Abstract
Purpose
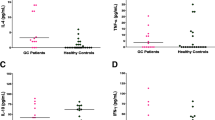
T cell immunoglobulin and mucin domain-containing molecule 3 (TIM-3) could weaken the Th1-mediated anti-tumor responses and accelerate the tumor cell proliferation by inhabiting the production of IL-2 or IFN-γ. This study was to assess the association between TIM-3 genetic variations and the development of gastric cancer.
Patients and methods
Five polymorphisms located in the promoter or encoding region of TIM-3 gene were genotyped in 212 gastric cancer patients and 252 controls who matched with the patients on the frequency of age, gender, smoking, and drinking. Logistic regression was used to determine whether the inherited variations within TIM-3 gene were associated with gastric cancer risk. Linkage disequilibrium and Haplotype analyses were performed by using SHEsis program.
Results
By the individual genotype analysis, three polymorphisms (−574G/T, −882C/T, and −1516G/T) within TIM-3 gene were significantly associated with gastric cancer in the study population [ORs (95% CIs): 2.74 (1.21–6.20), 3.19 (1.29–7.91), and 2.03 (1.15–3.59); respectively]. Among the gastric cancer patients, the relationship between the −1516 polymorphic genotype and the distant metastasis of tumor was found (OR = 2.21, 95% CI = 1.05–4.63). Under the analysis of haplotypes, an even stronger association with haplotype TTGCT was observed in gastric cancer risk (OR = 5.57, 95% CI: 1.04–29.80, P = 0.024).
Conclusion
These results indicated that the three genetic variations within the TIM-3 gene promoter may be associated with the increased susceptibility to gastric cancer, especially among the haplotypes with the risk.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer is one of the most common causes of cancer-related deaths worldwide, causing an estimated 700,000 deaths annually. In recent decades, although the incidence and mortality rates have been declining in most European countries, gastric cancer is still one of the most commonly diagnosed malignancies in China [1, 2]. Both genetic and environmental factors, including Helicobacter pylori infection, have been implicated in the genesis of this deadly disease [2, 3]. However, the detailed genetic mechanisms involved in this lethal cancer have not yet been fully elucidated. TIM-3, which was identified as a specific cell surface marker of Th1 CD4+ T cells and was preferentially expressed on fully differentiated Th1 lymphocytes, but not on Th2 cells, was one of the Ig superfamily members [4]. In this process, TIM-3 was expressed at a late stage, suggesting that TIM-3 might not contribute to the T cell differentiation, but might perform a critical function in the Th1 cells transportation [5]. The inhibitor activity of TIM-3 was first described in a series of autoimmune diseases. In the autoimmune disease model, the administration of TIM-3 antibody would result in the activation and expansion of macrophage population, and then the more severe clinical symptom would be occurred [4]. Another research showed that the administration of TIM-3-Ig in vivo would result in T cell hyperproliferation and would abrogate the induction of tolerance during the development of immune response [6]. When TIM-3 was interacted with its ligand, galectin-9, the Th1 responses would be blockade by promoting the death of IFN-γ-inducing Th1 cells [7]. In the previous functional study, experimental data demonstrated that the soluble form of TIM-3 would reduce the antigen-specific T cell responses and down-regulate the anti-tumor immunity in vivo by inhibiting the Th1 responses [8]. Despite TIM-3 had been usually thought to express in T-cells, it was not limited. In a series of normal tissue cells and malignant epithelial tissues, the TIM-3 expression was detected and had been proved to accelerate the tumor cell proliferation by inhabiting the production of IL-2 and IFN-γ which were the pivotal cytokines to induce the CTL and NK cells differentiation [9]. Moreover, the agonism of TIM-3 would significantly exacerbate Th1-mediated pathology, suggesting that it possessed a potential negative regulatory role for the endogenous Tim3–Tim3 ligand in vivo [8]. These data suggested that the TIM-3–TIM-3 ligand pathway would contribute to the attenuation of Th1-mediated anti-tumor responses and would be in charge of the development of cancer. In this study, we investigated the five polymorphisms located in the promoter region (−574G/T, −882C/T, −1516G/T, and −1541C/T) and in the encoding region +4259T/G (amino acid substitution: arginine to leucine) of TIM-3 gene to assess whether the genetic variants would be involved in the gastric cancer susceptibility in the Chinese population.
