverse T cell exhaustion and restore anti-tumor immunity. J Exp Med. 2010;207:2187–94.
Zhuang X, Zhang X, Xia X, Zhang C, Liang X, Gao L, et al. Ectopic expression of TIM-3 in lung cancers: a potential independent prognostic factor for patients with NSCLC. Am J Clin Pathol. 2012;137(6):978–85.
Gao X, Zhu Y, Li G, Huang H, Zhang G, Wang F, et al. TIM-3 expression characterizes regulatory T cells in tumor tissues and is associated with lung cancer progression. PLoS One. 2012. doi:10.1371/journal.pone.0030676.
Groome PA, Bolejack V, Crowley JJ, Kennedy C, Krasnik M, Sobin LH, et al. The IASLC Lung Cancer Staging Project: validation of the proposals for revision of the T, N, and M descriptors and consequent stage groupings in the forthcoming (seventh) edition of the TNM classification of malignant tumors. J Thorac Oncol. 2007;2:694–705.
Chae SC, Song JH, Pounsambath P, Yuan HY, Lee JH, Kim JJ, et al. Molecular variations in Th1-specific cell surface gene Tim-3. Exp Mol Med. 2004;36:274–8.
Frisancho-Kiss S, Nyland JF, Davis SE, Barrett MA, Gatewood SJ, Njoku DB, et al. Cutting edge: T cell Ig mucin-3 reduces inflammatory heart disease by increasing CTLA-4 during innate immunity. J Immunol. 2006;176:6411–5.
Chae SC, Park YR, Shim SC, Yoon KS, Chung HT. The polymorphisms of Th1 cell surface gene Tim-3 are associated in a Korean population with rheumatoid arthritis. Immunol Lett. 2004;95:91–5.
Chae SC, Park YR, Lee YC, Lee JH, Chung HT. The association of TIM-3 gene polymorphism with atopic disease in Korean population. Hum Immunol. 2004;65:1427–31.
Cao B, Zhu L, Zhu S, Li D, Zhang C, Xu C, et al. Genetic variations and haplotypes in TIM-3 gene and the risk of gastric cancer. Cancer Immunol Immunother. 2010;59:1851–7.
Boenisch O, D’Addio F, Watanabe T, Elyaman W, Magee CN, et al. TIM-3: a novel regulatory molecule of alloimmune activation. J Immunol. 2010;185:5806–19.
Wang L, Pino-Lagos K, de Vries VC, Guleria I, Sayegh MH, et al. Programmed death 1 ligand signaling regulates the generation of adaptive Foxp3+CD4+ regulatory T cells. Proc Natl Acad Sci U S A. 2008;105:9331–6.
Seki M, Oomizu S, Sakata KM, Sakata A, Arikawa T, et al. Galectin-9 suppresses the generation of Th17, promotes the induction of regulatory T cells, and regulates experimental autoimmune arthritis. Clin Immunol. 2008;127:78–88.
Zhuang X, Zhang X, Xia X, Zhang C, Liang X, et al. Ectopic expression of TIM-3 in lung cancers: a potential independent prognostic factor for patients with NSCLC. Am J Clin Pathol. 2012;137:978–85.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Jianwen Bai and Xiaoyan Li contributed equally to this work.
Rights and permissions
About this article
Cite this article
Bai, J., Li, X., Tong, D. et al. T-cell immunoglobulin- and mucin-domain-containing molecule 3 gene polymorphisms and prognosis of non-small-cell lung cancer. Tumor Biol. 34, 805–809 (2013). https://doi.org/10.1007/s13277-012-0610-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-012-0610-1