Patients and methods
Patients and controls
This hospital-based case–control study was performed in Chinese Han population, which comprised of 212 gastric cancer cases and 252 normal controls. All of the subjects were of unrelated Han nationality. Patients with gastric cancer were confirmed histopathologically by endoscopic biopsy or surgical specimen and consecutively recruited from Beijing Friendship Hospital and Shandong Qianfushan Hospital between 2005 and 2010. The individuals with secondary, recurrent malignancies, and who accepted blood transfusion from non-self were excluded. The tumor location showed that there were 7 gastric cancers located in cardia, 116 in non-cardia, 5 in upper third, 46 in middle third, and 38 in lower third. During the same time of case collection, cancer-free controls were selected amongst inpatients from the same hospital. The recruited criteria included non-neoplastic diseases, and matched to gastric cancer cases by gender and age (within 5 years). Control subjects with severe clinical symptoms, previous diagnosis of cancer, and genetic disease were excluded. In the two groups, individuals who formerly or currently smoked ten cigarettes per day on average were defined as smokers, and who consumed wine or liquor 150 ml per day on average were defined as drinkers. The mean age of the gastric cancer and control group was 41.78 ± 13.20 and 40.02 ± 14.67 (mean ± standard deviation), respectively. There were 92 females and 120 males in the gastric cancer group, and 116 females and 136 males in the control group. Detailed information on lifetime tobacco use, alcohol consumption, and demographic background was recorded during a personal interview and exhibited in Table 1. Every participant had signed an informed consent approved by the Local Committee on Clinical Investigation. After the written consent was obtained, 2 ml peripheral intravenous blood was obtained from all the participants.
SNP selection and genotyping methods
As previous study described [10], genomic DNA was extracted from blood samples using sodium dodecyl sulfate lysis and proteinase K digestion, followed by the standard phenol–chloroform extraction. The five polymorphisms within TIM-3 gene promoter and encoding region were identified by the polymerase chain reaction–restriction fragment length polymorphism (PCR–RFLP) assay [11]. The primers and PCR conditions were as described previously and listed in Table 2 [12–14]. PCR was performed in a 20 μl total reaction volume containing 2 μl 10× PCR buffer (Qiagen Inc., Hilden, Germany), 1.5 mM MgCl2, 0.5 μM each primer (shown in Table 2), 0.2 mM dNTP, 1.2 U Taq polymerase (Qiagen Inc., Hilden, Germany) and 200 ng of genomic DNA. After an initial denaturation at 95°C for 5 min, the DNA was amplified for 35 cycles at 94°C for 30 s, 55–60°C for 40 s (detailed annealing temperatures shown in Table 2), and 72°C for 45 s, with a final elongation at 72°C for 10 min on the Gene-Amp PCR System 9700 (PE Applied Biosystems, Foster City, CA, USA). PCR products were purified using a MultiScreen-PCR purifying plate (Millipore Company, Billerica, MA, USA). The purified PCR products including the five polymorphic sites were then digested with restriction enzymes Taqα I, BsoB I, Bsl I, BsaJ I, and Pst I (New England Biolabs, Beverly, MA, USA), respectively, by using the conditions recommended in the manufacturer’s instructions. The digested PCR products were fractionated on 2% agarose Tris–borate–EDTA gel (Agarose 1000; Gibco BRL, Rockville, MD, USA) and stained with ethidium bromide (product size after digestion shown in Table 2). All assays were conducted blindly by two researchers without the knowledge of the case or control status. About 10% of the samples were randomly selected and retested, and the results were 100% concordant. Additionally, DNA direct sequencing was determined to confirm the RFLP result in ten patients and controls for all of the five polymorphic sites.
Statistical analysis
The Pearson chi-square test was used to assess the difference in the distributions of categorical variables and allele frequencies between cases and controls. Distribution of age variable was compared by the Mann–Whitney U test. Hardy–Weinberg equilibrium in cases and controls was assessed by using the chi-square test. For all genotypes or allele, the homozygote of the common allele was used as the referent. Unconditional logistic regression was used to calculate the odds ratios (ORs) and 95% confidence intervals (CIs) for the association between genotypes and gastric cancer adjusted by age, gender, cigarette smoking, and alcohol consumption. The statistical analyses were conducted by using the SPSS/Win statistical package (version 11.5.1 for Windows; SPSS Inc, Chicago, IL, USA). All tests were two-sided at the 0.05 significance level. The SHEsis program was used to assess the pair-wise linkage disequilibrium (LD) among the polymorphisms within TIM-3 gene [15]. Haplotypes were reconstructed from genotype data and were statistically analyzed by the SHEsis program (http://analysis.bio-x.cn/myAnalysis.php).
Results
Demographic information and polymorphism associated analysis
As expected, cases and controls did not differ with respect to age at enrolment (the continuous data was in normal distribution), gender distribution, cigarette smoking, and alcohol consume, all of which were matching variables (for all, P > 0.05; data shown in Table 1). The results for the five polymorphic alleles or genotypes of the TIM-3 gene in cases and controls were summarized in Table 1. All the genotype distributions among cases and controls were in coincidence with Hardy–Weinberg equilibrium (for all, P value > 0.05). Among the five examined polymorphisms, three polymorphisms −574G/T, −882C/T, and −1516G/T which were located in the TIM-3 gene promoter were significantly associated with gastric cancer risk (Table 1). Compared to the carriers of all copies of the common genotype, carriage of one or two copies of the minor allele were associated with a risk of gastric cancer after adjusting for age, gender, alcohol consumption, and smoking status. The ORs and 95% CIs were 2.74 (1.21–6.20), 3.19 (1.29–7.91), and 2.03 (1.15–3.59), respectively (all P value < 0.05). However, the participants who possessed −1541C/T polymorphism in the promoter region and +4259T/G polymorphism in the encoding region were not observed to increase the risk for gastric cancer (for both, P value > 0.05). The association between the clinical pathological characteristics such as the tumor size, degree of differentiation, TNM stage, lymph or distant metastasis of the gastric cancer and the polymorphism distributions were shown in Table 3. Under the multivariate unconditional logistic regression analysis, the statistical results eventually revealed that only the TIM-3 −1516 (G/T) polymorphism was more closely associated with the tumor distant metastasis (P = 0.036, OR = 2.21, 95% CI: 1.05–4.63).
Association between TIM-3 haplotype and gastric cancer
To illustrate whether the specific TIM-3 gene alleles might be associated with gastric cancer development, the haplotypes were constructed by using SHEsis program (http://analysis.bio-x.cn/myAnalysis.php) [15]. Haplotypes were derived from the patient group, the control group and the combined patient–control cohort. The analysis results assessed by SHEsis program demonstrated that −574G/T, −882C/T, −1516G/T, −1541C/T, and +4259T/G polymorphisms were in tight LD in case and control populations. As expected, we observed pronounced differences in the LD map, depending on the LD statistic data (Fig. 1). When LD was estimated with |D′|, the five contiguous polymorphisms were clustering in block, the strong LD was observed among the five polymorphisms (D′ > 0.80). Meanwhile, only a few infirm LD was represented (0.50 < D′ < 0.80). These data were shown in Table 4. Because no complete linkage disequilibrium (D′ = 1) was detected among the five polymorphisms, all of them were included in haplotype analysis by using the same program. A total of 15 haplotypes whose frequency was more than 0.1% were obtained, and two haplotypes significantly presented the relation to gastric cancer (Table 5). The frequency of haplotypes GCGCT and TTGCT was notably greater in the patients than in the controls (OR = 2.25, 95% CI: 1.44–3.52, P < 0.001; OR = 5.57, 95% CI: 1.04–29.80, P = 0.024, respectively).
Discussion
TIM-3 gene located on human chromosome 5q33.2 (GenBank accession nos. NT 023133.12) was described as a transmembrane protein gene and expressed preferentially on Th1 cells. The mature human TIM-3 protein consisted of a 181 amino acid extracellular domain, a 21 aa transmembrane segment, and a 78 aa cytoplasmic tail [4]. TIM-3 protein was expressed on the surface of activated Th1 cells, dendritic cells, or malignant epithelial tissues, and could trigger the apoptosis pathway of Th1 cells by binding galectin-9 to the TIM-3 extracellular domain [7, 9]. The higher TIM-3 level might result in an elevated CD80 expression of cell, and then the CD80 would preferentially interact with the inhibitory molecule CTLA-4 (cytotoxic T lymphocyte-associated antigen-4). This would eventually lead to a local immunosuppression [16]. The elevated expression of TIM-3 in cancer cells would reduce its adhering capacity and would contribute to the development of cancer. Therefore, TIM-3 probably functioned as a pivotal immunoregulatory molecule in carcinogenesis. As soluble TIM-3 had been shown to bind to TIM-3 ligand on CD4+ T cells and could inhibit the anti-tumor immunity, the increased numbers of full-length TIM-3 molecule on mast cells could also exert a similar effect [6]. Being a regulative factor of autoimmunity, TIM-3 should lead to be a specific induction of nuclear factor κB (NF-κB), which was confirmed to be an inductive molecule of the transcription factor cascade [6, 17, 18]. When the blocking anti-TIM-3 was used, the 75% inhibition of galectin-9-mediated TNF-α secretion from human monocytes was observed [18]. Above of these evidences, TIM-3 was speculated that it would contribute to the expansion of tumors.
In our present study, we performed a polymorphic screening experiment of TIM-3 gene in the Chinese population to detect the association with gastric cancer development. As a result, the genotypes and alleles of these three polymorphisms, −574G/T, −882C/T, and −1516G/T located in TIM-3 promoter region, were significantly greater in the patients than in the controls, suggesting that the three polymorphisms might be associated with the increased risk of gastric cancer. Moreover, under the clinical pathological characteristics analysis such as the tumor size, degree of differentiation, TNM stage, lymph node status and the distant metastasis in the gastric cancer group, the relationship between the −1516 polymorphic genotype and the distant metastasis was prominently found. In the next haplotype analysis, although 15 haplotypes were preliminarily constructed by the five polymorphisms and revealed significant frequency difference between the two groups, only the haplotypes GCGCT and TTGCT were much more in gastric cancer patients than in the controls, suggesting that it might be a genetic risk factor for gastric cancer development. These findings also strongly supported the previous conclusions that the common variations in TIM-3 gene were associated with the immune deficiency disease or malignancy in the Chinese or other European population [14, 19–22].
In summary, the present study was for the first time to determine the polymorphisms within TIM-3 gene as a risk factor for gastric cancer development in the Chinese population. Our statistical results suggested that the risk genetic variants located in TIM-3 gene promoter region would be involved in the susceptibility of gastric cancer. These findings were preliminary due to the small sample size in our present study. Further research should be performed in larger samples and different ethnic groups. It will be helpful to elucidate the molecular epidemiology of gastric cancer. Meanwhile, it should be essential to demonstrate the functional mechanisms of these genetic variants in vitro.
References
Karim-Kos HE, de Vries E, Soerjomataram I, Lemmens V, Siesling S, Coebergh JW (2008) Recent trends of cancer in Europe: a combined approach of incidence, survival and mortality for 17 cancer sites since the 1990s. Eur J Cancer 44:1345–1389. doi:10.1007/s00262-004-0645-2
Forman D, Burley VJ (2006) Gastric cancer: global pattern of the disease and an overview of environmental risk factors. Best Pract Res Clin Gastroenterol 20:633–649. doi:10.1016/j.bpg.2006.04.008
Tahara E Jr, Tahara H, Kanno M, Naka K, Takeda Y, Matsuzaki T, Yamazaki R, Ishihara H, Yasui W, Barrett JC, Ide T, Tahara E (2005) G1P3, an interferon inducible gene 6–16, is expressed in gastric cancers and inhibits mitochondrial-mediated apoptosis in gastric cancer cell line TMK-1 cell. Cancer Immunol Immunother 54:729–740. doi:10.1007/s00262-004-0645-2
Monney L, Sabatos CA, Gaglia JL, Ryu A, Waldner H, Chernova T, Manning S, Greenfield EA, Coyle AJ, Sobel RA, Freeman GJ, Kuchroo VK (2002) Th1-specific cell surface protein Tim-3 regulates macrophage activation and severity of an autoimmune disease. Nature 415:536–541. doi:10.1038/415536a
Wang Y, Meng J, Wang X, Liu S, Shu Q, Gao L, Ju Y, Zhang L, Sun W, Ma C (2008) Expression of human TIM-1 and TIM-3 on lymphocytes from systemic lupus erythematosus patients. Scand J Immunol 67:63–70. doi:10.1111/j.1365-3083.2007.02038.x
Sabatos CA, Chakravarti S, Cha E, Schubart A, Sanchez-Fueyo A, Zheng XX, Coyle AJ, Strom TB, Freeman GJ, Kuchroo VK (2003) Interaction of Tim-3 and Tim-3 ligand regulates T helper type 1 responses and induction of peripheral tolerance. Nat Immunol 4:1102–1110. doi:10.1038/ni988
Zhu C, Anderson AC, Schubart A, Xiong H, Imitola J, Khoury SJ, Zheng XX, Strom TB, Kuchroo VK (2005) The Tim-3 ligand galectin-9 negatively regulates T helper type 1 immunity. Nat Immunol 6:1245–1252. doi:10.1038/ni1271
Geng H, Zhang GM, Li D, Zhang H, Yuan Y, Zhu HG, Xiao H, Han LF, Feng ZH (2006) Soluble form of T cell Ig mucin 3 is an inhibitory molecule in T cell-mediated immune response. J Immunol 176:1411–1420. doi:176/3/1411
van de Weyer PS, Muehlfeit M, Klose C, Bonventre JV, Walz G, Kuehn EW (2006) A highly conserved tyrosine of Tim-3 is phosphorylated upon stimulation by its ligand galectin-9. Biochem Biophys Res Commun 351:571–576. doi:10.1016/j.bbrc.2006.10.079
Naumova E, Mihaylova A, Stoitchkov K, Ivanova M, Quin L, Toneva M (2005) Genetic polymorphism of NK receptors and their ligands in melanoma patients: prevalence of inhibitory over activating signals. Cancer Immunol Immunother 54:172–178. doi:10.1007/s00262-004-0575-z
Qi P, Chen YM, Wang H, Fang M, Ji Q, Zhao YP, Sun XJ, Liu Y, Gao CF (2009) 509C > T polymorphism in the TGF-beta1 gene promoter, impact on the hepatocellular carcinoma risk in Chinese patients with chronic hepatitis B virus infection. Cancer Immunol Immunother 58:1433–1440. doi:10.1007/s00262-009-0660-4
Chae SC, Song JH, Pounsambath P, Yuan HY, Lee JH, Kim JJ, Lee YC, Chung HT (2004) Molecular variations in Th1-specific cell surface gene Tim-3. Exp Mol Med 36:274–278.
Zhang CC, Wu JM, Cui TP, Wang P, Pan SX (2006) Study on relationship between polymorphism sites of TIM-3 and allergic asthma in a population of adult Hans from Hubei province of China. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 23:74–77.
Du WT, Zhao HF, Xu JH, Gu DS, Xue F, Ge J, Dong XW, Chen ZP, Zhou ZP, Yang RC (2009) The role of T-cell immunoglobulin- and mucin-domain-containing molecule-3 polymorphisms in idiopathic thrombocytopenic purpura. Hum Immunol 70:398–402. doi:10.1016/j.humimm.2009.03.013
Li Z, Zhang Z, He Z, Tang W, Li T, Zeng Z, He L, Shi Y (2009) A partition–ligation–combination–subdivision EM algorithm for haplotype inference with multiallelic markers: update of the SHEsis (http://analysis.bio-x.cn). Cell Res 19:519-523. doi:10.1038/cr.2009.33
Zheng Y, Manzotti CN, Liu M, Burke F, Mead KI, Sansom DM (2004) CD86 and CD80 differentially modulate the suppressive function of human regulatory T cells. J Immunol 172:2778–2784.
Sanchez-Fueyo A, Tian J, Picarella D, Domenig C, Zheng XX, Sabatos CA, Manlongat N, Bender O, Kamradt T, Kuchroo VK, Gutierrez-Ramos JC, Coyle AJ, Strom TB (2003) Tim-3 inhibits T helper type 1-mediated auto- and alloimmune responses and promotes immunological tolerance. Nat Immunol 4:1093–1101. doi:10.1038/ni987
Anderson AC, Anderson DE, Bregoli L, Hastings WD, Kassam N, Lei C, Chandwaskar R, Karman J, Su EW, Hirashima M, Bruce JN, Kane LP, Kuchroo VK, Hafler DA (2007) Promotion of tissue inflammation by the immune receptor Tim-3 expressed on innate immune cells. Science 318:1141–1143. doi:10.1126/science.1148536
Nagahara K, Arikawa T, Oomizu S, Kontani K, Nobumoto A, Tateno H, Watanabe K, Niki T, Katoh S, Miyake M, Nagahata S, Hirabayashi J, Kuchroo VK, Yamauchi A, Hirashima M (2008) Galectin-9 increases Tim-3+ dendritic cells and CD8+ T cells and enhances antitumor immunity via galectin-9-Tim-3 interactions. J Immunol 181:7660–7669. doi:181/11/7660
Anderson DE (2007) TIM-3 as a therapeutic target in human inflammatory diseases. Expert Opin Ther Targets 11:1005–1009. doi:10.1517/14728222.11.8.1005
Page NS, Jones G, Stewart GJ (2006) Genetic association studies between the T cell immunoglobulin mucin (TIM) gene locus and childhood atopic dermatitis. Int Arch Allergy Immunol 141:331–336. doi:10.1159/000095459
Wu QW, Cai PC, Wang L, Li YR, Kong LL, Hu LH (2009) Family-based association study of Tim-1 and Tim-3 gene polymorphisms with childhood asthma in Chinese trios. Int Arch Allergy Immunol 150:252–260. doi:10.1159/000222677
Acknowledgments
This study was supported by grants from the National High Technology Research and Development Program of China (2007AA02Z4Z4 to Shutian Zhang) and the Beijing Municipal Natural Science Foundation (7092103 to Bangwei Cao).
Author information
Authors and Affiliations
Corresponding author
Additional information
B. Cao and L. Zhu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Cao, B., Zhu, L., Zhu, S. et al. Genetic variations and haplotypes in TIM-3 gene and the risk of gastric cancer. Cancer Immunol Immunother 59, 1851–1857 (2010). https://doi.org/10.1007/s00262-010-0910-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00262-010-0910-5